Abstract
Paired-like homeodomain transcription factor 1 (PITX1) has been implicated as a tumor suppressor in various cancers. However, the biological and clinical significance of PITX1 in osteosarcoma has not been fully elucidated. Here, we studied the expression and clinical significance of PITX1 in 6 normal lower limb bone tissue specimens and 35 osteosarcoma tissue samples by immunohistochemistry. PITX1 was expressed in all normal tissues (6/6, 100 %) and in 85.7 % (30/35) of tumor tissues (P > 0.05). In addition, all normal tissue specimens showed high PITX1 expression (6/6, 100 %) while only 23.3 % (7/30) osteosarcoma tissue specimens had high PITX1 expression (P < 0.05). Patients with median overall survival (OS) >12 months had significantly higher PITX1 levels compared with those whose median OS was less than or equal to 12 months (P < 0.05 or 0.001). Furthermore, patients with lung metastasis had significantly lower PITX1 levels than patients without lung metastasis. In conclusion, PITX1 expression is downregulated in osteosarcoma and correlates with patient survival and lung metastasis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Osteosarcoma is derived from primitive mesenchymal cells and originates from the bone and only rarely from soft tissue. It is the most frequent primary solid malignancy of the bone, and if untreated, osteosarcoma runs a dismal course with local and often metastatic progression. Before the introduction of chemotherapy, nine in ten patients with osteosarcoma succumbed to pulmonary metastases [1]. Although osteosarcoma only accounts for 5–6 % of all childhood tumors, because of its high propensity to metastasize, osteosarcoma ranks among the most frequent causes of cancer-related death [2]. The low prevalence of osteosarcoma and its considerable heterogeneity make it difficult to obtain meaningful progress in patient survival. Although the 5-year survival rate may reach 70 % in patients who have received multimodal therapy, a significant proportion of them have an unfavorable prognosis due to lack of effective treatment options [3]. Therefore, it is imperative to develop new approaches for accurate diagnosis and improved treatment of osteosarcoma.
Paired-like homeodomain transcription factor 1 (PITX1) was first identified as a bicoid-related transcription factor and is involved in proopiomelanocortin gene expression [4]. It plays a critical role in specifying hindlimb morphology in vertebrates and enriches in the hindlimb [5]. Accumulating evidence has implicated PITX1 as a tumor suppressor in many parenchymal tumors. However, little is known about the tumor-inhibitory roles of PITX1 in osteosarcoma. Current studies show that cancer develops as a result of multiple genetic and epigenetic alterations [6, 7]. The delineation of patient response to chemotherapy and stratification of metastasis risk are important for a more precise and effective treatment. Aberrant expressions of oncogenes and tumor suppressor genes have been identified in most cancers. Better knowledge of changes in gene expression that occur in osteosarcoma may lead to improvements in diagnosis, treatment, and prevention. We hypothesized that PITX1 expression in osteosarcoma tissues may be dysregulated and could be of predictive significance for patient outcome.
In the current study, we determined PITX1 expression in osteosarcoma tissues and further investigated the correlation between PITX1 expression and clinicopathologic variables of osteosarcoma patients.
Patients and methods
Patient samples
We collected archival formalin-fixed, paraffin-embedded tissues from 35 patients with osteosarcoma at the First Affiliated Hospital of Shantou University Medical College between January 2001 and December 2013. All patients underwent potentially curative surgery without preoperative chemotherapy or radiotherapy. Archival formalin-fixed, paraffin-embedded tissues from lower limb normal bone were also obtained from six subjects.
The study was approved by the ethical review committee of the Medical College of Shantou University. Patient consent was not required because of the retrospective nature of the study.
Immunohistochemistry
An S-P Kit (Gene Tech, Shanghai) was used for immunohistochemical analysis. In brief, paraffin sections (4 μm thick) were deparaffinized in xylene and rehydrated in gradient alcohol. Antigen retrieval was performed by microwave treatment in 0.01 mol/L sodium citrate buffer (pH = 6.0) for 15 min, cooled to room temperature for 1 h, and then washed with phosphate-buffered saline (PBS). Endogenous peroxidase activity was inactivated with 0.3 % hydrogen peroxide for 30 min and non-specifically blocked with 5 % normal goat serum for 1 h. The sections were then incubated overnight with rabbit polyclonal PITX1 antibodies (Abcam, Cambridge, MA, USA) diluted to 1:200 in PBS at 4 °C in a humidified chamber followed by incubation with biotinylated anti-rabbit IgG at 37 °C for 30 min. After wash with PBS, the sections were developed in 0.05 % freshly prepared diamino-benzidine solution and counterstained with 0.1 % hematoxylin. Intracytoplasmic and/or intranuclear buffy granular staining was regarded as positive. Ten visual fields were randomly selected from each section. Omission of primary antibodies acted as a negative control. Staining intensity was evaluated by Image-Pro Plus software version 6.0 and expressed as the mean integral optical density (IOD). Staining intensity not above background level, slightly above background level, and significantly above background level were considered negative, weak, and strong expression, respectively. A mean IOD that was 50 % higher than the mean IOD of normal bone tissue was considered high expression, and 50 % lower than the mean IOD of normal bone tissue was considered low expression. Low and high expressions were considered positive.
Statistical analysis
Data were expressed as mean ± SD and analyzed using the SPSS software version 17.0 (SPSS Inc., Chicago, IL). Overall survival (OS) was calculated from the day of diagnosis to the last day of follow-up or death. Kaplan–Meier method was utilized to calculate survival. A correlation between positive and negative and low and high PITX1 expression and patient demographic and clinicopathologic variables were analyzed by Fisher’s exact test, and comparisons for multiple groups were made using the Student–Newman–Keuls multiple range test. Individual pair-wise comparisons were made using the unpaired two-tailed Student’s test. Significance was defined at <0.05.
Results
Patient demographic and baseline characteristics
Thirty-five patients were eligible for the study, including 18 men and 17 women. Their median age was 20 years (range, 10~50 years). Patient demographic and baseline characteristics are shown in Table 1. Follow-up data was available for 35 (100 %) patients, and the patients were followed up for median duration of 9 months (range, 1 to 36 months). Eight (22.9 %) patients developed metastases, and most (32/35, 91.4 %) patients died during the follow-up period.
PITX1 expression was downregulated in human osteosarcoma tissues
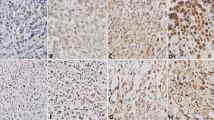
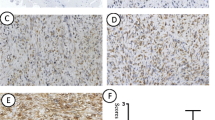
We determined the expression of PITX1 in the osteosarcoma tissues and normal bone tissues by immunohistochemistry. Representative images of PITX1 expression in normal bone tissue and osteosarcoma tissue samples are shown in Fig. 1a–d. Immunohistochemistry for PITX1 showed strong and constant membranous and cytoplasmic staining in positive cases (Fig. 1a–d). The mean IOD of PITX1 in normal bone tissues was 0.15 ± 0.01 and significantly higher than that of osteosarcoma tissues (0.05 ± 0.01) (P < 0.001) (Fig. 1e). Furthermore, PITX1 was expressed in all normal tissues (6/6, 100 %) and in 85.7 % (30/35) of tumor tissues (P < 0.05) (Table 2). In addition, all normal tissue specimens showed high PITX1 expression (6/6, 100 %) while only 23.3 % (7/30) osteosarcoma tissue specimens had high PITX1 expression (P < 0.05). These findings indicate that PITX1 expression was markedly downregulated in human osteosarcoma tissues compared with normal bone tissues.
PITX1 expression is downregulated in human osteosarcoma tissues. PITX1 expression in normal bone tissue (a) and osteosarcoma tissue (b–d) samples was examined by immunohistochemistry. a Strong positive PITX1 expression, b negative PITX1 expression, c weak positive PITX1 expression, d strong positive PITX1 expression; ×400. e The mean intensity of PITX1 expression in osteosarcoma tissue sample is significantly lower than that of normal bone tissue. ***P < 0.001 vs. normal tissue
Correlation of PITX1 expression and patient clinicopathologic variables
We then investigated whether PITX1 expression correlated with patient demographic and clinicopathologic parameters. PITX1 expression did not differ significantly by age, gender, or location (P > 0.05). The patients were categorized into four groups by survival, those whose survival was <6 months (median 3 months) (n = 13), between 6 and 12 months (median 8.6 months) (n = 12), between 12.1 and 24 months (median 16.7 months) (n = 7), and between 24.1 and 36 months (median 35.3 months) (n = 3). We found that patients with median OS >12 months had significantly higher PITX1 levels compared with those whose median OS was less than or equal to 12 months (P < 0.05 or 0.001) (Table 3). Furthermore, to investigate whether the lack of expression of PITX1 in osteosarcoma is associated with metastasis of lung cancer, we compared expression of PITX1 in osteosarcoma tissues between patients with lung metastasis (n = 8) and without lung metastasis (n = 27). We found that patients with lung metastasis had significantly lower PITX1 levels than patients without lung metastasis (**P < 0.01) (Fig. 2). Patients with low PITX1 expression exhibited four times higher risk of developing lung metastasis compared with those with high PITX1 expression (HR = 4.08, 95 % CI 1.50–11.0; P = 0.006). The Kaplan–Meier (KM) method is used to analyze ‘time-to-event’ data, which is shown in Fig. 3a (median, 9 months; range, 1 to 36 months). Furthermore, patients with high PITX1 expression had significantly longer OS (median, 20 months; range, 10 to 36 months) than patients with low PITX1 expression (median, 6.5 months; range, 1 to 22 months) (P < 0.05) (Fig. 3b). Patients with high PITX1 expression were more likely to survive longer (~16.6 months) and have reduced (~18.5 %) death risk compared with patients with low PITX1 expression (HR = 0.185, 95 % CI 0.060–0.567; P = 0.003) (Table 4).
Patients with lung metastasis have significantly lower PITX1 levels than patients without lung metastasis. The mean intensity of PITX1 expression in osteosarcoma tissues of patients with lung metastasis (Yes, n = 8) is significantly lower than that without lung metastasis (No, n = 27). **P < 0.01 vs. Yes
Patients with high PITX1 expression have longer survival. a The Kaplan-Meier survival curve of all osteosarcoma patients (n = 35). b The Kaplan-Meier survival curve of osteosarcoma patients stratified by low and high PITX1 expressions. Patients with high PITX1 expression are more likely to survive longer (~16.6 months) and have reduced (~18.5 %) death risk compared with patients with low PITX1 expression (HR = 0.185, 95 % CI 0.060–0.567; P = 0.003)
Discussion
Osteosarcoma is a rare but highly malignant tumor that originates from bone mesenchymal cells [8]. The 5-year survival rate of osteosarcoma patients has risen from about 20 % before chemotherapy was introduced to 65–70 % currently [9]. However, the previous two decades have seen no further improvements in terms of survival of almost one quarter of patients with clinically detectable metastatic disease at the time of initial diagnosis, hovering around 20–30 % [10]. Approximately half of the bone tumor patients eventually succumb to the disease.
Despite progress in the treatment of osteosarcoma, the prognosis of osteosarcoma patients remains poor, which inspires us to look for new therapeutic strategy. Recently, some translational studies in osteosarcoma have identified new molecular targets [11, 12]. These promote us to focus on addressing more novel targets that may be used in future clinical trials such as finding a reliable biomarker that can be used for targeting strategies or early diagnosis, to perform molecular profiling in order to identify recurrent genetic changes, or even to answer the basic question which cell type is the cause of osteosarcoma [13, 14]. The identification of novel biomarkers is important for better risk mitigation and treatment stratification of osteosarcoma patients. However, molecular analysis of surgical specimens is hampered by the fact that most resected specimens have experienced neoadjuvant chemotherapy and are thus not treatment naïve. The current study investigated archival surgical specimens from a rare cohort of treatment-naïve osteosarcoma patients and demonstrated that PITX1 expression was downregulated in osteosarcoma tissues and low PITX1 levels were associated with a poor survival outcome of osteosarcoma patients.
Numerous studies have shown that PITX1 is critical to vertebrate hindlimb development and an important morphological determinant of muscle, tendon, and bones of the hindlimb [15]. On the other hand, PTX1 is a tumor-suppressor gene in multiple human cancer types. In Barrett’s esophagus, PITX1 expression is downregulated, which is significantly accentuated during progression to Barrett’s associated adenocarcinoma [16]. Suppression of PITX1 expression is also observed in lung, gastric, and colon cancer cells as well as tumor tissues of the prostate and bladder [17–21]. In addition, Qi et al. demonstrated that PITX1 is a negative regulator of telomerase by repressing the TERT promoter transcriptionally [22]. These findings provide evidence that dysfunctions of the PITX1 gene play an important role in the development of various types of human cancers.
Despite a complete search that included all publications on PITX1, there is no clear link between the expression patterns and functions of PITX1 and osteosarcoma. To investigate the effect of PITX1 in cases of human osteosarcoma, we studied the expression of PITX1 in lower limb normal bone tissues and cancer tissue samples from 35 osteosarcoma patients by immunohistochemistry. Consistently, our results provide the first piece of evidence PITX1 expression was dysregulated in osteosarcoma. More importantly, PITX1 expression was significantly associated with the OS and lung metastasis of osteosarcoma patients. Since homeobox genes are known to regulate key cellular processes and be associated with differentiation and carcinogenesis, we speculate that downregulation of PITX1 may be a frequent molecular event in carcinogenesis of the bone. PITX1 may have a tumor suppressive function in osteosarcoma development and progression and serve as a potential biomarker for diagnosis and a therapeutic target for osteosarcoma.
We found no objective data linking expression of PITX1 with carcinogenesis of osteosarcoma. The relatively small number of patient samples included in this analysis may have contributed to lack of statistical significance for clinicopathologic variables except survival and lung metastasis. However, to the best of our knowledge, this is the first study to reveal that PITX1 expression was downregulated in osteosarcoma and inversely correlated with lung metastasis and OS. The mechanism(s) of modulating PITX1 expression in human osteosarcoma is still unknown. Further studies will be required to elucidate transcriptional regulation and biological functions of PITX1 in human osteosarcoma. The further identification of the cell type of osteosarcoma might help to clarify the molecular mechanisms of PITX1 expression. The predictive significance of PITX1 as a novel biomarker needs to be further examined in future studies involving a larger patient population. These will provide a basis for personalized medicine and molecular diagnosis using PITX1 as a biomarker.
In summary, the present study demonstrates that PITX1 expression is downregulated in osteosarcoma and correlates with patient survival and lung metastasis. PITX1 can be further explored as a new diagnostic biomarker and therapeutic target in osteosarcoma.
References
Arndt CA, Rose PS, Folpe AL, Laack NN. Common musculoskeletal tumors of childhood and adolescence. Mayo Clin Proc. 2012;87(5):475–87.
Linabery AM, Ross JA. Trends in childhood cancer incidence in the U.S. (1992–2004). Cancer. 2008;15(2):416–32.
Bielack SS, Kempf-Bielack B, Delling G, Exner GU, Flege S, Helmke K, et al. Prognostic factors in high-grade osteosarcoma of the extremities or trunk: an analysis of 1,702 patients treated on neoadjuvant cooperative osteosarcoma study group protocols. J Clin Oncol Off J Am Soc Clin Oncol. 2002;20(3):776–90.
Lamonerie T, Tremblay JJ, Lanctot C, Therrien M, Gauthier Y, Drouin J. Ptx1, a bicoid-related homeo box transcription factor involved in transcription of the pro-opiomelanocortin gene. Genes Dev. 1996;10(10):1284–95.
Infante CR, Park S, Mihala AG, Kingsley DM, Menke DB. Pitx1 broadly associates with limb enhancers and is enriched on hindlimb cis-regulatory elements. Dev Biol. 2013;374(1):234–44.
Yasui W, Yokozaki H, Fujimoto J, Naka K, Kuniyasu H, Tahara E. Genetic and epigenetic alterations in multistep carcinogenesis of the stomach. J Gastroenterol. 2000;35(12):111–5.
Ushijima T, Sasako M. Focus on gastric cancer. Cancer Cell. 2004;5(2):121–5.
Campanacci M, Baldini N, Boriani S, Sudanese A. Giant-cell tumor of bone. J Bone Joint Surg Am. 1987;69(1):106–14.
Allison DC, Carney SC, Ahlmann ER, Hendifar A, Chawla S, Fedenko A, et al. A meta-analysis of osteosarcoma outcomes in the modern medical era. Sarcoma. 2012;2012:704872.
Miller BJ, Cram P, Lynch CF, Buckwalter JA. Risk factors for metastatic disease at presentation with osteosarcoma: an analysis of the SEER database. J Bone Joint Surg Am. 2013;95(13):e89.
Schwab JH, Springfield DS, Raskin KA, Mankin HJ, Hornicek FJ. What’s new in primary bone tumors. J Bone Joint Surg Am. 2012;94(20):1913–9.
Yang J, Zhang W. New molecular insights into osteosarcoma targeted therapy. Curr Opin Oncol. 2013;25(4):398–406.
Mohseny AB, Szuhai K, Romeo S, Buddingh EP, Briaire-de Bruijn I, de Jong D, et al. Osteosarcoma originates from mesenchymal stem cells in consequence of aneuploidization and genomic loss of Cdkn2. J Pathol. 2009;219(3):294–305.
Ng AJ, Mutsaers AJ, Baker EK, Walkley CR. Genetically engineered mouse models and human osteosarcoma. Clin Sarcoma Res. 2012;2(1):19.
DeLaurier A, Schweitzer R, Logan M. Pitx1 determines the morphology of muscle, tendon, and bones of the hindlimb. Dev Biol. 2006;299(1):22–34.
Lord RV, Brabender J, Wickramasinghe K, DeMeester SR, Holscher A, Schneider PM, et al. Increased CDX2 and decreased PITX1 homeobox gene expression in Barrett’s esophagus and Barrett’s-associated adenocarcinoma. Surgery. 2005;138(5):924–31.
Ramaswamy S, Tamayo P, Rifkin R, Mukherjee S, Yeang CH, Angelo M, et al. Multiclass cancer diagnosis using tumor gene expression signatures. Proc Natl Acad Sci U S A. 2001;98(26):15149–54.
Singh D, Febbo PG, Ross K, Jackson DG, Manola J, Ladd C, et al. Gene expression correlates of clinical prostate cancer behavior. Cancer Cell. 2002;1(2):203–9.
Kolfschoten IG, van Leeuwen B, Berns K, Mullenders J, Beijersbergen RL, Bernards R, et al. A genetic screen identifies PITX1 as a suppressor of RAS activity and tumorigenicity. Cell. 2005;121(6):849–58.
Chen Y, Knosel T, Ye F, Pacyna-Gengelbach M, Deutschmann N, Petersen I. Decreased PITX1 homeobox gene expression in human lung cancer. Lung Cancer. 2007;55(3):287–94.
Chen YN, Chen H, Xu Y, Zhang X, Luo Y. Expression of pituitary homeobox 1 gene in human gastric carcinogenesis and its clinicopathological significance. World J Gastroenterol: WJG. 2008;14(2):292–7.
Qi DL, Ohhira T, Fujisaki C, Inoue T, Ohta T, Osaki M, et al. Identification of PITX1 as a TERT suppressor gene located on human chromosome 5. Mol Cell Biol. 2011;31(8):1624–36.
Acknowledgments
None.
Declaration
This work has not been published or presented, wholly or in part, in any other publication or at any conference or meeting.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Additional information
Gengbin Kong, Zhaoyong Liu and Kezhou Wu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Kong, G., Liu, Z., Wu, K. et al. Strong expression of paired-like homeodomain transcription factor 1 (PITX1) is associated with a favorable outcome in human osteosarcoma. Tumor Biol. 36, 7735–7741 (2015). https://doi.org/10.1007/s13277-015-3512-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-3512-1