Abstract
Morphological traits and three molecular markers techniques: start codon targeted (SCoT), inter-simple sequence repeat (ISSR) and directly amplified minisatellite DNA (DAMD) markers were compared for fingerprinting of 40 landraces chickpea genotypes collected from different geographical locations of north-west of Iran. Variance analysis of ten measured morphological traits showed significant differences existed between genotypes. Cluster analysis based on morphological traits, divided genotypes in three distinct clusters. Average polymorphism information content (PIC) for ISSR, DAMD and SCoT markers was 0.216, 0.232 and 0.232, respectively, and this revealed that SCoT markers were more informative, followed by ISSRs marker, than other markers for the assessment of diversity amongst genotypes. Cluster analysis for three different molecular types revealed that genotypes taken for the analysis can be divided in three and four distinct clusters. Accessions from same geographical regions mostly showed more genetic similarities than those from origins far isolated apart. These results suggest that efficiency of SCOT, DAMD and ISSR markers was relatively the same in fingerprinting of genotypes but SCOT and DAMD analysis are more effective in fingerprinting of chickpea genotypes. To our knowledge, this is the first detailed report of a comparison of performance among two targeted DNA region molecular markers (SCoT and DAMD) and the ISSR technique on a set of samples of chickpea. Overall, our results indicate that SCOT, ISSR and DAMD fingerprinting could be used to detect polymorphism for genotypes of chickpea.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The studies on genetic diversity was carried out in the region located between South east Turkey and north-west of Iran and the adjoining areas of Syria which has been proposed to be the Vavilovian center of origin for the cultivated chickpea (Cicer arietinum L.) (van der Maesen 1987; Talebi et al. 2008a). India and Ethiopia have been proposed as secondary centers of diversity of cultivated chickpea (Harlan 1992). However, Zeven and de Wet (1982) suggested that chickpea has different secondary centers of diversity located in at least four regions; the Near East Region (comprising the Fertile Crescent), Hindustani Region (basically the current India and East Pakistan), Central Asian Region (with Afghanistan, Western Pakistan, Iran and the south of the former USSR) and the Mediterranean Region (including Lebanon and Palestine). Assessment of the extent of genetic variability within chickpea is fundamental and important for chickpea breeding and conservation of genetic resources, and is particularly useful as a general guide in the choice of parents for breeding hybrids (Talebi et al. 2008b). Today it is realized that the use of genetically different varieties is an effective strategy in order to minimized genetic vulnerability (Fatehi et al. 2011). Depending on their geographical regions, landraces had specific genetic background that can be used in genetic research program (Harlan 1975). In addition, landraces are important genetic resources that improve gene pools of modern cultivars by introducing new alleles (Nevo and Payne 1987). Criteria for the estimation of the genetic diversity can be different, which include morphological traits, biochemical or molecular markers (Carvalho 2004; de Vicente et al. 2005). However, it is generally believed that the use of molecular markers is more reliable and repeatable as compared to characterization based on morphological characters (Carvalho 2004). A number of DNA based markers are now available for the effective quantification of genetic variation in plant species. Collectively, techniques such as AFLP, ISSR and RAPD, have been termed as arbitrarily amplified dominant (AAD) markers (Wolfe and Liston 1998) and are increasingly used for diagnostic genomic fingerprinting, genetic and qualitative trait loci (QTL) mapping and population genetic studies in chickpea (Iruela et al. 2002; Talebi et al. 2008a,b; Mahmood et al. 2011). More recently, the microsatellite sequences have been used as primers in polymerase chain reaction, where repeat motifs are anchored either at 5′ or 3′ end with one or few specific nucleotides and amplify the sequences between the two microsatellite loci referred to as inter simple sequence repeat (ISSR) markers. In addition, ISSRs can be targeted towards particular sequences, which are reported to be abundant in the genome and can overcome the technical difficulties of RFLP and RAPD (Rajesh et al. 2002). In recent years, many new alternative and promising marker techniques have been developed in line with the rapid growth of genomic research (Gupta and Rustgi 2004). With initiating a trend away from random DNA markers towards gene-targeted markers, a novel marker system called Start codon targeted (SCoT) (Collard and Mackill 2009) was developed based on the short conserved region flanking the ATG start codon in plant genes. SCoT markers are generally reproducible, and it is suggested that primer length and annealing temperature are not the sole factors determining reproducibility. Minisatellite DNAs are tandemly repeated regions of genomes, many of which show high levels of length differences due to variations in the number of repeat units (Jeffreys et al. 1985). This technique was first reported by Heath et al. (1993) and termed as directed amplification of minisatellite-region DNA (DAMD). It is speculated that minisatellites and the DNA sequences flanking them are involved in inversions, which results in their distribution on both strands in opposite orientations. From a few morphological, AFLP and RAPD marker-based studies in Iran, it has been reported that there exists high morphological but low biochemical and molecular diversity (Talebi et al. 2008a; Saeed et al. 2011; Naghavi and Jahansouz 2005). Another study was also conducted based on SSR markers but was limited to a few Iranian landrace accessions and additional varieties and wild spp., from other countries (Saeed et al. 2011). Here, the use of SCoT polymorphism and DAMD marker based techniques for studying genetic diversity was reported for the first time in chickpea genotypes. Objectives of the present study are as follows: (1) to determine the potential of these methodologies to generate polymorphic markers in chickpea; (2) to investigate whether SCoT and DAMD markers could be effectively used in determining genetic relationships among chickpea genotypes compared to ISSR markers data; and (3) to identify the relationships of different types of molecular fingerprinting with morphological-based clustering of genotypes.
Materials and methods
Plant materials and field evaluation
Forty landraces chickpea (Cicer arietinum L.) accessions from different geographical location of Kurdistan province, North-west of Iran, were investigated for the study of genetic variation both at morphological and molecular level (Table 1, Fig. 1). All chickpea landraces were sown on 15 February 2010, in well-prepared seed beds, using randomized block design with three replications. Seeds were hand drilled and each genotype was sown in three rows of 4 m, with row to row distance of 0.40 m. The trials were hand-weeded twice and were grown under rain-fed conditions. The grain yields (g m−2) were measured by harvesting each plot at crop maturity. Six plants were randomly chosen from each plot to measure the number of seeds per plant, number of pods per plant, Plant height, number of primary and secondary branches and 100-seed weight. For plant biomass, three randomly selected plants from each plot at crop maturity were weighted for dry plant biomass. Flowering time were recorded when 50 % of plants in each plant flowered. Leaf relative water content (RWC) was determined according to the methods of Barrs and Weatherley (1962), based on the following equation:
where FW is leaf fresh weight, DW is dry weight of leaves after drying at 85 °C for 3 days, and SW is the turgid weight of leaves after soaking in water for 4 h at room temperature (approximately 20 °C). Half of the third (from the top) fully expanded leaf was used.
Genomic DNA extraction
DNA was extracted from 2 g of young leaves collected from 10 days old seedling plants of each accession using the CTAB method (Lassner et al. 1989) with the modification described by Torres et al. (1993). The final concentration of DNA was determined by agarose-gel electrophoresis using a known concentration of uncut λ DNA as a standard.
ISSR and DAMD markers analysis
For ISSR analysis, a set of 10 primers representing di, tri, tetra and pentamer repeats (UBC set # 9) was procured from the Biotechnology Laboratory, University of British Columbia, Canada (Table 2). Following the optimization of PCR conditions and pre-screening of the first 10 primers on genotypes, seven primers providing clear and informative amplicon profiles across species were selected to survey ISSR variation in the accessions (Table 5). DAMD-PCR markers (originally derived from the repeat elements of weed rice) (Kang et al. 2002) were used in this study. Out of the ten primers screened (Table 2), seven with a GC content of 50–60 % were selected to generate the DNA fingerprint profiles of all the accessions at 48 °C annealing temperature (Table 5). PCR amplification was performed in 20 μl reaction containing 1× PCR buffer, 30 ng sample DNA, 2.5 μM primer, 200 μM of each dNTP, 1.5–2.5 mM MgCl2 and 1.5 unit of Taq DNA polymerase (Cinnagene, Iran). All amplification were carried out in a Eppendorf thermocycler as follows: 94 °C for 3 min, followed by 35 cycles of denaturation at 93 °C for 45 s, annealing at optimum Tm for 45 s, and extension at 72 °C for 90 s. A final extension cycle at 72 °C for 10 min followed. PCR products were separated on 1.2 % agarose gels, stained with ethidium bromide and scored for the presence or absence of bands.
SCoT PCR
Primer sequences employed in the present study were designed by Collard and Mackill (2009) based on the consensus sequences of translation initiation codon region in higher plants with ATG codon at positions +1, +2, +3; ‘G’ at position +4; and ‘A,’ ‘C,’ and ‘C’ at positions +7, +8, and +9, respectively (Table 2). PCR was optimized for 15 primers as described previously by Collard and Mackill (2009). PCR amplification was performed in 20 μl reaction containing 1× PCR buffer, 50 ng sample DNA, 2.5 μM primer, 200 μM of each dNTP, 3 mM MgCl2 and 1.5 unit of Taq DNA polymerase (Cinnagene, Iran). All amplification were carried out in a Eppendorf thermocycler as follows: 94 °C for 3 min, followed by 35 cycles of denaturation at 93 °C for 1 min, annealing at 48 °C for 1 min, and extension at 72 °C for 2 min. A final extension cycle at 72 °C for 10 min was followed. PCR products were separated on 1.2 % agarose gels, stained with ethidium bromide and scored for the presence or absence of bands.
Data analysis
For morphological traits, variance analysis for grain yield and other related characteristics were performed over trails after verifying the homogeneity of trail variance error using Bartlett’s test. Least significant difference (LSD) values were calculated at the 5 % probability level. The SAS (SAS Institute, Inc. 1996) procedures and programs were used for these calculations. Amplified bands obtained with all the molecular markers were scored visually for the presence (1) and absence (0) of bands for all the 40 accessions. Nei,s genetic distance (Nei 1973) was determined among the genotypes and used for grouping of the genotypes by UNJ (Un-weighted Neighbor Joining) cluster method (Perrier et al. 2003). The fit of dendrograms obtained were checked by bootstrapping using 100 replications. NTSYS ver 2.02 (Rohlf 1998) and DARwin ver 5.0 (Perrier and Jacquemoud-Collet 2006) were used for clustering. Mantel statistic was used to compare the dissimilarity matrices as well as the dendrograms produced by the ISSR, DAMD and SCoT techniques through NTSYS software. Polymorphic information content (PIC) values were calculated for each ISSR, DAMD and SCoT primers according to the formula:
where Pij is the frequency of the ith pattern revealed by the jth primer summed across all patterns revealed by the primers (Botstein et al. 1980).
Results
Diversity of morphological and agronomic traits
In variance analysis, the genotypic variance was significant for eight of the ten observed traits (Table 3). This indicated that differences existed between the accessions for these traits. Genotypes show high variation for number of pods per plant, days to flowering, number of secondary and primary branches. The morphological traits measured and their ratios were variable. There were significant differences among accessions (ANOVA, P < 0.05) for ten measured traits (Table 4). The traits like number of pods per plant, number of seeds per plant and biomass showed higher range of variation compared to other traits (Table 4). Forty accessions were grouped into three clusters (Fig. 2). Cluster I included seven genotypes that divided in to two sub-clusters. These genotypes originated from different zones that showed moderately high plant height, 100-seed weight and consequently moderate high yield. Sub-clusters of cluster II obtained 6 genotypes. Genotypes included in this cluster showed high yield compared to other clusters and most of them originated from ‘Sanandaj’ and ‘Marivan’. Remaining genotypes were grouped in cluster III. Genotypes belonged to this cluster showed moderately high 100-seed weight, plant height and seed yield. In general, genotypes included in cluster II and III showed close distance with each other and significantly separated from cluster I (Fig. 2).
Molecular markers analysis
DNA fingerprint database has been reported using the three different PCR-based molecular markers (ISSR, DAMD and SCoT) systems for 40 landraces chickpea accessions belonged to different geographical areas of Kurdistan province, north-west of Iran. Our results indicated that primers which were obtained from the different regions of genomic DNA, successfully amplified accessions template DNAs. All the three molecular markers used in this study were able to distinguish and identify each of 40 accessions to referred geographic region. Salient features of fingerprint database obtained using different markers are given below:
ISSR analysis
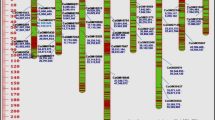
ISSR analysis revealed a distinct scorable fragment per primer and in total, 49 bands, both polymorphic and monomorphic were obtained using seven primers in 40 chickpea genotypes (Table 5). The number of amplified fragments varied from 5 to 11, with an average 7 bands per primer. Out of 49 bands, 32 bands were polymorphic and the polymorphism percentage averaged to 0.65 across all the genotypes. Maximum number of polymorphic bands (7 of the 11 bands) were obtained for UBC807. The average polymorphic information content (PIC) was 21.6, ranging from 0.019 to 0.311. The highest PIC value obtained by UBC878 (0.311) (Table 5). From the dendrogram constructed by DARwin using un-weighted neighbour joining method, it was discernible in to three major clusters (Fig. 3). Cluster I and III contained maximum number of genotypes. Some of genotypes in these clusters were grouped close to each that observed in morphological cluster analysis, but relatively low homology between these clusters and morphological-based cluster were observed. Cluster I mostly contained genotypes from Sanandaj, while most of the genotypes that originated from Qorveh grouped in cluster III. Cluster II contained genotypes for both Sanandaj and Marivan (Fig. 3).
DAMD analysis
A polymorphic chain reaction (PCR) based approach involving the direct amplification of minisatellite region DNA (DMAD) with seven monisatellite core sequences as primers was used to diversity analysis in 40 chickpea genotypes. Analysis indicated that the PCR profile and the optimized chemical concentrations resulted in reproducible and reliable DNA amplification. In the chickpea genotypes the number of amplified DAMD products varied from 8 to 12 fragments depending on the primers used. Total number of bands scored was 71 of which 49 were polymorphic (Table 5). PIC values ranged from 0.087 to 0.337, with an average value of 0.232 per locus. Cluster analysis grouped chickpea genotypes in four clusters (Fig. 4). Cluster I and IV contained maximum number of genotypes. Cluster II included genotypes that all of them originated from ‘Sanandaj’ and these genotypes showed moderately low yield potential compare to genotypes belonged to other clusters. Cluster III contained six genotypes that mostly originated from ‘Marivan’ and this cluster showed similar grouping pattern with those obtained by ISSR data.
SCoT Analysis
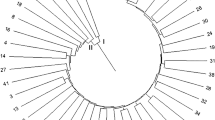
A set of 15 SCoT primers were used to fingerprint 40 chickpea accessions. Only primers that exhibited unambiguous and reproducible band patterns were selected for further analysis. Thus, a total of 10 primers that exhibit distinct and reliable band patterns were utilized for bands scoring, following genetic similarity analysis and cluster analysis. A total of 95 bands were detected among 40 chickpea genotypes using 10 SCoT markers out of which 65 were polymorphic (Table 5). Number of bands were ranged from 5 (SCoT35) to 14 (SCoT2) with an average 9.5 per primers. Percent polymorphism ranged from 33 % to as high as 100 % with an average polymorphism of 68 % across all accessions. PIC values ranged from 0.142 to 0.366, with an average value of 23.2 per primer (Table 5). Based on un-weighted neighbour-joining method, a dendrogram demonstrating the genetic relationships among the chickpea accessions were constructed. The 40 chickpea accessions fell under three major groups (Fig. 5). Cluster II and III divided into two sub-clusters. Cluster I contained maximum numbers of genotypes and showed relatively similar grouping pattern with cluster I that obtained by ISSR and DAMD markers. Similarity between genotypes clustering in SCoT analysis and morphological based clustering was relatively higher than those obtained by ISSR and DAMD markers. This finding also observed with relatively higher coefficient similarity obtained by mantel test (Table 6). The general dendrogram (Fig. 6) that was constructed using the combined data of the three sets of molecular markers divided in three distinct clusters was relatively similar to those obtained separately with each marker. The dendrogram from ISSR and SCoT data was most congruent with the general dendrogram.
Correlation between the similarity values measured using three marker systems
Cophenetic coefficient were acceptable in all three molecular markers systems (ISSR = 0.89; SCoT = 0.84 and DAMD = 0.81) indicating good fit for clustering. The values of mantel test correlation showed a positive correlation between the three marker types. The correlation coefficient (r) was 0.53 between ISSR and DAMD (significant P > 0.05), 0.42 between ISSR and SCoT (significant P > 0.05) and 0.48 (significant P > 0.05) between DAMD and SCoTs. All three molecular marker types showed positive but non-significant correlation with morphological cluster (Table 6). In order to estimate the genetic distance among genotypes, the dissimilarity matrix was computed. The mean value of genetic distance obtained by ISSR, DAMD and SCoT markers were 0.42, 0.44 and 0.37, respectively.
Discussion
Although chickpea has a narrow genetic base, with the discovery of diverse molecular markers from different targets regions of the genome, it is now possible to conduct extensive molecular diversity study in this important crop to identify genetically diverse germplasm with beneficial traits for use in crop improvement programs. The assessment of genetic diversity is important not only for crop improvement but also for efficient management and conservation of germplasm resources. DNA-based molecular markers such as random amplified polymorphic DNA (RAPD; Iruela et al. 2002; Talebi et al. 2008a) and RFLP (Powell et al. 1996) were unable to address the reliable genetic variation within chickpea (Saeed et al. 2011). Genetic fingerprinting in chickpea, for a long time hampered by the little variability in chickpea’s genome, can be facilitated by highly polymorphic functional markers such as SCoT and DAMD. In this study, we have compared the marker data-sets produced using morphological and three different marker systems, ISSR, DAMD and SCoT, to define genetic relationships within a set of 40 genotypes representing different geographical areas of north-west of Iran, and to know if these marker systems can be effectively used in breeding programme. We found relatively acceptable genetic diversity within available Iranian landrace chickpea accessions which is contrary to the findings of Iruela et al. (2002), who reported low level of genetic diversity within C. arietinum compared to the wild species. In present study, the genotypes were selected based on the presence or absence of field resistance to Ascochyta blight, cold (data not presented), drought and some morphological traits and differences in source of collection. The relationship observed using molecular markers may provide information on the history and biology of genotypes, but it does not necessarily reflect what may be observed with respect to agronomic traits (Métais et al. 2000). Previous studies have shown that the measurements of genetic divergence obtained from morpho-agronomical traits are not completely similar to molecular-based results (Talebi et al. 2008a; Saeed et al. 2011). We found that there was no strong relationship between morphological divergence and geographical origins because accessions from one place entered into more than one cluster. Conversely, accessions from different geographical origins were relatively unique and tend to be clustered in one part of the dendrogram, which information suggests that the morphological variation analyzed is determined not only by environmental differences but also by genetic factors. In our analysis, we were able to define groups of accessions (cluster I and cluster II) that were significantly different from each other for characters of interest. These finding are in agreement with previous studies on Iranian chickpea accessions using morphological traits by Naghavi and Jahansouz (2005). Considerable genetic variation which exists between chickpea genotypes in present study can be used efficiently for gene tagging and genome mapping of crosses to introgress the favorable traits such as high yield potential into the cultivated genotypes. Characters such as 100-seed weight, days to flowering and plant height showed low CVs, which appears to limit the scope of selection for these traits in the present accessions of chickpea. Hence, other sources of chickpea are needed in order to find high variation for these traits. A high level of polymorphism (146 polymorphic bands) were detected using 7 ISSR, 7 DAMD and 10 SCoT markers, with an average of 4.57, 7 and 9.5 bands per primer, respectively. These results suggested the presence of a considerable polymorphism at studied molecular markers and revealed a high level of genetic diversity in the existing chickpea germplasm (Table 5). Base on Mantel test results, the rate of genetic diversity for genotypes was nearly the same using SCoT, ISSR and DAMD markers (Table 6). Although the rate of diversity for the three marker techniques was approximately equal; we anticipate that the source of detected diversity is different, as each technique targets different regions of the genome. This indicates that the relative genetic diversity of the genotypes is similar when SCoT, ISSR and DAMD markers are used. The slightly lower level of correlation between these markers in the present study could probably reflect that these markers are known to target different genomic fractions involving repeat and/or unique sequences, which may have differentially evolved or been preserved during the course of natural or artificial selection. In this study, the average genetic distance obtained by ISSR, DAMD and SCoT (0.42, 0.44 and 0.37, respectively) was the same and this relatively high genetic dissimilarity among studied genotypes indicating the present of a considerable genetic diversity in these genotypes which can be considered for future parental selection in breeding programme. We may use more distant genotypes for future hybridization to improve the degree of genetic polymorphism available in Iran. The clustering pattern obtained with each type of markers showed some common groups and clustered some of the genotypes according to the location where they were collected. This can be helpful during strategy development for germplasm collections and evaluations. SCoT and DAMD markers showed more percent polymorphism and higher PIC value than ISSR markers. These techniques were more informative than previously used biochemical and molecular methods to study variation and genetic relationships in chickpea such as isozymes, storage proteins (Ahmad and Slinkard 1992; Labdi et al. 1996), AFLP markers (Talebi et al. 2008b) and RAPD markers (Talebi et al. 2008a). Based on the independent replications of DAMD, we observed that reproducible DNA markers were amplified and also noted that all DAMD primers used in this study produced RAPD like results but the numbers of bands were sharp and clear. The relatively high PCR stringencies in DAMD application effectively limited the PCR artifacts which commonly occur in RAPDs (Karaca et al. 2002; Ince et al. 2009). It was evident from the results that the dendrogram based on molecular markers was not completely in accordance with the dendrogram based on morphological traits (Table 6), as reported for other crops (Fernandez et al. 2002; Talebi et al. 2008b; Maric et al. 2004). There are several possible explanations for such results: some of them connected with nature and structure of different molecular markers that designed from various regions of genome. Another problem was the possibility of overestimating genetic similarity because fragments with the same size could have different origins (Talebi et al. 2008a). Our results demonstrate that high genetic diversity exists between the investigated accessions belonged to different and relatively close geographical locations. As the genotypes examined are diploid and they have relatively similar background, this range of diversity is interesting. SCoT, ISSR and DAMD generated high number of polymorphic markers which can be used in diagnostic fingerprinting studies of chickpea. Based on the average percentage polymorphism and PIC, the efficiency of SCoT and DAMD for fingerprinting of genotypes was more than ISSR marker. The efficiency of SCoT, ISSR and DAMD markers for fingerprinting of genotypes is relatively the same. In general, these three techniques could be used in conjunction with each other for diagnostic fingerprinting of chickpea. The present study showed existence of high genetic diversity in Iranian landrace chickpea germplasm accessions. The magnitude and pattern of genetic variation detected in this study can be useful for more systematic germplasm management and utilization in breeding programs (Tanya et al. 2011). The exploitation of crosses between genetically distant parents (e.g., recombinants parents from the local accessions and the introduced genotypes) and those from diverse local sources may produce higher heterosis, better genetic recombination and segregation in their progenies and result in varieties with broad genetic base (Chahal and Gosal 2002). The selection of genotypes for this study is primarily based on different geographic origin and also the important role of north-west of Iran in chickpea production for poor farmers. Therefore, we believe that there is a need for molecular markers based studies as a complementary study for the morphological traits in the field. It will reduce the amount of materials for study as well as the costs of experiments. Future germplasm collection and utilization strategies should take into consideration the magnitude and pattern of genetic diversity established by the present investigation. It is expected that when such diverse genotypes are involved in breeding programs, as a result of reshuffling of the alleles due to recombination, there are better chances for the appearance of transgressive segregation with beneficial traits that can be selected to extract high yielding lines with desirable trait combination (Sofalian et al. 2008). This study has implications not just for the origin of this crop, but also for the management of genetic resources and their uses in applied breeding programmes, particularly for the development of a core collection. Information about current genetic diversity permits the classification of our available germplasm into various/heterotic groups, which is particularly important to hybrid/cross-breeding programs in chickpea. The previous crossing programs in most of the research institutes of Iran were based on only phenotypic characters. The current study confirmed the importance of molecular studies (cheap, fast and informative markers) beside the morphological data in detecting genetic variation among genotypes in selecting diverse parents to carry out a new crossing program successfully.
Abbreviations
- DAMD:
-
Directly amplified minisatellite DNA
- ISSR:
-
Inter simple sequence repeat
- PCR:
-
Polymerase chain reaction
- PIC:
-
Polymorphism information content
- SCoT:
-
Start codon targeted
References
Ahmad F, Slinkard AE (1992) Genetic relationships in the genus Cicer L. as revealed by polyacrylamide gel electrophoresis of seed storage proteins. Theor Appl Genet 84:688–692
Barrs HD, Weatherley PE (1962) A re-examination of the relative turgidity techniques for estimating water deficits in leaves. Aust J Biol Sci 15:413–428
Botstein D, With RL, Skolnick M, Davis RW (1980) Construction of genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Carvalho MA (2004) Germplasm characterization of Arachis pintoi Krap. and Greg. (Leguminosae). PhD Thesis, University of Florida, USA.
Chahal GS, Gosal SS (2002) Principles and procedures of plant breeding: biotechnological and conventional approaches. Narosa, New Delhi
Collard BCY, Mackill DJ (2009) Start Codon Targeted (SCOT) polymorphism: a simple novel DNA marker technique for generating gene-targeted markers in plants. Plant Mol Biol Rep 27:86–93
de Vicente MC, Guzmán FA, Engels J, Rao VR (2005) Genetic characterization and its use in decision making for the conservation of crop germplasm. The role of biotechnology. In: International workshop the role of biotechnology for the characterization and conservation of crop, forestry, animal and fishery genetic resources. FAO, FOBIOTECH, ECOGENE, SIGA, Turin, pp 121–128
Fatehi R, Talebi R, Fayyaz F (2011) Characterization of Iranian landrace wheat accessions by inter simple sequence repeat (ISSR) markers. J Appl Env Biol Sci 1(10):423–436
Fernandez MN, Figueiras AM, Benito C (2002) The use of ISSR and RAPD markers for detection DNA polymorphism, genotype identification and genetic diversity among barley cultivars with known origin. Theor Appl Genet 104:845–851
Gupta PK, Rustgi S (2004) Molecular markers from the transcribed/expressed region of the genome in higher plants. Funct Integr Geonomics 4:139–162
Harlan JR (1975) Our vanishing genetic resources. Science 188:618–621
Harlan JR (1992) Crops and man. Am Soc Agron Crop Sci, Madison
Heath DD, Iwana GK, Delvin RH (1993) PCR primed with VNTR core sequences yield species specific patterns and hypervariable probes. Nucleic Acids Res 21:5782–5785
Ince AG, Karaca M, Onus N (2009) Development and utilization of diagnostic DAMD-PCR markers for Capsicum accessions. Genet Resour Crop Evol 56:211–221
Iruela M, Rubio J, Cubero JI, Gil J, Milan T (2002) Phylogenetic analysis in the genus Cicer and cultivated chickpea using RAPD and ISSR markers. Theor Appl Genet 104:643–651
Jeffreys AJ, Wilson V, Thein SL (1985) Hypervariable ‘minisatellite’ regions in human DNA. Nature 314:67–73
Kang HW, Park DS, Go SJ, Eun MY (2002) Fingerprinting of diverse genomes using PCR with universal rice primers generated from repetitive sequence of Korean weedy rice. Mol Cells 13:281–287
Karaca M, Saha S, Zipf A, Jenkins JN, Lang DJ (2002) Genetic diversity among forage bermudagrass (Cynodon spp.): evidence from chloroplast and nuclear DNA fingerprinting. Crop Sci 42:2118–2127
Labdi M, Robertson LD, Singh KB, Charrier A (1996) Genetic diversity and phylogenetic relationships among the annual Cicer species as revealed by isozyme polymorphisms. Euphytica 88:181–188
Lassner MW, Peterson P, Yoder JI (1989) Simultaneous amplification of multiple DNA fragments by polymerase chain reaction in the analysis of transgenic plants and their progeny. Plant Mol Biol Rep 7:116–128
Mahmood Z, Athar M, Khan MA, Ali M, Saima S, Dasti AA (2011) Analysis of genetic diversity in chickpea (Cicer arietinum L.) cultivars using random amplified polymorphic DNA (RAPD) markers. Afr J Biotech 10(2):140–145
Maric S, Bolaric S, Martincic J, Pejic I, Kozumplik V (2004) Genetic diversity of hexaploid wheat cultivars estimated by RAPD markers, morphological traits and coefficients of parentage. Plant Breed 123:366–369
Métais I, Aubry C, Hamon B, Jalouzot R (2000) Description and analysis of genetic diversity between commercial bean lines (Phaseolus vulgaris L.). Theor Appl Genet 101:1207–1214
Naghavi MR, Jahansouz MR (2005) Variation in the agronomic and morphological traits of Iranian chickpea accessions. J Integ Plant Biol 47:375–379
Nei (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Nevo E, Payne PL (1987) Wheat storage proteins: diversity of HMW glutenin subunits in wild emmer from Israel. Geographical patterns and ecological predictability. Theor Appl Genet 74:827–836
Perrier X, Jacquemoud-Collet JP (2006) DARwin software, http://darwin.cirad.fr/darwin
Perrier X, Flori A, Bonnot F (2003) Data analysis methods. In: Hamon P, Seguin M, Perrier X, Glaszmann JC (eds) Genetic diversity of cultivated tropical plants. Science Publishers, Enfield, pp 43–76
Powell W, Morgante M, Andre C, Hanafey M, Vogel J, And TS, Rafalski A (1996) The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2:225–238
Rajesh PN, Sant VJ, Gupta VS, Muehlbauer FJ, Ranjekar PK (2002) Genetic relationships among annual and perennial wild species of Cicer using inter simple sequence repeat (ISSR) polymorphism. Euphytica 129:15–23
Rohlf FJ (1998) NTSYS-pc numerical taxonomy and multivariate analysis system. Version 2.02. Exeter Publications, Setauket
Saeed A, Hovsepyan H, Darvishzadeh R, Imtiaz M, Panguluri SK, Nazaryan R (2011) Genetic diversity of iranian accessions, improved lines of Chickpea (Cicer arietinum L.) and their wild relatives by using simple sequence repeats. Plant Mol Biol Rep 29:848–858
SAS Institute Inc (1996) SAS/STAT software, change and enhancements. Release 6.11. SAS Institute Inc, North Carolina
Sofalian M, Chaparzadeh N, Javanmard A, Hejazi MS (2008) Study the genetic diversity of wheat landraces from northwest of Iran based on ISSR molecular markers. Int J Agri Bio 10(3):466–468
Talebi R, Fayaz R, Mardi M, Pirsyedi SM, Naji AM (2008a) Genetic relationships among chickpea (Cicer arietinum) elite lines based on RAPD and agronomic markers. Int J Agric Biol 10:301–305
Talebi R, Naji AM, Fayaz F (2008b) Geographical patterns of genetic diversity in cultivated chickpea (Cicer arietinum L.) characterized by amplified fragment length polymorphism. Plant Soil Environ 54:447–452
Tanya P, Taeprayoon P, Hadkam Y, Srinives P (2011) Genetic diversity among Jatropha and Jatropha-related species based on ISSR markers. Plant Mol Biol Rep 29:252–264
Torres AM, Weeden NF, Martin A (1993) Linkage among isozyme, RFLP and RAPD markers in Vicia faba. Theor Appl Genet 85:935–945
Van der Maesen LJG (1987) Origin, history and taxonomy of chickpea. In: Saxena MC, Singh K (eds) The Chickpea. CAB Inter, Wallingford, pp 11–34
Wolfe AD, Liston A (1998) Contribution of PCR based methods to plant systematic and evolutionary biology. In: Soltis PS, Soltis DE, Doyle JJ (eds) Molecular systematic of plants II. Chapman and Hall, New York, pp 43–86
Zeven AC, de Wet JMJ (1982) Dictionary of cultivated plants and their regions of diversity. Centre for Agricultural Publishing and Documentation, Wageningen
Acknowledgments
We would like to thank as this study was supported by Islamic Azad University, Sanandaj branch. Also, we wish to present our special thanks to Iranian National Gene Bank for kindly supplying seeds of genotypes.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pakseresht, F., Talebi, R. & Karami, E. Comparative assessment of ISSR, DAMD and SCoT markers for evaluation of genetic diversity and conservation of landrace chickpea (Cicer arietinum L.) genotypes collected from north-west of Iran. Physiol Mol Biol Plants 19, 563–574 (2013). https://doi.org/10.1007/s12298-013-0181-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-013-0181-7