Abstract
In recent years, several associations between common chronic human disorders and altered gut microbiome composition and function have been reported1,2. In most of these reports, treatment regimens were not controlled for and conclusions could thus be confounded by the effects of various drugs on the microbiota, which may obscure microbial causes, protective factors or diagnostically relevant signals. Our study addresses disease and drug signatures in the human gut microbiome of type 2 diabetes mellitus (T2D). Two previous quantitative gut metagenomics studies of T2D patients that were unstratified for treatment yielded divergent conclusions regarding its associated gut microbial dysbiosis3,4. Here we show, using 784 available human gut metagenomes, how antidiabetic medication confounds these results, and analyse in detail the effects of the most widely used antidiabetic drug metformin. We provide support for microbial mediation of the therapeutic effects of metformin through short-chain fatty acid production, as well as for potential microbiota-mediated mechanisms behind known intestinal adverse effects in the form of a relative increase in abundance of Escherichia species. Controlling for metformin treatment, we report a unified signature of gut microbiome shifts in T2D with a depletion of butyrate-producing taxa3,4. These in turn cause functional microbiome shifts, in part alleviated by metformin-induced changes. Overall, the present study emphasizes the need to disentangle gut microbiota signatures of specific human diseases from those of medication.
Similar content being viewed by others
Main
T2D is a disorder of elevated blood glucose levels (hyperglycaemia) primarily due to insulin resistance and inadequate insulin secretion, with rising global prevalence. Genetic and environmental risk factors are known, the latter including dietary habits and a sedentary lifestyle5. Gut microbiota involvement is also increasingly recognized3,4,6,7, although findings diverge between studies8; for example, Qin et al.3 report several Clostridium species enriched in T2D, whereas Karlsson et al.4 instead report enrichment of several lactobacilli species (see Supplementary Discussion). Treatment involves medication and lifestyle intervention, which may confound reported gut dysbiosis. Many T2D patients receive metformin, an oral blood-glucose-lowering non-metabolizable compound whose primary and dominant metabolic effect is the inhibition of liver glucose production9. At least 30% of patients report adverse effects including diarrhoea, nausea, vomiting and bloating, with underlying mechanisms poorly understood. Studies in animals10 and humans11 suggest that some beneficial effects of metformin on glucose metabolism may be microbially mediated. Here, we built a multi-country T2D metagenomic data set, starting with gut microbial samples from a nondiabetic Danish cohort of 277 individuals within the MetaHIT project12 and additional novel Danish MetaHIT metagenomes from 75 T2D and 31 type 1 diabetes (T1D) patients, sequenced using the same protocols (samples abbreviated as MHD). Treatment information was obtained for all MHD samples, as well as for samples from a previously reported4 cohort of 53 female Swedish T2D patients, along with 92 nondiabetic individuals (43 with normal glucose tolerance, 49 with impaired glucose tolerance) (SWE) and a subgroup of 71 Chinese T2D patients with available information on antidiabetic treatment as well as 185 nondiabetic Chinese individuals3 (CHN). For these 784 gut metagenomes (Supplementary Table 1), taxonomic and functional profiles were determined (see Methods), verifying our meta-analysis framework to be appropriate and robust in the context of theoretical considerations and through simulations (Supplementary Discussion 1 and Extended Data Fig. 1a), as well as characterizing differences between the data sets (Extended Data Fig. 2). Initial analysis unstratified for treatment but controlling for demographic and technical variation between data sets (Supplementary Discussion 2 and Supplementary Table 2) recovered a majority of previously reported associations (Supplementary Discussion 2 and Supplementary Table 3) but with large divergence between data sets. Suspecting confounding treatments, we tested for influence of diet and antidiabetic medications (Supplementary Discussion 3, Supplementary Table 4 and Extended Data Fig. 1b), finding an effect resulting only from use of metformin. As the fraction of medicated patients (denoted as T2D metformin+) varied strongly (21% CHN, 38% SWE and 77% MHD), samples were stratified on metformin treatment status. Multivariate analysis showed significant (permutational multivariate analysis of variance (PERMANOVA) false discovery rate (FDR) < 0.005) differences in gut taxonomic composition between metformin-untreated T2D (T2D metformin−) (n = 106) patients and nondiabetic controls (ND control) (n = 554), consistent with a broad-range dysbiosis in T2D (Fig. 1a and Supplementary Table 5; see also Extended Data Table 1a and Supplementary Discussion 3 for an analysis of variances broken down by source). While metformin treatment status could be reliably recovered from microbial composition using support vector machines, metformin-untreated T2D status itself could not (Fig. 1b and Supplementary Table 6). In contrast, in all three cohorts, drug-treatment-blinded T2D samples could be separated from ND control samples with similar accuracy as previously reported3,4, suggesting that the T2D metformin+ classifier robustly outperforms T2D metformin− classifiers across data sets (Supplementary Table 7).
Major treatment effects are seen in multivariate analysis and in classifier performance. a, Projection of genus-level gut microbiome samples from Danish, Chinese and Swedish studies constrained by diabetic state and metformin treatment. Multivariate analysis (dbRDA plot based on Canberra distances between bacterial genera) reveals a T2D dysbiosis, which overlaps only in part with taxonomic changes in metformin-treated patients. The ordination projects all T2D metformin+ (n = 93, dark red), T2D metformin− (n = 106, orange) and ND control (n = 554, teal) gut metagenomes, with confounding country effect adjusted for. Bacterial genera that show significant effects of metformin treatment and T2D status compared to ND control, respectively (limited to top five for each), are interpolated into the plane of maximal separation based on their abundances across all samples. Marginal box/scatter plots show the separation of the constrained projection coordinates (boxes show medians/quartiles, error bars extend to most extreme value within 1.5 interquartile ranges). The T2D separation is significant (PERMANOVA FDR < 0.005) in the joint data set and independently significant in CHN and MHD samples. The metformin separation is significant (PERMANOVA FDR < 0.1; Canberra distances) in MHD and SWE samples. bp, butyrate-producing. b, Classifying T2D and metformin treatment status based on gut microbiome profiles. Support vector machine classifiers were used to separate T2D metformin+ (n = 93), T2D metformin− (n = 106) and ND control (n = 554) gut metagenomes from each other based on genus-level gut microbiome taxonomic composition. Bold curves represent mean performance in hold-out testing of 1 out of 5 of the data each time, with separate tests shown as dashed curves. Error bars show ±1 s.d. Metformin-treated T2D samples can be well separated from controls (using Intestinibacter abundance as the only feature), whereas distinguishing T2D metformin− samples from ND control samples works poorly even in the best case, requiring 63 distinct microbial features to achieve this separation. ROC-AUC, area under the receiving operating characteristic curve.
We further explored T2D gut microbiome alterations in 106 metformin-untreated T2D compared with 554 ND control samples through univariate tests of microbial taxonomic and functional differences, with significant trends shown in Fig. 2a. Metformin-untreated T2D was associated with a decrease in genera containing known butyrate producers such as Roseburia spp., Subdoligranulum spp. and a cluster of butyrate-producing Clostridiales spp. (Supplementary Table 8), consistent with previous indications3,4. More fine-grained taxonomic analysis indicated some driver species (Supplementary Discussion 4 and Supplementary Table 9), and further found changes in abundance of several unclassified Firmicutes, often reduced or reversed under metformin treatment (see Supplementary Discussion 4). Although an increase in Lactobacillus spp. was seen in treatment-unstratified T2D samples (as previously found experimentally13), this trend was eliminated or reversed when controlling for metformin. Functionally, we found enrichment of catalase (conceivably a response to increased peroxide stress under inflammation) and modules for ribose, glycine and tryptophan amino acid degradation, but a decrease in threonine and arginine degradation, and in pyruvate synthase capacity (Supplementary Table 10). While these functional differences could result from strain-level composition changes or be a compound effect of subtle enrichment/depletion of larger ecological guilds, the abundance of most of these modules correlated with abundance of the significantly altered microbial genera (Fig. 2a).
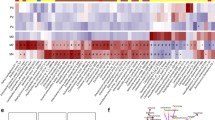
Differences between healthy controls and T2D patients contrasted against T1D as an alternative form of dysglycaemia. a, Taxonomic and functional microbiome signatures of metformin-naive T2D. The heat maps show bacterial genera (horizontal axis) and microbial gene functions (vertical axis) that are significantly (study-source-adjusted Kruskal–Wallis test and post-hoc Mann–Whitney U-test, markers in innermost marginal heat maps indicating *FDR < 0.05, +FDR < 0.1) different in abundance (nonparametric enrichment scores shown as intensity of innermost marginal heat maps; red–green colour scale) between T2D metformin− (n = 106) and ND control (n = 554) gut metagenomes, revealing a robust diabetic signature across data sets. None of these features is significantly different in a comparison of T1D (n = 31) with ND control (n = 277) gut metagenomes (outermost marginal heat maps, same notation as above), implying that they are not direct effects of dysglycaemia. The central heat map shows Spearman correlations (purple–red colour scale) between abundance of bacterial taxa and microbial gene modules (Spearman test FDR scores: *FDR < 0.05, ***FDR < 0.001). b, Elevated gene richness in adult type 1 diabetes samples. Comparing MHD samples only, T1D (n = 31) gut metagenomes show significantly (Mann–Whitney U-test, +FDR < 0.1, *FDR < 0.05) higher gut microbiome richness (that is, gene count) than all other sample subsets (n = 277 ND control, n = 58 T2D metformin+, n = 17 T2D metformin− gut metagenomes). Sample median richness is shown as horizontal black bars.
To interpret our findings on T2D gut microbiota shifts further, we compared them with 31 adult T1D patients (Supplementary Table 1; for further discussion of this sub-cohort, see also Supplementary Discussion 5 and Supplementary Tables 6 and 11). This group is dysglycaemic like T2D patients, allowing us to separate purely glycaemic phenotype effects from T2D-specific microbial features. Gene richness was significantly increased in the T1D microbiomes (Wilcoxon rank sum test FDR < 0.1) (Fig. 2b), but was reduced in T2D (Supplementary Table 10), as reported previously6. Features found to distinguish metformin-untreated T2D from ND control microbiomes did not replicate when comparing T1D to ND control. Instead, most differences between metformin-untreated T2D samples and ND controls were reversed in adult T1D patients. In contrast, some microbial functions differentially abundant between metformin-untreated T2D and controls showed similar trends in T1D samples (Fig. 2a), although not significantly, possibly owing to lower statistical power. We therefore conclude that the majority of gut microbiota shifts visible in metformin-untreated T2D are not simply effects of dysglycaemia, but rather directly or indirectly associated with the causes or progression of T2D.
Suspecting microbial mediation of some of the therapeutic effects of metformin, we next compared T2D metformin-treated (n = 93) and T2D metformin-untreated (n = 106) samples to characterize the treatment effect in more detail. Multivariate contrasts of T2D metformin-treated with T2D metformin-untreated samples appeared weaker than those between T2D metformin-untreated and ND control samples, the former only significant at the bacterial family level (PERMANOVA FDR < 0.1), suggesting that the effects of metformin treatment on gut microbial composition are poorly captured by multivariate analysis. Univariate tests of the effects of metformin treatment showed a significant increase of Escherichia spp. and a reduced abundance of Intestinibacter spp., the latter fully consistent across the different country data sets (Fig. 3a), whereas the former is not seen in the CHN cohort where both diabetic individuals and controls are enriched in Escherichia spp. relative to Scandinavian controls. Correcting for differences in gender, body mass index and fasting levels of plasma glucose or serum insulin (some of which were significantly different between data sets, Supplementary Table 12) retained these differences as significant (Supplementary Table 13). Fasting serum concentrations of metformin were obtained for the MHD cohort and correlated significantly with abundances of both genera (Fig. 3b). Amplicon-based analysis of an independent T2D cohort likewise validated an increase of Escherichia spp. and a reduced abundance of Intestinibacter spp. in metformin-treated patients (Extended Data Fig. 1c, Extended Data Table 1b and Supplementary Discussion 6). The metformin-associated changes might derive from taxon-specific resistance/sensitivity to the bacteriostatic or bactericidal properties of the drug14. The genus Intestinibacter was defined only recently15 and includes the human isolate Clostridium bartletti16, since reclassified as Intestinibacter bartlettii. Little is known about its role in the gut ecosystem and how it might affect human health. However, I. bartlettii abundances were lower in pigs susceptible to colonization by enterotoxigenic Escherichia spp.17, consistent with the pattern seen here following metformin treatment. Analysis of the SEED (see Supplementary Discussion 7) and GMM (see Methods) functional annotations linked to Intestinibacter shows it to be resistant to oxidative stress and able to degrade fucose, indicative of an indirect involvement in mucus degradation. It also appears to possess the genetic potential for sulfite reduction, including part of an assimilatory sulfate reduction pathway. Analysis of gut microbial functional potential more generally suggested that indirect metformin treatment effects (Fig. 3c), including reduced intestinal lipid absorption18 and lipopolysaccharide (LPS)-triggered local inflammation, can provide a competitive advantage to Escherichia species19, possibly triggering a positive feedback loop that further contributes to the observed taxonomic changes. At the same time, metformin may reverse T2D-associated changes, as several gut microbial genera were more similar in abundance to ND control levels under metformin treatment, notably Subdoligranulum and to some extent Akkermansia. The latter was previously shown to reduce insulin resistance in murine models when increased in abundance through prebiotics20, and has been shown to similarly increase in abundance under metformin treatment10,21. In human samples, however, the trend was inconsistent between country subsets, and only MHD samples show a similar response (Extended Data Fig. 3). With respect to microbiota-mediated impact on host glucose regulation, the functional analyses demonstrated significantly enhanced butyrate and propionate production potential in metformin-treated individuals (Fig. 3c and Supplementary Table 14). Interestingly, recent studies in mice have shown that an increase in colonic production of these short-chain fatty acids triggers intestinal gluconeogenesis (IGN) via complementary mechanisms. Butyrate activates IGN gene expression through a cAMP-dependent mechanism in enterocytes, whereas propionate, itself a substrate of IGN, activates IGN gene expression via the portal nervous system and the fatty acid receptor FFAR3 (refs 22, 23). In rodents, the net result of increased IGN is a beneficial effect on glucose and energy homeostasis with reductions in hepatic glucose production, appetite and body weight. Taken together, our characterization of a metformin-associated human gut microbiome suggests novel mechanisms contributing to the beneficial effects of the drug on host metabolism.
Characterization of the microbially mediated therapeutic and adverse effects of metformin. a, Gut microbial shifts under metformin treatment. Metformin treatment significantly (study-source-adjusted Kruskal–Wallis test and post-hoc Mann–Whitney U-test, *FDR < 0.05, ***FDR < 0.001) increases Escherichia and lowers Intestinibacter abundance. Box plots show median/quartile abundances, whiskers extend to 1.58× interquartile range/ , for T2D metformin+ (nCHN = 15, nMHD = 58, nSWE = 20), T2D metformin− (nCHN = 56, nMHD = 17, nSWE = 33) and ND control (nCHN = 185, nMHD = 277, nSWE = 92) gut metagenome samples. b, Correlations between serum levels of metformin and gut microbiota in Danish MetaHIT samples, including short-chain fatty acid production modules. Serum metformin levels of T2D patients (n = 75 gut metagenomes) are significantly (Spearman FDR < 0.1) positively correlated with Escherichia abundance, and in significant negative correlation with Intestinibacter abundance. Bacterial gene function modules for butyrate and propionate production increase in abundance as serum metformin levels increase. Dot markers are shown for all MHD samples for which serum metformin concentration was measured. Metformin-untreated T2D samples (serum concentration <10 mg ml−1) are shown in orange, treated samples in dark red. Spearman coefficients (ρ; calculated for treated samples only) and FDRs (Q) are shown. c, Microbial shifts under metformin treatment contribute to improved glucose control and to adverse effects. Schematic illustration of gut microbial changes and their impact on host health. Observed associations (orange lines) between microbial taxa abundances (orange ellipses), microbial functional potential (orange boxes), and blood values (filled orange boxes) and metformin treatment are linked with literature-derived metformin- or microbiota-induced host physiological effects (blue boxes and arrows; dashed arrows indicate hypothesized causality). Drug–host–microbiota interactions can contribute to previously described therapeutic (green triangles) and side (red triangles) effects of metformin treatment.
, for T2D metformin+ (nCHN = 15, nMHD = 58, nSWE = 20), T2D metformin− (nCHN = 56, nMHD = 17, nSWE = 33) and ND control (nCHN = 185, nMHD = 277, nSWE = 92) gut metagenome samples. b, Correlations between serum levels of metformin and gut microbiota in Danish MetaHIT samples, including short-chain fatty acid production modules. Serum metformin levels of T2D patients (n = 75 gut metagenomes) are significantly (Spearman FDR < 0.1) positively correlated with Escherichia abundance, and in significant negative correlation with Intestinibacter abundance. Bacterial gene function modules for butyrate and propionate production increase in abundance as serum metformin levels increase. Dot markers are shown for all MHD samples for which serum metformin concentration was measured. Metformin-untreated T2D samples (serum concentration <10 mg ml−1) are shown in orange, treated samples in dark red. Spearman coefficients (ρ; calculated for treated samples only) and FDRs (Q) are shown. c, Microbial shifts under metformin treatment contribute to improved glucose control and to adverse effects. Schematic illustration of gut microbial changes and their impact on host health. Observed associations (orange lines) between microbial taxa abundances (orange ellipses), microbial functional potential (orange boxes), and blood values (filled orange boxes) and metformin treatment are linked with literature-derived metformin- or microbiota-induced host physiological effects (blue boxes and arrows; dashed arrows indicate hypothesized causality). Drug–host–microbiota interactions can contribute to previously described therapeutic (green triangles) and side (red triangles) effects of metformin treatment.
Both on a compositional and functional level, we found significant microbiome alterations that are consistent with well-known side-effects of metformin treatment (Fig. 3c). Most of these metformin-associated functional shifts, including enrichment of virulence factors and gas metabolism genes, could be attributed to the significantly increased abundance of Escherichia species (Supplementary Discussion 7 and Supplementary Tables 14 and 15).
In conclusion, our results suggest partial gut microbial mediation of both therapeutic and adverse effects of the most widely used antidiabetic medication, metformin, although further validation is required to conclude causality and to clarify how such mediation might occur. Our study of T2D illustrates the need to disentangle specific disease dysbioses from effects of treatment on the human-associated microbiota. The importance of this point was further shown by the fact that the previously reported high accuracy3,4 of gut microbial signatures for identifying patients with treatment-unstratified T2D decreased markedly when considering a large set of metformin-naive patients only, highlighting a general need to bear treatment regimens in mind both when developing and applying microbiome-based diagnostic and prognostic tools for common disorders or their pre-morbidity states.
Methods
No statistical methods were used to predetermine sample size.
Danish MetaHIT diabetic study
Patient recruitment, enrolment and processing. Patients with T2D were either recruited from the Inter99 study population24 or from the out-patient clinic at Steno Diabetes Center, Gentofte, Denmark. Patients with known T2D were included if the patient had clinically defined T2D on the day of examination according to the WHO definition25. Inclusion criteria were fasting serum C-peptide above 200 pmol l−1 and negative testing for serum glutamic acid decarboxylase (GAD) 65 antibodies (to exclude T1D, latent autoimmune diabetes in adults), no secondary forms of diabetes like chronic pancreatitis diabetes or syndromic diabetes, no antibiotic treatment 2 months before inclusion, and no known gastro-intestinal diseases, no previous bariatric surgery or medication known to affect the immune system.
All patients with T1D were recruited from the out-patient clinic at Steno Diabetes Center, Gentofte, Denmark (n = 31). Inclusion criteria were dependence on insulin treatment from time of diagnosis, fasting serum C-peptide below 200 pmol l−1, glycated haemoglobin (HbA1c) above 8.0% (64 mmol l−1) to ensure current hyperglycaemia, T1D duration and dependence on insulin treatment > 5 years, no antibiotic treatment at least 2 months before inclusion, and no known gastrointestinal diseases. All study participants were of North European ethnicity.
The study participants were examined on 2 days that were approximately 14 days apart. On the first day, study participants were examined after an over-night fast. Height was measured without shoes to the nearest 0.5 cm, and weight was measured without shoes and wearing light clothes to the nearest 0.1 kg. Hip and waist circumference was measured using a non-expandable measuring tape to the nearest 0.5 cm. Waist circumference was measured midway between the lower rib margin and the iliac crest. Hip circumference was measured as the largest circumference between the waist and the thighs. Blood pressure was assessed while the participant was lying in an up-right position after at least 5 min of rest using a cuff of appropriate size (A&D, UA-787 plus digital or A&D, UA-779). Blood pressure was measured at least twice and the average of the measurements was calculated. On the second day of examination, all participants provided a stool sample which was immediately frozen after home collection and stored at −80 °C.
Information on medication status was obtained by questionnaire and interview on the first day of examination. Of the 75 T2D patients, 10 patients (13%) received no hyperglycaemic medications and 58 patients (77%) received the biguanide metformin; of these 75 TD2 patients, 28 patients (37%) received metformin as the only anti-hyperglycaemic medication, 10 patients (13%) received sulfonylurea alone or in combination with metformin, 14 patients (19%) received a combination of oral antidiabetic drugs and insulin treatment and 4 patients (5%) were on insulin treatment only. Eleven patients (15%) received dipeptidyl peptidase-4 (DPP4) inhibitors or glucagon-like peptide-1 (GLP1), all of them in combination with metformin. Patients were reported as receiving anti-hypertensive treatment if at least one of the following drugs was reported: spironolactone, thiazides, loop diuretics, beta blockers, calcium channel blockers, moxonidine or drugs affecting the renin–angiotensin system (n = 55 for T2D (73%) and n = 23 (74%) for T1D). Patients receiving statins, fibrates and/or ezetimibe were reported as receiving lipid-lowering medication (n = 56 for T2D (75%; all on statin treatment), and n = 24 for T1D (77%; 74% on statin treatment)). All T1D patients were on insulin treatment as their only blood glucose lowering treatment.
All biochemical analyses were performed on blood samples drawn in the morning after an over-night fast of at least 10 h. Plasma glucose was analysed by a glucose oxidase method (Granutest, Merck) with a detection limit of 0.11 mmol l−1 and intra- and interassay coefficients of variation (CV) of <0.8% and <1.4%, respectively. HbA1c was measured on G7 HPLC Analyzer (Tosoh) by ion-exchange high-performance liquid chromatography. Serum C-peptide was measured using a time-resolved fluoroimmunoassay with the AutoDELFIA C-peptide kit (PerkinElmer, Wallac), with a detection limit of 5 pmol l−1 and intra- and interassay CV of <4.7% and <6.4%, respectively. Serum insulin (excluding des and intact proinsulin) was measured using the AutoDELFIA insulin kit (PerkinElmer, Wallac) with a detection limit of 3 pmol l−1 and with intra- and interassay CV of <3.2% and <4.5%, respectively. Plasma cholesterol, plasma high-density lipoprotein cholesterol and plasma triglycerides were all measured on Vitros 5600 using reflect-spectrophotometrics. Plasma low-density lipoprotein cholesterol was calculated using Friedewald’s equation. Blood leukocytes and white blood cell differential count were measured on Sysmex XS 1000i using flow cytometrics. Plasma metformin was determined by high performance liquid chromatography followed by tandem mass spectrometry. Briefly, the proteins were precipitated with acetonitrile containing the deuterated internal standard, metformin-d6, hydrochloride and the supernatant diluted by acetonitrile. The analysis was performed on a Waters Acquity UPLC I-class system connected to a Xevo TQ-S tandem mass spectrometer in electrospray positive ionization mode. Separation was achieved on a Waters XBridgeT BEH Amide 2.5-μm column and gradient elution with 100 mM ammonium formate (pH 3.2), and with acetonitrile. The multiple reaction monitoring transitions used for metformin and metformin-d6 were 130.2 > 71.0 and 136.2 > 60.0. Calibrators were prepared by spiking drug-free serum with metformin to a concentration of 2,000 ng ml−1. B12 was measured using Vitros Immunodiagnostic Products. GAD65 was measured on serum samples by a sandwich ELISA (RSR ltd.). Inter- and intra-assay CV were < 16.6% and < 6.7% respectively, and with a detection limit of 0.57 Uml−1.
Stool samples were obtained at the homes of each participant and samples were immediately frozen by storing them in their home freezer. Frozen samples were delivered to Steno Diabetes Center using insulating polystyrene foam containers, and then they were stored at −80 °C until analysis. The time span from sampling to delivery at the Steno Diabetes Center was intended to be as short as possible and no more than 48 h.
A frozen aliquot (200 mg) of each faecal sample was suspended in 250 μl of guanidine thiocyanate, 0.1 M Tris, pH 7.5, and 40 μl of 10% N-lauroylsarcosine. Microbial DNA extraction was then performed as previously described12. The DNA concentration and its molecular size were estimated using nanodrop (Thermo Scientific) and agarose gel electrophoresis.
Generation and availability of metagenomic samples
Already available Danish metagenomic samples were those reported in ref. 26 and references therein (excluding 14 samples removed due to average read length below 40 nucleotides, and with 5 Chinese and 21 Swedish samples with less than the rarefaction threshold of 7 million reads in total excluded from functional profile or diversity analyses), with newly sequenced samples deposited in the European Bioinformatics Institute Sequence Read Archive under accession ERP004605.
All information on Swedish samples was retrieved from previously published data4. In addition to published data on Chinese individuals3, we retrieved information on metformin treatment in a subset of 71 Chinese T2D patients. One-hundred and twelve samples from ref. 3 lacked metformin treatment metadata and were therefore discarded, except for measuring differences between the country data sets disregarding treatment or diabetic status. Characteristics of all study participants included in the present protocol are given in Supplementary Table 1.
Validation cohort recruitment and sample processing
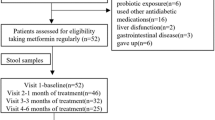
Additional Danish T2D patients were recruited at the Novo Nordisk Foundation Center for Basic Metabolic Research, University of Copenhagen throughout 2014 as a part of the ongoing MicrobDiab study (http://metabol.ku.dk/research-project-sites/microbdiab/). T2D patients were included in the study if the time of T2D diagnosis was less than 5 years ago, they were between 35 and 75 years of age, Caucasian and they had not received antibiotics within the past 4 months of inclusion. In total, 30 T2D patients (21 male and 9 female) were identified. Faecal samples were collected at the home of the patients, followed by immediate freezing of samples in home freezers, and transport of samples to the hospital stored on dry ice. The samples were stored at −80 °C until DNA extraction. Information of medication was obtained from questionnaires. In total, 21 (70%) of the T2D patients received metformin.
Ethics statement
All individuals in both the Danish MetaHIT study and the Danish validation study gave written informed consent before participation in the studies. Both studies were approved by the Ethical Committees of the Capital Region of Denmark (MetaHIT study: HC-2008-017; validation study: H-3-2013-102). Both studies were conducted in accordance with the principles of the Declaration of Helsinki.
Construction of a non-redundant metagenomic reference gene catalogue.
Illumina shotgun sequencing was applied to DNA extracted from 620 faecal samples originating from the MetaHIT project (Supplementary Table 1). Raw sequencing data were processed using the MOCAT (version 1.1) software package27. Reads were trimmed (option read_trim_filter) using a quality and length cut-off of 20 and 30 bp, respectively. Trimmed reads were subsequently screened against a custom database of Illumina adapters (option screen_fastafile) and the human genome version 19 using a 90% identity cut-off (option screen). The resulting high-quality reads were assembled (option assembly) and assemblies revised (option assembly revision). Genes were predicted on scaftigs with a minimum length of 500 bp (option gene_prediction).
Predicted protein-coding genes with a minimum length of 100 bp were clustered at 95% sequence identity using Cd-hit (version 4.6.1)28 with parameters set to: -c 0.95, -G 0 -aS 0.9, -g 1, -r 1. The representative genes of the resulting clusters were ‘padded’ (that is, extended up to 100 bp at each end of the sequence using the sequence information available from the assembled scaftigs), resulting in the final reference gene catalogue used in this study.
The reference gene catalogue was functionally annotated using SmashCommunity29 (version 1.6) after aligning the amino acid sequence of each gene to the KEGG30 (version 62) and eggNOG31 (version 3) databases.
Profiling of metagenomic samples
Raw insert (sequenced fragments of DNA represented by single or paired-end reads) count profiles were generated using MOCAT27 by mapping high-quality reads from each metagenome to the reference gene catalogue (option screen) using an alignment length and identity cut-off of 45% and 95%, respectively. For each gene, the number of inserts that matched the protein-coding region was counted. Counts of inserts that mapped with the same alignment score to multiple genes were distributed equally among them. Taxonomic abundances were computed at the level of metagenomic operational taxonomic units (mOTUs)32, normalized to the length of the concatenated marker genes for each mOTU to yield the abundances used for the study, and subsequently binned at broader taxonomic levels (genus, family, class, etc.).
Rarefaction of metagenomic data and microbial diversity measurements
For all metagenome-derived measures except the mOTU taxonomic assignments, read counts were ‘rarefied’ in order to avoid any artefacts of sample size on low-abundance genes. Rarefied matrices were obtained as follows. Data matrices were rarefied to 7 million reads per sample. This threshold was chosen to include most samples, but 5 Chinese and 21 Swedish samples were excluded due to having less than 7 million reads per sample. Rarefactions were performed using a C++ program developed for the Tara project33. In total we performed 30 repetitions, and in each of these we measured the richness, evenness, chao1 and Shannon diversity metrics within a rarefaction. The median value of these was taken as the respective diversity measurement for each sample. The first of 30 rarefactions of each sample were used to create a rarefied gene abundance matrix and KEGG orthologue abundance profiles were calculated by summing the rarefied abundance of genes annotated to the respective KEGG orthologue gene.
Metagenomic species (MGS) construction
Clustering of the catalogue genes by co-abundance, as described in ref. 34, defined 10,754 co-abundance gene groups (CAGs) with very high correlations (Pearson correlation coefficient > 0.9). The 925 largest of these, with more than 700 genes, were termed metagenomic species (MGS). The abundance profiles of the CAGs and MGSs were determined as the medium gene abundance (downsized to 7 million reads per sample) throughout the samples. Furthermore, the CAGs and MGS were taxonomically annotated by sequence similarity to known reference genomes.
Functional annotation/binning of metagenomes
To avoid drawing false conclusions about gut microbial functions from high abundance of single genes remotely homologous to members of a functional pathway, we used an approach that required presence of multiple pathway members. Functional pathway abundance was calculated from gene catalogue KEGG orthologue annotation and MGS abundances per sample. Thus KEGG orthologues present in each MGS were used to determine for that CAG/MGS which functional modules were represented within its genetic repertoire. This required that >90% of KEGG orthologues necessary for the completion of a reaction pathway should be present, when also taking alternative enzymatic pathways into account. The module abundance within a sample was calculated from CAG abundance in each respective sample, summing over all CAGs which had the module present. Rarefied median coverages of CAG/MGS were used, so no further normalization of the module abundance matrix was required. Abundance of genetic potential falling under the same higher-order functional levels was calculated by summing up all abundances of the lower-level functional modules within each sample.
Existing functional annotation databases cover gut metabolic pathways relatively poorly. To account for this, a number of additional bacterial gene functional modules were curated and annotated, extending the KEGG system; these are referred to in result tables as GMMs (gut microbial modules) and were previously described in ref. 12.
16S amplicon processing
16S amplicons from frozen samples were sequenced 300 bp and 200 bp paired-end reads using an Illumina miSeq machine. We used the LotuS35 pipeline in short amplicon mode with default quality filtering, clustering and denoising operational taxonomic units (OTUs) with UPARSE36, removing chimaeric OTUs against the RDP reference database (http://drive5.com/uchime/rdp_gold.fa) with uchime37, merging reads with FLASH38 and assigning a taxonomy against the SILVA 119 rRNA database39, and further refined by BLAST searches against the NCBI rRNA database40 to identify Intestinibacter OTUs, using the following LotuS command line options: ‘-p miSeq -refDB SLV -doBlast blast -amplicon_type SSU -tax_group bacteria -derepMin 2 -CL 2 -thr 14’.
Univariate tests of taxonomic or functional abundance differences
Microbial taxa where mean abundance over all samples was less than 30 reads, or that were present in less than 3 samples, were excluded from univariate and classifier analyses. All abundances were normalized by total sample sum. For module tables, no feature filters were used except requiring the module to be present in at least 20 samples. Filtered data tables were made available online (http://vm-lux.embl.de/~forslund/t2d/).
Univariate testing for differential abundances of each taxonomic unit between two or more groups was tested using Mann–Whitney-U or Kruskal–Wallis tests, respectively, corrected for multiple testing using the Benjamini–Hochberg false discovery rate control procedure (Q values)41. Post-hoc statistical testing for significant differences between all combinations of two groups was conducted only for taxa with abundances significantly different at P < 0.2. Wilcoxon rank-sum tests were calculated for all possible group combinations and corrected for multiple testing again using the Benjamini–Hochberg false discovery rate, as implemented in R. When controlling for potential confounders such as source study, we used blocked ‘independence_test’ function calls with options ‘ytrafo = rank, teststat=scalar’ for blocked WRST and ‘ytrafo = rank, teststat=quad’ for blocked Kruskal–Wallis test, as implemented in the COIN software package42 for R. Similarly, we applied these independence tests in the framework of post-hoc testing as described above.
Analysis of correlations between taxonomic or functional features, community diversity indices and sample metadata variables were conducted using Spearman correlation tests as implemented in R, and corrected for multiple tests using the Benjamini–Hochberg false discovery rate control procedure. To control for confounders such as source study in univariate correlation analyses, blocked Spearman tests as implemented in COIN (settings ‘independence_test’, options ytrafo = rank, xtrafo = rank, distribution = asymptotic) were used.
In some analyses, taxa were corrected for the influence of a continuous confounder variable such as microbial community richness; in these cases, the residual of a linear model between normalized log-transformed taxa abundances and overall sample gene richness was used to correct for the confounding variable. Power analysis was conducted by randomly subsampling to a given sample number, repeated 5 times to achieve robust results.
Ordinations and multivariate tests
All ordinations (NMDS, dbRDA) and subsequent statistical analyses were calculated using the R package vegan43 using Canberra distances on normalized taxa abundance matrices, then visualized using the ggplot2 R package44. Community differences were calculated using a permutation test on the respective NMDS reduced feature space, as implemented in vegan.
Furthermore, we calculated intergroup differences for the microbiota using PERMANOVA45 as implemented in vegan. This test compares the intragroup distances to the intergroup distances in a permutation scheme and from this calculates a P value. For all PERMANOVA tests, we used 2 × 105 randomizations and a normalized genus-level mOTU abundance matrix, using Canberra intersample distances. PERMANOVA post-hoc P values were corrected for multiple testing using the Benjamini–Hochberg false discovery rate control procedure. Analysis of variance broken down by cohort, treatment and disease status was conducted by fitting these distances to a linear model of sample metadata distances, as further described in Supplementary Discussion 3.2.
Classifier construction and evaluation
To create classifiers for separating samples from different subsets, an L1 restricted LASSO using the R glmnet package46 was carried out to test for an optimal value of lambda (number of features to be used in the final predictor) in a fivefold cross-validated and internally fourfold cross-validated LASSO run on all data. After this, the previously determined value of lambda was manually controlled for number of features used against the root mean square error of the classifier. In a fivefold cross-validation, an independent LASSO classifier was trained on 4/5 of the data using the previously determined value of lambda, and response values were predicted on 1/5 of the data. LASSO models with a Poisson response type were used in all cases.
Binary classifications between T2D and ND control samples were performed with an R reimplementation of the robust recursive feature elimination support vector machine (rRFE-SVM)47 procedure. The SVM was performed in an outer cross-validation scheme on 4/5 of the data. Of these, 90% were randomly selected 200 times in each cross-validation for the RFE, to create a feature ranking from an average over these runs. Classifier performance was validated on the remaining 1/5 of samples using the pre-established feature ranking. In case of several cohorts, the area under the receiver operating characteristic curve (ROC-AUC) scores were measured for each cohort separately.
Code availability
The MGS technology has previously been described34 and is available online (http://git.dworzynski.eu/mgs-canopy-algorithm/wiki/Home). The mOTU resource has been made publically available (http://www.bork.embl.de/software/mOTU/) and was analysed using MOCAT27 which is also publically available (http://vm-lux.embl.de/~kultima/MOCAT/). The 16S pipeline LotuS35 is freely available online (http://psbweb05.psb.ugent.be/lotus). The novel gene catalogue has been deposited online (http://vm-lux.embl.de/~kultima/share/gene_catalogs/620mhT2D/), as have the raw amplicon sequences (http://vm-lux.embl.de/~forslund/t2d/). Statistical analysis and data visualization was conducted using freely available R libraries: vegan, COIN and ggplot2 and is described in more details elsewhere48,49. Data matrices and R source code for replicating the central tests conducted on the data have been deposited online (http://vm-lux.embl.de/~forslund/t2d/).
Evaluation of dietary habits
A subset of the Danish study participants answered a validated food frequency questionnaire in order to obtain information on the habitual dietary habits. A complete data set was obtained for 66% of the nondiabetic individuals and 88% of T2D patients. When evaluating the dietary data, the consumed quantity was determined by multiplying portion size by the corresponding consumption frequency reported. Standard portion sizes for women and men, separately, were used in this calculation50,51. All food items in the questionnaire were linked to food items in the Danish Food Composition Databank52. Estimation of daily intake of macro- and micronutrients for each participant was based on calculations in the software program FoodCalc version 1.353.
Accession codes
Primary accessions
European Nucleotide Archive
Sequence Read Archive
Data deposits
Raw nucleotide data can be found for all samples used in the study in the Sequence Read Archive (accession numbers: SRA045646 and SRA050230, CHN samples) and the European Nucleotide Archive (accession numbers: ERP002469, SWE samples; ERA000116, ERP003612, ERP002061 and ERP004605, MHD samples).
References
Shreiner, A. B., Kao, J. Y. & Young, V. B. The gut microbiome in health and in disease. Curr. Opin. Gastroenterol. 31, 69–75 (2015)
Cho, I. & Blaser, M. J. The human microbiome: at the interface of health and disease. Nature Rev. Genet. 13, 260–270 (2012)
Qin, J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490, 55–60 (2012)
Karlsson, F. H. et al. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature 498, 99–103 (2013)
Schellenberg, E. S., Dryden, D. M., Vandermeer, B., Ha, C. & Korownyk, C. Lifestyle interventions for patients with and at risk for type 2 diabetes: a systematic review and meta-analysis. Ann. Intern. Med. 159, 543–551 (2013)
Larsen, N. et al. Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PLoS ONE 5, e9085 (2010)
Zhang, X. et al. Human gut microbiota changes reveal the progression of glucose intolerance. PLoS ONE 8, e71108 (2013)
de Vos, W. M. & Nieuwdorp, M. Genomics: A gut prediction. Nature 498, 48–49 (2013)
Pernicova, I. & Korbonits, M. Metformin–mode of action and clinical implications for diabetes and cancer. Nat. Rev. Endocrinol. 10, 143–156 (2014)
Shin, N. R. et al. An increase in the Akkermansia spp. population induced by metformin treatment improves glucose homeostasis in diet-induced obese mice. Gut 63, 727–735 (2014)
Napolitano, A. et al. Novel gut-based pharmacology of metformin in patients with type 2 diabetes mellitus. PLoS ONE 9, e100778 (2014)
Le Chatelier, E. et al. Richness of human gut microbiome correlates with metabolic markers. Nature 500, 541–546 (2013)
Sato, J. et al. Gut dysbiosis and detection of “live gut bacteria” in blood of Japanese patients with type 2 diabetes. Diabetes Care 37, 2343–2350 (2014)
Cabreiro, F. et al. Metformin retards aging in C. elegans by altering microbial folate and methionine metabolism. Cell 153, 228–239 (2013)
Gerritsen, J. et al. Characterization of Romboutsia ilealis gen. nov., sp. nov., isolated from the gastro-intestinal tract of a rat, and proposal for the reclassification of five closely related members of the genus Clostridium into the genera Romboutsia gen. nov., Intestinibacter gen. nov., Terrisporobacter gen. nov. and Asaccharospora gen. nov. Int. J. Syst. Evol. Microbiol. 64, 1600–1616 (2014)
Song, Y. L., Liu, C. X., McTeague, M., Summanen, P. & Finegold, S. M. Clostridium bartlettii sp. nov., isolated from human faeces. Anaerobe 10, 179–184 (2004)
Messori, S., Trevisi, P., Simongiovanni, A., Priori, D. & Bosi, P. Effect of susceptibility to enterotoxigenic Escherichia coli F4 and of dietary tryptophan on gut microbiota diversity observed in healthy young pigs. Vet. Microbiol. 162, 173–179 (2013)
Czyzyk, A., Tawecki, J., Sadowski, J., Ponikowska, I. & Szczepanik, Z. Effect of biguanides on intestinal absorption of glucose. Diabetes 17, 492–498 (1968)
Winter, S. E. et al. Host-derived nitrate boosts growth of E. coli in the inflamed gut. Science 339, 708–711 (2013)
Everard, A. et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Natl Acad. Sci. USA 110, 9066–9071 (2013)
Lee, H. & Ko, G. Effect of metformin on metabolic improvement and gut microbiota. Appl. Environ. Microbiol. 80, 5935–5943 (2014)
De Vadder, F. et al. Microbiota-generated metabolites promote metabolic benefits via gut-brain neural circuits. Cell 156, 84–96 (2014)
Croset, M. et al. Rat small intestine is an insulin-sensitive gluconeogenic organ. Diabetes 50, 740–746 (2001)
Jørgensen, T. et al. A randomized non-pharmacological intervention study for prevention of ischaemic heart disease: baseline results Inter99. Eur. J. Cardiovasc. Prev. Rehabil. 10, 377–386 (2003)
WHO. Definition, Diagnosis and Classification of Diabetes Mellitus and its Complications. Part 1: Diagnosis and Classification of Diabetes Mellitus. Report No. WHO/NCD/NCS/99.2 (World Health Organization, 1999)
Li, J. et al. An integrated catalog of reference genes in the human gut microbiome. Nature Biotechnol. 32, 834–841 (2014)
Kultima, J. R. et al. MOCAT: a metagenomics assembly and gene prediction toolkit. PLoS ONE 7, e47656 (2012)
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006)
Arumugam, M., Harrington, E. D., Foerstner, K. U., Raes, J. & Bork, P. SmashCommunity: a metagenomic annotation and analysis tool. Bioinformatics 26, 2977–2978 (2010)
Kanehisa, M. et al. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 36, D480–D484 (2008)
Powell, S. et al. eggNOG v3.0: orthologous groups covering 1133 organisms at 41 different taxonomic ranges. Nucleic Acids Res. 40, D284–D289 (2012)
Sunagawa, S. et al. Metagenomic species profiling using universal phylogenetic marker genes. Nature Methods 10, 1196–1199 (2013)
Sunagawa, S. et al. Structure and function of the global ocean microbiome. Science 348, (2015)
Nielsen, H. B. et al. Identification and assembly of genomes and genetic elements in complex metagenomic samples without using reference genomes. Nature Biotechnol. 32, 822–828 (2014)
Hildebrand, F. et al. LotuS: an efficient and user-friendly OTU processing pipeline. Microbiome 2, 30 (2014)
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nature Methods 10, 996–998 (2013)
Edgar, R. C. et al. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011)
Magoč, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011)
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013)
Madden, T. in The NCBI Handbook [Internet]. (eds, McEntyre J. & Ostell J. ) Ch. 16 (National Center for Biotechnology Information, 2002) http://www.ncbi.nlm.nih.gov/books/NBK21097/
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. A Stat. Soc. 57, 289–300 (1995)
Hothorn, T., Hornik, K., van de Wiel, M. A. & Zeileis, A. A Lego system for conditional inference. Am. Stat. 60, 257–263 (2006)
Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 14, 927–930 (2003)
Wickham H. ggplot2: Elegant Graphics for Data Analysis. (Springer, 2009)
Anderson, M. J. A new method for non-parametric multivariate analysis of variance. Austral. Ecol. 26, 32–46 (2001)
Friedman, J. et al. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010)
Abeel, T., Helleputte, T., Van de Peer, Y., Dupont, P. & Saeys, Y. Robust biomarker identification for cancer diagnosis with ensemble feature selection methods. Bioinformatics 26, 392–398 (2010)
Hildebrand, F. et al. A comparative analysis of the intestinal metagenomes present in guinea pigs (Cavia porcellus) and humans (Homo sapiens). BMC Genomics 13, 514 (2012)
Hildebrand, F. et al. Inflammation-associated enterotypes, host genotype, cage and inter-individual effects drive gut microbiota variation in common laboratory mice. Genome Biol. 14, R4 (2013)
Haraldsdóttir, J. et al. Portionsstorleker - Nordiska standardportioner av mat och livsmedel (Nordisk Ministerråd, 1998)
Biltoft-Jensen, A. et al. Danskernes kostvaner 2000–2002. DFVF publication No. 11 (Danmarks Fødevareforskning, Afdeling for Ernæring, 2005)
Møller, A. et al. Fødevaredatabanken version 5.0. Fødevareinformatik, Institut for Fødevaresikkerhed og Ernæring, Fødevaredirektoratethttp://www.foodcomp.dk (2002)
Lauritsen, J. FoodCalc. www.ibt.ku.dk/jesper/FoodCalc/ (2004)
Acknowledgements
The authors wish to thank A. Forman, T. Lorentzen, B. Andreasen, G. J. Klavsen and M. J. Nielsen for technical assistance, and T. F. Toldsted and G. Lademann for management assistance. J. Nielsen and F. Bäckhed are thanked for providing access to T2D metagenome data and metformin treatment status before publication4. V. Benes and the GeneCore facility of EMBL Heidelberg are thanked for their assistance with the metformin signature validation experiments, as is Y. Yuan for assistance with computer infrastructure. This research has received funding from European Community’s Seventh Framework Program (FP7/2007-2013): MetaHIT, grant agreement HEALTH-F4-2007-201052, MetaCardis, grant agreement HEALTH-2012-305312, International Human Microbiome Standards, grant agreement HEALTH-2010-261376, as well as from the Metagenopolis grant ANR-11-DPBS-0001, from the European Research Council CancerBiome project, contract number 268985, and from the European Union HORIZON 2020 programme, under Marie Skłodowska-Curie grant agreement 600375. Additional funding came from The Lundbeck Foundation Centre for Applied Medical Genomics in Personalized Disease Prediction, Prevention and Care (LuCamp, http://www.lucamp.org), the Novo Nordisk Foundation (grant NNF14CC0001), and the European Molecular Biology Laboratory (EMBL). The Novo Nordisk Foundation Center for Basic Metabolic Research is an independent Research Center at the University of Copenhagen partially funded by an unrestricted donation from the Novo Nordisk Foundation (http://www.metabol.ku.dk). Additional funding for the validation experiments was provided by the Innovation Fund Denmark through the MicrobDiab project.
Author information
Authors and Affiliations
Consortia
Contributions
O.P., S.D.E. and P.B. devised the project, designed the study protocol and supervised all phases of the project. T.N., T.H., T.J., H.V., J.L. and O.P. carried out patient phenotyping and clinical data analyses. T.N. and F.L. performed sample collection and DNA extraction. J.D. supervised DNA extraction, J.W., K.K. supervised DNA sequencing and gene profiling, A.Y.V. and R.H. performed additional microbial DNA extraction and amplicon sequencing. J.R., H.B.N., S.B., S.D.E., P.B. and O.P. designed and supervised the data analyses. K.F., F.H., G.F., E.L.C., S.S., E.P., S.S.-V., V.G., H.K.P, M.A., P.I.C., J.R.K. and H.B.N performed the data analyses. K.F., F.H., T.N., P.B, S.D.E. and O.P. wrote the paper. All authors contributed to data interpretation, discussions and editing of the paper. All authors are members of the MetaHIT consortium. Additional consortium members contributed to the design and execution of the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A list of participants and their affiliations appears in the Supplementary Information.
Extended data figures and tables
Extended Data Figure 1 Validation of meta-analysis pipeline on simulated data.
a, As a positive control for the meta-analysis pipeline, true signal was removed from the data by randomly reshuffling sample labels. Artificial contrast was thereafter introduced between random groups containing as many such reshuffled samples as were in the original sets of T2D metformin+ (nCHN = 15, nMHD = 58, nSWE = 20) and T2D metformin− (nCHN = 56, nMHD = 17, nSWE = 33) samples in each original study subset, using the genus Akkermansia as an example feature. Samples randomly assigned to the sets of fake ‘metformin-treated’ and ‘control’ categories had their Akkermansia genus abundances adjusted to match the scale of the metformin effect on Escherichia genus abundance reported here (metformin-treated samples were roughly 150% as likely to have non-zero abundance, with a roughly threefold higher abundance where present), while retaining their data set origin labels. The full meta-analysis pipeline (study set blocked Kruskal–Wallis test, post-hoc Wilcoxon rank-sum test) was applied to these samples. Benjamini–Hochberg-corrected P values (FDR scores/Q values) from testing for a metformin effect on Akkermansia abundance are plotted in logarithmic scale on the vertical axis for 100 randomizations of the entire shuffled data set, either without (left box plot) or with (right box plot) the artificial Akkermansia metformin signal added after shuffling the data to remove original signal. Box plot borders show medians and quartiles, with points outside this range shown as vertical whisker lines and point markers. Whiskers extend to 1.58× interquartile range/ . Horizontal guide lines are shown for ease of visualization corresponding to different false discovery rate thresholds. For randomly reshuffled data, no significant contrast is detected as expected, whereas the artificially introduced signal is reliably detected, roughly matching expectations from the definition of the false discovery rate itself. b, To investigate statistical power for the other medications tracked, five random sub-samplings were made of pairs of medicated and non-medicated samples at each increasing number of included sample pairs and the overall analysis was replicated for each. We tested each genus for significantly differential abundance between cases and controls (Kruskal–Wallis test followed by post-hoc Wilcoxon rank-sum test) at different Benjamini–Hochberg FDR significance cut-offs, which are represented by different colours. Of the total number of samples for which medication status was known, equal numbers (n) of medicated and unmedicated samples were chosen randomly in repeated iterations. This number n was varied up to its largest possible value (smallest of either number of medicated or unmedicated samples in the overall data set) and is shown on the x axis. The y axis shows the number of significant features relative to each cut-off. Error bars show ±1 s.d. of each set of five randomized samples. c, The graphs show Intestinibacter and Escherichia median and quartile abundances as box plots, whiskers extend to 1.58× interquartile range/
. Horizontal guide lines are shown for ease of visualization corresponding to different false discovery rate thresholds. For randomly reshuffled data, no significant contrast is detected as expected, whereas the artificially introduced signal is reliably detected, roughly matching expectations from the definition of the false discovery rate itself. b, To investigate statistical power for the other medications tracked, five random sub-samplings were made of pairs of medicated and non-medicated samples at each increasing number of included sample pairs and the overall analysis was replicated for each. We tested each genus for significantly differential abundance between cases and controls (Kruskal–Wallis test followed by post-hoc Wilcoxon rank-sum test) at different Benjamini–Hochberg FDR significance cut-offs, which are represented by different colours. Of the total number of samples for which medication status was known, equal numbers (n) of medicated and unmedicated samples were chosen randomly in repeated iterations. This number n was varied up to its largest possible value (smallest of either number of medicated or unmedicated samples in the overall data set) and is shown on the x axis. The y axis shows the number of significant features relative to each cut-off. Error bars show ±1 s.d. of each set of five randomized samples. c, The graphs show Intestinibacter and Escherichia median and quartile abundances as box plots, whiskers extend to 1.58× interquartile range/ , with samples that are extreme relative to the interquartile range shown as point markers, and with samples below detection threshold (DT) plotted at y = 0, in 21 additional T2D metformin+ and 9 additional T2D metformin− samples. Differences in abundance between sample categories are significant (Wilcoxon rank-sum test, Benjamini–Hochberg FDR < 0.1). All samples in which Intestinibacter was detected fall among the 9 out of 30 untreated rather than the 21 out of 30 metformin-treated samples, consistent with severe depletion under treatment; whereas Escherichia abundances increase under treatment, likewise consistent with observations from the main data set.
, with samples that are extreme relative to the interquartile range shown as point markers, and with samples below detection threshold (DT) plotted at y = 0, in 21 additional T2D metformin+ and 9 additional T2D metformin− samples. Differences in abundance between sample categories are significant (Wilcoxon rank-sum test, Benjamini–Hochberg FDR < 0.1). All samples in which Intestinibacter was detected fall among the 9 out of 30 untreated rather than the 21 out of 30 metformin-treated samples, consistent with severe depletion under treatment; whereas Escherichia abundances increase under treatment, likewise consistent with observations from the main data set.
Extended Data Figure 2 Differences in physiological variables and microbiome characteristics between gut metagenome sample sets.
Chinese (n = 368), Danish MetaHIT (n = 383) and Swedish (n = 145). a, Several participant metadata variables are significantly different between cohorts. A subselection is shown as box plots displaying median and quartiles, with samples outside this range shown as point markers and whiskers. Whiskers extend to 1.58× interquartile range/ . b, In a principal coordinates analysis ordination of Bray–Curtis distances between samples on bacterial family level, clear differences between samples from the different cohorts become apparent. These are largely explained by taxonomic differences as summarized at the phylum level. c, Box plots for gut microbial taxa show medians and quartiles of log-transformed read counts for mOTUs summarized at the level of bacterial genera for the three country subsets across sample categories, with samples outside this range shown as point markers and whiskers. Whiskers extend to 1.58× interquartile range/
. b, In a principal coordinates analysis ordination of Bray–Curtis distances between samples on bacterial family level, clear differences between samples from the different cohorts become apparent. These are largely explained by taxonomic differences as summarized at the phylum level. c, Box plots for gut microbial taxa show medians and quartiles of log-transformed read counts for mOTUs summarized at the level of bacterial genera for the three country subsets across sample categories, with samples outside this range shown as point markers and whiskers. Whiskers extend to 1.58× interquartile range/ . For all box plots, tests for significant differences (Kruskal–Wallis test adjusted for study source) were performed, with P values shown at the head of each figure. Asterisks denote statistical significance of tests done for each country subset separately (***P < 0.001).
. For all box plots, tests for significant differences (Kruskal–Wallis test adjusted for study source) were performed, with P values shown at the head of each figure. Asterisks denote statistical significance of tests done for each country subset separately (***P < 0.001).
Extended Data Figure 3 Microbiome taxonomic composition comparison between gut metagenomes with particular focus on possible taxonomic restoration under metformin treatment for certain taxa.
T2D metformin− (n = 106), T2D metformin+ (n = 93) and ND control (n = 554). Box plots show medians and quartiles log-transformed read counts for mOTUs summarized at the level of bacterial genera, for the three country subsets across sample categories, with samples outside this range shown as point markers and whiskers. Whiskers extend to 1.58× interquartile range/ . Tests for significant differences (Kruskal–Wallis test adjusted for study source) were performed, with P values shown at the head of each figure. Asterisks denote statistical significance of tests for each country subset separately (*P < 0.05; **P < 0.01; ***P < 0.001).
. Tests for significant differences (Kruskal–Wallis test adjusted for study source) were performed, with P values shown at the head of each figure. Asterisks denote statistical significance of tests for each country subset separately (*P < 0.05; **P < 0.01; ***P < 0.001).
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion, full legends for Supplementary Tables 1-16, Supplementary References and a list of additional MetaHIT consortium members. (PDF 669 kb)
Supplementary Tables
This file contains Supplementary Tables 1-16 – see Supplementary Information document for legends. (ZIP 465 kb)
Rights and permissions
About this article
Cite this article
Forslund, K., Hildebrand, F., Nielsen, T. et al. Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature 528, 262–266 (2015). https://doi.org/10.1038/nature15766
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15766
- Springer Nature Limited
This article is cited by
-
Intestinal microbiota and metabolome perturbations in ischemic and idiopathic dilated cardiomyopathy
Journal of Translational Medicine (2024)
-
Integration of polygenic and gut metagenomic risk prediction for common diseases
Nature Aging (2024)
-
Gut Microbiome Composition in Polycystic Ovary Syndrome Adult Women: A Systematic Review and Meta-analysis of Observational Studies
Reproductive Sciences (2024)
-
A Dual Therapeutic Approach to Diabetes Mellitus via Bioactive Phytochemicals Found in a Poly Herbal Extract by Restoration of Favorable Gut Flora and Related Short-Chain Fatty Acids
Applied Biochemistry and Biotechnology (2024)
-
Gut microbiota in relationship to diabetes mellitus and its late complications with a focus on diabetic foot syndrome: A review
Folia Microbiologica (2024)