Abstract
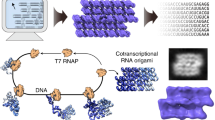
RNA nanostructures can be used as scaffolds to organize, combine, and control molecular functionalities, with great potential for applications in nanomedicine and synthetic biology. The single-stranded RNA origami method allows RNA nanostructures to be folded as they are transcribed by the RNA polymerase. RNA origami structures provide a stable framework that can be decorated with functional RNA elements such as riboswitches, ribozymes, interaction sites, and aptamers for binding small molecules or protein targets. The rich library of RNA structural and functional elements combined with the possibility to attach proteins through aptamer-based binding creates virtually limitless possibilities for constructing advanced RNA-based nanodevices.
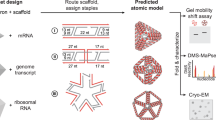
In this chapter we provide a detailed protocol for the single-stranded RNA origami design method using a simple 2-helix tall structure as an example. The first step involves 3D modeling of a double-crossover between two RNA double helices, followed by decoration with tertiary motifs. The second step deals with the construction of a 2D blueprint describing the secondary structure and sequence constraints that serves as the input for computer programs. In the third step, computer programs are used to design RNA sequences that are compatible with the structure, and the resulting outputs are evaluated and converted into DNA sequences to order.
The original version of this chapter was revised. The erratum to this chapter is available at: DOI 10.1007/978-1-4939-6454-3_20
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Jaeger L, Chworos A (2006) The architectonics of programmable RNA and DNA nanostructures. Curr Opin Struct Biol 16:531–543
Fu TJ, Seeman NC (1993) DNA double-crossover molecules. Biochemistry 32:3211–3220
Grabow WW, Jaeger L (2014) RNA self-assembly and RNA nanotechnology. Acc Chem Res 47:1871–1880
Geary C, Rothemund PW, Andersen ES (2014) A single-stranded architecture for cotranscriptional folding of RNA nanostructures. Science 345:799–804
Geary C, Andersen ES (2014) Design principles for single-stranded RNA origami structures. Lecture notes in computer science: DNA computing and molecular programming (DNA20)
Dietz H, Douglas SM, Shih WM (2009) Folding DNA into twisted and curved nanoscale shapes. Science 325:725–730
Ikeda RA, Richardson CC (1986) Interactions of the RNA polymerase of bacteriophage T7 with its promoter during binding and initiation of transcription. Proc Natl Acad Sci U S A 83:3614–3618
Grabow WW et al (2011) Self-assembling RNA nanorings based on RNAI/II inverse kissing complexes. Nano Lett 11:878–887
Severcan I et al (2010) A polyhedron made of tRNAs. Nat Chem 2:772–779
Baklanov MM, Golikova LN, Malygin EG (1996) Effect on DNA transcription of nucleotide sequences upstream to T7 promoter. Nucleic Acids Res 24:3659–3660
Afonin KA et al (2010) In vitro assembly of cubic RNA-based scaffolds designed in silico. Nat Nanotechnol 5:676–682
Acknowledgement
This work was supported by a Sapere Aude Starting Grant from the Danish Council for Independent Research (DFF-0602-01772) and the Centre for DNA Nanotechnology (http://cdna.au.dk/) funded by the Danish National Research Foundation (DNRF81).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media New York
About this protocol
Cite this protocol
Sparvath, S.L., Geary, C.W., Andersen, E.S. (2017). Computer-Aided Design of RNA Origami Structures. In: Ke, Y., Wang, P. (eds) 3D DNA Nanostructure. Methods in Molecular Biology, vol 1500. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6454-3_5
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6454-3_5
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6452-9
Online ISBN: 978-1-4939-6454-3
eBook Packages: Springer Protocols