Abstract
The ability of different Lactobacillus strains to produce conjugated linoleic acid (CLA) from linoleic acid was evaluated. Preliminary experiments revealed that L. plantarum among the screened strains had the highest CLA-producing potential (95.25 μg/mL). The cell growth of this bacterium was studied in three media of MRS broth, skim milk and skim milk supplemented with yeast extract and glucose. Results showed that the use of yeast extract and glucose could significantly increase the cell growth and CLA production. Response surface methodology (RSM) was applied to investigate the effects of three independent variables; linoleic acid (LA), yeast extract concentrations and inoculum size on the CLA formation. A second-order polynomial model with high R 2 value (0.981) was developed using multiple linear regression analysis. The optimum conditions to achieve the highest CLA production (240.69 μg/mL) was obtained using 3 mg/mL LA, 4 g/L yeast extract and inoculum size of 4 % v/v. CLA concentration of the optimal sample was analyzed by Gas Chromatography (GC). The cis-9, trans-11 CLA was the major CLA isomer of total CLA detected.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fatty acids or carboxylic acids with long aliphatic chains are formed by the triglycerides hydrolysis (Scrimgeour 2005). Linoleic acid (LA; cis, cis, 9, 12-octadecadienoic acid) is one of the unsaturated fatty acid (UFA) which can transfer to its conjugated form namely conjugated linoleic acid (CLA). CLA also is a generic name for a mixture of LA isomers in which the double bonds are conjugated (Lin et al. 2003). These double bonds can be at positions 8 and 10, 9 and 11, and 10 and 12 or 11 and 13. Each of these positional conjugated isomers can occur in cis-trans, trans-cis and cis-cis or trans-trans geometrical configurations (Li et al. 2013). It was reported that these isomers have many health benefits such as reduced risk of carcinogenesis, atherosclerosis and obesity, improved hyperinsulinemia and prevention of catabolic effects of the immune system (Yadav et al. 2007).
Meat, poultry, seafood, cheese, butter, milk and vegetable oils are the principal sources of CLA in the human diet. For example, the CLA concentration in cow’s milk was variable and ranged from 2.4 to 21.8 mg of CLA/g of fat. In the dairy products, the cis-9, trans-11 CLA isomer with 90 % of the conjugated dienes was also shown to be the major CLA isomer with the most biologically active (Ye et al. 2013).
In contrast to the natural conversion, other methods including lipase-catalyzed (enzymatic) synthesis and alkali isomerization from LA using the various solvents and conditions appear to be complex mixtures of isomers. Therefore, microbial synthesis of CLA may be an attractive and promising method to produce it in an industrial-scale (Das et al. 2005). CLA is produced as an intermediate during the bio-hydrogenation of LA by many microorganisms. Butyrivibrio fibrisolvens is one of the most common bacteria able to synthesize this functional component (Van Nieuwenhove et al. 2007). Some strains of lactic acid bacteria (LAB) such as Bifidobacterium, Lactobacillus, and Streptococcus and some dairy starter cultures such as Propionobacterium were able to convert LA to CLA. Among these strains, Lactobacillus species especially L. plantarum, L. casei, L. acidophilus, L. rhamnosus, L. salivarius, L. paracasei subsp. paracasei, L. brevis, L. pentosus, and L. bulgaricus were found to be the most suitable for increasing the CLA content during fermentation processes (Ogawa et al. 2005).
Alonso et al. (2003) noted that the addition of 0.02 % LA to the medium for a 24 h incubation led to an effective CLA production by L. acidophilus L1. Kishino et al. (2002) used the washed cell of L. plantarum AKU1009a after its screening from 14 genera of LAB to achieve the highest CLA amount (20 mg/mL) during fermentation process. Lin et al. (2003) found that the use of mixed culture of L. acidophilus CCRC 10479 and yoghurt starter can significantly increase the cis-9, trans-11 C18:2 amounts in non-fat yoghurt supplemented with 0.1 % LA and 5 % fructooligosaccharides. Lin (2000) earlier showed that inoculation of L. acidophilus into skim milk (SM) medium with 60 g/L sweeteners or 10 g/L sodium chloride was effective to promote the CLA formation. Kim and Liu (2002) also demonstrated that CLA content of milk supplemented with sunflower oil (SO) was increased by the lactic fermentation of L. lactis subsp. lactis I-01.
Response surface methodology (RSM) is a collection of mathematical and empirical techniques useful for establishing models, and for optimizing processes even in the presence of complex interactions. Central composite rotatable design (CCRD) is one of the successful factorial designs for the parameters optimization with a limited number of experimental trials, development time and overall cost, and estimates the response surface (Gharibzahedi et al. 2013). Although many studies concerning the fermentation of CLA have been performed, but, to the best of our knowledge, there is few investigations on the optimization of CLA production from Lactobacillus species based on an inexpensive carbon source such as skim milk. Therefore, the objective of this research was to optimize the effective parameters on CLA production by the best screened lactic strain using RSM-CCRD.
Materials and methods
Chemicals
Hexane, chloroform, methanol, Tween 80, urea, yeast extract and D-glucose were purchased from Merck Chemical Co (Darmstadt, Germany). CLA standard was purchased from sigma chemical Co (St. Louis, MO, USA). Ethanol (96 %) was provided by Hamoon Teb (Mashhad, Iran). All the other chemicals used in this study were in analytical grade.
Microorganism and culture medium
All the used LAB strains including L. rhamnosus (DSM 20021), L. acidophilus (DSM 20179), L. casei (DSM 20011), L. plantarum (DSM 20179), L. paracasei subsp. Paracasei (DSM 20207), L. delbrueckii (DSM 20081), L. helveticus (DSM 20075), and L. reuteri (DSM 20016) were provided by Bioprocess Engineering Laboratory (BPEL, University of Tehran, Iran). de Man, Rogosa and Sharpe (MRS) broth was also provided from Merck Chemical Co (Darmstadt, Germany).
Extraction of free fatty acids
Extraction of free fatty acids (FFAs) was carried out according to the applied method by Wu et al. (2008). Briefly, FFAs were formed from sunflower oil (SO) by saponifying with 10 % KOH aqueous ethanol, acidifying to pH of 3.0 and then treating with hexane. The obtained FFAs were mixed with urea in 95 % aqueous ethanol and heated at 60 °C. The urea-fatty acid adduct was crystallized at 0 °C for 3 h and the resulted crystals separated from the liquid by fast filtration. The liquid was diluted with water and acidified to pH = 2–3. An equal volume of hexane was subsequently added and the FFAs were extracted at 65 °C using a rotary evaporator (Heidoplh, model Laborota4000, Germany). FFA from the urea complexing fraction was recovered after adding water, 6 N HCl and hexane in a similar manner.
Screening of Lactobacilli cultures for CLA production
All the lactobacillus strains were activated in MRS broth for 24 h at 37 °C. These strains were cultivated in MRS medium containing 0.06 % LA for 24 h at 28 °C with a shaking rate of 120 rpm. The cells were harvested by centrifugation (8000×g, 10 min), washed twice with 0.85 % NaCl, centrifuged again, and then used as the washed cells for the reactions. The cultivation was carried out in test tubes with 10 mL of MRS containing 1 mg/mL of LA, 1 mg/mL of Tween 80 and 5 % (v/v) of the washed cells for 24 h at 37 °C and 120 rpm. Only the strain showing the highest ability to produce CLA was selected for the next steps.
Manipulation of fermentation medium
A modification was conducted to increase the bacteria cells and CLA amounts. For this reason, the growth of selected strain was studied in skim milk (SM) (10 % w/v), SM supplemented with 5 g/L yeast extract and 7 g/L D-glucose and MRS broth for 48 h at 37 °C and 120 rpm. The growth of selected strain was determined by measuring values of optical density at 600 nm (Li et al. 2013).
Fermentation process
The fermentation process was done according to three following steps: in the first step, the selected strain was activated in MRS broth at 37 °C for 24 h. In the second one, 5 % (v/v) of the prepared inoculum was pre cultured in 10 % (w/v) skim milk at the same conditions (37 °C for 24 h) with a shaking rate of 120 rpm and finally this transferred culture was inoculated in 20 mL culture medium shacked at 120 rpm and 37 °C for 48 h. The reaction mixture contained 20 mL SM in Erlenmeyer flask with 1 mg/mL Tween 80, 7 g/L D-glucose and different amounts of LA, yeast extract and cell concentration. The amounts of these variables are given in Table 1. For the investigation of effects of LA, yeast extract and cell concentration on CLA production, the reactions were essentially carried out under the same conditions as described above with varying target parameters. These compounds were filter-sterilized through 0.45 μm syringe filter in order to ensure sterility of LA and Tween 80.
Lipid extraction
The lipids were extracted according to the method of Bligh and Dyer as previously described by Wang et al. (2007). Briefly, the bacterial suspensions (10 mL) were mixed with 20 mL of chloroform: methanol (1:2, v/v), followed by vigorously shaking for 30 s, and centrifuged at 4500 rpm for 5 min at 4 °C. The bottom phase to obtain the extracted lipids was recovered and dehydrated with anhydrous Na2SO4.
LA quantification
The LA amounts were determined based on the method described by Rosson and Grunde (2001). In brief, the organic phase was scanned from 200 to 400 nm in 1 cm quartz cuvettes using an UV-visible spectrophotometer (CECIL-CE 1020, England) at 233 nm. Standard curve was constructed for the absorbance at 233 nm versus the different concentrations of CLA (0–12 μg/mL).
Identification and quantification of CLA isomers
The method of Chin et al. (1994) was applied to methylate the samples with BF3-methanol. The FFA (up to 10 mg) reacted with BF3-methanol (1 mL; 14 %) for 30 min at ambient temperature. The methylated sample was mixed with 2 mL hexane: water (1:1 v/v). The upper phase dried over anhydrous sodium sulfate, and then the solvent evaporated without heating in a stream of nitrogen for further quantification of CLA isomers by gas chromatography (GC) system.
The solvent was analyzed for the CLA isomers using a GC system (Agilent 7890, Agilent Technologies, Santa Clara, CA, USA) equipped with an FID detector and fused silica capillary column HP-88 (100 m, 0.32 mm, 0.25 μm). The temperature of the GC oven was programmed at 110 °C for 2 min and from 110 to 230 °C at the rate of 10 °C/min and held for 5 min. The flow rate of make-up gas was 1.5 L/min. The split ratio was set at 1:50. The amount of CLA isomer in each sample was expressed as μg/mL.
Experimental method and statistical analysis
Twenty treatments as determined by central composite rotatable design (CCRD, Table 1) were conducted to study the effects of three parameters viz yeast extract (1.95–12.05 g/L w/w, X 1), LA (0.99–3.51 mg/mL, X 2), and inoculum size (2.98–8.02 v/v, X 3) on the concentration of CLA (Y 1). The Design Expert (Trial Version 7.1.3, Stat-Ease Inc., Minneapolis, MN, USA) was applied for regression and graphical analyzes of the obtained data. Experiments were randomized in order to minimize the effects of unexplained variability in the observed responses due to extraneous factors. Our preliminary studies demonstrated that the addition of the concentration levels investigated to the culture medium resulted in desirable amounts of LA by the selected strain. For statistical calculations, the relation between the coded values and actual values are described by Eq. (1):
where x i , the coded value of the variable, X i , the actual value of the variable, X 0, the actual value of X i at the center point, and ΔX, the step change value of the variables.
The generalized regression second-order polynomial model proposed for predicting the response variables is given as (Eq. 2):
where Y is the predicted response; β k0, β ki , β kii and β kij represent regression coefficients; and X i , X j are the coded independent factors.
The quality of the fit of polynomial model was expressed by the coefficient of determination R 2, adjusted R 2 (R 2adj ) in Eqs. (3)–(4), respectively (Gharibzahedi et al. 2012):
Adequate precision (AP) compares the range of the predicted values at the design points to the average prediction error. The AP definition is in Eqs. (5) and (6):
In Eqs. (3)–(6), SS is the sum of squares, DF is the degrees of freedom, \( \overline{y} \) is the predicted value, p is the number of model parameters, σ 2 is the residual mean square from analysis of variance (ANOVA) table, and n is the number of experiments.
Results and discussion
Selection of the best LAB strain
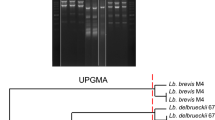
The ability of used LAB strains including L. rhamnosus, L. acidophilus, L. casei, L. plantarum, L. paracasei subsp. paracasei, L. delbrueckii, L. helveticus and L. reuteri to produce CLA were assayed. All the strains of lactobacilli tested, except L. helveticus produced CLA in MRS broth medium. L. plantarum showed the highest potential for CLA-producing (95.25 μg/mL) and was thus selected for the following experiments (Fig. 1). The amount of CLA produced by L. rhamnosus and L. casei were 79.95 and 78.40 μg/mL, respectively. However, there was a significant difference in total amount of CLA formed by these lactic strains and L. plantarum (P < 0.05).
Liu et al. (2011) screened 43 LAB strains with different CLA-producing abilities from the Chinese pickles in MRS broth medium and reported that L. plantarum strain lp15 had the highest ability to synthesize CLA (26.1 μg/mL). Yang et al. (2014) assessed the CLA production capability of some strains of food-derived lactobacilli from LA. They found that L. plantarum ZS2058 was the most efficient CLA producer in MRS broth with more than 50 % LA conversion to c9, t11-CLA and t9, t11-CLA as dominant isomers. These results appear to be in contrast of Jiang et al. (1998), who examined 19 strains of lactobacilli, lactococci, streptococci and propionibacteria to find their ability for producing CLA in MRS broth. They found that none of the tested LAB was able to produce CLA from LAs. Two strains of Propionibacterium freudenreichii ssp. freudenreichii and one strain of Pr. freudenreichii ssp. shermanii were found to be able to convert LA to CLA in high level (265 μg/mL) (Jiang et al. 1998). Numbers of LAB were sensitive to unsaturated fatty acids such as LA, oleic acid, linolenic acid and arachidonic acid, etc. (Li et al. 2013). The ability of CLA production by some strains of lactobacilli, in particular L. plantarum might be due to the FFA detoxification mechanism in these strains (Yang et al. 2014).
Selection of the most suitable culture medium
A modification in culture medium was studied to increase the growth of bacterial cells and CLA production. The measurement of L. plantarum biomass and its microbial growth profile using an UV-visible spectrophotometer (at 600 nm) were assessed in three media including MRS broth, skim milk (SM) and SM supplemented with 5 g/L yeast extract and 7 g/L glucose (Fig. 2). As can be seen in this figure, the growth level of L. plantarum in SM supplemented with yeast extract and glucose surprisingly was more than two other media. This fact shows that the biomass and CLA productions were highly affected by yeast extract and glucose.
Yeast extract as the most commonly used nitrogen source provides convenient growth factors for microbial growth by its high buffering capacity because it is a nutritional complex medium containing amino acids, peptides, nucleotides, some carbohydrates, trace elements and group-B vitamins (Gharibzahedi et al. 2012). Furthermore, enhancement of bacterial growth and CLA production in the glucose presence can be mainly due to the activation of bacterial cell growth by supplementing an easily used energy source (Kim and Liu 2002).
The results also showed that SM was better than MRS for the CLA synthesis. This fact can be probably due to the neutralization of the inhibitory effect of fatty acids by milk proteins such as α-lactalbumin and β-lactoglobulin and the oxidation prevention in SM by the binding to CLA (Shantha and Decker 1993). These results were in agreement with findings of Kim and Liu (2002), who studied the ability of LAB strains to produce CLA in SM. They noted that CLA formation can enhance in dry milk powder due to the inhibition of CLA oxidation by free radical reactions (Kim and Liu 2002).
Model fitting
RSM showed that a second-order polynomial mathematical model can be an effective equation to fit the experimental data obtained from the microbial production of CLA (Eq. 7):
ANOVA revealed that the constructed model had a high probability (p < 0.0001) and non-significant lack of fit (p < 0.05). A high value of R 2 (0.9884) is an indication that the fitted model can be used for prediction with reasonable precision. The sample variation of 98.84 % for CLA synthesis might be linked to the independent variables and only 1.16 % of the total variation could not be explained by the model (Table 2). Adjusted R 2 (0.9768) is a modification of R 2 that adjusts for the number of explanatory terms in a model. Unlike R 2, the adjusted R 2 increases only if the new term improves the model more than would be expected by chance (Ghasemlou et al. 2012). The AP measures the signal-to-noise ratio, with a ratio greater than 4 being desirable (Gharibzahedi et al. 2012). The AP of 35.34 indicated that this model could be used to navigate the design space (Table 2). The comparison between the actual response values obtained from experimental data and the predicted response values based on the polynomial regression model also proves that the fitted model cover the experimental range of studies sufficiently (Table 1).
Optimization of CLA production
Table 2 shows that the linear and quadratic effects of LA content, yeast extract concentration and inoculum size were significant (P < 0.0001; P < 0.001; P < 0.01) on CLA produced by the selected L. plantarum. Also, all the mutual interaction between independent variables were found to be significant (Table 2). As shown in Fig. 3, the variation in the CLA yield could be explained as a nonlinear function of the LA and yeast extract contents and inoculum density. The most significant (p < 0.05) effect on CLA synthesis was shown to be the interaction effect of LA concentration and inoculum size followed by the linear effect of LA content and interaction effect of yeast extract concentration and inoculum density (Table 2).
The hydrolyzed sunflower oil (0.99–3.51 mg/mL) as an inducer was added to SM medium. The results showed that CLA production can significantly increase by adding 1 to 3 mg/mL of the substrate (Fig. 3a–b and e–f). The presence of LA isomerase and its activity in the culture can be a reliable reason for improving CLA production by adding LA to the medium. L. plantarum strain in the SM medium showed a relatively high tolerance to sunflower oil due to the detoxification mechanism in this strain. Ogawa et al. (2001) also pointed out that high production amount of CLA from LA by L. acidophilus was due to the enzyme system induced by LA. Puniya et al. (2008) similarly found that L. lactis and L. viridescens could tolerate the increased concentrations of LA to produce high amount of CLA. Lin et al. (1999) examined ability of five LAB strains to produce CLA with different levels of LA in SM medium. They found that the CLA yield after lactic fermentation was primarily dependent on the LA addition and reported the highest level of CLA could be obtained with adding 5 mg/mL LA. An increase in CLA content in probiotic dahi by lipolysis of natural milk fat during fermentation of L. acidophilus and L. casei was observed even without addition of external linoleic acid during storage (Yadav et al. 2007). Maybe, these lactic strains utilized LA synthesized during the lipolysis of milk fat as an ideal substrate for CLA synthesis in probiotic dahi.
It appeared that addition of more than 3 mg/mL LA exerted an inhibitory effect on the CLA production and cell growth (Fig. 3). The antimicrobial effect of free LA at high concentrations on propionic acid bacteria (PLB) was previously reported by Boyaval et al. (1995) and Jiang et al. (1998). In contrast, Li et al. (2013) by investigating the effect of LA concentration on CLA production by L. acidophilus F0221 found that the conversion rate gradually decreased by adding free LA from 16.80 % at 0.5 mg/mL LA to 6.15 % at 2.0 mg/mL LA. Nieman (1954) reported that FFAs generally disturbed the permeability of cytoplasmic membranes of LAB and hence reduced CLA production. It was noted that the CLA synthesis in the mixtures was significantly influenced by the cell and substrate concentrations. In this study, an increase in concentration of LA increased CLA production, and simultaneously cell viability decreased (Dubey et al. 2012). It might be due to the fact that the high level of LA concentration by changing the enzyme structure could reduce the contact areas between enzyme and substrate (Zhao et al. 2011).
The biohydrogenation of LA is a multiple-step process in which CLA is found as an intermediate product. In this process, unsaturated fatty acid is first converted into conjugated form and further hydrogenated into its monoenoic form depending on the linoleate isomerase (Kim et al. 2000). Additionally, it was also found that membrane fraction from the dead or lysed cells retained their ability to convert LA into CLA (Kim et al. 2000), and thus supports the results found and it might be the reason why the CLA was increased while the cell survivability declined (Dubey et al. 2012).
In general, the inoculum size affects cell growth and CLA production rate by delaying the lag-phase length during the bacterial growth profile. A significant increase in produced CLA amount could be obtained by applying inoculum size of 4 % for L. plantarum (Fig. 3c–f). Lower inoculum levels of 4 % might be insufficient for initiating the cell growth and CLA synthesis. Moreover, increasing the inoculum size more than 4 % because of nutrients depletion at the enhanced biomass led to a gradual decrease in CLA production (Fig. 3c–f). In agreement to the results obtained in this study, Ye et al. (2013) reported that the CLA production can significantly be improved with increasing the inoculum size from 2 to 5 %.
Further increase in the inoculum size led to a decrease in CLA production. Liong and shah (2005) noted that the growth of L. acidophilus can considerably increase by increasing inoculum size from 1.20 to 2.27 % (w/v), while further increase can decrease the production amount due to the substrate limitation with increased cells number (Liong and shah 2005). Yeast extract is one of the most important factors for microbial growth because it is a rich source of amino acids, short peptides, B-vitamin complexes, nitrogen, minerals and trace ingredients. In this study, it was added to the culture in the range of 2–12 g/L. There was a significant elevation in total amount of CLA formation with addition 2–4 g/L yeast extract, but its higher concentrations had no effect on the CLA production (Fig. 3a–d). The results were in accordance with those obtained by Li et al. (2012), who investigated the optimal conditions for CLA production by L. plantarum in the MRS medium and found that there was a significant increase in CLA production (0.112 g/L) using 4 g/L yeast extract. Hennessy et al. (2009) also investigated the effect of different levels of yeast extract on the bioconversion of LA to CLA by Bifidobacterium breve NCIMB 702258 in SM medium. They reported that yeast extract in a concentration range of 0–10 g/L affected an increase in CLA production, but, further increase in the amount of this component had no significant effect (Hennessy et al. 2009).
Optimization and validation procedure
An optimum set level of independent variables leading to the desirable response goal namely the highest CLA production by the selected L. plantarum was evaluated according to the multiple optimization procedure using the Design-Expert software. The optimization method proved that the overall optimum region with the highest overall desirability (D = 1.00) was found by setting the experiment at 4.01 g/L yeast extract, 3.00 mg/mL LA and inoculum density of 4.09 % v/v. Under the recommended optimum condition, maximum CLA yield was predictably found to be 240.979 μg/mL. The adequacy of response surface equation was checked by the comparison of experimental and fitted values predicted by the response regression model. The mean experimental value for four replicates to produce CLA under the optimal point was 242.32 ± 8.46 μg/L which was very close to the predicted value. Figure 4 shows a typical GC chromatogram of the extracted oil phase from L. plantarum culture. The result indicated the c9,t11-CLA was the major CLA isomer with more than 90 % of total CLA detected. GC analysis for the optimal sample also revealed that the CLA was the most abundant FFA in the culture. Therefore, there was no significant difference (P > 0.05) between experimental and predicted values, suggesting that the response surface equations had high capability to determine the medium formulation for maximizing CLA.
Partial GC chromatogram obtained from a oil phase extracted from L. plantarum culture b CLA isomer standards. For GC column and conditions see Materials and methods
Conclusion
In the present study, eight Lactobacillus strains were compared to produce CLA by adding LA as substrate. L. plantarum exhibited the highest level of CLA production. The results also showed that the growth level of this bacterium in SM supplemented with yeast extract and glucose was more than MRS broth and SM media. Thus, the effects of three independent variables including LA content, inoculum size and yeast extract concentration with RSM-CCRD were optimized in the selected medium to obtain the highest amount of CLA synthesis. The RSM analysis showed a significant quadratic regression equation with high R 2 and non-significant lack-of-fit. The most important variable in this study was LA concentration. Optimization analysis also showed that the maximum CLA production (240.69 μg/mL) can be obtained at 3.00 mg/mL LA, 4.01 g/L yeast extract and inoculum density of 4.09 % v/v.
References
Alonso L, Cuesta E, Gilliland S (2003) Production of free conjugated linoleic acid by Lactobacillus acidophilus and Lactobacillus casei of human intestinal origin. J Dairy Sci 86:1941–1946
Boyaval P, Corre C, Dupuis C, Roussel E (1995) Effects of free fatty acids on propionic acid bacteria. Lait 75:17–29
Chin SF, Storkson JM, Liu W, Albright KJ, Pariza MW (1994) Conjugated linoleic acid (9, 11-and 10, 12-octadecadienoic acid) is produced in conventional but not germ-free rats fed linoleic acid. J Nutr 124:694–701
Das S, Holland R, Crow V, Bennett R, Manderson G (2005) Effect of yeast and bacterial adjuncts on the CLA content and flavour of a washed-curd, dry-salted cheese. Int Dairy J 15:807–815
Dubey V, Ghosh AR, Mandal BK (2012) Appraisal of conjugated linoleic acid production by probiotic potential of Pediococcus spp. GS4. Appl Biochem Biotechnol 168:1265–1276
Gharibzahedi SMT, Razavi SH, Mousavi SM, Moayedi V (2012) High efficiency canthaxanthin production by a novel mutant isolated from Dietzia natronolimnaea HS-1 using central composite design analysis. Ind Crop Prod 40:345–354
Gharibzahedi SMT, Razavi SH, Mousavi SM (2013) Ultrasound-assisted formation of the canthaxanthin emulsions stabilized by arabic and xanthan gums. Carbohydr Polym 96:21–30
Ghasemlou M, Khodaiyan F, Jahanbin K, Gharibzahedi SMT, Taheri S (2012) Structural investigation and response surface optimisation for improvement of kefiran production yield from a low-cost culture medium. Food Chem 133:383–389
Hennessy A, Ross R, Devery R, Stanton C (2009) Optimization of a reconstituted skim milk based medium for enhanced CLA production by bifidobacteria. J Appl Microbiol 106:1315–1327
Jiang J, Björck L, Fonden R (1998) Production of conjugated linoleic acid by dairy starter cultures. J Appl Microbiol 85:95–102
Kim Y, Liu R (2002) Increase of conjugated linoleic acid content in milk by fermentation with lactic acid bacteria. J Food Sci 67:1731–1737
Kim YJ, Liu RH, Bond DR, Russell JB (2000) Effect of linoleic acid concentration on conjugated linoleic acid production by Butyrivibrio fibrisolvens A38. Appl Environ Microbiol 66:5226–5230
Kishino S, Ogawa J, Omura Y, Matsumura K, Shimizu S (2002) Conjugated linoleic acid production from linoleic acid by lactic acid bacteria. J Am Oil Chem Soc 79:159–163
Li H, Liu Y, Bao Y, Liu X, Zhang H (2012) Conjugated linoleic acid conversion by six Lactobacillus plantarum strains cultured in MRS broth supplemented with sunflower oil and soymilk. J Food Sci 77:330–336
Li J, Zhang L, Han X et al (2013) Effect of incubation conditions and possible intestinal nutrients on cis-9, trans 11 conjugated linoleic acid production by Lactobacillus acidophilus F0221. Int Dairy J 29:93–98
Lin TY (2000) Conjugated linoleic acid concentration as affected by lactic cultures and additives. Food Chem 69:27–31
Lin TY, Lin C-W, Lee C-H (1999) Conjugated linoleic acid concentration as affected by lactic cultures and added linoleic acid. Food Chem 67:1–5
Lin T, Lin C-W, Wang Y-J (2003) Production of conjugated linoleic acid by enzyme extract of Lactobacillus acidophilus CCRC 14079. Food Chem 83:27–31
Liong M, Shah N (2005) Acid and bile tolerance and cholesterol removal ability of lactobacilli strains. J Dairy Sci 88:55–66
Liu P, Shen S-R, Ruan H, Zhou Q, Ma L-L, He G-Q (2011) Production of conjugated linoleic acids by Lactobacillus plantarum strains isolated from naturally fermented Chinese pickles. J Zhejiang Univ Sci 12:923–930
Nieman C (1954) Influence of trace amounts of fatty acids on the growth of microorganisms. Bacteriol Rev 18:147–163
Ogawa J, Matsumura K, Kishino S, Omura Y, Shimizu S (2001) Conjugated linoleic acid accumulation via 10-hydroxy-12-octadecaenoic acid during microaerobic transformation of linoleic acid by Lactobacillus acidophilus. Appl Environ Microbiol 67:1246–1252
Ogawa J, Kishino S, Ando A, Sugimoto S, Mihara K, Shimizu S (2005) Production of conjugated fatty acids by lactic acid bacteria. J Biosci Bioeng 100:355–364
Puniya AK, Chaitanya S, Tyagi A, De S, Singh K (2008) Conjugated linoleic acid producing potential of lactobacilli isolated from the rumen of cattle. J Ind Microbiol Biotechnol 35:1223–1228
Rosson RA, Grunde AD (2001) Linoleate Isomerase: World Patent, 100846
Scrimgeour C (2005) Chemistry of fatty acids. In: Shahidi F (ed) Baileys industrial oil and fat products, 6th edn. Wiley, New York
Shantha N, Decker E (1993) Conjugated linoleic acid concentrations in processed cheese containing hydrogen donors, iron and dairy-based additives. Food Chem 47:257–261
Van Nieuwenhove C, Oliszewski R, González S, Perez Chaia A (2007) Conjugated linoleic acid conversion by dairy bacteria cultured in MRS broth and buffalo milk. Lett Appl Microbiol 44:467–474
Wang L-M, Lv J-P, Chu Z-Q, Cui Y-Y, Ren X-H (2007) Production of conjugated linoleic acid by Propionibacterium freudenreichii. Food Chem 103:313–318
Wu X, Zhang L, Gurley E et al (2008) Prevention of free fatty acid-induced hepatic lipotoxicity by 18β‐glycyrrhetinic acid through lysosomal and mitochondrial pathways. Hepatology 47:1905–1915
Yadav H, Jain S, Sinha P (2007) Production of free fatty acids and conjugated linoleic acid in probiotic dahi containing Lactobacillus acidophilus and Lactobacillus casei during fermentation and storage. Int J Dairy J 17:1006–1010
Yang B, Chen H, Gu Z et al (2014) Synthesis of conjugated linoleic acid by the linoleate isomerase complex in food‐derived lactobacilli. J Appl Microbiol
Ye S, Yu T, Yang H et al (2013) Optimal culture conditions for producing conjugated linoleic acid in skim-milk by co-culture of different Lactobacillus strains. Ann Microbiol 63:707–717
Zhao H-W, Lv J-P, Li S-R (2011) Production of conjugated linoleic acid by whole-cell of Lactobacillus plantarum A6-1F. Biotechnol Biotec Equip 25:2266–2272
Acknowledgments
Authors would like to thank the University of Tehran and Center of Excellence for Application of Modern Technology for Producing Functional Foods and Drinks, for providing the laboratory facilities and financial support for this project.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Khosravi, A., Safari, M., Khodaiyan, F. et al. Bioconversion enhancement of conjugated linoleic acid by Lactobacillus plantarum using the culture media manipulation and numerical optimization. J Food Sci Technol 52, 5781–5789 (2015). https://doi.org/10.1007/s13197-014-1699-6
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13197-014-1699-6