Abstract
Background
By gating image acquisition in myocardial perfusion SPECT (MPS) to ECG, left ventricular (LV) volumes and function can be determined. Several previous studies have shown that existing MPS software packages underestimate LV volumes compared to cardiac magnetic resonance (CMR). The aim of this study was therefore to develop a new LV segmentation algorithm for gated MPS using CMR as reference standard.
Methods and Results
A total of 126 patients with suspected coronary artery disease, who underwent both gated MPS and CMR were retrospectively included. The proposed LV segmentation algorithm (Segment) was trained in 26 patients, and tested in 100 patients in comparison to four commercially available MPS software packages (QGS, MyoMetrix, ECTb, and Exini) using CMR as reference standard. Mean bias ± SD between MPS and CMR was for EDV −5% ± 12%, −43% ± 8%, −40% ± 8%, −42% ± 9%, −32% ± 7%, for ESV 0% ± 17%, −41% ± 16%, −34% ± 15%, −54% ± 13%, −41% ± 10%, for EF −2% ± 13%, −1% ± 14%, −7% ± 15%, 17% ± 16%, 10% ± 17% for Segment, QGS, MyoMetrix, ECTb, and Exini, respectively, and for LVM 3% ± 18%, 33% ± 25%, 37% ± 24% for Segment, QGS, and ECTb, respectively. Correlation between MPS by Segment and CMR were for EDV R 2 = 0.89, for ESV R 2 = 0.92, for EF R 2 = 0.69, and for LVM R 2 = 0.72, with no difference compared to the correlation between the other MPS software packages and CMR (EDV R 2 = 0.86-0.92, ESV R 2 = 0.91-0.93, EF R 2 = 0.64-0.65, and LVM R 2 = 0.68-0.70).
Conclusion
The Segment software quantifies LV volumes and EF by MPS with similar correlation and a low bias compared to other MPS software packages, using CMR as reference standard. Hence, the Segment software shows potential to provide clinically relevant volumes and functional values from MPS.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Myocardial perfusion SPECT (MPS) is an established non-invasive imaging technique for detection and quantification of myocardial perfusion defects. When image acquisition is gated to ECG it also allows for simultaneous assessment of left ventricular (LV) volumes and function,1 which are important diagnostic and prognostic parameters for patients with coronary artery disease.2 The combined information of perfusion and function has shown to improve accuracy for detecting coronary artery disease.3
Accurate segmentation of myocardial borders is a requirement for quantification of LV volumes and function. Automatic segmentation methods are superior to manual delineation for observer-independence and reproducibility. However, it is important that the software provides possibility to perform manual correction of the segmentation if necessary. Today there are a number of automated algorithms for quantification of perfusion and LV volumes and function by MPS.4-8 Several previous studies have compared LV volumes quantified by those algorithms to the reference standard, cardiac magnetic resonance (CMR).9 In the majority of the studies the end-diastolic volume (EDV) and end-systolic volume (ESV) were underestimated by MPS.10-15 EDV by MPS has also been shown to range from overestimation to underestimation compared to CMR depending on the software package used.16-18 Only one study has shown an overestimation in both EDV and ESV by MPS.19 The primary aim of this study was therefore to develop a new LV segmentation algorithm in gated MPS by using CMR for optimization. A second aim was to test the performance of the new algorithm in comparison to existing algorithms for MPS analysis using CMR as reference standard.
Materials and Methods
Study Population and Design
All patients provided written informed consent to participate in the study and the study was approved by the regional ethics committee. A total of 126 patients, referred for MPS imaging due to known or suspected coronary artery disease, were retrospectively included in the study. The patients had also undergone CMR imaging either due to clinical indications or as a part of a research protocol, when the CMR was performed the same day as the MPS. The time between the two studies was in median 1 day (range 0-57). Exclusion criteria were cardiac events between the MPS and CMR acquisitions, poor image quality in the CMR image or more than 2 months between the MPS and CMR acquisition. Patients were randomly divided into training set (26 studies; 17 men, 9 women) and test set (100 studies; 58 men, 42 women). Mean age was 61 ± 9 years (range, 44-78 years) in the training set and 60 ± 12 years (range, 21-81 years) in the test set. The training set was used for optimization of parameters in the automatic LV segmentation algorithm in MPS and the test set was used to validate the automatic LV segmentation. In the test set, 31% had a clinical history of coronary artery disease, 21% had a clinical history of prior myocardial infarction, 49% had hypertension, and 7% had diabetes mellitus. Clinical characteristics concerning perfusion defect size and affected coronary artery territory are presented in Figure 1. Eighty-eight of the patients in this study were also included in a previous study comparing LV volumes in MPS to CMR.13
Patient characteristics for the test population quantified by the software QPS. A Distribution of summed rest score, which quantifies perfusion within the myocardium. B Distribution of perfusion defects according to coronary artery territories. A patient could be represented in more than one group due to multivessel coronary artery disease. LAD, Left anterior descending coronary artery; RCA, right coronary artery; LCx, left circumflex coronary artery
MPS Acquisition and Analysis
MPS was performed according to established clinical 2 days protocols using a dual head camera (ADAC, Milpitas, CA). Gated MPS images were acquired at rest for each patient, after injection with 400-800 MBq 99mTc tetrofosmin (Amersham Health, Buckinghamshire, UK) depending on bodyweight. The patient was placed in supine position and imaged in steps of 5.6° using a 64 × 64 matrix with a pixel size of 5 × 5 mm and a slice thickness of 5 mm. Images were gated to electrocardiogram using 8 frames per cardiac cycle. Image acquisition time was approximately 15 minutes. Iterative reconstruction using maximum likelihood-expectation maximization (MLEM) was performed with a low-pass Butterworth filter. Twelve MLEM iterations were used with filtered-back projection image as the starting condition and a cutoff frequency set to 0.55 of Nyquist and order 5.0. No attenuation or scatter correction was applied. Short-axis images were reconstructed semi-automatically with manual adjustments using the program AutoSPECT Plus (Philips Pegasys software version 5.01).
Reconstructed MPS images were loaded into each of five software packages; Segment (version 1.8R1554; Medviso AB, Lund, Sweden), Quantitative Gated SPECT (QGS) (version 4.0; Cedar-Sinai Medical Centre, Los Angeles, CA),4 MyoMetrix (Xeleris version 2.1220; GE Healthcare),5 Emory Cardiac Toolbox (ECTb) (version 3.05; Emory University Medical Centre, Atlanta, GA),6 and Exini heart (version 3.1; Exini Diagnostics AB, Lund, Sweden).7 Default settings without manual interactions were used for the fully automatic LV segmentation. For each of the five MPS software packages, EDV, ESV, and ejection fraction (EF) were calculated based on the LV segmentation. The left ventricular mass (LVM) could only be quantified by Segment, QGS, and ECTb, and was therefore only presented for those software packages. Perfusion within the myocardium was automatically analyzed by the software Quantitative Perfusion SPECT (QPS) (version 4.0; Cedar-Sinai Medical Centre, Los Angeles, CA)20 for the purpose of illustrating patient characteristics. Uptake was graded in each of the 17 segments of the LV on a 5-point scale (0-4). The summed rest score was defined as sum of the scores in all segments. Presence of a perfusion defect in a coronary artery territory was defined by a score greater than or equal to 4 in that territory.
To compare base and apex definition between the MPS LV segmentation algorithms, 20 patients were randomly selected from the test set. In this subset, apical-basal distance (mean of end-diastole and end-systole) and atrioventricular-plane (AV-plane) displacement were compared between Segment and the four other MPS software packages.
CMR Acquisition and Analysis
CMR image data were acquired in both short-axis and long-axis projections with a 1.5 T scanner (Intera, Philips Medical Systems, Best, the Netherlands). Short-axis imaging covering the entire LV was undertaken using a retrospectively triggered balanced steady-state free precession (SSFP) sequence. Typical imaging parameters were TR/TE: 2.9/1.5 ms, flip angle 60°, 30 time frames per cardiac cycle, pixel resolution 1.4 × 1.4 mm, slice thickness 8 mm and slice gap 0 mm.
The CMR image analysis was performed in the software Segment. Endo- and epicardium of the LV were manually traced by experienced observers in each short-axis image slice in both end-diastole and end-systole. The LV base and apex were defined by comparison with the long-axis images using established methods.21 Trabecular and papillary muscles which were not contiguous to the myocardial wall were excluded and the endo- and epicardium were adjusted to preserve LVM throughout the cardiac cycle. Based on the final LV segmentation, EDV, ESV, EF, and LVM were calculated. The delineation was performed in consensus by two observers. To determine interobserver variability, a third observer, blinded to the first delineation manually delineated the LV in a subset of 10 out of the 100 patients in the test set. The observers were physicians with 4, 11, and 11 years of both clinical and research experience of CMR, respectively. The delineations were performed blinded to MPS data and clinical data for the patients.
Segmentation Algorithm for LV
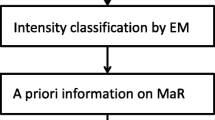
The proposed algorithm for segmentation of the LV in gated MPS images is implemented in the software Segment (http://segment.heiberg.se). Segment is a software platform where both manual and automatically analysis of cardiovascular images from different imaging modalities can be performed.22 The proposed algorithm is fully automatic and is an extension of the LV segmentation algorithm previously described for ungated MPS images.23 A summary of the LV segmentation algorithm for gated image stacks are presented below and in Figure 2. A more detailed description of the algorithm is given in Appendix A. Threshold and parameter values used in the algorithm were all optimized in the training set by comparing the result from MPS to CMR.
Flow scheme for the LV segmentation algorithm by the Segment software. The labels P, U, and G stand for preparation, ungated and gated, respectively. Each step in the algorithm is more extensive described in Appendix A
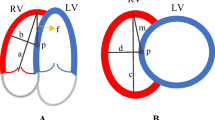
The first step for the algorithm was calculating an ungated image stack, as mean over all time frames. In this ungated image stack the LV base and apex were defined by short-axis slices satisfying both volume and counts thresholds. Within the LV, the midmural line was estimated by the position representing peak count in radial direction. On each side of the midmural line, the myocardial borders were defined by the position representing 90% of radial peak count with restriction on wall thickness and were refined with spline fitting. In the last step the basal outflow tract was defined and LV volumes were calculated. The steps above correspond to P1-U9 in Figure 2.
The LV base and apex in the gated image stack were defined using the LV base and apex definition from the ungated image stack and restriction on AV-plane displacement. Within the LV, the midmural line was defined by search for peak count in radial direction and was refined by spline fitting. On each side of the midmural line, the myocardial borders were defined by minimizing the difference between the ungated image stack and the gated image stack regarding LVM and mean LV lumen volume. In the optimization, the LVM was preserved over the heart cycle and a lumen expansion parameter was used in order to compensate for underestimation of the LV lumen radius caused by the limited spatial resolution. From the LV segmentation, end-diastole and end-systole were defined as the time frame with largest and smallest lumen, respectively, and LV volumes and functional values were calculated. The steps above correspond to G1-G5 in Figure 2.
Statistical Analysis
Values are presented as mean ± SD unless otherwise stated. The error for the LV volumes and EF was calculated as ((value by MPS) − (value by CMR))/(value by CMR). To investigate potential sources of error in LV segmentation in MPS by Segment, a linear regression analysis of the relation between error in LV volumes/EF and four different LV parameters was performed. The four LV parameters were wall thickness by CMR, apical-basal distance by CMR, mid-ventricular lumen radius by CMR, and perfusion defect size by MPS. The impact of the myocardial border threshold in LV segmentation in MPS by Segment was tested by varying the threshold from 80% to 99% (default was 90%). Pearson’s linear regression analysis was performed to calculate the relationship between two data sets were normal distribution could be assumed. The differences in bias for LV volumes and EF, apical-basal distance and AV-plane displacement were tested by the one-way analysis of variance (ANOVA). If the ANOVA test on the level 0.05 was consider significantly different for a parameter, Student’s paired t test using the Bonferroni correction was performed to test the difference between two methods. Differences in correlation coefficients were tested by a Chi-square test for correlated correlation coefficients.24 If the Chi-square test on the level 0.05 was considered significantly different for a parameter, a pairwise Z-test using the Bonferroni correction was performed to test if the correlation coefficients for the Segment algorithm differed from the correlation coefficients for the other MPS software packages. Intraobserver variability was calculated as mean ± SD of the difference between observations. A P value <.05 was considered statistically significant.
Results
Comparison Between MPS by Segment and CMR
The LV volumes and EF for the test set by MPS and CMR are presented in Table 1 and Figure 3. There was no significant difference between MPS by Segment and CMR for EDV, ESV, EF, and LVM, respectively.
Linear regression analysis of the relationship between error in the LV volumes/EF, and the four LV parameters is presented in Table 2. The result shows that wall thickness was a strong predictor for error in LVM (R 2 = 0.73), and hence a source of error in LVM quantification. The mean end-diastolic wall thickness in MPS by Segment was 6.3 ± 0.4 mm, compared to 6.1 ± 1.2 mm for CMR. The mean end-diastolic LV dimensions for the manually outlined CMR images were 97 ± 11 mm for apical-basal distance and 28 ± 3.2 mm for mid-ventricular lumen radius. The interobserver variability for CMR was −2% ± 2% for EDV, −1% ± 2% for ESV, 0% ± 1% for EF, and 2% ± 6% for LVM.
By varying the myocardial border threshold in the LV segmentation algorithm by Segment, a significant change in mean LVM was found for a threshold ≤88 or ≥92 when compared to using the default threshold value of 90%. Mean EDV and ESV were significantly changed for a threshold ≤81 or ≥96 and mean EF for a threshold ≤84 or ≥95.
Comparison Between MPS Software Packages
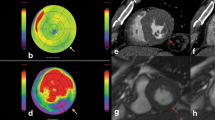
The comparison between the five MPS software packages is presented in Table 1 and Figure 4. The bias in EDV, ESV, and LVM was lower with Segment than the four other MPS software packages. The bias in EF was similar with Segment, QGS, MyoMetrix, and Exini, but higher with ECTb. The correlation coefficients for LV volumes and EF were not significantly different between Segment and the other MPS software packages. In Figure 5, the LV segmentation by CMR and all five MPS software packages is illustrated for a patient for whom the LV volume errors were close to mean bias for all MPS software packages.
Illustration of LV segmentation in one representative patient in both end-diastole (ED) and endsystole (ES). Images are shown in mid-ventricular short-axis (SA), vertical long-axis (VLA), and horizontal long-axis (HLA) projection. Top panel shows the manual segmentation in CMR and the following panels the segmentation by the five automatic LV segmentation algorithms in MPS. The same reconstructed short-axis image stack was used in the segmentation process in all MPS software packages, and the slight variation in noise/smoothness is attributable to different visualization settings. Note the higher agreement in LV volumes between CMR and MPS by Segment than between CMR and the four other MPS software packages
In the subset of 20 patients, there was no significant difference in apical-basal distance between the MPS software packages (ANOVA P = .75). Furthermore, there was no difference in AV-plane displacement between Segment, and MyoMetrix, ECTb, and Exini. QGS defines AV-plane displacement with a mean bias of 6.3 mm lower than Segment (P < .01).
Discussion
The major finding of this study was that LV volumes and EF in MPS by Segment show no bias compared to CMR over a wide range of clinically relevant LV volumes. Compared to four commercially available MPS software packages, Segment had lower bias for quantification of EDV, ESV, and LVM and lower or equal bias for EF, using CMR as reference standard.
Comparison Between MPS by Segment and CMR
There was a good agreement between CMR and MPS by Segment, with no difference in mean values, for LV volumes and EF. However, they should not be used interchangeably due to the relative high variance between the methods. Plots of the differences between MPS by Segment and CMR yielded no trend for EDV, ESV, and EF (Figure 3B, D, and F). LVM, however, yielded a visible trend of overestimation of small hearts (low LVM) and underestimation of large hearts (high LVM), as shown in Figure 3H. This trend was correlated to wall thickness by CMR (Table 2).
The study population consisted of patients with and without perfusion defects. As presented in Table 2, no or very weak correlations between error in LV volumes/EF and defect size were found. Thus, in this study presence of perfusion defects had little effect on the accuracy of the LV segmentation by Segment.
Comparison Between MPS Software Packages
The correlation coefficients between MPS and CMR for LV volumes and EF were similar between Segment and the four other MPS software packages, and in line with results reported in earlier studies.10-19 In the current study the EDV and ESV were underestimated by MPS, as compared to CMR, when using QGS, MyoMetrix, ECTb, and Exini. This underestimation has been observed in previous studies, with the same magnitude12,13 or with a lower magnitude.10,11,15 The lower magnitude of the underestimation may be explained by older turbo gradient echo CMR sequences in those studies compared to the current standard SSFP imaging that was used in the current study. It has been shown that turbo gradient echo imaging shows significantly smaller EDV and ESV compared to SSFP imaging.25 In contrast to the findings in the present study, EDV and ESV by QGS, ECTb, and Exini were shown to range from underestimated to overestimated compared to CMR.16,17,19 The lower bias between MPS and CMR in Lipke et al16 and Winz et al17 compared to this study are explained by similar mean LV volumes by MPS but lower mean LV volumes by CMR. Since the MPS analysis is fully automatic a possible explanation for the difference in bias is different approaches for the LV delineation in CMR. The most critical part of the LV delineation is the definition of the most basal short-axis slice and inclusion or exclusion of the trabecular and papillary muscles in the LV. The overestimation by MPS in the study by Faber et al19 may be explained by the use of older turbo gradient echo CMR imaging in that study compared to the SSFP imaging that was used in the current study.
In previous studies, the underestimation of EDV and ESV was mainly explained by the limited temporal and spatial resolution in MPS. This limited resolution causes blurring of the short end-systolic phase, low contrast between blood and myocardium, a non-visible outflow tract, and merging of the papillary muscles into the myocardium. The LV segmentation algorithm by Segment attempted to overcome the issue of limited spatial resolution by optimizing the myocardial border threshold from CMR and using a lumen expansion parameter. The result was a lower bias in EDV and ESV between Segment and CMR than between the four other MPS software packages and CMR.
The LV segmentation algorithm in Segment was overall similar to the four other LV segmentation algorithms, but differs with regards to computational approaches. The three main steps in the algorithms were definition of base and apex, midmural line, and myocardial borders. Even though the approaches for finding the base and apex differ between the algorithms, the LV segmentation algorithms resulted in similar base and apex definitions. The midmural line in Segment was defined by spline fitting followed by a lumen expansion compensation. The other algorithms used ellipsoid fitting (QGS and MyoMetrix), a statistical heart shape model (Exini), or purely the location of radial maximum count (ECTb). Using splines instead of an ellipsoid or a heart shape model, results in a more flexible model which adapts to the shape of the present heart. Finally, the myocardial borders were defined on each side of the midmural line by either using a relative count threshold (Segment 90% and Exini 75%), a fixed number of SDs below the myocardial maximum (QGS and MyoMetrix), or a fixed wall thickness of 10 mm in end-diastole (ECTb). The approach for defining the myocardial borders by Segment is based on the training set in this study, which has a mean end-diastolic wall thickness of 6.4 mm. The other MPS software packages assume an end-diastolic wall thickness baseline around 10 mm. This difference in assumption of wall thickness baseline may be explained by using SSFP imaging in the current study. As showed in a previous study by Plein et al, the mean wall thickness by CMR was significantly lower in the current standard SSFP imaging compared to older turbo gradient echo imaging.26 As presented above, the threshold for the myocardial border definition affects the LV volumes and EF by Segment. Varying the threshold affects the LVM measurement more than the EDV, ESV, and EF measurements. Altogether, the different computational approaches resulted in a larger LV lumen volume and lower myocardial mass in Segment than the other MPS software packages, as illustrated in the patient example in Figure 5.
Study Limitations
The CMR and MPS imaging were not performed simultaneously, and the physiological states of the patients can therefore differ between the two studies. To reduce this risk, only patients without cardiac events between the studies were included, and a majority of the studies (69%) were performed within 1 day of each other. The different MPS software packages have their own recommendations for how to reconstruct the images. This study only includes MPS data reconstructed by an iterative approach according to a normal clinical protocol. The parameters in the LV segmentation by Segment were optimized for the image reconstruction method used in this study, and the parameters dependence on pixel size and reconstruction method for the MPS images was not investigated.
Conclusion
The Segment software quantifies LV volumes and EF by MPS with similar correlation and a low bias compared to other MPS software packages, using CMR as the reference standard. Hence, the Segment software shows potential to provide clinically relevant volumes and functional values from MPS.
References
Abidov A, Germano G, Hachamovitch R, Berman DS. Gated SPECT in assessment of regional and global left ventricular function: Major tool of modern nuclear imaging. J Nucl Cardiol 2006;13:261-79.
Sharir T, Germano G, Kavanagh PB, Lai S, Cohen I, Lewin HC, et al. Incremental prognostic value of post-stress left ventricular ejection fraction and volume by gated myocardial perfusion single photon emission computed tomography. Circulation 1999;100:1035-42.
Lima RS, Watson DD, Goode AR, Siadaty MS, Ragosta M, Beller GA, et al. Incremental value of combined perfusion and function over perfusion alone by gated SPECT myocardial perfusion imaging for detection of severe three-vessel coronary artery disease. J Am Coll Cardiol 2003;42:64-70.
Germano G, Kiat H, Kavanagh PB, Moriel M, Mazzanti M, Su HT, et al. Automatic quantification of ejection fraction from gated myocardial perfusion SPECT. J Nucl Med 1995;36:2138-47.
MyoMetrix White Paper. GE Medical Systems; 2006.
Faber TL, Cooke CD, Folks RD, Vansant JP, Nichols KJ, DePuey EG, et al. Left ventricular function and perfusion from gated SPECT perfusion images: An integrated method. J Nucl Med 1999;40:650-9.
Lomsky M, Richter J, Johansson L, El-Ali H, Astrom K, Ljungberg M, et al. A new automated method for analysis of gated-SPECT images based on a three-dimensional heart shaped model. Clin Physiol Funct Imaging 2005;25:234-40.
Ficaro EP, Lee BC, Kritzman JN, Corbett JR. Corridor4DM: The Michigan method for quantitative nuclear cardiology. J Nucl Cardiol 2007;14:455-65.
Pennell DJ, Sechtem UP, Higgins CB, Manning WJ, Pohost GM, Rademakers FE, et al. Clinical indications for cardiovascular magnetic resonance (CMR): Consensus Panel report. J Cardiovasc Magn Reson 2004;6:727-65.
Bax JJ, Lamb H, Dibbets P, Pelikan H, Boersma E, Viergever EP, et al. Comparison of gated single-photon emission computed tomography with magnetic resonance imaging for evaluation of left ventricular function in ischemic cardiomyopathy. Am J Cardiol 2000;86:1299-305.
Bavelaar-Croon CD, Kayser HW, van der Wall EE, de Roos A, Dibbets-Schneider P, Pauwels EK, et al. Left ventricular function: Correlation of quantitative gated SPECT and MR imaging over a wide range of values. Radiology 2000;217:572-5.
Chan J, Jenkins C, Khafagi F, Du L, Marwick TH. What is the optimal clinical technique for measurement of left ventricular volume after myocardial infarction? A comparative study of 3-dimensional echocardiography, single photon emission computed tomography, and cardiac magnetic resonance imaging. J Am Soc Echocardiogr 2006;19:192-201.
Hedeer F, Palmer J, Arheden H, Ugander M. Gated myocardial perfusion SPECT underestimates left ventricular volumes and shows high variability compared to cardiac magnetic resonance imaging—A comparison of four different commercial automated software packages. BMC Med Imaging 2010;10:10.
Mistry N, Halvorsen S, Hoffmann P, Muller C, Bohmer E, Kjeldsen SE, et al. Assessment of left ventricular function with magnetic resonance imaging vs. echocardiography, contrast echocardiography, and single-photon emission computed tomography in patients with recent ST-elevation myocardial infarction. Eur J Echocardiogr 2010;11:793-800.
Persson E, Carlsson M, Palmer J, Pahlm O, Arheden H. Evaluation of left ventricular volumes and ejection fraction by automated gated myocardial SPECT versus cardiovascular magnetic resonance. Clin Physiol Funct Imaging 2005;25:135-41.
Lipke CS, Kuhl HP, Nowak B, Kaiser HJ, Reinartz P, Koch KC, et al. Validation of 4D-MSPECT and QGS for quantification of left ventricular volumes and ejection fraction from gated 99mTc-MIBI SPET: Comparison with cardiac magnetic resonance imaging. Eur J Nucl Med Mol Imaging 2004;31:482-90.
Winz OH, Meyer PT, Knollmann D, Lipke CS, Kuhl HP, Oelve C, et al. Quantification of left ventricular volumes and ejection fraction from gated 99mTc-MIBI SPECT: MRI validation of the EXINI heart software package. Clin Physiol Funct Imaging 2009;29:89-94.
Stegger L, Lipke CS, Kies P, Nowak B, Schober O, Buell U, et al. Quantification of left ventricular volumes and ejection fraction from gated 99mTc-MIBI SPECT: Validation of an elastic surface model approach in comparison to cardiac magnetic resonance imaging, 4D-MSPECT and QGS. Eur J Nucl Med Mol Imaging 2007;34:900-9.
Faber TL, Vansant JP, Pettigrew RI, Galt JR, Blais M, Chatzimavroudis G, et al. Evaluation of left ventricular endocardial volumes and ejection fractions computed from gated perfusion SPECT with magnetic resonance imaging: Comparison of two methods. J Nucl Cardiol 2001;8:645-51.
Slomka PJ, Nishina H, Berman DS, Akincioglu C, Abidov A, Friedman JD, et al. Automated quantification of myocardial perfusion SPECT using simplified normal limits. J Nucl Cardiol 2005;12:66-77.
Pennell DJ. Ventricular volume and mass by CMR. J Cardiovasc Magn Reson 2002;4:507-13.
Heiberg E, Sjogren J, Ugander M, Carlsson M, Engblom H, Arheden H. Design and validation of Segment—Freely available software for cardiovascular image analysis. BMC Med Imaging 2010;10:1.
Soneson H, Ubachs JF, Ugander M, Arheden H, Heiberg E. An improved method for automatic segmentation of the left ventricle in myocardial perfusion SPECT. J Nucl Med 2009;50:205-13.
Meng XL, Rosenthal R, Rubin DB. Comparing correlated correlation-coefficients. Psychol Bull 1992;111:172-5.
Alfakih K, Plein S, Thiele H, Jones T, Ridgway JP, Sivananthan MU. Normal human left and right ventricular dimensions for MRI as assessed by turbo gradient echo and steady-state free precession imaging sequences. J Magn Reson Imaging 2003;17:323-9.
Plein S, Bloomer TN, Ridgway JP, Jones TR, Bainbridge GJ, Sivananthan MU. Steady-state free precession magnetic resonance imaging of the heart: Comparison with segmented k-space gradient-echo imaging. J Magn Reson Imaging 2001;14:230-6.
Mortelmans L, Nuyts J, Vanhaecke J, Verbruggen A, De Roo M, De Geest H, et al. Experimental validation of a new quantitative method for the analysis of infarct size by cardiac perfusion tomography (SPECT). Int J Cardiac Imaging 1993;9:201-12.
Piegl L, Tiller W. The Nurbs book, 2nd ed. New York: Springer-Verlag; 1997.
Acknowledgments
The authors would like to thank technicians Ann-Helen Arvidsson and Christel Carlander for invaluable help with data acquisition. Financial support was provided by Swedish Heart Lung Foundation, Lund University Faculty of Medicine, the Swedish Research Council (grant 2008-2461, 2008-2949), Knowledge Foundation (grant 2009-0080), and Region of Scania.
Author information
Authors and Affiliations
Corresponding author
Appendices
Appendix A
LV Segmentation Algorithm
The labels in the following sections refer to the labeled steps in Figure 2.
Preparation of image stack
-
P1
An ungated image stack was calculated as the mean count values over time. In the ungated image stack the LV was segmented and LV values, used in the LV segmentation in the gated image stack, were calculated.
LV segmentation in the ungated image stack
-
U1
To minimize influence of extra-cardiac activity in the definition of base and apex, only pixels within a circular region of interest (ROI), with diameter equal to image size, were considered. Base and apex were defined as the most basal and the most apical slice, respectively, with a large enough area of high counts. For the base, the area was 350 mm2 and the count threshold was 34% of maximal count. For the apex, the area was 80 mm2 and the count threshold was 42% of maximal count. The threshold values were derived from the training set by minimizing the difference in apical-basal distance between CMR and MPS.
-
U2
The LV center was estimated in the LV short-axis slices in three steps. First, a circle was fitted to high counts in each mid-ventricular slice. The middle of these circles was used as a first estimation of the mid-ventricular LV center points. This estimation was then refined by fitting a first degree polynomial to the points. Finally, this line was extrapolated in basal and apical direction, and thereafter defines the LV center through the whole LV.
-
U3
When the LV center was defined the ROI was centered in the LV center and the diameter was set to 90 mm. The diameter was derived from the maximal epicardial diameter in the CMR training set. Base and apex were then redefined as described in U1.
-
U4
To calculate the mean wall thickness, myocardial borders were estimated in the mid-ventricular slice with highest total counts within the ROI. The midmural line was estimated as the location of the radial peak count, and myocardial borders were defined as the location of 90% of radial maximum, on each side of this line. The border threshold was derived in the training set by minimizing the difference in LVM between CMR and MPS. Wall thickness was then defined as the mean distance between endo- and epicardium.
-
U5
The definition of the myocardial borders in the whole LV starts by estimation of the midmural line in each LV short-axis slice. The line was defined as the location of the peak count in the radial direction from the LV center. To exclude outliers, the line was refined using a cost-minimization algorithm.23
-
U6
The radius of the midmural line was used to identify incorrect apex definition, often caused by apical defects. A midmural radius in apex that exceeds 34% of the maximum midmural radius defines incorrect apex definition. This threshold was derived from the relation between midmural radius in apex and maximum midmural radius in the CMR training set. To correct the apex definition, an ellipsoid was fitted to the midmural points in a least square sense. The position of the most apical edge of the ellipsoid redefines apex. In the potentially newly included apical slices, the midmural line was derived from the fitted ellipsoid.
-
U7
On each side of the midmural line myocardial borders were estimated by searching for the line closest to the border threshold (90% of maximum count in the radial direction) with a restriction that the wall thickness must be within 80% to 120% of mean wall thickness. Myocardial borders were then refined by fitting a 2 dimensional spline to the myocardial points as described in Appendix B.
-
U8
The last step was to correct the basal LV segmentation by searching for the outflow tract in the basal part. The basal part was defined as the 25% most basal slices, and was derived from the CMR training set. The outflow tract was defined as a continuous region in the septal wall with counts falling below mean count of the LV lumen. The wall thickness in the outflow tract was then set to zero.
-
U9
The LVM and LV lumen volume were calculated based on the final myocardial borders. The LV center, maximal epicardial radius, base and apex, lumen volume, outflow tract region, and LVM from the ungated image stack were then used as reference values (therefore referred to as ungated reference values below) in the segmentation of LV in the gated image stack.
LV segmentation in the gated image stack
-
G1
The segmentation of LV in the gated image stack starts by using the ungated reference LV center as definition of the LV center in all time frames.
-
G2
Thereafter, base and apex were estimated separately in each time frame, as described in U1 with a ROI centered in the LV center and a radius derived from the ungated reference maximal epicardial radius. The base estimation in the first time frame was restricted to be within one slice from the ungated reference base, and the AV-plane displacement to be within 5 to 20 mm (1-4 slices). The apex estimation was restricted to be within one slice from the ungated reference apex, and the apex displacement to be within 0 to 5 mm (0-1 slices). The estimation of base and apex were then adjusted to be smooth over the cardiac cycle. The displacement thresholds were derived from the CMR training set.
-
G3
In the LV slices, the midmural line was estimated as the location of the peak count in the radial direction from the LV center. This estimation was refined by fitting a 3 dimensional spline, as described in Appendix B.
-
G4
On each side of the midmural line, myocardial borders were defined simultaneously for all time frames by an optimization algorithm. The algorithm minimizes the differences in LVM and mean LV lumen volume (LVV) between the ungated image stack and the gated image stack. In the minimization, the LVM was preserved over the heart cycle. The objective function (F) was
$$ F = \frac{{\mathop \sum \nolimits_{t = 1}^{T} \left| {\frac{{{\text{LVM}}_{t} - {\text{LVM}}_{\text{ungated}} }}{{{\text{LVM}}_{\text{ungated}} }}} \right|}}{T} + \left| {\frac{{{\text{LVV}}_{\text{ungated}} - l \cdot \frac{{\mathop \sum \nolimits_{t = 1}^{T} {\text{LVV}}_{t} }}{T}}}{{{\text{LVV}}_{\text{ungated}} }}} \right| $$where t is the time frame in the gated image stack, T is the number of time frames, and l = 0.73, is a lumen expansion parameter. The lumen expansion parameter compensated for the underestimation of the LV lumen radius. This was a consequence of the limited spatial resolution and the convexity of the myocardium, which produced a shift of the activity towards the LV center.27 The value of the lumen expansion parameter were derived from the training set by minimizing the difference in EDV and ESV between MPS and CMR.
-
G5
Finally the outflow tract was defined according to the ungated reference outflow tract. Based on the LV segmentation, the end-diastole was defined as the time frame with largest lumen and end-systole as the time frame with smallest lumen. Thereafter, EDV, ESV, EF, and LVM were calculated.
Appendix B
Surface Refinement by Spline Fitting
To have a smooth surface representation and exclude data points outliers, the myocardial and midmural surfaces were refined by fit a spline to the estimated surface points. For non-gated image stacks the resulting spline surface was 2 dimensional, and for gated image stacks 3 dimensional. The spline was fitted to the data points by a least square surface approximation.28 The surface was build up by fitting curves across the data in one direction at a time, starting in the temporal direction, thereafter in the longitudinal direction and finally in the circumferential direction. The surface approximation used a fixed number of control points in each direction and a fixed degree of the fitted curves. The number of control points and the degree of the fitted curves were derived from the MPS training set in order to generate a smooth surface. The number of control points and the degree of the fitted curve were, respectively, 10 and 2 in the circumferential direction, 3 and 3 in longitudinal direction, and 4 and 2 in temporal direction.
In each direction the m given data points, Q, were approximated in a least square sense by a nonrational curve C of degree p, according to
where the corner points were generated exactly
and the remaining Q k were approximated by
with respect to the n control points P i . N was the B-spline basis functions, and \( \tilde{u}_{k} \) parameter values assigned to each Q k . \( \tilde{u}_{k} \) was calculated by
where
and
The knots, u, were calculated by
where
where \( \left| {\_x\_} \right| \) rounds the element x down to the nearest integer.
Rights and permissions
About this article
Cite this article
Soneson, H., Hedeer, F., Arévalo, C. et al. Development and validation of a new automatic algorithm for quantification of left ventricular volumes and function in gated myocardial perfusion SPECT using cardiac magnetic resonance as reference standard. J. Nucl. Cardiol. 18, 874–885 (2011). https://doi.org/10.1007/s12350-011-9427-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12350-011-9427-3