Abstract
Background
The influence of the microbiota on hypoglycemic agents is becoming more apparent. The effects of metformin, a primary anti-diabetes drug, on gut microbiota are still not fully understood.
Research design and methods
This prospective cohort study aims to investigate the longitudinal effects of metformin on the gut microbiota of 25 treatment-naïve diabetes patients, each receiving a daily dose of 1500 mg. Microbiota compositions were analyzed at baseline, and at 1, 3, and 6 months of medication using 16S rRNA gene sequencing.
Results
Prior to the 3-month period of metformin treatment, significant improvements were noted in body mass index (BMI) and glycemic-related parameters, such as fasting blood glucose (FPG) and hemoglobin A1c (HbA1c), alongside homeostasis model assessment indices of insulin resistance (HOMA-IR). At the 3-month mark of medication, a significant reduction in the α-diversity of the gut microbiota was noted, while β-diversity exhibited no marked variances throughout the treatment duration. The Firmicutes to Bacteroidetes ratio. markedly decreased. Metformin treatment consistently increased Escherichia-Shigella and decreased Romboutsia, while Pseudomonas decreased at 3 months. Fuzzy c-means clustering identified three longitudinal trajectory clusters for microbial fluctuations: (i) genera temporarily changing, (ii) genera continuing to decrease (Bacteroides), and (iii) genera continuing to increase(Lachnospiraceae ND3007 group, [Eubacterium] xylanophilum group, Romboutsia, Faecalibacterium and Ruminococcaceae UCG-014). The correlation matrix revealed associations between specific fecal taxa and metformin-related clinical parameters HbA1c, FPG, Uric Acid (UA), high-density lipoproteincholesterol (HDL-C), alanine aminotransferase (ALT), hypersensitive C-reactive protein (hs-CRP), triglyceride (TG) (P < 0.05). Metacyc database showed that metformin significantly altered 17 functional pathways. Amino acid metabolism pathways such as isoleucine biosynthesis predominated in the post-treatment group.
Conclusions
Metformin’s role in glucose metabolism regulation may primarily involve specific alterations in certain gut microbial species rather than an overall increase in microbial species diversity. This may suggest gut microbiota targets in future studies on metabolic abnormalities caused by metformin.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Type 2 diabetes mellitus (T2DM) is a prevalent chronic disease associated with life-threatening complications, disabilities, and substantial financial burdens, concurrently reducing life expectancy [1]. This disorder, marked by insulin resistance and β-cell dysfunction, is linked to inflammation induced by overnutrition and environmental factors [2]. According to the International Diabetes Federation (IDF), the prevalence of T2DM is projected to increase by 46% by 2045, affecting an estimated 783 million adults [3]. The gut microbiome is recognized as a modifiable “novel organ”, intricately shaping T2DM’s metabolic and immunological functions [4]. The complex interaction of the intestinal microbiota is theorized to play a pivotal role in the pathophysiology of obesity, T2DM, and associated metabolic disorders. Changes in the gut microbiome’s composition and functionality, termed intestinal dysbiosis, may diminish short-chain fatty acid (SCFA) synthesis [5], potentially compromising gut barrier integrity, impacting pancreatic β-cell proliferation, and attenuating insulin biosynthesis, thereby accelerating T2DM’s progression toward insulin resistance.
The interaction between gut microbes and commonly used drugs in T2DM patients is intricate and bidirectional [6]. Metformin, the first-line drug for T2DM, is chosen for its cost-effectiveness, safety, and protective impact on cardiovascular disease mortality [7]. Metformin enhances intestinal glucose uptake, increases lactate production, elevates glucagon-like peptide 1 (GLP-1) concentrations, influences the bile acid pool, and induces microbiota alterations [8]. Metformin, with approximately 50% bioavailability, results in nearly equal exposure in both the intestine and plasma; however, metformin accumulates in the intestine at a rate 300 times greater than in the plasma [9]. In contrast to oral dosing, intravenously administered metformin does not improve glucose metabolism [10], suggesting that the gastrointestinal tract may be this drug’s principal site of action. A delayed-release oral metformin targeting the ileum shows comparable or superior glucose-lowering effects with reduced systemic exposure compared to immediate or extended-release formulations [11]. Moreover, concurrent oral administration of a broad-spectrum antibiotic with metformin in mice nullifies its glucose-lowering effect, emphasizing the probable origin of metformin’s efficacy in the lower bowel [12].
Although a substantial body of literature exists on the gut microbiota and metformin, existing research spans treatment durations of 24 h to 12 months, with the majority centered on four months. Nevertheless, a lack of consensus exists regarding the results of these investigations [13]. Mueller et al. [14] conducted a randomized trial to examine the effects of metformin treatment and behavioral weight loss on gut microbiota composition over 12 months in overweight and obese adults. Despite the extensive observation period, the study did not include an analysis of the longitudinal dynamics of the gut microbiota over time. Furthermore, most study designs include cross-sectional investigations or case-control trials [4, 15,16,17,18,19,20,21], with noted disparities in basic clinical data between experimental and control groups. Due to significant individual differences in gut microbiota composition, attributing observed variations to individual specificity is challenging. Addressing these limitations necessitates establishing a longitudinal cohort study. This study aims to investigate gut microbiota changes during different treatment stages (1 month, 3 months, and 6 months) of metformin therapy and their correlation with glycemic and lipid metabolism indicators. Through longitudinal data analysis, we intend to provide novel insights into the longer-term effects of metformin on gut microbiota, potentially paving the way for personalized T2DM management strategies.
Materials and methods
Study subjects and sample collection
This prospective, longitudinal study was performed at Peking Union Medical College Hospital (PUMCH, Beijing, China). We prospectively recruited participants from the outpatient department of the endocrinology center of PUMCH between April 2021 and October 2023. Twenty-five patients were included. The detailed flow chart of the study design is shown in Fig. 1.
Inclusion criteria were: (1) Clinical diagnosis of T2DM based on the 2020 American Diabetes Association (ADA) criteria, with no antidiabetic medication in the past three months. (2) Age between 18 and 70 years, with no gender restrictions. (3) Not previously subjected to any antidiabeic treatment. Exclusion criteria included: (1) Other types of diabetes. (2) Acute diabetic complications in the last 6 months, such as diabetic ketoacidosis, lactic acidosis, and hyperosmolar hyperglycemic state. (3) Severe chronic diabetes complications, e.g., foot lesions, chronic ulcers, end-stage renal disease. (4) History of infectious diseases or recent occurrences within the last 6 months. (5) Significant abnormalities in liver function or impaired kidney function. (6) History of gastrointestinal diseases or surgery. (7) History of dental or oral diseases. (8) Mental disorders, tumors, alcohol dependence, substance abuse history, lactating, or participation in other clinical studies in the preceding three months. (9) Cardiovascular diseases, including coronary heart disease, cerebrovascular diseases, intermittent claudication, stroke, primary myocarditis, and other diagnosed cardiovascular diseases. (10) History of autoimmune or genetic diseases. (11) Long-term antacids, anti-inflammatory drugs, or proton pump inhibitors use. (12) Recent use of antibiotics or supplements with prebiotics, probiotics, and live bacteria in the last three months. (13) Allergic disposition or known allergy to multiple drugs, as deemed unsuitable by the investigator.
Fecal sampling
Participants were provided with a fecal sampling kit and instructions by outpatient medical staff. The kit included sterile sampling paper, a collection tube, sterile gloves, sample labels, and a collection bag. Following the provided instructions, participants collected approximately 5 g of fecal material from various points on the surface, middle, and depth of their fecal specimens at home. Samples were coded as follows: MET0 – pre-metformin(MET) treatment, MET1 – 1 month after the start of treatment, MET2 – 3 months into therapy, and MET3 – 6 months of metformin treatment. The collected sample was placed in the collection tube, securely capped, inverted for thorough mixing, and promptly delivered to the hospital on the same day. Within 24 h, participants stored the samples in a −80 °C ultra-low-temperature freezer.
Clinical data collection
We collected the baseline information of patients, including age, BMI, duration of diabetes, complications, family history, and health status. Demographic and clinical information for these patients was collected by an experienced neurologist from the patient review and/or medical records.
Participants underwent clinical examination four times during the trial: at baseline (visit 1), 1 month (visit 2), 3 months (visit 3), 6 months (visit 4) of metformin. Gastrointestinal symptoms, diet, and physical activity were collected at all four visits. Participants were examined with height, weight, and blood pressure at each visit. All participants were instructed to conduct at-home blood glucose monitoring four times, encompassing fasting and postprandial (2-hour) measurements, with an emphasis on maintaining dietary consistency throughout the monitoring period. No venous blood sampling was scheduled at the 1-month time point of medication administration. At visits 1, 3 and 4, respectively, blood samples were collected after participants had fasted for 10 h overnight. Fasting plasma glucose (FPG), Uric Acid (UA), alanine aminotransferase (ALT), aspartate aminotransferase (AST), hypersensitive C-reactive protein (hs-CRP), total cholesterol (TC), triglyceride (TG), high-density lipoprotein cholesterol (HDL-C), low-density lipoprotein Cholesterol (LDL-C), urine albumin-to-creatinine ratio (ACR), free fatty acid (FFA), Apolipoprotein B (ApoB) were analyzed by full-automatic blood biochemical analyzer. Fasting insulin (FINS) and C-Peptide (C-P) were measured using chemiluminescence immunoassay. Hemoglobin A1c (HbA1c) was measured using liquid chromatography. We calculated homeostasis model assessment indices (HOMA) of β-cell function (HOMA-β) and of insulin resistance (HOMA-IR) as follows: HOMA-IR = FPG (mmol/L)*FINS (mIU/L)/22.5. HOMA-β = 20*FINS (mIU/L)/(FPG (mmol/L) − 3.5). BMI = weight(kg)/height(m)2.
Bacterial DNA extraction and 16S rRNA gene sequencing
DNA extraction utilized the DNeasy® PowerSoil® Pro Kit (QIAGEN, Germany) following the manufacturer’s protocols. The V3–V4 region of the 16S rRNA gene was Polymerase Chain Reaction (PCR)-amplified, and Illumina HiSeq 2500 platform sequenced the PCR products. Paired-end reads were merged using FLASH [22]. Quality filtering, performed by the fastq quality filter in the FASTX-Toolkit 0.0.14, retained only high-quality reads (Q value ≥ 25) with a base ratio of ≥90%. Removal of chimeric reads was carried out using USEARCH 64-bit v8.0.1517. Read normalization, based on the smallest sample size, was achieved by random subtraction.
Bioinformatics analysis of sequencing data
Operational taxonomic units (OTUs) were aligned via the UCLUST algorithm with a 97% identity and taxonomically classified using the SILVA 16S rRNA database v128 [23]. Microbial alpha diversity was assessed using the Chao1, Shannon and Simpson indexs for richness and evenness. Beta diversity was computed using weighted UniFrac distance metrics, visualized through principal coordinate analysis (PCoA). The UniFrac distance metric measures the phylogenetic tree overlap between sample pairs, while the PCoA plot arranges samples based on community similarity. Analysis of similarities (ANOSIM) was used to assess the significance of separation between the tested groups of samples.
Statistical analysis
Data was presented as mean ± standard error of the mean. The paired two-tailed Student’s t test utilized differences in mean values for parametric variables between two groups. ANOVA with Tukey post hoc test was employed for comparing means among four groups, while the Mann-Whitney U test evaluated differences for nonparametric variables. Linear discriminant analysis effect size (LefSE) was utilized to identify taxonomic bacterial biomarkers and genes, setting an linear discriminant analysis (LDA) log score cutoff of 2 [24]. LDA, a statistical method used for dimensionality reduction and classification, was used to identify biomarkers significantly explaining differences between microbial taxa. LDA > 2 indicates significant differences, and when this ratio is >3, the differences become more pronounced. Our main focus is on taxa with LDA scores greater than 3. The DESeq2 package calculated differentially expressed OTUs [25]. Spearman correlation analyses assessed associations between taxa abundance and glycemic/inflammatory traits. To classify the longitudinal pattern of taxonomic changes in the gut microbiota, we calculated the within-cluster sum of squares for different cluster numbers and used the elbow method to obtain the optimal number of clusters. The results were assessed using the R package NbClust. Fuzzy c-means clustering, using the Mfuzz R package, classified gut microbiota trajectories [26]. Multivariable linear regression models, including BMI as covariates, were computed to understand the bacterial genera and temporal correlations in each cluster. PICRUSt2 [27] evaluated differences in community functional attributes, identifying significantly differential KEGG Orthologys (KOs) through LEfSe. Metabolic pathways were verified in the MetaCyc database. GraphPad Prism (v10.1.0) and R software(v4.3.2) were used for analysis and generate images. Significance was considered at P < 0.05.
Results
Characteristic of the study subjects
This study comprised 25 treatment-naïve T2DM patients, including 15 males and 10 females, with an average age of 46.80 ± 11.42 years (range: 25–69 years). The median duration of diabetes was 12 months (range: 1–120 months). Nineteen individuals underwent an 8-week intervention involving diet and exercise control, and 17 patients had a family history of diabetes. Concerning complications, three had diabetic nephropathy, and two presented with mild diabetic retinopathy. Eight patients were concurrently using antihypertensive medications, while 6 were using lipid-lowering medications. The overall rate of intervention-related side effects was 8%. Two patients experienced gastrointestinal adverse reactions, persisting in one individual but resolving in the other after one week of medication. No participants withdrew due to severe side effects. Clinical and biochemical data at baseline and at 3 and 6 months of medication are presented in Table 1. At 1 month of metformin treatment, patients exhibited significant reductions in weight (82.12 ± 16.40 vs. 80.45 ± 15.87, P < 0.001) and BMI (28.10 ± 4.41 vs. 27.53 ± 4.27, P < 0.001) compared to pre-treatment levels. Weight continued to decrease with ongoing treatment, with a more pronounced decrease observed within the first 3 months of treatment (P < 0.001), and weight tended to stabilize between months 3 and 6 of treatment. Regarding glycemic and lipid metabolism indicators, significant improvements were observed at 1 month of metformin therapy, with self-monitoring of blood glucose levels showing a significant decrease in fasting blood glucose (FBG) (8.64 ± 2.15 vs. 6.35 ± 0.89, P < 0.001) and 2-h postprandial blood glucose (2 h PBG) (12.91 ± 4.80 vs. 7.72 ± 1.89, P < 0.001). At 3 months of treatment, both FPG and HbA1c significantly decreased compared to pre-treatment levels (Table 1). At 3 months of treatment, only 4 patients still had HbA1c levels exceeding 6.5%. By 6 months of treatment, HOMA-IR had decreased compared to pre-treatment levels, with a significant statistical difference in the decrease. Lipid and inflammation indicators showed decreases compared to pre-treatment levels, but without significant statistical differences.
Differences in microbial diversity
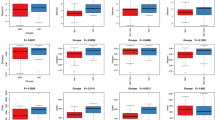
This study examined differences in α diversity indices among different groups, encompassing the Chaol index for species richness assessment, Shannon and Simpson indices for a comprehensive reflection of richness and evenness within species; and the observed species index reflecting the number of observed OTU. According to the results of the inter-group diversity index analysis (Fig. 2a), after treatment with metformin, there was an overall decrease in species richness and diversity of the intestinal microbiota. Specifically, the observed species(P = 0.005), Shannon index (P = 0.004), and Simpson index (P = 0.043) at 3 months of treatment were significantly lower than pre-treatment levels, indicating statistically significant differences (Table 2). β-diversity in subjects before metformin treatment and at 1 month, 3 months, and 6 months of treatment did not exhibit significant separation trends along the principal component 1 (PC1) and principal component 2 (PC2) directions (Fig. 2b, ANOSIM analysis, P = 0.066). We observed no significant differences in β diversity estimates among the four groups.
The alteration of phyla richness and composition structure in T2DM patients for metformin exposure. a Comparison of α-diversity in microbiota of the study groups based on the Shannon index, Simpson index, and Chao1 analysis. b Principal coordinates analysis of the weighted Unifrc dissimilarity matrix shows no significant differences in the beta diversity (the composition structure of fecal microbiota) between groups. c The percentage composition of the top-ranking phyla by relative abundance across four different groups, highlighting Firmicutes and Bacteroidetes as the dominant phyla. d The percentage contribution of the top-ranking genera to the total genera abundance across four different groups, indicating the highest proportions are: Bacteroides, Prevotella 9 and Faecalibacterium genus
Changes in taxonomic profile induced by metformin treatment
In total, 100 stool samples were obtained from 25 T2DM individuals. A total of 43,582,493 DNA sequencing reads were left after demultiplexing and quality control filtering. A rarefaction depth of 435,825 reads per sample was selected, and 25,712 OTUs were identified across 100 samples. We next tested for differences in the number of observed OTUs across the groups of participants.
As shown in Fig. 2c, the phylum Bacteroidetes accounted for 40.09%, 48.42%, 47.04%, and 47.31% of the total relative abundance in the MET0, MET1, MET2 and MET3 groups, respectively. The phylum Firmicutes represented 46.56%, 36.29%, 31.26% and 34.69% of the total relative abundance in the four groups, respectively. From the boxplot, we can see that the fecal microbiome of metformin-treated patients showed a trend for a decrease in the ratio of phyla Firmicutes to Bacteriodetes (F/B ratio) (ANOVA: P = 0.037), and there was a statistically significant difference between the paired two groups (MET0 vs. MET1, 1.734 ± 2.153 vs. 0.81 ± 0.373, P = 0.049; MET0 vs. MET2, 1.734 ± 2.153 vs. 0.694 ± 0.266, P = 0.028; MET0 vs. MET3, 1.734 ± 2.153 vs. 0.757 ± 0.269, P = 0.039). At the genus level, the predominant genera of gut microbiota in the three groups are as follows (Fig. 2d): Bacteroides genus: accounted for 25.35%, 34.57%, 31.62%, and 35.92% in the MET0, MET1, MET2, and MET3 groups, respectively. Prevotella 9 genus: represented 12.51%, 10.50%, 11.95%, and 8.10% in the MET0, MET1, MET2, and MET3 groups, respectively. Faecalibacterium genus: constituted 7.89%, 5.89%, 4.69% and 3.64% in the MET0, MET1, MET2, and MET3 groups, respectively. There were no significant alterations in the ranking of genera with high proportions, yet specific genera exhibited changes in abundance before and after medication. To observe in-depth changes in the gut microbiome composition, we used DESeq2 and evaluated the statistical significance of differential absolute abundance of OTU groups between time points. Metformin treatment for 1, 3, and 6 months resulted in significant alterations in the abundance of 6, 13, and 19 bacterial strains, respectively, with a false-discovery rate (FDR) < 0.05. At the genera level, at one month, when compared with pre-treatment, a significant increase in Citrobacter (Proteobacteria| Enterobacteriaceae) was observed (adj. P < 0.05, Fig. 3a). A significant decrease in Romboutsia (Firmicutes| Peptostreptococcaceae) and Clostridium sensu stricto 1(Firmicutes| Clostridiaceae) was found. The Romboutsia difference remained significant the most in 3 months compared with the group before metformin (Fig. 3b). Pseudomonas (Proteobacteria| Pseudomonadaceae) and Escherichia-Shigella (Proteobacteria| Enterobacteriaceae) increased significantly. Alloprevotella (Bacteroidetes| Prevotellaceae) increased at three months of treatment. Ruminiclostridium (Firmicutes| Ruminococcaceae) decreased most, and Tyzzerella 3 (Firmicutes| Lachnospiraceae) increased most in 6-month patient subgroups compared with pre-treatment, respectively (adj. P < 0.05, Fig. 3c).
The statistical significance of differential absolute abundance of OTU groups between time points by Volcano plot: (a) MET0 versus MET1, (b) MET0 versus MET2, (c) MET0 versus MET3. LDA scores (log10) of the OTUs. LDA was used to identified differentially abundant OTU features based on LDA effect size (LEfSe), with LDA score >2.0 defined as significantly greater abundance for comparisons of the (d) MET0 versus MET1, (e) MET0 versus MET2, (f) MET0 versus MET3. MET0 – pre-metformin treatment, MET1 – 1 month after the start of treatment, MET2 – 3 months into therapy, and MET3 – 6 months of metformin treatment
In addition, for graphic representation of differentially abundant taxa as well as their effect sizes and phylogenetic relationship, the LEfSe method was performed (Fig. 3d–f). This method detected 17 differentially abundant taxonomic clades, which mainly matched those found with edgeR analysis. At the phylum level, we found that at the final point, subjects from the 6-month group had a higher abundance of Bacteroidetes (LDA > 3), as well as its family Bacteroidaceae (LDA > 4) and its genus Bacteroides (LDA > 4), while subjects in the pre-treatment group presented a higher abundance of Firmicutes (LDA > 4), and its family Ruminococcaceae (LDA > 2), Peptostreptococcaceae (LDA > 3) and its genus Ruminiclostridium (LDA > 2), Faecalibacterium (LDA > 3), and Romboutsia (LDA > 2).
In order to verify the findings from previous publications reporting that metformin increased the abundance of Akkermansia (Verrucomicrobia| Akkermansiaceae) Verrucomicrobia (P = 0.757), Prevotella (Bacteroidetes| Prevotellaceae) (P = 0.551), and Fusobacterium (Fusobacteria| Fusobacteriaceae) (P = 0.270), a continuous growth trend was noted in these bacterial species, but these changes were not statistically significant. This particular microbiota can produce SCFAs. SCFAs are directly utilized as an energy source by the intestinal mucosal cells or transferred to the systemic circulation to generate an important source of energy for the host and have the ability to behave as signaling molecules [28].
Metformin treatment led to distinct changes in different gut microbiota subgroups over time
To identify the different longitudinal trajectories of the gut microbiota in patients treated with metformin, we calculated the within-cluster sum of squares for different cluster numbers based on the mean relative abundance of the microbial genera. Three clusters of longitudinal trajectories for microbial fluctuation were identified via the “elbow” method (Supplementary Fig. S1). Then, the longitudinal trajectories of the three clusters were constructed using fuzzy c-means clustering, including (i) genera temporarily changing from pre-treatment to 6 months (Fig. 4a); (ii) genera continuing to decrease from pre-treatment to 6 months (Fig. 4b), and (iii) genera continuing to increase from pre-treatment to 6 months (Fig. 4c). Detailed information on the gut microbiota of each cluster is shown in Figure S2. Genera in all clusters were selected to construct multivariate linear regression models to verify the correlation between gut microbiota and time. After adjusting for BMI, six genera were found to have changed significantly over time (Fig. 4d). The abundance of the Lachnospiraceae ND3007 group (P = 0.006), [Eubacterium] xylanophilum group (P = 0.014), Romboutsia (P = 0.013), Faecalibacterium (P = 0.003), and Ruminococcaceae UCG-014 (P = 0.008) decreased significantly over time. Conversely, the abundance of Bacteroides (P = 0.049) increased over time.
Metformin treatment leads to distinct changes in different subgroups of the gut microbiota over time. Fuzzy c-means clustering was used to classify the gut microbiota into different clusters based on longitudinal tracks (a–c). The mean relative abundance of bacteria at the genus level was used to establish the longitudinal trajectories of three clusters. d A horizontal bar plot shows the standard coefficients for different genera
Clinical parameters correlated with the gut microbiota
Metformin treatment was associated with significantly improved treatment efficacy and changes of 15 clinical metabolic parameters. We further investigated whether these improvements correlate with the change in microbiota compositions and clinical metabolic parameters by using a correlation matrix based on the Spearman correlation distance test. Heatmaps showing correlations between clinical factors and gut microbiota at baseline and after metformin treatment (Fig. 5a, b illustrated the correlation at phylum and genus level, respectively) Phylum Firmicutes positively correlated with HbA1c and FPG, whereas Bacteroidetes exhibited the opposite correlation. At the genus level, Bacteroides and Escherichia_Shigella positively associated with decreased HbA1c and FPG; Ruminococcus, Lachnospira, Blautia, and Lachnospiraceae UCG_008 were negatively correlated with elevated HbA1c, FINS, and HOMA; Paraprevotella was positively correlated with elevated LDL-C, while Parasutterella was negative with it. Lachnospiraceae UCG_008 was correlated with elevated FPG, while Escherichia_Shigella exhibited the opposite correlation with the change of FPG. However, we noticed that these correlations were not statistically significant.
Spearman correlation analysis of the relationships among related clinical metabolic parameters and the relative abundance of altered genera at (a) phylum and (b) genus level, respec-tively. Asterisks indicate statistically significant Spearman correlations (*P < 0.05, **P < 0.01, ***P < 0.001). ALT alanine transaminase, hsCRP hypersensitive C-reactive protein, FPG Fasting plasma glucose, UA Uric Acid, AST aspartate aminotransferase, TC total cholesterol, TG triglyceride, HDL-C high-density lipoprotein cholesterol, LDL-C low-density lipoprotein Choles-terol, ACR albumin-to-creatinine ratio, FFA free fatty acid, ApoB Apolipoprotein B, FINS fasting serum lisulin, C-P C-Peptide, HbA1c Hemoglobin A1c, HOMA homeostasis model assessment indices
Functional analysis
To investigate the functional role of the gut microbiota in immunotherapy, we further annotated the metagenomic sequencing data in the KEGG database and identified 28 KOs enriched in the MET0 group, 4 KOs enriched in the after-treatment group by LEfSe (LDA > 2, P < 0.05) (Fig. 6a). The most abundant KOs in the pre-treatment group and post-treatment group were energy metabolism pathways and amino acid metabolism pathways, respectively (Fig. 6b). At the pathway level, the MET2 group was significantly enriched with KOs that played essential functions in several KEGG pathways, including cationic antimicrobial peptide (CAMP) resistance (map01503), pertussis(map05133), betalain biosynthesis (map00965), peroxisome (map04146) and longevity regulating pathway(map04211). Moreover, the MAPK signaling pathway - fly (map04013) was enriched in the MET1 group (Supplementary Fig. S3). Several after-treatment group-associated KOs, Bacteroides, Achromobacter, and Parabacteroides, which belong to the Bacteroidetes and Proteobacteria phylum, were found to be the dominant species enriched (Fig. 6c, d). Furthermore, comparisons of differential metabolic pathways in the KEGG database showed that amino acid metabolism pathways, such as isoleucine biosynthesis, predominated in the post-treatment group. In the MetaCyc database, we found that metformin significantly altered 17 functional pathways (Supplementary Figs. S4 and 5), including ten pathways of biosynthesis, four pathways of L-arginine, vanillate or ornithine degradation, a polymyxin resistance pathway, and two pathways in metabolism: superpathway of chorismate metabolism and ketogluconate metabolism.
Functional annotation of the gut microbiome metagenomic sequencing data. a Different KOs enriched in the four groups identified by LEfSe (LDA > 2, P < 0.05). b Functional pathways associated with the differentially-enriched KOs identified by LEfSe (LDA > 2, P < 0.05). c The top 10 bacterial species contributed to starvation-inducible iron complex multidrug efflux system (K03085) enrichment in the after-treatment group. d The top 10 bacterial species contributed to starvation-inducible iron complex outermembrane recepter protein (K02014) enrichment in the after-treatment group
Discussion
Our study, the first of its kind, utilized a longitudinal design to investigate the long-term effects of metformin therapy on treatment-naive patients. Analyzing 100 samples collected before treatment and at 1, 3 and 6 months of metformin treatment, we aimed to understand the temporal dynamics of the gut microbiota. Unlike single-time-point studies, longitudinal designs offer crucial insights into periodic patterns, interdependencies, and temporal variations within the microbiome [29].
Results revealed a decrease in intestinal microbiota species diversity at 3 months of metformin treatment compared to pre-treatment, while β-diversity remained unchanged. This reduction in diversity may stem from the drug’s direct impact on microbial species or dietary control initiated by patients post-T2DM diagnosis, leading to a more uniform dietary structure. Similar findings have been observed in healthy individuals taking metformin, with some studies noting a significant reduction in microbial community diversity within 24 h of metformin administration [30]. The prevailing perspective suggests lower species diversity in the gut microbiota of individuals with disease states compared to healthy individuals [31]. However, conflicting findings exist; a recent systematic review summarizing 13 clinical studies revealed varying impacts of metformin on microbial diversity, with 8 studies reporting no significant effect, 3 showing an increasing trend, and 2 indicating an apparent decrease in α-diversity [13].
In terms of species composition at the phylum level, Firmicutes, Bacteroidetes, and Proteobacteria were the top three dominant bacterial phyla. Metformin administration led to a gradual increase in Bacteroidetes, a decline in Firmicutes, and a significant decrease in the Firmicutes/Bacteroidetes (F/B) ratio. Our study, alongside previous research on metformin’s impact on the gut microbiota in T2DM patients, suggest a significant decrease in the F/B ratio after metformin intake. A study by Carbajal et al. in 2020 found a nearly 50% decrease in the relative abundance of Firmicutes in patients with T2DM who had been orally administered metformin for at least a year [32]. Additionally, a study in an Estonian population of 38 individuals suggested that a reduction in certain anaerobic bacteria in the gut, such as Bacteroidetes, could predict hyperglycemia [33]. The F/B ratio’s association with increased plasma glucose following an oral glucose load aligns with previous studies, although conflicting trends exist [34, 35].
Our study identified three longitudinal trajectories for the altered gut microbiota. Cluster 1 comprised genera that exhibited transient changes, returning to baseline levels over time. Clusters 2 and 3 exhibited a relatively stable change trend. Bacteroides showed enrichment in patients after metformin treatment, which was strongly linked to notable improvements in glucose metabolism. While previous studies reported a significant decrease in Bacteroides after metformin treatment, which correlated with reduced HbA1c [6], our findings align with Sun et al.‘s discovery of that short-term metformin treatment was associated with a lowered abundance of Bacteroides fragilis, resulting in improved glucose tolerance [19]. Despite our 16S rRNA gene-based amplicon sequencing not being capable of perfectly assigning individual species among the genus Bacteroides, we observed inconsistent results were observed. Clinical studies have reported a decrease in the abundance of Bacteroides [36, 37], as well as in C57BL/6 mice fed with a high-fat diet for 24 weeks of metformin [34]. Furthermore, metformin tended to increase the trend in the abundance of Akkermansia and Prevotella, both known for producing SCFAs to improve metabolic dysfunction, including hyperglycemia.
The bacterial genera Lachnospiraceae ND3007 group, [Eubacterium] xylanophilum group, Romboutsia, Faecalibacterium, and Ruminococcaceae UCG-014 exhibited a decreasing trend over time, with only Romboutsia showing a significant change. [Eubacterium] xylanophilum ferments complex phytochemicals to produce SCFAs, including butyrate, which has been found to be antiobesogenic in human studies [38, 39]. The [Eubacterium] xylanophilum group, known for fermenting complex phytochemicals to produce SCFAs, including butyrate, showed potential antiobesogenic effects in human studies [40]. Ruminococcaceae UCG-014 can increase nitric oxide (NO) synthase activity by regulating the MyD88 pathway, thereby upregulating NO levels and inhibiting inflammatory responses [41]. Faecalibacterium is a fiber-degrading bacteria capable of fermenting fiber and producing SCFAs [42]. Interestingly, the bacteria mentioned above, all producers of SCFAs, showed a notable increase in abundance at 6 months, despite a decrease observed in 1 and 3 months. It is noteworthy that most clinical studies reveal that metformin enhances the bacteria-producing SCFAs, mediating therapeutic effects through improvement in SCFA production, bile acid (BA) pool composition alteration, or reduction in lipopolysaccharide (LPS) production [5]. SCFAs exhibit various beneficial effects on animal performance and intestinal health [43], though some studies suggest their effects may not be sustained through 12 months, potentially due to compensatory use during prolonged negative energy balance [14]. Prevotella and Bifidobacterium, also SCFA-producing bacteria, exhibited a continuous but statistically nonsignificant increase over time. Above results indicated that not all SCFA-producing bacteria consistently rise during metformin treatment. The cause of the waning effect on SCFAs and its relevance to health require further research, and our study will enhance SCFA examination to provide more clarity in the future.
Conversely, Romboutsia, Ruminococcus, and Clostridium sensu stricto exhibited positive correlations with body weight and serum lipids, and gradually decreased with improved weight and blood lipid profiles due to metformin treatment. The mechanism of action for the Lachnospiraceae_ND3007_group remains unclear. GMrepo, a database linking human phenotypes and gut microbiota [44], associates Lachnospiraceae with Crohn’s disease, colitis, and obesity. This partly explains the reduction in this bacterium with metformin treatment due to its positive correlation with obesity-related parameters.
In addition to the longitudinal fluctuations observed in certain taxa over the treatment period, our cross-sectional analysis conducted before and during medication revealed significant reductions in Pseudomonas, Ruminiclostridium and Clostridium sensu stricto, alongside notable increases in Escherichia-Shigella and Tyzzerella 3 as shown in Fig. 3. Pseudomonas exhibited a notable decline after three months of medication, aligning with improved blood glucose levels. However, its overall trend showed an increase, particularly in non-controlled blood glucose. This Gram-negative opportunistic pathogen might contribute to T2DM pathogenesis through endotoxin production inducing inflammatory factors [45]. Pseudomonas remained more prominent in non-controlled blood glucose levels, indicating its potential relevance to disease management. Several studies have indicated a correlation between the genus Ruminiclostridium and obesity. Obese mice have higher levels of Ruminococcus in their fecal microbiota compared to lean mice [46, 47]. Furthermore, a positive correlation between Ruminococcus abundance and blood IgM levels in obese mice suggests a potential role in inflammation and dyslipidemia [47]. Clostridium sensu stricto 1 is an opportunistic pathogen [48, 49], which can cause intestinal inflammation and decrease the content of SCFAs. We did observe a gradual increase in Escherichia-Shigella abundance over time following medication, consistent with previous studies [16, 50, 51]. This increase is commonly attributed to gastrointestinal side effects associated with metformin intake, with reports indicating a positive correlation between its abundance and the reduction in BMI after metformin administration [36, 52]. Simultaneously, an increase in Escherichia has been linked to increases in serum acetate, correlating with decreases in fasting insulin, anti-inflammation, and insulin resistance in individuals with T2DM [14, 53]. The relationship between Tyzzerella 3 and glucose metabolism is unclear, but studies have found significantly lower abundance of this bacterium in individuals with T2DM compared to healthy individuals [54]. In our study, we observed a significant increase in Tyzzerella 3 abundance at 6 months of metformin treatment. Therefore, it is speculated that the decrease in this bacterium in T2DM patients may be associated with high blood glucose levels.
Functional analysis revealed that amino acid metabolism pathways, such as isoleucine biosynthesis, predominated in the post-treatment group based on comparisons of differential metabolic pathways in the KEGG database. In the MetaCyc database, we discovered that metformin significantly altered 17 functional pathways, including pathways of biosynthesis, amino acid degradation, and metabolism. These findings suggested that the underlying mechanisms of the gut microbiome influencing metformin’s benefit in glycometabolism might be driven by specific bacterial species involved in different metabolic pathways.
Our study’s strengths lie in its longitudinal design, which enables the exploration of the lasting effects of metformin therapy. Additionally, our cohort consisted of individuals recently diagnosed with T2DM, reducing confounding variables by excluding those not on other T2DM medications, with only a minority using statins or antihypertensive therapy. However, it is essential to acknowledge specific limitations. Firstly, from a technological perspective, the study’s reliance on 16S rRNA gene amplicon sequencing is acknowledged; although suitable for genus-level signals, it has limitations at the species level. Secondly, the investigation did not evaluate individual dietary habits or consider the potential influence of diet on the gut microbiota. Thirdly, a long-term study inherently carries the unavoidable risk of the impact of common environmental factors that permanently shape gut microbiota composition. Fourthly, there was a selection bias as study participants were recruited exclusively from the Han Chinese population, raising uncertainties regarding the generalizability of our findings to other ethnic groups. Last, all participants in the study exhibited satisfactorily controlled glucose levels, hindering the extrapolation of results to individuals with more severe forms of diabetes. Previous studies have found that the baseline microbiota composition can predict the short-term efficacy of treatment in newly diagnosed type 2 diabetes patients [18]. Observing changes in the microbiota within the first seven days of treatment to predict blood glucose control at three months is a promising approach. In our study, we had only one patient who showed poor response to metformin (HbA1c reduction of less than 1% at 6 months), which was insufficient for subgroup analysis. In future research, expanding the sample size will enable us to further investigate this aspect.
Conclusions
In conclusion, our prospective cohort study contributes valuable insights into the longitudinal dynamics of metformin’s impact on the gut microbiota. Consistent with shorter-term trials, metformin-induced alterations in microbiota composition, including increased Escherichia-Shigella and decreased Romboutsia. However, the link between microbial changes and long-term clinical outcomes remains unknown and warrants investigation in more extensive studies. Nevertheless, our study sheds light on the biological mechanisms through which metformin influences health outcomes and identifies potential microbial targets for preventing metabolic disorders or gastrointestinal symptoms affecting drug tolerability. Future well-designed trials, encompassing larger groups of T2DM patients, multi-omics data, and placebo control, are essential to unravel the intricacies of drug-microbiota interactions, comprehend the role of baseline gut microbiota, and advance microbiome-based applications in diabetes treatment.
Data availability
Supplementary data are available online at https://pan.baidu.com/s/1uL5Mm-eRUyp3aLlKIaOdJg?pwd=t4ph.
References
Collaborators GBDD, Global, regional, and national burden of diabetes from 1990 to 2021, with projections of prevalence to 2050: a systematic analysis for the Global Burden of Disease Study 2021. Lancet 402(10397), 203–234 (2023). https://doi.org/10.1016/S0140-6736(23)01301-6
U. Galicia-Garcia, A. Benito-Vicente, S. Jebari, A. Larrea-Sebal, H. Siddiqi, K.B. Uribe et al. Pathophysiology of Type 2 Diabetes Mellitus. Int. J. Mol. Sci. 21(17), 6275 (2020). https://doi.org/10.3390/ijms21176275
H. Sun, P. Saeedi, S. Karuranga, M. Pinkepank, K. Ogurtsova, B.B. Duncan et al. IDF Diabetes Atlas: Global, regional and country-level diabetes prevalence estimates for 2021 and projections for 2045. Diabetes Res. Clin. Pract. 183, 109119 (2022). https://doi.org/10.1016/j.diabres.2021.109119
H. Tilg, A.R. Moschen, Microbiota and diabetes: an evolving relationship. Gut 63(9), 1513–1521 (2014). https://doi.org/10.1136/gutjnl-2014-306928
A.L. Cunningham, J.W. Stephens, D.A. Harris, Gut microbiota influence in type 2 diabetes mellitus (T2DM). Gut. Pathog. 13(1), 50 (2021). https://doi.org/10.1186/s13099-021-00446-0
R. Li, F. Shokri, A.L. Rincon, F. Rivadeneira, C. Medina-Gomez, F. Ahmadizar, Bi-Directional Interactions between Glucose-Lowering Medications and Gut Microbiome in Patients with Type 2 Diabetes Mellitus: A Systematic Review. Genes. 14(8) (2023) https://doi.org/10.3390/genes14081572
M. Foretz, B. Guigas, B. Viollet, Metformin: update on mechanisms of action and repurposing potential. Nat. Rev. Endocrinol. 19(8), 460–476 (2023). https://doi.org/10.1038/s41574-023-00833-4
L.J. McCreight, C.J. Bailey, E.R. Pearson, Metformin and the gastrointestinal tract. Diabetologia 59(3), 426–435 (2016). https://doi.org/10.1007/s00125-015-3844-9
C.J. Bailey, C. Wilcock, J.H. Scarpello, Metformin and the intestine. Diabetologia 51(8), 1552–1553 (2008). https://doi.org/10.1007/s00125-008-1053-5
E. Bonora, M. Cigolini, O. Bosello, C. Zancanaro, L. Capretti, I. Zavaroni et al. Lack of effect of intravenous metformin on plasma concentrations of glucose, insulin, C-peptide, glucagon and growth hormone in non-diabetic subjects. Curr. Med. Res. Opin. 9(1), 47–51 (1984). https://doi.org/10.1185/03007998409109558
J.B. Buse, R.A. DeFronzo, J. Rosenstock, T. Kim, C. Burns, S. Skare et al. The Primary Glucose-Lowering Effect of Metformin Resides in the Gut, Not the Circulation: Results From Short-term Pharmacokinetic and 12-Week Dose-Ranging Studies. Diabetes Care 39(2), 198–205 (2016). https://doi.org/10.2337/dc15-0488
N.R. Shin, J.C. Lee, H.Y. Lee, M.S. Kim, T.W. Whon, M.S. Lee et al. An increase in the Akkermansia spp. population induced by metformin treatment improves glucose homeostasis in diet-induced obese mice. Gut 63(5), 727–735 (2014). https://doi.org/10.1136/gutjnl-2012-303839
P. Pavlo, I. Kamyshna, A. Kamyshnyi, Effects of metformin on the gut microbiota: A systematic review. Mol. Metab. 77, 101805 (2023). https://doi.org/10.1016/j.molmet.2023.101805
N.T. Mueller, M.K. Differding, M. Zhang, N.M. Maruthur, S.P. Juraschek, E.R. Miller 3rd et al. Metformin Affects Gut Microbiome Composition and Function and Circulating Short-Chain Fatty Acids: A Randomized Trial. Diabetes Care 44(7), 1462–1471 (2021). https://doi.org/10.2337/dc20-2257
J. de la Cuesta-Zuluaga, N.T. Mueller, V. Corrales-Agudelo, E.P. Velasquez-Mejia, J.A. Carmona, J.M. Abad et al. Metformin Is Associated With Higher Relative Abundance of Mucin-Degrading Akkermansia muciniphila and Several Short-Chain Fatty Acid-Producing Microbiota in the Gut. Diabetes Care 40(1), 54–62 (2017). https://doi.org/10.2337/dc16-1324
K. Forslund, F. Hildebrand, T. Nielsen, G. Falony, E. Le Chatelier, S. Sunagawa et al. Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature 528(7581), 262–266 (2015). https://doi.org/10.1038/nature15766
F. Zhang, M. Wang, J. Yang, Q. Xu, C. Liang, B. Chen et al. Response of gut microbiota in type 2 diabetes to hypoglycemic agents. Endocrine 66(3), 485–493 (2019). https://doi.org/10.1007/s12020-019-02041-5
I. Elbere, I. Silamikelis, I.I. Dindune, I. Kalnina, M. Ustinova, L. Zaharenko et al. Baseline gut microbiome composition predicts metformin therapy short-term efficacy in newly diagnosed type 2 diabetes patients. PLoS One 15(10), e0241338 (2020). https://doi.org/10.1371/journal.pone.0241338
L. Sun, C. Xie, G. Wang, Y. Wu, Q. Wu, X. Wang et al. Gut microbiota and intestinal FXR mediate the clinical benefits of metformin. Nat. Med. 24(12), 1919–1929 (2018). https://doi.org/10.1038/s41591-018-0222-4
H. Nakajima, F. Takewaki, Y. Hashimoto, S. Kajiyama, S. Majima, H. Okada et al. The Effects of Metformin on the Gut Microbiota of Patients with Type 2 Diabetes: A Two-Center, Quasi-Experimental Study. Life 10(9), 195 (2020). https://doi.org/10.3390/life10090195
A. Napolitano, S. Miller, A.W. Nicholls, D. Baker, S. Van Horn, E. Thomas et al. Novel gut-based pharmacology of metformin in patients with type 2 diabetes mellitus. PLoS One 9(7), e100778 (2014). https://doi.org/10.1371/journal.pone.0100778
T. Magoc, S.L. Salzberg, FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27(21), 2957–2963 (2011). https://doi.org/10.1093/bioinformatics/btr507
C.A. Dennison, A.J. Eslinger, R.A. Reimer, Preconception Prebiotic and Sitagliptin Treatment in Obese Rats Affects Pregnancy Outcomes and Offspring Microbiota, Adiposity, and Glycemia. Front. Endocrinol. 8, 301 (2017). https://doi.org/10.3389/fendo.2017.00301
N. Segata, J. Izard, L. Waldron, D. Gevers, L. Miropolsky, W.S. Garrett et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12(6), R60 (2011). https://doi.org/10.1186/gb-2011-12-6-r60
M.I. Love, W. Huber, S. Anders, Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15(12), 550 (2014). https://doi.org/10.1186/s13059-014-0550-8
R.L. Cannon, J.V. Dave, J.C. Bezdek, Efficient Implementation of the Fuzzy c-Means Clustering Algorithms. IEEE Trans. Pattern Anal. Mach. Intell. 8(2), 248–255 (1986). https://doi.org/10.1109/tpami.1986.4767778
G.M. Douglas, V.J. Maffei, J.R. Zaneveld, S.N. Yurgel, J.R. Brown, C.M. Taylor et al. PICRUSt2 for prediction of metagenome functions. Nat. Biotechnol. 38(6), 685–688 (2020). https://doi.org/10.1038/s41587-020-0548-6
A. Koh, F. De Vadder, P. Kovatcheva-Datchary, F. Backhed, From Dietary Fiber to Host Physiology: Short-Chain Fatty Acids as Key Bacterial Metabolites. Cell 165(6), 1332–1345 (2016). https://doi.org/10.1016/j.cell.2016.05.041
S.Y. Park, A. Ufondu, K. Lee, A. Jayaraman, Emerging computational tools and models for studying gut microbiota composition and function. Curr. Opin. Biotechnol. 66, 301–311 (2020). https://doi.org/10.1016/j.copbio.2020.10.005
I. Elbere, I. Kalnina, I. Silamikelis, I. Konrade, L. Zaharenko, K. Sekace et al. Association of metformin administration with gut microbiome dysbiosis in healthy volunteers. PLoS One 13(9), e0204317 (2018). https://doi.org/10.1371/journal.pone.0204317
S. Mrozinska, P. Radkowski, T. Gosiewski, M. Szopa, M. Bulanda, A.H. Ludwig-Galezowska et al. Qualitative Parameters of the Colonic Flora in Patients with HNF1A-MODY Are Different from Those Observed in Type 2 Diabetes Mellitus. J. Diabetes Res. 2016, 3876764 (2016). https://doi.org/10.1155/2016/3876764
A. Chavez-Carbajal, M.L. Pizano-Zarate, F. Hernandez-Quiroz, G.F. Ortiz-Luna, R.M. Morales-Hernandez, A. De Sales-Millan et al. Characterization of the Gut Microbiota of Individuals at Different T2D Stages Reveals a Complex Relationship with the Host. Microorganisms 8(1), 94 (2020). https://doi.org/10.3390/microorganisms8010094
E. Sepp, H. Kolk, K. Loivukene, M. Mikelsaar Higher blood glucose level associated with body mass index and gut microbiota in elderly people. Microb. Ecol. Health Dis. 25 (2014). https://doi.org/10.3402/mehd.v25.22857
P.M. Ryan, E. Patterson, I. Carafa, R. Mandal, D.S. Wishart, T.G. Dinan et al. Metformin and Dipeptidyl Peptidase-4 Inhibitor Differentially Modulate the Intestinal Microbiota and Plasma Metabolome of Metabolically Dysfunctional Mice. Can. J. Diabetes 44(2), 146–55.e2 (2020). https://doi.org/10.1016/j.jcjd.2019.05.008
N. Larsen, F.K. Vogensen, F.W. van den Berg, D.S. Nielsen, A.S. Andreasen, B.K. Pedersen et al. Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PLoS One 5(2), e9085 (2010). https://doi.org/10.1371/journal.pone.0009085
H.S. Ejtahed, R.Y. Tito, S.D. Siadat, S. Hasani-Ranjbar, Z. Hoseini-Tavassol, L. Rymenans et al. Metformin induces weight loss associated with gut microbiota alteration in non-diabetic obese women: a randomized double-blind clinical trial. Eur. J. Endocrinol. 180(3), 165–176 (2019). https://doi.org/10.1530/EJE-18-0826
X. Niu, P. Lu, L. Huang, Y. Sun, M. Jin, J. Liu, et al. The effect of metformin combined with liraglutide on gut microbiota of Chinese patients with type 2 diabetes. Int. Microbiol. (2023). https://doi.org/10.1007/s10123-023-00380-y
L. Crovesy, D. Masterson, E.L. Rosado, Profile of the gut microbiota of adults with obesity: a systematic review. Eur. J. Clin. Nutr. 74(9), 1251–1262 (2020). https://doi.org/10.1038/s41430-020-0607-6
C.K. Chakraborti, New-found link between microbiota and obesity. World J. Gastrointest. Pathophysiol. 6(4), 110–119 (2015). https://doi.org/10.4291/wjgp.v6.i4.110
T. Tavella, S. Rampelli, G. Guidarelli, A. Bazzocchi, C. Gasperini, E. Pujos-Guillot et al. Elevated gut microbiome abundance of Christensenellaceae, Porphyromonadaceae and Rikenellaceae is associated with reduced visceral adipose tissue and healthier metabolic profile in Italian elderly. Gut Microbes 13(1), 1–19 (2021). https://doi.org/10.1080/19490976.2021.1880221
J.L. Kubinak, C. Petersen, W.Z. Stephens, R. Soto, E. Bake, R.M. O’Connell et al. MyD88 signaling in T cells directs IgA-mediated control of the microbiota to promote health. Cell Host Microbe 17(2), 153–163 (2015). https://doi.org/10.1016/j.chom.2014.12.009
Q. Zeng, D. Li, Y. He, Y. Li, Z. Yang, X. Zhao et al. Discrepant gut microbiota markers for the classification of obesity-related metabolic abnormalities. Sci. Rep. 9(1), 13424 (2019). https://doi.org/10.1038/s41598-019-49462-w
P. Guilloteau, L. Martin, V. Eeckhaut, R. Ducatelle, R. Zabielski, F. Van Immerseel, From the gut to the peripheral tissues: the multiple effects of butyrate. Nutr. Res. Rev. 23(2), 366–384 (2010). https://doi.org/10.1017/S0954422410000247
S. Wu, C. Sun, Y. Li, T. Wang, L. Jia, S. Lai et al. GMrepo: a database of curated and consistently annotated human gut metagenomes. Nucleic Acids Res. 48(D1), D545–D553 (2020). https://doi.org/10.1093/nar/gkz764
M. Tong, X. Li, L. Wegener Parfrey, B. Roth, A. Ippoliti, B. Wei et al. A modular organization of the human intestinal mucosal microbiota and its association with inflammatory bowel disease. PLoS One 8(11), e80702 (2013). https://doi.org/10.1371/journal.pone.0080702
P. Wang, J. Gao, W. Ke, J. Wang, D. Li, R. Liu et al. Resveratrol reduces obesity in high-fat diet-fed mice via modulating the composition and metabolic function of the gut microbiota. Free Radic. Biol. Med. 156, 83–98 (2020). https://doi.org/10.1016/j.freeradbiomed.2020.04.013
Q. Luo, D. Cheng, C. Huang, Y. Li, C. Lao, Y. Xia et al. Improvement of Colonic Immune Function with Soy Isoflavones in High-Fat Diet-Induced Obese Rats. Molecules 24(6), 1139 (2019). https://doi.org/10.3390/molecules24061139
P. Fan, P. Liu, P. Song, X. Chen, X. Ma, Moderate Dietary Protein Restriction Alters the Composition of Gut Microbiota and Improves Ileal Barrier Function in Adult Pig Model. Sci. Rep. 7, 43412 (2017). https://doi.org/10.1038/srep43412
W.Y. Yang, Y. Lee, H. Lu, C.H. Chou, C. Wang, Analysis of Gut Microbiota and the Effect of Lauric Acid Against Necrotic Enteritis in Clostridium Perfringens and Eimeria Side-by-Side Challenge Model. PloS One 14(5), e0205784 (2019). https://doi.org/10.1371/journal.pone.0205784
H. Wu, E. Esteve, V. Tremaroli, M.T. Khan, R. Caesar, L. Manneras-Holm et al. Metformin alters the gut microbiome of individuals with treatment-naive type 2 diabetes, contributing to the therapeutic effects of the drug. Nat. Med. 23(7), 850–858 (2017). https://doi.org/10.1038/nm.4345
Q. Zhang, X. Xiao, J. Zheng, M. Li, M. Yu, F. Ping et al. Featured article: Structure moderation of gut microbiota in liraglutide-treated diabetic male rats. Exp. Biol. Med. 243(1), 34–44 (2018). https://doi.org/10.1177/1535370217743765
M. Million, E. Angelakis, M. Maraninchi, M. Henry, R. Giorgi, R. Valero et al. Correlation between body mass index and gut concentrations of Lactobacillus reuteri, Bifidobacterium animalis, Methanobrevibacter smithii and Escherichia coli. Int. J. Obes. 37(11), 1460–1466 (2013). https://doi.org/10.1038/ijo.2013.20
C. Pedersen, U.Z. Ijaz, E. Gallagher, F. Horton, R.J. Ellis, E. Jaiyeola et al. Fecal Enterobacteriales enrichment is associated with increased in vivo intestinal permeability in humans. Physiol. Rep. 6(7), e13649 (2018). https://doi.org/10.14814/phy2.13649
Y. Que, M. Cao, J. He, Q. Zhang, Q. Chen, C. Yan et al. Gut Bacterial Characteristics of Patients With Type 2 Diabetes Mellitus and the Application Potential. Front. Immunol. 12, 722206 (2021). https://doi.org/10.3389/fimmu.2021.722206
Author information
Authors and Affiliations
Contributions
Conceptualization, W.Z. T.Y., and B.Z.; methodology, B.Z., W.Z. and Y.G.; software, N.L. and Y.G.; validation, T.Z., and N.L.; formal analysis, S.L., N.L., T.Z. and Y.G.; investigation, S.L., T.Z., Y.F. and Y.G.; resources, S.L., T.Z., N.L. and Y.G.; data curation, S.L., T.Z. and Y.G.; writing—original draft preparation, Y.G.; writing—review and editing, W.Z., T.Y., and Y.G.; visualization, N.L. and Y.G.; supervision, Y.F., T.Y., and W.Z; project administration, W.Z. and B.Z. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Consent for publication
The authors affirm that all individual participants provided informed consent for the publication of the data. Additional informed consent was obtained from all individual participants for whom identifying information was included in this article. Written informed consent has been obtained from the patients to publish this paper.
Ethics approval and consent to participate
The study was conducted in accordance with the Declaration of Helsinki, and approved by the Institutional Review Board of PUMCH (JS-2579/202009.22). Informed consent was obtained from all subjects involved in the study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gao, Y., Zhao, T., Lv, N. et al. Metformin-induced changes of the gut microbiota in patients with type 2 diabetes mellitus: results from a prospective cohort study. Endocrine 85, 1178–1192 (2024). https://doi.org/10.1007/s12020-024-03828-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12020-024-03828-x