Abstract
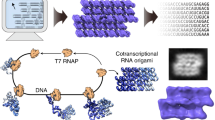
We have recently introduced an experimental method for the design and production of RNA-origami nanostructures that fold up from a single strand while the RNA is being enzymatically produced, commonly referred to as cotranscriptional folding. To realize a general and scalable architecture we have developed a theoretical framework for determining RNA crossover geometries, long-distance interactions, and strand paths that are topologically compatible with cotranscriptional folding. Here, we introduce a simple parameterized model for the A-form helix and use it to determine the geometry and base-pair spacing for the five types of RNA double-crossover molecules and the curvature resulting from crossovers between multiple helices. We further define a set of paranemic loop-loop and end-to-end interactions compatible with the design of folding paths for RNA structures with arbitrary shape and programmable curvature. Finally, we take inspiration from space-filling curves in mathematics to design strand paths that have high-locality, programmed folding kinetics to avoid topological traps, and structural repeat units that might be used to create infinite RNA ribbons and squares by rolling circle transcription.
Access provided by Autonomous University of Puebla. Download to read the full chapter text
Chapter PDF
Similar content being viewed by others
References
Afonin, K.A., Cieply, D.J., Leontis, N.B.: Specific RNA self-assembly with minimal paranemic motifs. Journal of the American Chemical Society 130, 93–102 (2008)
Andersen, E.S.: Prediction and design of DNA and RNA structures. New Biotechnology 27, 184–193 (2010)
Arnott, S., Hukins, D.W., Dover, S.D., Fuller, W., Hodgson, A.R.: Structures of synthetic polynucleotides in the A-RNA and A’-RNA conformations: x-ray diffraction analyses of the molecular conformations of polyadenylic acid–polyuridylic acid and polyinosinic acid–polycytidylic acid. Journal of Molecular Biology 81, 107–122 (1973)
Costa, M., Michel, F.: Rules for RNA recognition of GNRA tetraloops deduced by in vitro selection: comparison with in vivo evolution. The EMBO Journal 16, 3289–3302 (1997)
Delebecque, C.J., Lindner, A.B., Silver, P.A., Aldaye, F.A.: Organization of intracellular reactions with rationally designed RNA assemblies. Science 333, 470–474 (2011)
Dickerson, R.E., Drew, H.R., Conner, B.N., Wing, R.M., Fratini, A.V., Kopka, M.L.: The anatomy of A-, B-, and Z-DNA. Science 216, 475–485 (1982)
Drew, H.R., Wing, R.M., Takano, T., Broka, C., Tanaka, S., Itakura, K., Dickerson, R.E.: Structure of a B-DNA dodecamer: conformation and dynamics. Proceedings of the National Academy of Sciences of the United States of America 78, 2179–2183 (1981)
Ennifar, E., Walter, P., Ehresmann, B., Ehresmann, C., Dumas, P.: Crystal structures of coaxially stacked kissing complexes of the HIV-1 RNA dimerization initiation site. Nat. Struct. Biol. 12, 1064–1068 (2001)
Fu, T.J., Seeman, N.C.: DNA double-crossover molecules. Biochemistry 32, 3211–3220 (1993)
Geary, C., Baudrey, S., Jaeger, L.: Comprehensive features of natural and in vitro selected GNRA tetraloop-binding receptors. Nucleic Acids Research 36, 1138–1152 (2008)
Geary, C., Chworos, A., Jaeger, L.: Promoting RNA helical stacking via A-minor junctions. Nucleic Acids Research 39, 1066–1080 (2011)
Geary, C.W., Rothemund, P.W.K., Andersen, E.S.: A single-stranded architecture for cotranscriptionally folded RNA tiles. Accepted in Science (2014)
Grabow, W.W., Jaeger, L.: RNA Self-Assembly and RNA Nanotechnology. Accounts of Chemical Research (2014)
Hao, C., Li, X., Tian, C., Jiang, W., Wang, G., Mao, C.: Construction of RNA nanocages by re-engineering the packaging RNA of Phi29 bacteriophage. Nature Communications 5, 3890 (2014)
Jaeger, L., Westhof, E., Leontis, N.B.: TectoRNA: modular assembly units for the construction of RNA nano-objects. Nucleic Acids Research 29, 455–463 (2001)
Jossinet, F., Ludwig, T.E., Westhof, E.: Assemble: an interactive graphical tool to analyze and build RNA architectures at the 2D and 3D levels. Bioinformatics 26, 2057–2059 (2010)
Ko, S.H., et al.: Synergistic self-assembly of RNA and DNA molecules. Nature Chemistry 2, 1050–1055 (2010)
Larson, M.H., et al.: A pause sequence enriched at translation start sites drives transcription dynamics in vivo. Science 344, 1042 (2014)
Lee, A.J., Crothers, D.M.: The solution structure of an RNA loop-loop complex: the ColE1 inverted loop sequence. Structure 6, 993–1007 (1998)
Lee, J.B., Hong, J., Bonner, D.K., Poon, Z., Hammond, P.T.: Self-assembled RNA interference microsponges for efficient siRNA delivery. Nature Materials 11, 316–322 (2012)
Lin, C., Wang, X., Liu, Y., Seeman, N.C., Yan, H.: Rolling circle enzymatic replication of a complex multi-crossover DNA nanostructure. Journal of the American Chemical Society 129, 14475–14481 (2007)
Rothemund, P.W.: Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006)
Severcan, I., Geary, C., Chworos, A., Voss, N., Jacovetty, E., Jaeger, L.: A polyhedron made of tRNAs. Nature Chemistry 2, 772–779 (2010)
Sherman, W.B., Seeman, N.C.: Design of minimally strained nucleic Acid nanotubes. Biophysical Journal 90, 4546–4557 (2006)
Shih, W.M., Quispe, J.D., Joyce, G.F.: A 1.7-kilobase single-stranded DNA that folds into a nanoscale octahedron. Nature 427, 618–621 (2004)
Wang, J.C.: Helical repeat of DNA in solution. Proceedings of the National Academy of Sciences of the United States of America 76, 200–203 (1979)
Woo, S., Rothemund, P.W.: Programmable molecular recognition based on the geometry of DNA nanostructures. Nature Chemistry 3, 620–627 (2011)
Yan, H., Zhang, X., Shen, Z., Seeman, N.C.: A robust DNA mechanical device controlled by hybridization topology. Nature 415, 62–65 (2002)
Yoffe, A.M., Prinsen, P., Gelbart, W.M., Ben-Shaul, A.: The ends of a large RNA molecule are necessarily close. Nucleic Acids Research 39, 292–299 (2011)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer International Publishing Switzerland
About this paper
Cite this paper
Geary, C.W., Andersen, E.S. (2014). Design Principles for Single-Stranded RNA Origami Structures. In: Murata, S., Kobayashi, S. (eds) DNA Computing and Molecular Programming. DNA 2014. Lecture Notes in Computer Science, vol 8727. Springer, Cham. https://doi.org/10.1007/978-3-319-11295-4_1
Download citation
DOI: https://doi.org/10.1007/978-3-319-11295-4_1
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-11294-7
Online ISBN: 978-3-319-11295-4
eBook Packages: Computer ScienceComputer Science (R0)