Abstract
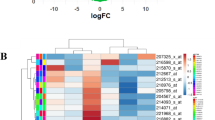
Acquired erlotinib resistance (AER) during cancer treatment remains a major clinical challenge that results in the recurrence and metastasis of cancers. Therefore, we sought to identify differentially expressed genes (DEGs) by performing a meta-analysis of AER-related microarray datasets and discover biomarkers by conducting a systemic in-silico analysis. Using the RankProd algorithm, we identified 775 DEGs (536 up-regulated and 239 down-regulated). Functional enrichment analyses of the total DEG s suggested that “cell adhesion” and “cytokine-cytokine receptor interactions” may be closely associated with AER process. Some DEGs shared target sites of the potential micro-RNA including miR-21, miR-200b/c, miR-429 and miR-9. Target sites of FOXJ1, NFAT, FOXO4, and JUN were also significantly enriched. From the proteinprotein interaction network, we clustered four functional modules by p-value and node density and found hub genes with many interacting neighbors. Finally, we identified seven candidate hub DEGs (TIMP3, SPARC, ITGA1, CCNA1, SOX2, KRT14, and PTPRZ1) for AER development.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Szakacs, G., Paterson, J. K., Ludwig, J. A., Booth-Genthe, C. & Gottesman, M. M. Targeting multidrug resistance in cancer. Nat Rev Drug Discov 5:219–234 (2006).
Chen, K. G. & Sikic, B. I. Molecular pathways: regulation and therapeutic implications of multidrug resistance. Clin Cancer Res 18:1863–1869 (2012).
Bareschino, M. A. et al. Erlotinib in cancer treatment. Ann Oncol 18 Suppl 6, vi35–41 (2007).
Xu, Y., Liu, H., Chen, J. & Zhou, Q. Acquired resistance of lung adenocarcinoma to EGFR-tyrosine kinase inhibitors gefitinib and erlotinib. Cancer Biol Ther 9:572–582 (2010).
Hrustanovic, G., Lee, B. J. & Bivona, T. G. Mechanisms of resistance to EGFR targeted therapies. Cancer Biol Ther 14:304–314 (2013).
Beloueche-Babari, M. et al. Acquired resistance to EGFR tyrosine kinase inhibitors alters the metabolism of human head and neck squamous carcinoma cells and xenograft tumours. Br J Cancer 112 Suppl, 1206–1214 (2015).
Nguyen, K. S., Kobayashi, S. & Costa, D. B. Acquired resistance to epidermal growth factor receptor tyrosine kinase inhibitors in non-small-cell lung cancers dependent on the epidermal growth factor receptor pathway. Clin Lung Cancer 10:281–289 (2009).
Nurwidya, F. et al. Acquired resistance of non-small cell lung cancer to epidermal growth factor receptor tyrosine kinase inhibitors. Respir Investig 52:82–91 (2014).
Lin, Y., Wang, X. & Jin, H. EGFR-TKI resistance in NSCLC patients: mechanisms and strategies. Am J Cancer Res 4:411–435 (2014).
Ramasamy, A., Mondry, A., Holmes, C. C. & Altman, D. G. Key issues in conducting a meta-analysis of gene expression microarray datasets. PLoS Med 5:e184 (2008).
Zhao, Y. et al. Identification of potential therapeutic target genes, key miRNAs and mechanisms in acute myeloid leukemia based on bioinformatics analysis. Med Oncol 32:572 (2015).
Giles, K. M. et al. Axl mediates acquired resistance of head and neck cancer cells to the epidermal growth factor receptor inhibitor erlotinib. Mol Cancer Ther 12:2541–2558 (2013).
Zhang, Z. et al. Activation of the AXL kinase causes resistance to EGFR-targeted therapy in lung cancer. Nat Genet 44:852–860 (2012).
Ahmad, A. et al. Inhibition of Hedgehog signaling sensitizes NSCLC cells to standard therapies through modulation of EMT-regulating miRNAs. J Hematol Oncol 6:77 (2013).
Cufi, S. et al. Silibinin suppresses EMT-driven erlotinib resistance by reversing the high miR-21/low miR-200c signature in vivo. Sci Rep 3:2459 (2013).
Bryant, J. L. et al. A microRNA gene expression signature predicts response to erlotinib in epithelial cancer cell lines and targets EMT. Br J Cancer 106:148–156 (2012).
Takeuchi, K., Shin-ya, T., Nishio, K. & Ito, F. Mitogenactivated protein kinase phosphatase-1 modulated JNK activation is critical for apoptosis induced by inhibitor of epidermal growth factor receptor-tyrosine kinase. FEBS J 276:1255–1265 (2009).
Boudjadi, S., Carrier, J. C. & Beaulieu, J. F. Integrin alpha1 subunit is up-regulated in colorectal cancer. Biomark Res 1:16 (2013).
Kanda, R. et al. Erlotinib resistance in lung cancer cells mediated by integrin beta1/Src/Akt-driven bypass signaling. Cancer Res 73:6243–6253 (2013).
Kaleagasioglu, F. & Berger, M. R. SIBLINGs and SPARC families: their emerging roles in pancreatic cancer. World J Gastroenterol 20:14747–14759 (2014).
Anania, M. C. et al. TIMP3 regulates migration, invasion and in vivo tumorigenicity of thyroid tumor cells. Oncogene 30:3011–3023 (2011).
Kitkumthorn, N. et al. Cyclin A1 promoter hypermethylation in human papillomavirus-associated cervical cancer. BMC Cancer 6:55 (2006).
Rivera, A., Mavila, A., Bayless, K. J., Davis, G. E. & Maxwell, S. A. Cyclin A1 is a p53-induced gene that mediates apoptosis, G2/M arrest, and mitotic catastrophe in renal, ovarian, and lung carcinoma cells. Cell Mol Life Sci 63:1425–1439 (2006).
Otsubo, T., Akiyama, Y., Yanagihara, K. & Yuasa, Y. SOX2 is frequently downregulated in gastric cancers and inhibits cell growth through cell-cycle arrest and apoptosis. Br J Cancer 98:824–831 (2008).
Karantza, V. Keratins in health and cancer: more than mere epithelial cell markers. Oncogene 30:127–138 (2011).
Xu, Y. & Fisher, G. J. Receptor type protein tyrosine phosphatases (RPTPs) -roles in signal transduction and human disease. J Cell Commun Signal 6:125–138 (2012).
Xia, J. et al. INMEX-a web-based tool for integrative meta-analysis of expression data. Nucleic Acids Res 41:W63–70 (2013).
Song, G. G. et al. Meta-analysis of differentially expressed genes in primary Sjogren’s syndrome by using microarray. Hum Immunol 75:98–104 (2014).
Fang, F., Pan, J., Xu, L., Li, G. & Wang, J. Identification of potential transcriptomic markers in developing ankylosing spondylitis: a meta-analysis of gene expression profiles. Biomed Res Int 2015:826316 (2015).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27:1739–1740 (2011).
Quan, Z. et al. Protein-protein interaction network and mechanism analysis in ischemic stroke. Mol Med Rep 11:29–36 (2015).
Rakshit, H., Rathi, N. & Roy, D. Construction and analysis of the protein-protein interaction networks based on gene expression profiles of Parkinson’s disease. PLoS One 9:e103047 (2014).
Nepusz, T., Yu, H. & Paccanaro, A. Detecting overlapping protein complexes in protein-protein interaction networks. Nat Methods 9:471–472 (2012).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Lee, Y.S., Kim, J.K., Park, T.H. et al. Systematic identification of novel biomarker signatures associated with acquired erlotinib resistance in cancer cells. Mol. Cell. Toxicol. 12, 139–148 (2016). https://doi.org/10.1007/s13273-016-0018-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13273-016-0018-x