Abstract
Investigation of genetic diversity is essential for the selection of parents for crop breeding and conservation of genetic resources. To estimate the genetic variability and population structure in the midst of 45 accessions of sponge gourd brought together from different geographical areas of India, morphological traits and two molecular markers, ISSR and SCoT markers were compared. Principal components analysis of 20 morphological traits showed 72.70% variability and significant positive correlations between fruit traits. All three marker techniques clustered all accessions into two groups with few outgroups. High level of polymorphism was observed among ISSR (74.6%) and SCoT (71.5%) primers. The Bayesian model revealed the hidden grouping and showed admixture type of population. The diversity pattern is influenced by genetic marker used, as different molecular markers have different polymorphism evaluation efficiency. This study can be helpful in amplifying the genetic base and selection of specific traits for breeding. Thus, ISSR and SCoT markers are potential marker for identification in sponge gourd and provide valuable data on its genetic correlation and structure.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sponge gourd (Luffa cylindrica) is an important cucurbit vegetable and cultivated worldwide in rainy and summer season. Its origin is subtropical Asia particularly India, considered the primary center of origin of sponge gourd (Kalloo 1993; Sirohi et al. 2005). It is a monoecious and cross-pollinated crop with diploid chromosome number 2n = 2x = 26. Its fruits are the rich source of vitamin A, vitamin C and iron (Yawalkar. 2004) and have a potent anti-inflammatory effect, anti-tumor, anti-viral activities and have protective role against fever, enteritis, diabetes, with role in removing toxins and regeneration of skin (Azeez et al. 2013; Lee and Yoo 2006). Dried fruits of sponge gourd are utilized worldwide in form of bath sponges, packaging material, manufacture of ethanol and as adsorptive of heavy metals (Papanicolaou et al. 2015). Little attention has been given on genetic diversity and genetic improvement of sponge gourd in India. Knowledge of genetic variability is essential for conservation and crop breeding (Talebi et al. 2008) and reduced genetic susceptibility (Fatehi et al. 2011). Genetic variability can be estimated through study of morphological characters, molecular and biochemical markers (Carvalho 2004; de Vicente et al. 2005).
Morphological characterization is employed before molecular characterization as this method estimate genetic diversity along with the activity of genotypes in given environmental conditions (Hoogendijk and Williams 2001). Molecular markers are considered more reliable and used extensively to investigate genetic differences as well as employed in plant breeding. ISSR is an easy, quick technique for determination of genetic diversity, genetic mapping, gene tagging and evolutionary analyses without previous information of the sequence of the genome (Biabani et al. 2013; Arcade et al. 2000; Sankar and Moore 2001). It encompass nearly all of the advantages of SSR and AFLP and the extensiveness of RAPD marker (Reddy et al. 2002). Recently, besides random DNA markers, many new gene-targeted markers have been developed. Start codon targeted (SCoT) marker is a new, gene-directed marker and used for cultivar identification, genetic investigation, and mapping (Luo et al. 2010; Collard and Mackill 2009).
Hitherto, few molecular studies have been conducted for luffa variability assessment. Thus, there is a need of more polymorphic markers. Earlier investigations have been conducted to illustrate the genetic variability of Luffa based on morphological variability (Prakash et al. 2013), molecular markers such as RAPD (Hoque and Rabbani 2009), morphological and RAPD (Junhui and Changping 2008), SSR and SRAP (Jun et al. 2010), morphological and SRAP (Tyagi et al. 2016), RAPD and ISSR (Rathod et al. 2015), morphological, ISSR and DAMD (Misra et al. 2017) and SSR (Pandey et al. 2017). To date, no data is available on comparative evaluation of genetic variability within sponge gourd using ISSR, SCoT and morphological markers. Here, the utilization of SCoT marker system for genetic variability study was illustrated for the first time in sponge gourd accessions.
Thus, the aim of present investigation was to estimate the efficiency of ISSR and SCoT along with morphological traits in ascertaining the genetic diversity and population structure of sponge gourd accessions.
Materials and methods
Plant material
Seeds of 45 accessions of L. cylindrica were collected for this study from 10 different states of India i.e. West Bengal, Uttar Pradesh, Punjab, Jharkhand, Maharashtra, Gujarat, New Delhi, Himachal Pradesh, Uttarakhand and Madhya Pradesh (Table 1). These were selfed and retained as pure lines in the fields of Horticulture department, IARI, New Delhi during the month of March–May with sub-humid conditions. The experiment was laid out during the spring–summer season of 2013 and 2014 in a randomized block design with three replications. The lines were sown in rows of 2.5 m with 75 cm spacing between the plants, with fifteen plants per replication. Observations were made on 10 randomly selected plants in each replication, and all 3 replications were analyzed.
Morphological analysis
The morphological measurements were recorded for 14 qualitative traits and 12 quantitative traits of 10 randomly chosen plants of each accession based on descriptors adopted and created by Joshi et al. (2004) with some modifications (Table 2).
DNA extraction
Total genomic DNA was extracted from young healthy leaves using CTAB procedure as described by Saghai-Maroof et al. (1984). The purity and yield of the extracted DNA was determined electrophoretically on 0.8% agarose gel.
ISSR analysis
A set of 60 primers were screened, out of which 20 primers were found reproducible and selected for profiling. The PCR was performed in a reaction volume of 25 µl containing 50 ng of genomic DNA, 10X PCR buffer, 0.2 mM of each dNTP, 1.5 mM MgCl2, 8 µM of primer and 0.5 U of Taq DNA polymerase (Thermo-Scientific). Bioer-Gene Pro-cycler was used for PCR amplification using the following conditions: 5 min at 94 °C; followed by 35 cycles of 1 min at 94 °C, 1 min at 48–60 °C, and 2 min at 72 °C, and a final extension of 7 min at 72 °C. The amplification products were separated on 2% (w/v) agarose gel using the 50 bp (Thermo-Scientific) and 1 kb ladders (Thermo-Scientific) as standards in 1X TBE buffer.
SCoT analysis
A set of 80 primers were screened, out of which 23 primers were found reproducible and selected for profiling. The PCR was performed in a reaction volume of 25 µl containing 50 ng of genomic DNA, 10X PCR buffer, 0.2 mM of each dNTP, 1.5 mM MgCl2, 8 µM of primer and 0.5 U of Taq DNA polymerase (Thermo-Scientific). Bioer-Gene Pro-cycler was used for PCR amplification using the following conditions: 5 min at 94 °C; followed by 35 cycles of 1 min at 94 °C, 1 min at 48–60 °C, and 2 min at 72 °C, and a final extension of 7 min at 72 °C. The amplification products were separated on 2% (w/v) agarose gel using the 50 bp (Thermo-Scientific) and 1 kb ladders (Thermo-Scientific) as standards in 1X TBE buffer.
Data analysis
For morphological traits, scoring was carried out by giving a series of numbers to each trait. Pearson’s correlation coefficient was calculated between quantitative traits. Principal component analysis (PCA) and total variance were calculated. All these calculations were performed using SPSS 16.0 software. The morphological data was subjected to cluster analysis based on Euclidean distance using NTSYS-pc version 2.1(Rohlf 2000).
Clear and distinct bands obtained with both molecular markers were scored as 0 (absence) and 1(presence) and entered into a binary matrix. The Jaccard similarity coefficient was calculated using binary matrices to determine similarity among accessions using NTSYS-pc version 2.1 (Rohlf 2000). The UPGMA based clustering was performed to infer genetic relatedness among accessions using NTSYS-pc software. The markers informativeness was investigated by calculating amplified bands (AB), polymorphic bands (PB) and polymorphism information content (PIC). Genetic diversity parameters such as polymorphic loci percentage (P%), Nei gene diversity (H) (Nei 1973) and Shannon information index (I) (Shannon and Weaver 1949) were calculated by Popgene software version 1.31(Yeh and Boyle 1997). Population structure was explored using the Bayesian model-based clustering using STRUCTURE 2.3.4 software (Pritchard et al. 2000), which cluster individuals into optimal number of populations (K) based on the multilocus genotypic data based on Markov Chain Monte Carlo (MCMC) algorithm (Falush et al. 2003) and also identify membership of accessions. The admixture model-based simulations were carried out by performing five independent runs for each K (from 1 to 10) with 1,00,000 burn-in period and 1,00,000 MCMC replications. The optimal number of populations was determined by calculating ΔK value (Evanno et al. 2005) through a change in likelihood function with respect to each K.
Results
Morphological analysis
The descriptive statistics results suggest that considerable variation was present among eleven quantitative traits (Table 3). The Pearson correlation was evaluated between nine important quantitative traits and significant positive correlation was observed among fruit traits (Table 4). The selection of such traits will save both time and labor. From the economic point of view, accessions with high yield and a large number of fruits per plant should be selected for future breeding. All morphological traits were found highly polymorphic except five qualitative characters which displayed no marked variation including flower color (yellow), stem shape (angular), peduncle shape (sharply angled), leaf margin (dented) and tendrils (present) therefore, not used in the principal component analysis. The leaf shape showed three morphological classes viz. ovate in 11 accessions, orbicular in 10 accessions and reniform in rest 24 accessions. Green leaf spot was found in the majority of accessions i.e. 21 followed by admixture type spot (16 accessions) and silver leaf spot in 8 accessions. 11 accessions had shallow leaf lobes and intermediate in 24 accessions. Clear variation was found in stem end shape (25 rounded, 14 pointed and 6 flattened) and blossom end shape (19 rounded, 10 pointed and 16 flattened). Highest polymorphism was observed in fruit traits as fruit color varied from green (16), light green (17) to dark green (12); fruit shape varied from elliptical (4), elongated (17) to elongated tapered (25); fruit ribs were absent in 2 accessions, deep in 12, intermediate in 12 and superficial in 19 accessions; fruit skin texture showed 2 morphological variants i.e. 8 were smooth and 37 were grainy.
The principal component analysis (PCA) was performed on 20 morphological characters and the first 7 components which had eigenvalues greater than 1 explained 72.70% of the total morphological variation (Table 5). PCA analysis revealed that traits like blossom end shape, stem end fruit shape, fruit width, fruit weight, seed length, seed weight and fruit yield were major contributors of morphological variations in sponge gourd germplasm. Therefore, the variation observed in these traits could be exploited by plant breeders. Dendrogram was constructed based on Euclidean distance and grouped accessions into two clusters (Fig. 1). Cluster I included six accessions i.e. VRSL12, VRSL13, DSG48, DSG98, DSG38 and NSG-1-11. Majority of these accessions showed orbicular leaf shape, green leaf spots, intermediate leaf lobes, rounded stem end shape and light green colored elongated tapered fruits with grainy texture of fruits. The rest other accessions were included in cluster II. No relationship was found between morphological diversity and geographical origin as admixture type of clustering observed.
ISSR analysis
Based on the reproducibility of scorable bands, 20 ISSR primers were used for characterization of genetic diversity. Amplification allowed the visualization of 182 clear and bright bands and size ranged between 225 and 3000 bp. Out of 182 DNA fragments, 137 were found diverse with an average of 6.85 variable fragments per primer (Table 6). The number of polymorphic fragments generated were 16 (ISSR2 and ISSR14) to 5 bands (ISSR821 and ISSR900). An average polymorphism of 74.6% was recorded. PIC is a good indicator of the primer efficiency in differentiating accessions and depends on the unique banding patterns. Thus, ISSR10 with 0.29 PIC value was the most efficient primer. On determining gene diversity (H), a mean diversity of 0.27 was observed with a range of 0.06–0.42. Another genetic variability parameter is Shannon information index (I) with 0.39 average value. Similarity coefficient value was found 0.81 based on the pair-wise comparison of the accessions. Dendrogram is formed which delineated all accessions into well defined two clusters and one outgroup (Fig. 2). Highest genetic similarity was observed between VRSL1 and DSG31. Cluster I contained two accessions viz. DSG98 (Uttarakhand) and NSG28 (Maharashtra). The second cluster comprised remaining accessions. Pusa Supriya was seen outside the whole dendrogram. Both major clusters were further subdivided into sub-clusters.
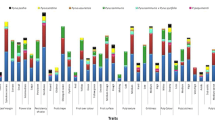
The genetic population structure was investigated using a bayesian model-based clustering to determine the most probable number of clusters in the population. The ΔK was maximum at K = 2 which implied the existence of two sub-populations and revealed admixture type of population (Fig. 3). There was no clear demarcation between accessions according to their geographical origin as observed earlier also in case of UPGMA clustering. This indicates that a clear geographical population structure differentiation could not be identified.
The population structure of sponge gourd based on the optimal value of K for ISSR marker. Each vertical line represents an individual, and the different colors represent populations. The length of the colored segment illustrates the estimated proportion of membership in corresponding clusters as calculated through Structure 2.3.4
SCoT analysis
Out of 80 SCoT primers, 23 were selected for further analysis based on unambiguous and reproducible fragments. A total of 212 DNA fragments were obtained with 151 polymorphic fragments with sizes between 200 and 3200 bp (Table 7). The amplified fragments ranged from 3 (S53) to 23 (25). The detected percentage of polymorphism was 71.5% with a range of 27 (S24) to 100% (S25, S26, S28 and S72). PIC values exhibited by primers were in range of 0.03 (S15 and S53) to 0.33 (S25). H value had a range of 0.11 (S30) to 0.33 (S15) as compared to I value which varied from 0.12 (S59) to 0.53 (S32) indicating lower variation among accessions.
All 212 DNA fragments were subjected for pairwise comparison of the accessions and similarity coefficient was calculated. A dendrogram was created utilizing coefficient values (Fig. 4). The coefficient of 0.94 was recorded between VRSL1 (West Bengal) and DSG7 (Uttar Pradesh) demonstrated the highest similarity. Dendrogram grouped all the accessions into two major clusters. Both major clusters were further subdivided into two sub-clusters. There was little grouping observed according to the geographical region in sub-cluster I of cluster II viz. West Bengal (VRSL1, VRSL2, VRSL3, VRSL5), Uttar Pradesh (DSG7, VRSL6, VRSL12, VRSL8, VRSL7, VRSL13, VRSL9, VRSL10, VRSL11, VRSL14, NDSG1, VRSL15) and Jharkhand (CHSG1 and CHSG2). Thus, results suggest little correlation geographic and genetic distances.
To identify the hidden population structure, the bayesian model-based clustering was performed as it grouped samples into genetically distinguishable groups. Sponge gourd population comprised of two sub-populations as K showed the peak value at 2. A clear pattern of ecological differentiation was not found, but spreading and mixing of the subpopulations was inferred (Fig. 5). Results were not in concordance with the UPGMA clustering. However, the resemblance was inferred between ISSR and SCoT results in both dendrogram and structure analysis.
The population structure of sponge gourd based on the optimal value of K for SCoT marker. Each vertical line represents an individual, and the different colors represent populations. The length of the colored segment illustrates the estimated proportion of membership in corresponding clusters as calculated through Structure 2.3.4
Discussion
Understanding genetic diversity and genetic relationships is necessary for conservation and sustainable management of species (Lynch et al. 1999). It facilitates selection of diverse parental combinations with maximum genetic variability and desirable gene introgression from diverse germplasm into the available genetic base which proved useful in crop breeding programs (Barrett and Kidwell 1998; Thompson et al. 1998).
In the current study, two different molecular marker systems ISSR and SCoT with morphological traits were utilized for genetic diversity estimation within 45 accessions of sponge gourd. The parallel use of these 3 data sets allowed precise assessment of the relationship of different accessions of sponge gourd. In our view, the molecular investigation should correspond with morphological traits analysis because along with phylogeny agronomic traits study is also essential which can be done with morphological analysis (Métais et al. 2000).
Higher morphological variation was recorded in leaf, fruit and seed traits. On performing correlation analysis among quantitative traits, a significant correlation was noticed between fruit and seed traits. Above results suggest the selection of such polymorphic traits for future breeding plans. As the selection of plant with high-quality fruit traits will automatically lead to the selection of plant of superior quality seed traits. Thus, such strategies will prove beneficial from both economical and time point of view. The first seven principal components explained 72.70% of total variation. Thus, selection of accessions which were found diverse in these 7 components could be done. The results were in conformity with earlier analysis where high genetic variability was reported in fruit and seed characters of Indian sponge gourd accessions (Prakash et al. 2013). Leaf traits were considered reliable in the evaluation of genetic relationships and taxonomic studies (Almajali et al. 2012). Hence, such traits could be used as a marker for ascertaining genetic relationships between accessions and distinguishing accessions.
Both molecular markers showed comparable genetic diversity values but a higher level of polymorphism represented by ISSR. Differences were noticed in genetic similarity coefficient values and PIC values. Therefore, we can say that the variation pattern is clearly controlled by the genetic marker used. According to Karp and Edwards (1995), different markers had different characteristics and may reveal different features of genetic variability. Variation in the chromosomal location of the markers has also influenced the diversity pattern (Kojima et al. 1998). Overall, the genetic background of sponge gourd is inferred narrow in the present study. These results are in agreement with other studies, which concluded moderate genetic variation in sponge gourd (Tyagi et al. 2016; Marr et al. 2005). But incongruence was found with others who reported high genetic diversity (Rathod et al. 2015; Prakash et al. 2014; Junhui and Changping 2008). The reasons can be a single place of domestication or use of few key varieties for hybridization (Marr et al. 2005; Ghaffari et al. 2014). Other reasons can be difference in the number of primers, type of plant variety used (Hajibarat et al. 2015).
Dendrogram and population structure analysis did not demonstrate the existence of a definite pattern of relationships amid ecological origin and genetic variability as the accessions from same geographical regions did not fall exclusively in a single or two clusters. However, little grouping was seen in 2–3 subclusters. This could be due to the difference in nature of each technique as markers designed from various regions of the genome, genome region coverage by each marker, polymorphism and loci number (Gorji et al. 2011; Pakseresht et al. 2013; Souframanien and Gopalakrishna 2004). Hence, it becomes crucial to broadening the genetic base of the sponge gourd to increase the yield. This can be achieved by introgression of genes from wild species which possess enormous variability.
In brief, the investigation had suggested considerable diversity and provides a pathway for future crop variety identification and conservation. In addition, SCoT and ISSR markers in being economical, fast and informative can be used in selecting diverse parents for crop improvement programs. SCoT marker is a gene-targeted marker which can be converted into functional markers after sequencing and would be more useful for variability analysis and linkage maps construction.
Abbreviations
- UPGMA:
-
Un-weighted pair group method with arithmetic average
- ISSR:
-
Inter simple sequences repeat
- PIC:
-
Polymorphism information content
- PCA:
-
Principal component analysis
- SCoT:
-
Start codon targeted
- PCR:
-
Polymerase chain reaction
References
Almajali D, Abdel-Ghani AH, Migdadi H (2012) Evaluation of genetic diversity among Jordanian fig germplasm accessions by morphological traits and ISSR markers. Sci Hort 147:8–19
Arcade A, Anselin F, Rampant PF, Lesage MC, Paques LE, Prat D (2000) Application of AFLP, RAPD and ISSR markers to genetic mapping of European and Japanese larch. Theor Appl Genet 100:299–307
Azeez MA, Bello OS, Adedeji AO (2013) Traditional and medicinal uses of Luffa cylindrica: a review. J Med Plant Stud 1:102–111
Barrett BA, Kidwell KK (1998) AFLP-based genetic diversity assessment among wheat cultivars from the Pacific northwest. Crop Sci 38:1261–1271
Biabani A, Rafii MY, Saleh GB, Latif MA (2013) Inter– and intra–population genetic variations in Jatropha curcas populations revealed by inter-simple sequence repeat molecular markers. Maydica 58:111–118
Carvalho MA (2004) Germplasm characterization of Arachis pintoi Krap. and Greg. (Leguminosae). Ph.D. Thesis, University of Florida, USA
Collard BCY, Mackill DJ (2009) Start Codon Targeted (SCOT) polymorphism: a simple novel DNA marker technique for generating gene-targeted markers in plants. Plant Mol Biol Rep 27:86–93
de Vicente MC, Guzmán FA, Engels J, Rao VR (2005) Genetic characterization and its use in decision making for the conservation of crop germplasm. The role of biotechnology. In: International workshop the role of biotechnology for the characterization and conservation of crop, forestry, animal and fishery genetic resources. FAO, FOBIOTECH, ECOGENE, SIGA, Turin, pp 121–128
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14:2611–2620
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure: extensions to linked loci and correlated allele frequencies. Genetics 164:1567–1587
Fatehi R, Talebi R, Fayyaz F (2011) Characterization of Iranian land-race wheat accessions by inter simple sequence repeat (ISSR) markers. J Appl Env Biol Sci 1(10):423–436
Ghaffari P, Talebi R, Keshavarzi F (2014) Genetic diversity and geographical differentiation of Iranian landrace, cultivars, and exotic chickpea lines as revealed by morphological and microsatellite markers. Physiol Mol Biol Plants 20:225–233. https://doi.org/10.1007/s12298-014-0223-9
Gorji AM, Poczai P, Polgar Z, Taller J (2011) Efficiency of arbitrarily amplified dominant markers (SCOT, ISSR and RAPD) for diagnos-tic fingerprinting in tetraploid potato. Am J Potato Res 88:226–237
Hajibarat Z, Saidi A, Hajibarat Z, Talebi R (2015) Characterization of genetic diversity in chickpea using SSR markers, start codon targeted polymorphism (SCoT) and conserved DNA-derived polymorphism (CDDP). Physiol Mol Biol Plants 21:365–373. https://doi.org/10.1007/s12298-015-0306-2
Hoogendijk M, Williams DE (2001) Characterizing the genetic diversity of home garden crops: some examples from the Americas. In: Watson JW, Eyzaguirre PB (eds) Proceedings of the second international home gardens workshop: contribution of home gardens and in situ conservation of plant genetic resources in farming systems, IPGRI, Germany, pp 34–40
Hoque S, Rabbani MG (2009) Assessment of genetic relationship among landraces of Bangladeshi ridge gourd (Luffa acutangula Roxb.) using RAPD markers. J Sci Res 1:615–623
Joshi BK, Hari BKC, Tiwari RK, Ghale M, Sthapit BR, Upadhyay MP (2004) Descriptors for sponge gourd (Luffa cylindrica). NARC, LIBIRD and IPGRI
Jun L, Mei-rong X, Zhi-wei Z, Wen-bo L, Huo-lin S (2010) Genetic diversity analysis of Luffa accessions by SSR and SRAP markers. China Cucurbits Veg 23:1–4
Junhui X, Changping X (2008) Analysis of genetic diversity in Luffa via morphological and RAPD markers. China Veg 10:21–25
Kalloo G (1993) Loofah-Luffa spp. In: Kalloo G, Bergh BO (eds) Genetic improvement of vegetable crops. Pergamon Press, Oxford, pp 265–266
Karp A, Edwards KJ (1995) Molecular techniques in the analysis of the extent and distribution of genetic diversity. In: IPGRI workshop on molecular genetic tools in Plant genetic resources 9–11 Oct, Rome, IPGRI
Kojima T, Nagaoka T, NodaK OY (1998) Genetic linkage map of ISSR and RAPD markers in einkorn wheat in relation to that of RFLP markers. Theor Appl Genet 96:37–45
Lee S, Yoo JG (2006) (Wo/2006/019205) method for preparing transformed Luffa cylindrica Roem. (world intellectual property organization). http://www.wipo.int/pctdb/en/wo.jsp?IA=KR2004002745&DISPLAY=STATUS. Accessed 27 July 2014
Luo C, He XH, Chen H, Ou SJ, Gao MP (2010) Analysis of diversity and relationships among mango cultivars using start codon targeted (SCoT) markers. Biochem Syst Ecol 38:1176–1184
Lynch M, Pfrender M, Spitze K, Lehman N, Hicks J, Allen D, Latta L, Ottene M, Bogue F, Colbourne J (1999) The quantitative and molecular genetic architecture of a sub-divided species. Evolution 53:100–110
Marr KL, Bhattarai NK, Xia YM (2005) Allozymic, morphological, and phenological diversity in cultivated Luffa acutangula (Cucurbitaceae) from China, Laos, and Nepal, and allozyme divergence between L. acutangula and L. aegyptiaca. Econ Bot 59:154–165
Métais I, Aubry C, Hamon B, Jalouzot R (2000) Description and analysis of genetic diversity between commercial bean lines (Phaseolus vulgaris L.). Theor Appl Genet 101:1207–1214
Misra S, Srivastava AK, Verma S, Pandey S, Bargali SS, Rana TS, Nair KN (2017) Phenetic and genetic diversity in Indian Luffa (Cucurbitaceae) inferred from morphometric, ISSR and DAMD marker. Genet Resour Crop Evol 64:995–1010
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3324
Pakseresht F, Talebi R, Karami E (2013) Comparative assessment of ISSR, DAMD and SCoT markers for evaluation of genetic diversity and conservation of landrace Chickpea (Cicer arietinum L.) genotypes collected from North-West of Iran. Physiol Mol Biol Plants 19(4):563–574
Pandey S, Ansari WA, Choudhary BR, Pandey M, Jena SN, Singh AK, Dubey RK, Singh B (2017) Microsatellite analysis of genetic diversity and population structure of hermaphrodite ridge gourd (Luffa hermaphrodita). 3 Biotech 8:17. https://doi.org/10.1007/s13205-017-1030-0
Papanicolaou GC, Psarra E, Anastasiou D (2015) Manufacturing and mechanical response optimization of epoxy resin/Luffa cylindrica composite. J Appl Polym Sci 132:41992. https://doi.org/10.1002/app.41992
Prakash K, Pandey A, Radhamani J, Bisht IS (2013) Morphological variability in cultivated and wild species of Luffa (Cucurbitaceae) from India. Genet Resour Crop Evol 60:2319–2329. https://doi.org/10.1007/s10722-013-9999-7
Prakash K, Pati K, Arya L, Pandey A, Verma M (2014) Population structure and diversity in cultivated and wild Luffa species. Biochem Syst Ecol 56:165–170. https://doi.org/10.1016/jbse.2014.05.012
Pritchard JK, Stephens P, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rathod RR, Mehta DR, Gajera HP, Delvadiya NA (2015) Molecular characterization of ridge gourd (Luffa acutangula L.) and sponge gourd (Luffa cylindrica L.) genotypes through PCR based molecular markers. Int J Agric Env Biotech 8:521–530. https://doi.org/10.5958/2230-732X.2015.00059.5
Reddy MP, Sarla MP, Siddiq EA (2002) Inter simple sequence repeat (ISSR) polymorphism and its application in plant breeding. Euphytica 128:9–17
Rohlf FJ (2000) NTSYS-PC, Numerical taxonomy and multivariate analysis system, version 2.1. Exeter Publications, New York
Saghai-Maroof MA, Soliman KM, Jorgesm RA, Allard RW (1984) Ribosomal DNA spacer length in barley: mendelian inheritance, chromosomal location and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Sankar AA, Moore GA (2001) Evaluation of inter–simple sequence repeat analysis for mapping in Citrus and extension of genetic linkage map. Theor Appl Genet 102:206–214
Shannon C, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Urbana
Sirohi PS, Munshi AD, Kumar G, Behera TK (2005) Cucurbits. In: Dhillon BS, Tyagi RK, Saxena S, Randhawa GJ (eds) Plant genetic resources: horticultural crops. Narosa Publishing House, New Delhi, pp 34–58
Souframanien J, Gopalakrishna T (2004) Acomparative analysis of genetic diversity in black gram genotypes using RAPD and ISSR markers. Theor Appl Genet 109:1687–1693
Talebi R, Fayaz R, Mardi M, Pirsyedi SM, Naji AM (2008) Genetic relationships among chickpea (Cicer arietinum) elite lines based on RAPD and agronomic markers. Int J Agric Biol 8:301–305
Thompson JA, Nelson RL, Vodkin LO (1998) Identification of diverse soybean germplasm using RAPD markers. Crop Sci 38:1348–1355
Tyagi R, Sharma V, Sureja AK, Munshi AD, Arya L, Verma M (2016) Deciphering population structure and diversity in Luffa cylindrica (L.) M. Roem. using morphological and sequence-related amplified polymorphism markers. Plant Genet Resour 14:234–237. https://doi.org/10.1017/S1479262115000258
Yawalkar KS (2004) Cucurbitaceous or vine crops, V ed. Vegetable Crops of India, Nagpur, pp 152–155
Yeh FC, Boyle TJB (1997) Population genetic analysis of codominant and dominant markers and quantitative traits. Belg J Bot 129:157
Acknowledgements
We are grateful to the Head, Division of Genomic Resources and the Director, National Bureau of Plant Genetic Resources for providing access to the facilities.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tyagi, R., Sharma, V., Sureja, A.K. et al. Genetic diversity and population structure detection in sponge gourd (Luffa cylindrica) using ISSR, SCoT and morphological markers. Physiol Mol Biol Plants 26, 119–131 (2020). https://doi.org/10.1007/s12298-019-00723-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-019-00723-y