Abstract
Computer-aided diagnosis systems for assisting the classification of various diseases have the potential to improve radiologists’ diagnostic accuracy and efficiency, as reported in several studies. Conventional systems generally provide the probabilities of disease types in terms of numerical values, a method that may not be efficient for radiologists who are trained by reading a large number of images. Presentation of reference images similar to those of a new case being diagnosed can supplement the probability outputs based on computerized analysis as an intuitive guide, and it can assist radiologists in their diagnosis, reporting, and treatment planning. Many studies on content-based medical image retrievals have been reported on. For retrieval of perceptually similar and diagnostically relevant images, incorporation of perceptual similarity data by radiologists has been suggested. In this paper, studies on image retrieval methods are reviewed with a special focus on quantification, utilization, and the evaluation of subjective similarities between pairs of images.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Content based image retrieval (CBIR) has been one of the active research topics in computer vision and medical image analysis for decades. In the era of big data and high-performance computing, interest in medical image retrieval is growing rapidly. There are two levels of “similarity” considered in medical image retrieval: (1) images are of the same imaging modality, the same body orientation (such as posteroanterior or lateral view and axial, sagittal or coronal slices), and the same body parts or organs under examination; and (2) the images depict the same pathologic condition. In the former situation, the purpose of the image retrieval can be image indexing. In the latter, on the other hand, the retrieved images are most likely to be used for computer-aided diagnosis (CAD) purposes. With advances in medical imaging devices, radiologists are exposed to a large amount of data from multimodality imaging systems. Providing an accurate diagnosis while maintaining efficiency is not an easy task. Images depicting a similar pathologic condition based on past studies can assist radiologists in their diagnosis, filing of radiologic reports, and treatment planning. Image retrieval systems can also be useful for educational purposes.
There have been several review papers about CBIR applied to the medical imaging field in the past 15 years [1,2,3,4,5]. Muller et al. published the first comprehensive review paper about CBIR in medical imaging [1]. Long et al. highlighted the status of CBIR and the problems yet to be solved in the implementation of CBIR systems based on the evaluation of example systems [2]. Akgul et al. reviewed features and similarity measures used in medical CBIR systems in the literature [3]. Kumar et al. placed a special focus on the application of CBIR to multidimensional and multimodality data [4]. Most recently, Li et al. introduced recent methodologies as well as challenges and opportunities in the context of big data [5]. These papers addressed the methodologies, status, and future direction of CBIR at the time of their writing. All of these papers discussed the semantic gap, which is the difference between the information expressed by the image features and the findings perceived by human observers, i.e., medical doctors.
The author has been studying the selection methods for similar images of breast lesions in CAD framework, with special emphasis on trying to fill the gap between perceptual and objective (computer-derived) similarities. In this paper, an overview of image retrieval studies, especially in the breast CAD framework, is presented. The review places a special focus on the quantification and utilization of subjective similarities of images for medical image retrieval.
2 Basic methodology of CBIR
Conventional image retrieval methods generally have two main components: feature extraction (off-line) and similarity determination/image selection (on-line) stages.
2.1 Image features
The image features employed in image retrieval systems are generally common to those used in computerized detection or classification schemes. They may include morphologic or shape features, gray-level or color features, and edge-characteristic features, depending on the target anatomy or the disease under study. In the examination of blob-like lesions such as breast masses on mammograms and ultrasonograms, lung nodules on computed tomography (CT) images, and tumors on PET images, the lesion shape is one of the important characteristics. Features such as the circularity, compactness, irregularity, eccentricity, and the major and minor axis ratio are some of the shape-related types. The size, number, and density (number per unit area) of lesions are other geometric features that may be employed for searching of images of some tumor types in which the size is important, or of images such as microcalcification clusters on mammograms and microaneurysms on retinal fundus images.
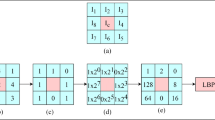
Gray level features may include the contrast, average and variance of pixel values, and various features based on pixel value histograms. Color features are employed mostly for pathologic images in medical images. These features based on pixel values are among the fundamental features representing perceptual similarity. Textural features can be particularly useful for images with characteristic patterns, such as CT images of diffuse lung diseases and pathologic images. Features based on co-occurrence matrix [6], gray level run length matrix [7], Gabor filter [8], Markov random field [9], and local binary patterns (LBP) [10] are some of the texture features often used in CBIR methods.
The edge gradient features can describe the boundary characteristics of lesions. One characteristic finding for breast cancer on mammograms and lung cancer on radiographs or CT is the presence of spicula. Edge features such as the radial gradient index (RGI) [11] and the vector convergence index [12] can describe boundary shapes and the distinctiveness of margins.
2.2 Similarity measures
The most simple and frequently used similarity measure is the Euclidean distance in feature space. It is based on a simple idea: the closer the feature values, the greater the similarity. In general, each feature is normalized, in which the Euclidean distance becomes equivalent to the Mahalanobis distance. However, with this measure, all features are treated equally. It is often the case with medical image diagnosis, however, that some findings are more important than others. In such case, the weighted distance is a possible index if appropriate weights corresponding to the relative contributions of the features can be determined.
An alternative approach to the selection of similar images is that of graph matching [13]. In graph matching, an image is represented by a graph, i.e., features and their relationships. In a study by Sharma et al. [14], the similarities of histologic images were determined using graph matching method. The histologic images were first segmented into regions that contained different tissue types. Features such as the area and perimeter of the regions as well as the relationship between the regions, such as the distance between the centroids and common boundary length, were determined. Based on this graph representation, the best-matching images were searched.
Similarly in a study by Kumar et al. [15], a graph-based approach was employed in the retrieval of PET/CT images. A graph was generated by segmenting of anatomic regions (lung in this case) from CT and tumors from PET images. The features based on the segmented regions and their relationships were determined. In the Kumer study, the gold standard of “similarity” was tumor localization; images that had the similar tumor distribution with respect to the organs were considered relevant. Therefore, the graph approach was considered effective in matching spatial arrangements of the tumors.
3 Retrieval of perceptually similar images
A large number of studies is related to content-based medical image retrieval and CAD. The application of CBIR includes, but has not been limited to breast masses [16,17,18,19,20,21,22,23,24,25,26,27] and microcalcification clusters [28,29,30] on mammograms, breast masses on ultrasound images [31, 32], lung nodules [33,34,35,36,37] and diffuse lung diseases [38, 39] in CT, focal liver lesions in CT [40,41,42,43,44], brain tumors on MRI [45, 46], brain hemorrhages in CT [47], diabetic retinopathy on retinal fundus images [48, 49], tumors on PET images [50], histopathologic images of breast [51, 52] and skin [53] cancers, skin lesions on dermoscopic images [54], and lesions in endomicroscopic video [55]. Not all of these studies can be described in this paper; instead, some of the early studies are introduced briefly, and studies involving quantification, utilization, and evaluation of subjective similarity of images are discussed in more detail in this section.
3.1 Early studies
One of the early studies on similar image retrieval for diagnosis of breast lesions on mammograms was reported by Qi and Snyder [16]. Their system determines simple features related to lesion shape, and the images with a small vector distance to a query image are retrieved. Giger et al. proposed a system called an intelligent workstation, which provides the likelihood of malignancy of a queried mass as well as the similar images selected on the basis of the closeness of a single feature, multiple features, or the likelihood of malignancy measure [17]. Sklansky et al. proposed a mapped-database system for mammographic regions of interest (ROIs) with microcalcifications, as shown in Fig. 1 [28]. An artificial neural network computes a relational map, which is a 2-dimensional map showing the distributions of benign and malignant ROIs in the database and the location of a query. The map also depicts the area where biopsy recommended cases are located. Based on this map, similar ROIs can be selected for display. The study indicated the usefulness of the proposed system for the diagnosis of benign and malignant clusters by aided radiologists in a receiver operating characteristic (ROC) study.
User interface of a mapped-database system proposed by Sklansky et al. [28]
Presentation of similar images was considered useful in assisting radiologists’ diagnosis; however, it was uncertain and difficult to evaluate whether retrieved images were visually similar. To select visually similar images, Li et al. proposed the use of a machine learning system, which was trained on subjective similarities of lesions as assessed by expert radiologists [34]. The similarity determined, called a psychophysical similarity measure, takes into account the image features and the perceptual similarity through iterative training.
3.2 Quantification of subjective similarity
The gold standard of similarity must be established for machine learning and evaluation of a system. Perceptual similarities assessed by a group of radiologists can be employed as the gold standard. Some of the challenges in obtaining such data are that there is a large variation in subjective similarities for image pairs of lesions/abnormalities, and radiologists/diagnosticians are not accustomed to assessing image similarities. Whether subjective similarities of image pairs can be determined consistently and reliably has been questioned by researchers.
Nishikawa et al. examined observers’ ability to make a similarity judgement for clustered microcalcifications on mammograms [56]. Thirty pairs of images were used in their experiment. First, each pair was rated for its similarity on a 5-point scale (called an absolute rating method). Next, all possible combinations of 2 pairs were judged as which pair was more similar than the other by use of a paired comparison method. Four observers, including 3 experienced radiologists and one experienced research technician, participated in the study, in which two of them completed the reading twice for intra-reader agreement analysis. The intra-reader agreements were 0.51 and 0.82 for the absolute and paired comparison methods, respectively, in terms of the intra-class correlation. The inter-reader agreements were 0.39 and 0.37, respectively. The Pearson correlation coefficient between the average ratings by the two methods was 0.77. The authors concluded that the readers were internally more consistent in the paired comparison than in the absolute rating; however, if the readers had different criteria for image similarity, agreement between readers would be reduced, even though each reader was internally consistent. Overall, the high correlation between the two methods indicated that observers can judge similarity in a consistent manner.
In a follow-up study by Wong et al. [57], 1000 pairs of microcalcification images were rated on a 10-point scale. Before and during the rating session, if requested, five anchor images for precalibration were provided, so that a uniform measure was established among the readers. The average inter-reader correlation coefficient among 5 radiologists was 0.489. Despite the variation among these individuals, the group of readers achieved a high level of consistency, as indicated by a correlation coefficient of 0.698 between the average scores for the 5 radiologists and for 5 non-radiologist readers.
Muramatsu et al. investigated the intra- and inter-observer variation as well as the intergroup correlation in the rating of subjective similarities for pairs of microcalcifications on mammograms [58]. One hundred fourteen pairs of clustered microcalcifications were rated on a continuous rating scale by 13 breast radiologists, 10 general radiologists, and 10 non-radiologists, of whom 1, 1, and 5 observers, respectively, repeated the study 5 times, whereas 8, 0, and 3, respectively, repeated it twice. Figure 2 shows the trend of the intraobserver correlation between two consecutive readings for 7 observers in 5 repeated reading sessions. When the time between two readings was very short, the correlations were generally increased, which could be due in part to an improvement in memory. The general trend was that, as the study is repeated, intracorrelations were improved slightly or stayed high. The authors expected that this result might be due to a training effect. The observers were likely to become familiar with the extraordinary task and established their own criteria for image similarity.
Trend in intraobserver correlation between consecutive readings with time elapsed from the first reading session. Data were obtained for seven observers who repeated the study five times [58]
The authors expected that averaging of the repeated reading data would reduce the inter-reader variation. The average interobserver correlations between the first and second readings and between the averages of two readings are listed in Table 1. Although the interobserver correlations were relatively low for the single readings, they were improved slightly when the average of the two readings was taken. Similarly, when the ratings were averaged for a group of observers, the intergroup correlation increased as the number of observers in each group increased, as shown in Fig. 3. The intergroup correlations between breast radiologists and general radiologists and between breast radiologists and non-radiologists were 0.846 [95% confidence interval (0.789, 0.888)] and 0.817 [0.747, 0.869], respectively, values which were significantly higher than those between single observers. These results indicate that multiple readings by single observers and ratings by multiple observers can increase the reliability of subjective similarity.
Effect of the numbers of observers in each group on the intergroup correlation, Two groups of observers were randomly sampled from 13 breast radiologists and the rating were averaged in each group. The random sampling process was repeated for 100 times [58]
Subjective similarities of pairs of mass images and pairs of microcalcification images based on the absolute rating and on paired comparison were compared in a study by Muramatsu et al. [59]. Pairs of masses and pairs of microcalcifications had been rated previously [19, 58] on an absolute scale by groups of radiologists. By the absolute rating method, 6 pairs of ratings were obtained simultaneously on one monitor by placement of an index case in the center and three comparison cases each on the right and left sides so that they could serve as scaling cases for each other. From these cases in the previous studies, 8 pairs of masses and 8 pairs of microcalcifications were selected for the paired comparison. The selection criteria were: (1) the absolute similarity ratings were approximately evenly distributed from 0 to 1, (2) the standard deviations of the ratings were relatively small, and (3) no image was included in more than one pair. Figure 4 shows the study cases. Using 2-alternative forced choice (2AFC, also known as paired comparison) method, a similarity rating in absolute scale cannot be determined; instead, pairs can be ranked for their relative similarities. Each pair was compared with seven other pairs in each group of 8 pairs one by one, and the number of times selected as more similar than the other was summed; the result was defined as the similarity ranking score, in which the highest possible score was 7. Ten observers, including four breast radiologists, one breast imaging fellow, two general radiologists, and three radiology residents, participated in the study. Two reading sessions were set up: in the first session, 8 pairs of masses and 8 pairs of microcalcifications were grouped separately, and in the second session, 4 odd-ranked masse pairs and 4 even-ranked calcification pairs were grouped, and vice versa (mixed groups).
As in the study by Nishikawa et al. [56], the observers in this study were very consistent in selecting the most similar pairs. Based on the first session, the average intraobserver correlations for the mass and microcalfication groups were 0.92 and 0.90, respectively, whereas the average interobserver correlations were 0.74 and 0.86, respectively. The correlation coefficients between the average absolute similarity ratings and the average ranking scores were 0.94 and 0.98 for the mass and the calcification pairs, respectively. The relationships between the average absolute similarity ratings and the average similarity ranking scores for the two sessions are shown in Fig. 5. The results indicate that radiologists can judge the similarities of pairs of lesions in a consistent manner. In the second session, it was questioned whether the similarity of a mass pair can be compared with that of a calcification pair. The correlations between the average absolute ratings and ranking scores were 0.92 and 0.96 for the two groups. The results indicate that observers have a basic concept of similarity and can quantify their impression of similarity in an absolute scale. Even if the lesion types are different, a mass pair with a similarity of 0.8, for example, can be compared with a calcification pair with a similarity of 0.4 in a consistent way.
Relationships between average absolute similarity ratings and ranking scores by 2AFC methods. Top left: for 8 pairs of massed; top right: for 8 pairs of microcalcifications; and bottom: for two sets of mixed 8 pairs [59]
This conclusion was confirmed in a study by Kumazawa et al. in which similarities of pairs of masses on mammograms and pairs of nodules on CTs were compared [60]. Even for the different diseases (breast abnormalities vs lung abnormalities) on different image modalities (mammography vs chest CT) read by different groups of observers (breast radiologists vs chest radiologists), similarity of images were assessed reliably proving that the image similarity is a sharable concept. While observers were more consistent in determining similarity using the 2AFC method, it is desirable to obtain similarities on an absolute scale because reading of all possible pairs in the 2AFC method is a demanding task for radiologists, and the ranking scores by the 2AFC method are dependent on the cases included in the study. The results of the above studies indicated that subjective similarities of lesions in an absolute scale can be determined reliably.
Tourassi et al. compared different data collection methods for obtaining subjective similarities of masses on mammograms [61]. Three methods were compared: a rating method in which a similarity score for a pair was obtained using a continuous scale; a preference method which is analogous to a paired comparison method in which three masses (e.g., A, B, and C) are shown at once and observers are asked to select the most similar pair (A and B, A and C, or B and C) or no particular pair; and a hybrid method, in which a query mass is placed in the center of a display and other masses are placed in a circular format around the query. The hybrid method is somewhat analogous to the method employed by Li et al. [34] and Muramatsu et al. [19, 58], in which observers provide rating scores while adjusting their judgment using all possible pairs in the display. Using the data collected, the authors developed individualized user models for predicting radiologists’ perceptual judgments. The result indicated that the hybrid method was the most accurate in constructing the user models, whereas the rating method was the most time-efficient. They concluded that the hybrid method provides an intuitive and efficient way of obtaining perceptual similarity data.
Faruque et al. performed a simulation study on perceptual similarity measures for focal liver lesions [62]. Similarity scores for 171 pairwise comparisons of 19 lesions on CT images were obtained from three radiologists. Based on their model, the number of readers required for achieving acceptable levels of similarity was estimated. The result indicated that an excellent estimate of a simulated ground truth of similarity scores could be obtained with a relatively small number of readers whose ratings exhibited moderate to good inter-reader agreement.
3.3 Incorporation of subjective data
For the selection of perceptually similar images, a similarity index that agrees well with the subjective similarity determined by radiologists is desired. In their study, Li et al. [34] employed an artificial neural network (ANN) with a single hidden layer to train the relationship between the image features and subjective ratings. Seven units corresponding to the diameter, CT values, and the RGI of the two nodules and the pixel difference were used as the input. For teacher data, subjective similarity scores from 0 to 3, allowing the fractional scores, for 240 pairs of nodules were determined by 10 radiologists. Using a leave-one-out cross validation method, the ANN was trained with 239 pairs of nodules, and the trained ANN provided the output, called a psychophysical similarity measure, for a test case. A relatively high correlation (0.72) between the subjective ratings and the psychophysical measure was achieved as compared with those by the conventional feature-distance-based method and the cross-correlation-based method.
Similarly, Muramatsu et al. employed ANNs for the determination of similarity measures for pairs of masses and pairs of microcalcifications on mammograms [30, 63]. In both studies, 300 pairs of lesions were examined by breast radiologists for obtaining subjective similarity ratings, and the average ratings were used as teacher data in the training of the ANNs. By incorporation of the subjective aspect of lesion similarities through machine learning, similarity measures that were in relatively good agreement with the radiologists’ perception on lesion similarity could be determined.
El-Naqa et al. investigated a machine learning approach with use of sequential networks [29]. In their method, the first network was used for triage to eliminate images that were not similar at all. In the first stage, a classifier such as a support vector machine (SVM) was employed for classifying a pair as sufficiently similar or not similar. In the second stage, a regression network, e.g., another SVM, was trained to estimate similarities of pairs. Thirty microcalcification clusters which constituted 435 pairwise comparisons were examined by 6 observers for providing a similarity score for each pair in terms of the spatial distribution of the calcifications on a 10-point scale. An additional 30 artificial pairs made of identical images with a similarity score of 10 were included in the study. Based on the cross-validation test, a higher retrieval precision was achieved using the two-stage network than using a single regression network or a conventional Euclidean metric.
Zheng et al. proposed a retrieval method that included an “interactive step” to improve the visual similarity of retrieved images for masses on mammograms [21]. The masses were subjectively rated from 1 to 9 for their margin spicularity, and similar images were retrieved from those which margin scores were within ± 1 of that of a query case.
Another type of two stage selection methods was investigated by Nakayama et al. [64], in which combinations of a distance-based measure and a psychophysical similarity measure were compared. They examined the subjective similarities of 20 pairs of masses and 20 pairs of microcalcifications, of which 5 pairs each were selected by 4 different methods: selection by the distance-based measure, selection by the psychophysical measure, a sequential selection by the distance measure followed by the psychophysical measure, and a sequential selection by the psychophysical measure followed by the distance-based measure. They discussed the potential utility of preselection by the distance measure with more refined selection by the psychophysical measure for retrieving perceptually similar images.
A machine learning method, in general, requires a large number of training samples with a variety of cases. However, it is not easy to prepare such a database with subjective data. In the study by Muramatsu et al. [19], pairs of spiculated masses had high similarity ratings as well as strong (very high or very low) feature values. These samples had a strong influence in training of an ANN, because the number of training samples was small. As a result, a trained ANN tends to yield high scores for a pair that includes a spiculated mass, causing bias during image retrieval. As a potential solution, a similarity space modeling method, rather than direct estimation of similarity for each pair, was investigated.
A subjective similarity space was modeled using a multidimentional scaling (MDS) [65] in a study by Muramatsu et al. [66]. Twenty-seven breast mass images of different pathologic types were selected, and subjective similarity ratings for 351 pairwise comparisons were obtained from eight experienced physicians who were certified for breast image reading. Figure 6 shows the sample mass images of different subtypes of breast lesion pathologies. A similarity map was obtained by application of the MDS to the average similarity (dissimilarity) ratings, as shown in Fig. 7, which reflected the readers’ intuition of similarities between lesions of these subtypes. Despite the small sample size, cysts and fibroadenomas, which are almost indistinguishable on mammograms, were clustered and located away from the typical malignant cases. Likewise, ductal carcinomas in situ, papilotubular carcinomas, and solid-tubular carcinomas were mapped close by, whereas scirrhous carcinomas and invasive lobular carcinomas were mapped close together. If such a perceptual similarity space can be reliably modeled and cases without subjective data can be projected to the space, perceptually similar images may be retrieved.
Sample mass images with different subtypes [66]
Similarity map obtained by MDS using the average subjective similarity ratings for mass pairs on mammograms [66]. FA fibroadenima, PT phyllodes tumor, DCIS ductal carcinoma in situ, PTC papilotubular carcinoma, STC solid tubular carcinoma, MC mucinous carcinoma, SC scirrhous carcinoma and ILC invasive lobular carcinoma
Similarity spaces spanned by MDS using subjective similarity ratings for mass pairs on mammograms and ultrasound images were reconstructed using 3 layered ANNs [67, 68]. The ANNs were trained with the image features as input and 3-dimensional coordinates of the MDS spaces based on 351 pairs of masses on mammograms and 666 pairs of masses on ultrasound images. Using a leave-one-case-out cross validation method, the perceptual similarity spaces were estimated. The similarity measures based on the distances in the reconstructed spaces correlated relatively well with the subjective similarity ratings. The performance of image retrieval was evaluated in terms of the precision, which is the fraction of relevant images in the retrieved images; images with the same pathology (benignity or malignancy) are considered to be relevant. High precisions above 0.8 for the independent test cases without subjective data were obtained for masses on mammograms and for masses on ultrasonograms.
The direct similarity estimation method and the similarity space modeling method have advantages and disadvantages. One such advantage of the space modeling method is that the ANN training is simpler. The ANN takes a feature vector of an image as input for the estimation of a coordinate in each dimension, which can be a more focused task than is the estimation of similarity ratings from two feature vectors for a pair. On the other hand, more abundant subjective data are generally required for modeling of the space with MDS, because all possible pairwise comparisons must be made. This could be partially solved using MDS analysis which allows missing data. When applying to an unknown case, it must be paired with all of the cases in the database for estimating similarities by the direct estimation method, whereas such a process can be avoided by projecting of the unknown case to the modeled similarity space. Preselection may be useful in both methods, especially when the database becomes very large. Further studies are needed for the evaluation of objective similarity and image retrieval methods.
3.4 Interactive/feedback methods
When similar images are retrieved, they may include cases that are very similar and useful, but also cases that are not very similar or useful for assisting radiologists in their diagnosis. If such information, whether the retrieved images are useful assessed by users, can be fed back, the image retrieval system can be improved. Several research groups have proposed such interactive methods or methods with relevance feedback. Oh et al. proposed a relevance feedback system based on incremental learning with SVM, which takes into account the feedback samples and already trained samples that are in the neighborhood of the feedback samples in the hyperplane of SVM [69]. They reported that the performance of image retrieval in terms of precision and recall curves was improved considerably with one feedback sample per case compared with the offline mode (no feedback), although the improvement became less with three and five feedback samples.
Wei et al. also proposed an interactive retrieval method for masses and microcalcifications on mammograms [70]. Images were first retrieved by the feature-based hierarchical selection method, in which features with greater importance were given larger weights in determining the similarity measure. After the first image retrieval, users may provide relevance feedback to an arbitrary number of images, which were used for training of an SVM for classification of relevant and irrelevant cases. Superior precision and recall curves were obtained for both mass and calcification cases when the relevance feedback mode was used.
Bugatti et al. proposed a CBIR system, which employs a relevance feedback system to refine the search through user profiles [39]. The concept of the system is to collect static and dynamic user profiles to maintain users’ preference for system utility. In their experiment, retrieval methods for diffuse lung diseases in CT images and breast lesions on mammograms were studied. After an initial search, feedbacks for retrieved images were obtained by asking users to select 5 relevant images in the order of perceived similarity. Based on the differences in the initial selection order and the perceived similarity order, the best distance function used for the similarity measure was selected.
Another interactive system with an adaptation module that integrates radiologists’ similarity ratings as a relevance feedback was proposed by Cho et al. [71]. An original feature vector of a query was modified by the sets of feature vectors of relevant images and irrelevant images so that the virtual query vector is moved toward the relevant samples. The virtual vector was computed as the weighted sum of the original vector, relevant-group vector, and irrelevant-group vector. In their experiment, 9 point similarity ratings by radiologists were employed as relevance feedback with a threshold, and balancing weights for the original, relevant, and irrelevant vectors were iteratively adjusted through training. A higher average similarity and a higher classification performance were obtained by the interactive system with retrieval of breast masses on ultrasonography.
4 Current trends
In the field of medical image analysis, deep learning based methods are rapidly replacing the conventional hand-crafted feature based methods. Several CBIR methods that use deep learning techniques have been proposed. Liu et al. proposed a method using a convolutional neural network (CNN) for retrieval of radiographs with the same image modality, body orientation, body region, and biological system examined [72]. The network was trained with radiographs from Image Retrieval in Medical Application (IRMA) database which includes images with more than 193 categories. Once the network was trained, features from the last full connection layer having 1000 units were extracted for obtaining a CNN code. This code was combined with a conventional radon barcode for image retrieval.
Similarly, Anavi et al. [73] extracted features from the last layers of a CNN which was pre-trained with the ImageNet [74] database. The CNN features were either used directly for the determination of a distance measure based on the intersection of the feature histograms or for training of an SVM for classification of 8 classes of diseases on chest radiographs. In the latter, 8 output probabilities for pairs of images were then employed for determination of a distance measure to retrieve similar images.
For retrieval of similar images of 24 classes of radiographs of different body parts, Qayyam et al. employed CNN features from the last three full connection layers [75]. The Euclidean distance metric was calculated with the feature vectors of a query and images in the database. In addition, a class label predicted by the CNN was used for limiting the search area in the database.
Khatami et al. employed a CNN for shrinking of the search space [76]. For retrieval of radiographs using the IRMA database, the classification result from the CNN was used for limiting the search space, followed by a second search-space shrinking with Radon projection vectors. The final selection was made with the LBP-based Manhattan distance measures.
The CNN features from the full connection layer-6 of the AlexNet [74] model were also employed for similarity measure determination by Pang et al. [77]. The image retrieval performance was evaluated with three different databases: the NEMA-CT database that includes different body parts (different levels of axial sections), the TCIA-CA database with different body parts, and the OASIS-MR database including images classified based on the shape of the ventricular. Deep features (CNN features) combined with a preference learning model obtained a high performance compared with the conventional feature based methods.
Most of the above methods employed a CNN as a feature extractor. Muramatsu et al. investigated the use of the CNN to determine the similarity measures directly for pairs of images [78] and to model the similarity space for image retrieval [79]. In the former, the network consisted of two input layers for taking a pair of images followed by a few sets of convolutional layers and pooling layers, a concatenation layer, another sets of convolutional layers and pooling layers, and full connection layers with a regression output layer. Sample pairs of images with subjective similarity ratings were used for training of the network. Because of the small sample size of training cases with the subjective data, the network was pre-trained for classification of benign and malignant lesions by entering the same image as two input images. Subsequently, the network was fine-tuned with the paired data for the similarity estimation by changing the last layer with the regression output. A schematic diagram is shown in Fig. 8.
In the similarity space modeling method, a regular network structure, such as the AlexNet and VGG-net, was employed, but with the regression outputs corresponding to 3-dimensional space coordinates. The network was pre-trained using the classification dataset as the direct estimation method, which was then fine-tuned for similarity space modeling. Figure 9 is a schematic diagram of the proposed method. In a preliminary investigation, a comparable performance was obtained using CNN and the conventional methods.
5 Commercial systems
There are a few commercial diagnostic support systems with a reference image retrieval feature. Quantitative Insights [80] is a company that provides CADx (computer aided classification) workstations which are based on technology developed by a research group at the University of Chicago. The company obtained the first FDA clearance for a machine-learning-driven cancer diagnosis system, which includes image retrieval of breast lesions on MRI (Fig. 10).
A medical imaging and information management system, called SYNAPSE, by Fujifilm allows a case match of lung cancer images [81]. Based on the seed point entered by a user, the system automatically segments the lesion and retrieves similar cases with confirmed diagnosis and radiologic report. Combined search with keywords is also allowed.
A similar case retrieval system by Panasonic selects similar cases of lung CT images with nodular and diffused opacities [82]. The system extracts keywords from diagnostic report and features from images, and it finds the best matched images from the database. They have incorporated a CNN in the classification of image patches into 12 disease categories, which results are used for case matching.
6 Conclusion
Conventional computer-aided classification systems generally provides the probabilities of diseases in question. Although such computer aids were reported to have potential utility, users, i.e., physicians, may question the basis of the results of computer analysis. Presentation of reference images that are perceptually similar and diagnostically relevant can supplement the numerical outputs in an intuitive way and sometimes provide different opinions.
There have been many studies on content based medical image retrieval for image indexing and diagnostic aid. For promoting the utility of reference images in assisting disease classification, the perceptual similarity of the retrieved images is one of the important factors. In this paper, studies on the quantification and incorporation of subjective similarity for retrieval of visually similar images were introduced. In these studies, the feasibility of determining subjective similarities for pairs of images with various abnormalities was discussed, and the result supported the fact that perceptual similarity is a robust concept that are shared by radiologists/physicians and can be quantified reliably. The experimental results on computerized determination of similarity measures and image retrieval indicated the potential usefulness of the similarity measures based on subjective data.
The field of computerized medical image analysis has entered an era of big data and high-performance computing, allowing deep learning and high-speed data mining. Effective utilization of a vast amount of information from accumulated medical data is imperative. However, at present, much of the valuable data supply is left unused. One way to make use of the data is to perform image retrieval. Although perceptual evaluation is important, acquisition of subjective data is a challenging task. A design for systematic and efficient acquisition of subjective similarity data or feedback is still needed.
Some studies suggested the use of metadata and combined information from multiple image modalities [83, 84]. Methods for the fusion of multidisciplinary information must be investigated for a multimodality reading environment. The size and variety of the database are essential for image retrieval and computerized image analysis. An automatic update of the database with and without truth marking remains necessary. When the database becomes exceedingly large, an exhaustive search could be time-consuming and the database may include some undesirable cases (outliers). Techniques for optimization of a reference library [85] could be a research topic of interest. Imaging systems and computer technology are continuously improving, and new cases are constantly obtained. Therefore, computer algorithms must also be improved continuously. Self-learning systems are one of the exciting topics that need to be investigated.
References
Muller H, Michoux N, Bandon D, Geissbuhler A. A review of content-based image retrieval systems in medical applications—clinical benefits and future directions. Int J Med Inf. 2004;73:1–23.
Long LR, Antani S, Deserno TM, Thoma GR. Content-based image retrieval in medicine: retrospective assessment, state of the art, and future directions. Int J Health Inf Syst Inform. 2009;4(1):1–16.
Akgul CB, Rubin DL, Napel S, Beaulieu CF, Hayit G, Acar B. Content-based image retrieval in radiology: current status and future directions. J Digit Imaging. 2011;24(2):208–22.
Kumar A, Dim J, Cai W, Fulham M, Feng D. Content-based medical image retrieval: a survey of applications to multidimensional and multimodality data. J Digit Imaging. 2013;26:1025–39.
Li Z, Zhang X, Muller H, Zhang S. Large-scale retrieval for medical image analytics: a comprehensive review. Med Image Anal. 2018;43:66–84.
Haralick RM, Shanmugam K, Dinstein I. Textural features for image classification. IEEE Trans Sys Manuf Cyber. 1973;SMC-3(6):610–21.
Tang X. Texture information in run-length matrices. IEEE Image Process. 1998;7(11):1602–9.
Daugman JG. Uncertainty relation for resolution in space, spatial frequency, and orientation optimized by two-dimensional visual cortical filters. J Opt Soc Am. 1985;2(7):1160–9.
Cross GR, Jain AK. Markov random field texture models. IEEE Trans Pat Anal Mach Intel. 1983; PAMI-5(1):25–39.
Ojala T, Pietikainen M, Maenpaa T. Multiresolution gray-scale and rotation invariant texture classification with local binary patterns. IEEE Trans Pat Anal Mach Intel. 2002;24(7):971–87.
Huo Z, Giger ML, Vyborny CJ, Bick U, Lu P, Wolverton DE, Schmidt RA. Analysis of spiculation in the computerized classification of mammographic masses. Med Phys. 1995;22:1569–79.
Kobatake H, Hashimoto S. Convergence index filter for vector fields. IEEE Trans Image Process. 1999;8(8):1029–38.
Bunke H, Irniger C, Neuhaus M. Graph matching – challenges and potential solutions. Int Conf Image Anal Process. 2005;LNCS3617:1–10.
Sharma H, Alekseychuk A, Leskovsky P, Hellwich O, Anand RS, Zerbe N, Hufnagl P. Determining similarity in histological images using graph-teoretic description and matching methods for content-based image retrieval in medical diagnostics. Diagn Pathol. 2012;7:134.
Kumar A, Kim J, Wen L, Fulham M, Feng D. A graph-based approach for the retrieval of multi-modality medical images. Med Image Anal. 2014;18:330–42.
Qi H, Snyder WE. Content-based image retrieval in picture archiving and communications systems. J Digit Imaging. 1999;12(2):81–3.
Giger ML, Huo Z, Vyborny CJ, Lan L, Bonta I, Horsch K, Nishikawa RM, Rosenbourgh I. Intelligent CAD workstation for breast imaging using similarity to known lesions and multiple visual prompt aids. Proc SPIE Med Imaging. 2002;4684:78–73.
Nakagawa T, Hara T, Fujita H, Iwase T, Endo T. Image retrieval system of mammographic masses by using local pattern matching technique. In: Peitgen HO, editor. Digital Mammography. Berlin: Springer; 2003. pp. 562–5.
Muramatsu C, Li Q, Suzuki K, Schmidt RA, Shiraishi J, Newstead GM, Doi K. Investigation of psychophysical measure for evaluation of similar images for mammographic masses: preliminary results. Med Phys. 2005;32:2295–304.
Alto H, Rangayyan RM, Desautels JEL. Content-based retrieval and analysis of mammographic masses. J Electron Imaging. 2005;14(2):023016.
Zheng B, Lu A, Hardesty LA, Sumkin JH, Hakim CM, Ganott MA, Gur D. A method to improve visual similarity of breast masses for an interactive computer-aided diagnosis environment. Med Phys. 2006;33:111–7.
Kinoshita SK, Marques PMA, Pereira R, Rodrigues JAH, Rangayyan RM. Content-based retrieval of mammograms using visual features related to breast density patterns. J Digit Imaging. 2007;20:172–90.
Nakayama R, Abe H, Shiraishi J, Doi K. Evaluatino of objective similarity measures for selecting similar images of mammographic lesions. J Digit Imaging. 2011;24:75–85.
Wei CH, Chen SY, Liu X. Mammogram retrieval on similar mass lesions. Comput Methods Prog Biomed. 2012;106(3):234–48.
Liu J, Zhang S, Liu W, Zhang X, Metaxas DN. Scalable mammogram retrieval using anchor graph hashing. In: IEEE International symposium on biomedical imaging, ISBI 2014; pp 898–901.
Jaing M, Zhang S, Li H, Metazas DN. Computer-aided diagnosis of mammographic masses using scalable image retrieval. IEEE Trans Biomed Eng. 2015;62(2):783–92.
Bedo MVN, Pereira dos Santos D, Ponciano-Silva M, Marques PMA, Ferreira de Carvalho APL, Traina C. Endowing a content-based medical image retrieval system with perceptual similarity using ensemble strategy. J Digit Imaging. 2016;29:22–37.
Sklansky J, Tao EY, Bazargan M, Ornes CJ, Murchison RC, Teklehaimanot S. Computer-aided, case-based diagnosis of mammographic regions of interest containing microcalcifications. Acad Radiol. 2000;7:395–405.
El-Naqa I, Yang Y, Galatsanos NP, Nishikawa RM, Wernick MN. A similarity learning approach to content-based image retrieval: Application to digital mammography. IEEE Trans Med Imaging. 2004;23(10):1233–44.
Muramatsu C, Li Q, Schmidt RA, Shiraishi J, Doi K. Investigation of psyhophysical similarity measures for selection of similar images in the diagnosis of clustered microcalcifications on mammograms. Med Phys. 2008;35:5695–702.
Kuo WJ, Chang RF, Lee CC, Moon WK, Chen DR. Retrieval technique for the diagnosis of solid breast tumors on sonogram. Ultrasound Med Biol. 2002;28(7):903–9.
Cho H, Hadjiiski L, Sahiner B, Chan HP, Helvie M, Paramagul C, Nees AV. Similarity evaluation in content-based image retrieval (CBIR) CADx system for characterization of breast masses on ultrasound images. Med Phys. 2011;38(4):1820–31.
Aisen AM, Broderick LS, Winer-Muram H, Brodley CE, Kak AC, Pavlopoulou C, Dy J, Shyu CR, Marchiori A. Automated storage and retrieval of thin-section CT images to assist diagnosis: System Description and preliminary assessment. Radiology. 2003;228:265–70.
Li Q, Li F, Shiraishi J, Katsuragawa S, Sone S, Doi K. Investigation of new psyhophysical measures for evaluation of similar images on thoracic CT for distinction between benign and malignant nodules. Med Phys. 2003;30:2584–93.
Kawata Y, Niki N, Ohmatsu H, Moriyama N. Example-based assisting approach for pulmonary nodule classification in three-dimensional thoracic computed tomography images. Acad Radiol. 2003;10:1402–15.
Lam MO, Disney T, Raicu DS, Furst J, Channin DS. BRISC—an open source pulmonary nodule image retrieval framework. J Digit Imaging. 2007;20:63–71.
Wei G, Cao H, Ma H, Qi S, Qian W, Ma Z. Content-based image retrieval for lung nodule classification using texture features and learned distance metric. J Med Syst. 2018;42:13.
Depeursinge A, Varagas A, Gaillard F, Platon A, Geissbuhler A, Poletti PA, Muller H. Case-based lung image categorization and retrieval for interstitial lung diseases clinical workflows. Int J CARS. 2012;7:97–110.
Bugatti PH, Kaster DS, Pociano-Silva M, Traina C Jr. Marques PMA, Traina AJM. PRosPer: perceptual similarity queries in medical CBIR systems through user profiles. Comput Bio Med. 2014;45:8–19.
Xu J, Faruque J, Beaulieu C, Rubin D, Napel S. A comprehensive descriptor of shape: Method and application to content-based retrieval of similar appearing lesions in medical images. J Digit Imaging. 2012;25:121–8.
Yang W, Lu Z, Yu M, Huang M, Feng Q, Chen W. Content-based retrieval of focal liver lesions using bag-of-visual-words representations of single- and multiphase contrast-enhanced CT images. J Digit Imaging. 2012;25:708–19.
Dankerl P, Cavallaro A, Tsymbal A, Costa MJ, Suehling M, Janka R, Uder M, Hammon M. A retrieval-based computer-aided diagnosis system for the characterization of liver lesions in CT scans. Acad Radiol. 2013;20:1526–34.
Roy S, Chi Y, Liu J, Venkatesh SK, Brown MS. Three-dimensional spatiotemporal features for fast content-based retrieval of focal liver lesions. IEEE Trans Biomed Eng. 2014;61(11):2768–78.
Spanier AB, Caplan N, Sosna J, Acar B, Joskowicz L. A fully automatic end-to-end method for content-based image retrieval of CT scans with similar liver lesion annotation. Int J CARS. 2018;13:165–74.
Yang W, Feng Q, Yu M, Lu Z, Gao Y. Content-based retrieval of brain tumor in contrast-enhanced MRI images using tumor margin information and learned distance metric. Med Phys. 2012;39(11):6929–42.
Faria AV, Oishi K, Yoshida S, Hillis A, Miller ML, Mori S. Content-based image retrieval for brain MRI: An image-searching engine and population-based analysis to utilize past clinical data for future diagnosis. NeuroImage Clin. 2015;7:367–76.
Zaki WMDW., Fauzi MFA, Besar R. Retrieval of intracranial hemorrhages in computed tomography brain images using binary coherent vector. J Electron Imaging. 2010;19(4):043021.
Chaum E, Karnowski TP, Covindasamy VP, Abdelrahman M, Tobin KW. Automated diagnosis of retinopathy by content-based image retrieval. Retina. 2008;28:1463–77.
Quellec G, Lamard M, Cazuguel G, Roux C, Cochener B. Case retrieval in medical databases by fusing heterogeneous information. IEEE Trans Med Imaging. 2011;30(1):108–18.
Kim J, Cai W, Feng D, Wu H. A new way for multidimensional medical data management: volume of interest (VOI)-based retrieval of medical images with visual and functional features. IEEE Trans Inf Technol Biomed. 2006;10(3):598–607.
Zheng X, Liu W, Dundar M, Badve S, Zhang S. Towards large-scale histopathological image analysis: Hashing-based image retrieval. IEEE Trans Med Imaging. 2015;34(2):496–506.
Zheng Y, Jiang Z, Zhang H, Xie F, Ma Y, Shi H, Zhao Y. Histopathological whole slide image analysis using context-based CBIR. IEEE Trans Med Imaging 2018 (in press).
Caicedo JC, Gonzalez FA, Romero E. Content-based histopathology image retrieval using a kernel-based semantic annotation framework. J Biomed Inf. 2011;44:519–28.
Baldi A, Murace R, Dragonetti E, Manganaro M, Guerra O, Bizzi S, Galli L. Definition of an automated content-based image retrieval (CBIR) system for the comparison f dermoscopic images of pigmented skin lesions. BioMed Eng Online. 2009;8:18.
Tafresh MK, Linard N, Andre B, Ayache N, Vercauteren T. Semi-automated query construction for content-based endomicroscopy video retrieval. In: Golland P, Hata N, Barillot C, Hornegger J, Howe R, editors. Medical image computing and computer-assisted intervention—MICCAI 2014 LNCS 8673, pp 89–96.
Nishikawa RM, Yang Y, Huo D, Wernick M, Sennett CA, Papioannou J, Wei L. Observers’ ability to judge the similarity of clustered calcifications on mammograms. Proc SPIE Med Imaging. 2004;5372:192–8.
Wang J, Jing H, Wernick MN, Nishikawa RM, Yang Y. Analysis of perceived similarity between pairs of microcalcification clustered in mammograms. Med Phys. 2014;41(5):051904.
Muramatsu C, Li Q, Schmidt R, Suzuki K, Shiraishi J, Newstead G, Doi K. Experimental determination of subjective similarity for pairs of clustered microcalcifications on mammograms: observer study results. Med Phys. 2006;33(9):3460–8.
Muramatsu C, Li Q, Schmidt RA, Shiraishi J, Suzuki K, Newstead GM, Doi K. Determination of subjective similarity for pairs of masses and pairs of clustered microcalcifications on mammograms: Comparison of similarity, ranking scores and absolute similarity ratings. Med Phys. 2007;34(7):2890–5.
Kumazawa S, Muramatsu C, Li Q, Li F, Shiraishi J, Caligiuri P, Schmidt RA, MacMahon H, Doi K. An investigation of radiologists’ perception of lesion similarity: observations with paired breast masses on mammograms and paired lung nodules on CT images. Acad Radiol. 2008;15:887–94.
Tourassi G, Yoon HJ, Xu S, Morin-Ducote G, Hudson K. Comparative analysis of data collection methods for individualized modeling of radiologists’ visual similarity judgments in mammograms. Acad Radiol. 2013;20:1371–80.
Faruque J, Rubin DL, Beaulieu CF, Napel S. Modeling perceptual similarity measures in CT images of focal liver lesions. J Digit Imaging. 2013;26:714–20.
Muramatsu C, Li Q, Schmidt RA, Shiraishi J, Doi K. Determination of similarity measures for pairs of mass lesion on mammograms by use of BI-RADS lesion descriptors and image features. Acad Radiol. 2009;16:443–9.
Nakayama R, Abe H, Shiraishi J. doi K. Evaluation of objective similarity measures for selecting similar images of mammographic lesions. J Digit Imaging. 2011;24(1):75–85.
Kruskal JB, Wish M. Multidimensional scaling. Beverly Hills: Sage; 1978.
Muramatsu C, Nishimura K, Endo T, Oiwa M, Shiraiwa M, Doi K, Fujita H. Representation of lesion similarity by use of multidimensional scaling for breast masses on mammograms. J Digit Imaging. 2013;26:740–7.
Nishimura K, Muramatsu C, Oiwa M, Shiraiwa M, Endo T, Doi K, Fujita H. Psychophysical similarity measure based on multi-dimensional scaling for retrieval of similar images of breast masses on mammograms. Proc SPIE Med Imaging. 2013;8670:86701R.
Muramatsu C, Takahashi T, Morita T, Endo T, Fujita H. Similar image retrieval of breast masses on ultrasonography using subjective data and multidimensional scaling. In: Tingberg A et al., editors. Proceedings of international workshop on breast imaging, IWDM 2016. Lecture notes in computer science, vol 9699. 2016; pp 43–50.
Oh JH, Yang Y, El-Naqa I. Adaptive learning for relevance feedback: application to digital mammography. Med Phys 201;37(8):4432–44.
Wei CH, Li Y, Huang PJ. Mammogram retrieval through machine learning within BI-RADS standard. J Biomed Inform. 2011;44:607–14.
Cho HC, Hadjiiski L, Sahiner B, Chan HP, Paramagul C, Helvie M, Nees AV, Cho HC. A similarity study of interactive content-based image retrieval scheme for classification of breast lesions. IEICE Trans Inf Syst. 2016; E99-D:1663–1670.
Liu X, Tizhoosh HR, Kofman J. Generating binary tags for fast medical image retrieval based on convolutional nets and Radon transform. In: Proceedings of the International Joint Conference on Neural Networks 2016.
Anavi Y, Kogan I, Gelbart E, Geva O, Greenspan H. Visualizing and enhancing a deep learning framework using patients age and gender for chest X-ray image retrieval. Proc SPIE Med Imaging. 2016;9785:978510.
Krizhevsky A, Sutskever I, Hinton G. Imagenet classification with deep convolutional neural networks. In: Proc advances in neural information processing systems 2012; pp 1097–105.
Qayyam A, Anwar SM, Awais M, Majid M. Medical image retrieval using deep convolutional neural network. Neurocomputing. 2017;266:8–20.
Khatami A, Babaie M, Tizhoosh HR, Khosravi A, Nguyen T, Nahavandi S. A wequential serach-space shrinking using CNN transfer learning and a radon projection pool for medical image retrieval. Expert Syst Appl. 2018;100:224–33.
Pang S, Orgun MA, Yu Z. A novel biomedical image indexing and retrieval system via deep preference learning. Comput Methods Programs Biomed. 2018;158:53–69.
Muramatsu C, Higuchi S, Morita T, Oiwa M, Fujita H. Similarity estimation for reference image retrieval in mammograms using convolutional neural network. Proc SPIE Med Imaging. 2018;10575:105752U.
Muramatsu C, Higuchi S, Morita T, Oiwa M, Kawasaki T, Fujita H. Retrieval of reference images of breast masses on mammograms by similarity space modeling. In: Proceedings of IWBI LNCS 2018. (in press).
Quantitative Insights, Inc. https://www.quantinsights.com. Accessed 10 April 2018.
Oosawa A, Hisanaga R, Inoue T, Hoshino T, Shimura K. Development and commercialization of “SYNAPSE Case Match” content-based image retrieval system for effectively supporting the interpretation of physician. Med Imag Tech. 2014;32:26–31. (in Japanese).
Kiyono M. Development of Similar case retrieval system by AI. Innervision 2017:32:46–49. (in Japanese).
Korenblum D, Rubin D, Napel S, Rodriguez C, Beaulieu C. Managing biomedical image metadata for search and retrieval of similar images. J Digit Imaging. 2011;24(4):739–48.
Takahashi T, Muramatsu C, Hiramatsu Y, Morita T, Hara T, Endo T, Fujita H. Similar image search of breast masses by combination of mammograms and ultrasound images—study of psychophysical similarity measure based on multi-dimensional scaling. IEICE Technical Report 2016; MI2015-107:161–164. (in Japanese).
Park SC, Sukthankar R, Mummert L, Satyanarayanan M, Zheng B. Optimization of reference library used in content-based medical image retrieval scheme. Med Phys. 2007;34(11):4331–9.
Acknowledgements
The author is grateful to past members of Dr. Doi’s group at the Kurt Rossmann Laboratory and past and present members of the breast imaging team in the Department of Radiology at the University of Chicago, past and present members of the Fujita-Hara-Zhou Laboratory at Gifu University, breast imaging physicians at the Nagoya Medical Center, and members of the Japan Central Organization on Quality Assurance of Breast Cancer Screening who contributed to the author’s studies on similar image retrieval for their participation and discussion. Special thanks to Prof. Kunio Doi and Prof. Hiroshi Fujita for their continuous support.
Funding
Studies by Chisako Muramatsu et al. introduced here are supported in part by a Grant-in-Aid for Scientific Research (C) (No.17K09061) by Japan Society for the Promotion of Science, a Grant-in-Aid for Scientific Research on Innovative Areas (No. 26108005) by Ministry of Education, Culture, Sports, Sciences and Technology in Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declare that she has no conflict of interest.
Ethical approval
Not applicable.
Informed consent
Not applicable.
About this article
Cite this article
Muramatsu, C. Overview on subjective similarity of images for content-based medical image retrieval. Radiol Phys Technol 11, 109–124 (2018). https://doi.org/10.1007/s12194-018-0461-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12194-018-0461-6