Abstract
A novel β-agarase AgaJ11 belonging to the glycoside hydrolase (GH) 16 family was identified from an agar-degrading bacterium Gayadomonas joobiniege G7. AgaJ11 was composed of 317 amino acids (35 kDa), including a 26-amino acid signal peptide, and had the highest similarity (44 % identity) to a putative β-agarase from an agarolytic marine bacterium Agarivorans albus MKT 106. The agarase activity of purified AgaJ11 was confirmed by zymogram analysis. The optimum pH and temperature for AgaJ11 activity were determined to be 4.5 and 40 °C, respectively. Notably, AgaJ11 is an acidic β-agarase that was active only at a narrow pH range from 4 to 5, and less than 30 % of its enzymatic activity was retained at other pH conditions. The K m and V max of AgaJ11 for agarose were 21.42 mg/ml and 25 U/mg, respectively. AgaJ11 did not require metal ions for its activity, but severe inhibition by several metal ions was observed. Thin layer chromatography and agarose-liquefying analyses revealed that AgaJ11 is an endo-type β-agarase that hydrolyzes agarose into neoagarohexaose, neoagarotetraose, and neoagarobiose. Therefore, this study shows that AgaJ11 from G. joobiniege G7 is a novel GH16 β-agarase with an acidic enzymatic feature that may be useful for industrial applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
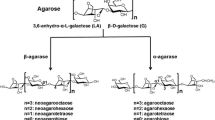
Agar is a heterogeneous polysaccharide composed of agarobiose and porphyrobiose, which is the main component of the cell wall of red algae, such as Gelidium and Gracilaria [1, 2]. Agarobiose consists of a linear chain of alternating residues of α-1,3 linked 3,6-anhydro-α-l-galactose and β-1,4 linked d-galactose [2]. Porphyrobiose is composed of alternating residues of β-1,4 linked d-galactose and α-1,3 linked l-galactose-6-sulfate [3, 4]. Agar-degrading bacteria identified from various environments are assigned to the genera Alteromonas [5], Pseudomonas [6], Vibrio [7, 8], Pseudoalteromonas [9], Streptomyces [10, 11], Flammeovirga [12], Zobellia [13], Catenovulum [14], Agarivorans [15], Cohnella [16], Bacillus [17], Thalassomonas [18], Stenotrophomonas [19], Simiduia [20], and Gayadomonas [21]. Agar can be specifically hydrolyzed by agarases found in these bacteria [1].
Agarases (EC 3.2.1.81), belonging to glycoside hydrolases (GH) family, can be classified into three groups based on the cleavage pattern: α-agarase, β-agarase, and β-porphyrannase [1]. α-Agarases hydrolyze α-1,3 glycosidic linkages of agarose to produce agarooligosaccharides with 3,6-anhydro-α-l-galactose at the reducing end, whereas β-agarases hydrolyze β-1,4 linkages to produce neoagarooligosaccharides with d-galactose residues at the reducing ends [1, 22]. β-Porphyranases hydrolyze the β-1,4 linkages of d-galactose and l-galactose-6-sulfate to produce oligosaccharides with d-galactose residues at the reducing ends [4]. Until now, many β-agarases have been identified in taxonomically diverse genera [1]. They mainly belong to four distinct families (GH16, GH50, GH86, and GH118) [1]. Unlike β-agarase, only two α-agarases and β-porphyranases were identified [4, 18, 23], and they belong to the GH96 and GH16 families, respectively. Cleaved neoagarobiose is hydrolyzed into two monosaccharide, d-galactose and 3,6-anhydro-α-l-galactose, by a neoagarobiose hydrolase or a neoagarooligosaccharide hydrolase [24–26]. The catabolic pathways for utilization of d-galactose are well determined in many bacteria such as Escherichia coli [27], whereas a catabolic pathway for utilization of 3,6-anhydro-α-l-galactose was recently determined in Vibrio sp. strain EJY3 [28]. 3,6-Anhydro-α-l-galactose can be further catabolized to 3,6-anhydrogalactonate and 2-keto-3-deoxy-galactonate, by an NADP+-dependent 3,6-anhydro-α-l-galactose dehydrogenase and an 3,6-anhydrogalactonate cycloisomerase, respectively [28]. Therefore, Vibrio sp. EJY3 was capable of catabolizing 3,6-anhydro-α-l-galactose as a sole carbon source [28]. These agarolytic enzymes can be used for bioconversion of agarose biomass into biofuels or industrial chemicals.
Gayadomonas joobiniege G7 is a novel agar-degrading bacterium recently isolated from a coastal seawater sample from Gaya Island [21]. In this report, a putative agarase (AgaJ11) of G. joobiniege G7 was purified and its biochemical characterization was elucidated. Based on its activity and pH stability, AgaJ11 is an acidic endo-type β-agarase.
Materials and Methods
Bacterial Strains and Culture Conditions
Plasmid pET-28a was used for cloning and expression in E. coli strain ER2566. E. coli was routinely grown in Luria-Bertani (LB) medium at 37 °C. Kanamycin (50 μg/ml) was added when required.
Molecular Cloning of the Agarase Gene agaJ11
The 873-bp gene fragment encoding AgaJ11 (NCBI Reference Sequence: WP_040459371.1) lacking the predicted secretion signal peptide (amino acids 1–26) was amplified by PCR from G. joobiniege G7 genomic DNA using a forward primer (5′- CGCGCGGCAGCCATATGCTAAATCATGAGCGTTTGGC-3′; NdeI site underlined) and a reverse primer (5′-GCTCGAATTCGGATCCGTTTTACGCATTATCTAGTT-3′; BamHI site underlined). Amplification reactions were performed in a Q-Cycler (Quanta Biotech, England) for 5 min at 95 °C, followed by 40 cycles of 95 °C for 20 s, 55 °C for 20 s, and 72 °C for 1 min/kb, concluding with extension at 72 °C for 5 min. After digestion, the NdeI- BamHI fragment was inserted into the corresponding sites of pET-28a to make pHis-AgaJ11.
Expression and Purification of Recombinant AgaJ11
E. coli ER2566 cells harboring pHis-AgaJ11 were grown in LB at 37 °C to an OD600 of 0.5 and induced with 0.1 mM (final concentration) isopropyl-β-d-thiogalactopyranoside (IPTG) for 3 h at 30 °C. Harvested cells were resuspended in binding buffer (50 mM Tris-HCl (pH 8.0), 300 mM NaCl) and then passed two times through a French pressure cell at 10,000 p.s.i. Remaining cells and cell debris were removed by centrifugation at 15, 000 g for 20 min at 4 °C. The soluble fraction was loaded onto the BD TALON™ metal affinity resin and bound proteins were eluted with binding buffer containing 200 mM imidazole. To remove imidazole, the eluent was dialyzed using 50 mM Tris-HCl (pH 8.0) containing 50 mM NaCl for 12 h. The purity of the protein preparation was estimated by SDS-PAGE with Coomassie brilliant blue staining, and the final concentration was determined by Bradford assay [29].
Agarase Assay by 3,5-Dinitrosalicylic Acid Method
The standard assay of agarase activity was determined by measuring the amount of released reducing sugar equivalent using the 3,5-dinitrosalicylic acid (DNS) method as described previously [9]. Briefly, 20 μl of the enzyme solution was mixed with 500 μl of 10 mM sodium citrate buffer (pH 4) containing 0.2 % agarose. After incubation at 40 °C for 30 min, 500 μl of DNS reagent solution (6.5 g of DNS, 325 ml of 2 N NaOH, and 45 ml of glycerol in 1 l distilled water) was added to the sample and boiled at 100 °C for 10 min, followed by cooling the sample in ice-cold water for 2 min. The amount of reducing sugars was monitored at 540 nm against a blank in which 20 μl of distilled water was used instead of the enzyme solution. The optimum pH condition was determined at 40 °C under various pH conditions: 3–6 (10 mM sodium citrate), 6–7 (10 mM MOPS), 7–9 (10 mM Tris-HCl), 9–10 (10 mM Glycine-NaOH). The optimum temperature was determined for temperatures ranging from 20 to 60 °C in 10 mM sodium citrate buffer (pH 4.5). The pH stability was tested under the defined optimal conditions (40 °C and pH 4.5) after pre-incubation of AgaJ11 at pH ranging from 3 to 10 at 30 °C for 1 h. The effect of metal ions on AgaJ11 activity was measured at 40 °C in 10 mM sodium citrate buffer (pH 4.5) containing various metal ions (KCl, ZnCl2, CuCl2, CaCl2, MnCl2, NiCl2, NaCl, CoCl2, and FeCl2) or EDTA at a final concentration of 5 mM.
Determination of Kinetic Parameters
The K m and V max kinetic parameters of purified AgaJ11 were determined in the defined optimal conditions (40 °C and pH 4.5) with various amounts of agarose ranging from 0.1 to 8 mg/ml. The samples in 10 mM sodium citrate buffer (pH 4.5) were incubated at 40 °C for 10 min, which was adequate to limit substrate utilization below 5 % [30]. The K m and V max values were calculated from the Lineweaver-Burk plot.
Zymogram Assay for AgaJ11 Activity on SDS-PAGE
The unboiled AgaJ11 protein sample was loaded onto a 10 % polyacrylamide gel containing 0.3 % agarose. After electrophoresis, the gel was immersed in 2 % Triton X-100 for 30 min, washed twice with 20 mM Tris-HCl (pH 8) for 30 min, and incubated in 20 mM sodium citrate buffer (pH 4.5) at 37 °C overnight. The gel was then stained with Lugol’s iodine solution.
Measurement of Viscosity
Kinetic viscosity of AgaJ11 was determined using a DV2T viscometer (Brookfield AMETEK, USA). The enzymatic reaction was carried out at 30 °C in 20 mM sodium citrate buffer (pH 4.5) containing 0.5 % agarose, and viscosity was measured at various reaction times ranging from 0 to 60 min.
Thin Layer Chromatography (TLC) Analysis of the Reaction Products
Purified AgaJ11 was incubated with 0.1 % agarose or neoagarooligosaccharides dissolved in 10 mM sodium citrate buffer (pH 4.5) at 40 °C. The enzymatic reaction was stopped by heating the mixture in boiling water for 5 min. After cooling on ice for 5 min, the sample was loaded onto a Silica Gel 60 plate (Merck, USA). The plate was developed with an n-butanol/acetic acid/water solution (2:1:1, by volume). Spots were visualized by spraying with 20 % H2SO4 in methanol and heating at 120 °C for 2 min [31].
Mass Spectrometer Analysis of Hydrolysis Products
The unstained regions corresponding to hydrolysis products on the TLC plates were scraped out with razor and dissolved in methanol. The dissolved sample was dried and its molecular mass was determined using a matrix-assisted laser desorption ionization time-of-flight (MALDI-TOF) mass spectrometer (Autoflex III, Germany) by using 2-(4′-hydroxybenzeneazo) benzoic acid (HABA) as the matrix.
Results
Identification of a Novel Agarase-Encoding Gene of G. joobiniege G7
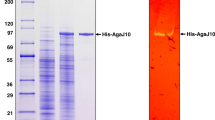
In the G. joobiniege genome, we searched for a gene encoding a putative β-agarase. The agaJ11 gene of G. joobiniege, encoding 317 amino acids (NCBI Reference Sequence: WP_040459371.1), was predicted to be a β-agarase belonging to the GH16 family with an E-value of 7.61 × e−73. Comparative sequence analyses using BLAST showed that this protein displayed 44 % identity (61 % similarity) with a putative β-agarase (GenBank accession No. GAD03772.1) from an agarolytic marine bacterium Agarivorans albus MKT 106 [32], and 39 % identity (54 % similarity) with a characterized β-agarase DagA (GenBank accession No. CAB61795.1) from Streptomyces coelicolor A3(2) [11]. Based on the SignalP program (http://www.cbs.dtu.dk/services/SignalP), the agaJ11 gene was predicted to have an N-terminal signal sequence peptide (amino acids 1–26). To characterize this putative β-agarase, the agaJ11 gene fragment, encoding a mature form lacking the predicted secretion signal peptide, was cloned into a pET28a vector to produce a His-tagged fusion recombinant protein. The hexahistidine-tagged recombinant AgaJ11 protein was expressed in E. coli ER2566 and purified using TALON™ metal affinity chromatography (Fig. 1a). The molecular weight of purified AgaJ11 was estimated to be approximately 35 kDa on SDS-PAGE, which is consistent with the calculated molecular weight (34,921 Da). Zymogram analysis showed that AgaJ11 has agarase activity (Fig. 1b).
Purification and zymogram analysis of AgaJ11. a The recombinant AgaJ11 was purified using metal affinity chromatography and separated on an SDS-PAGE gel after boiling for 5 min. Lane M, size marker; lane 1, cell extract of ER2566 cells containing pHis-AgaJ11 before induction; lane 2, cell extract of ER2566 cells containing pHis-AgaJ11 after induction with 1 mM IPTG; lane 3, AgaJ11 purified by metal affinity chromatography. b Zymogram analysis of purified AgaJ11. Purified AgaJ11 was loaded onto a 10 % polyacrylamide gel containing 0.3 % agarose. After the enzymatic reaction, the gel was stained with Lugol’s iodine solution
Enzymatic Properties
The effect of pH on AgaJ11 activity was determined by measuring relative activity within a pH range of 3 to 10 (Fig. 2a). AgaJ11 had maximum agarase activity at pH 4.5 in 10 mM sodium citrate buffer. The enzyme was active only at a narrow pH range from 4 to 5, and less than 30 % of enzymatic activity retained at other pH conditions, indicating that this enzyme is an acidic agarase. The effect of temperature on the agarase activity of AgaJ11 was examined by measuring the relative activity at various temperatures ranging from 20 to 60 °C (Fig. 2b). AgaJ11 showed a maximum activity at 40 °C and was active only at a narrow temperature range from 30 to 40 °C. The temperature stability was tested under the defined optimal conditions after pre-incubation at different temperatures for 1 h (Fig. 2b). AgaJ11 was stable at a temperature range from 20 to 30 °C, but its stability dramatically declined at 40 °C. The effect of metal ions on AgaJ11 activity was determined by measuring the relative activity in the presence of various metal ions (Fig. 2c). The agarase activity was strongly inhibited by several metal ions including CuCl2, FeCl2, MnCl2, ZnCl2, and CaCl2, whereas it was moderately inhibited by CoCl2 (40 % remaining) and NiCl2 (50 % remaining). EDTA showed a slight inhibitory effect, indicating that cofactors of metal ions are not necessary for the enzymatic activity of AgaJ11. The K m and V max of the dimeric AgaJ11 for agarose were 21.42 mg/ml and 25 U/mg, respectively (Fig. 3).
Effects of pH, temperature, and metal ions on AgaJ11 activity. a The enzymatic activity was determined at 40 °C under various pH conditions: diamonds, 3–6 (10 mM sodium citrate); squares, 6–7 (10 mM MOPS); triangles, 7–9 (10 mM Tris-HCl); circles, 9–10 (10 mM Glycine-NaOH). b The agarase activity of AgaJ11 was determined at temperatures ranging from 20 to 60 °C in 10 mM sodium citrate buffer (pH 4.5) (diamonds). The temperature stability was tested under the defined optimal conditions after pre-incubation at temperature ranging from 20 to 60 °C for 1 h (circles). c The effect of metal ions on AgaJ11 activity was measured at 40 °C in 10 mM sodium citrate buffer (pH 4.5) containing various metal ions or EDTA at a final concentration of 5 mM
Identification of AgaJ11 Hydrolysis Products
To determine whether AgaJ11 is an α- or β-agarase, we tested substrate specificity of AgaJ11 using two chromogenic substrates, p-nitrophenyl-α-d-galactopyranoside and p-nitrophenyl-β-d-galactopyranoside. Unexpectedly, AgaJ11 did not show any activity toward either substrate (data not shown). To address whether AgaJ11 has an endo- or exo-type agarase, we timed the change in viscosity of the agarose solution during the agarose hydrolysis reaction. The viscosity of the agarose solution rapidly decreased for the first 5 min and dropped slowly until the end of the reaction (Fig. 4a). This result implies that AgaJ11 is an endo-acting agarase.
Analysis of hydrolyzed products by the AgaJ11 agarase. a The enzymatic reaction of AgaJ11 was carried out at 40 °C in 10 mM sodium citrate buffer (pH 4.5) containing 0.5 % agarose, and kinetic viscosity was measured at various reaction times ranging from 0 to 60 min. b TLC analysis of hydrolyzed products of agarose depending on reaction time. The enzymatic reactions were performed at 40 °C for the indicated time in 10 mM sodium citrate buffer (pH 4.5) containing 0.1 % agarose. c The enzymatic reaction of AgaJ11 using various neoagarooligosaccharides as substrates. After the hydrolysis reactions of various neoagarooligosaccharides at 40 °C for 24 h, reaction products were separated on a silica gel 60 TLC plate. NA2 neoagarobiose, NA4 neoagarotetraose, NA6 neoagarohexaose
The hydrolysis products produced by AgaJ11 were determined by TLC (Fig. 4b). TLC results showed that agarose was rapidly degraded to neoagarobiose (NA2), neoagarotetraose (NA4), neoagarohexaose (NA6), and a trace amount of neooligosaccharides larger than NA6 during the late stage of the reaction. We also carried out TLC analysis using NA2, NA4, and NA6 as a substrate (Fig. 4c). NA2 and NA4 were not hydrolyzed by AgaJ11, whereas NA6 was entirely hydrolyzed to NA4 and NA2 by cleavage of the β-1,4 glycosidic bond after overnight incubation. The molecular masses of hydrolysis products were examined by mass spectrometer analysis (Fig. 5). The oligosaccharides corresponding to NA4 and NA6 were extracted from the TLC plates. MALDI-TOF analysis revealed that the hydrolysis products had a molecular ion at m/z of 653 (M + Na)+ and 959 (M + Na)+, corresponding to NA4 and NA6, respectively. Therefore, we conclude that AgaJ11 is an endo-type β-agarase producing NA2, NA4, and NA6 from agarose.
MALDI-TOF mass spectrum of hydrolysis products. NA4 (a) and NA6 (b) spots on the TLC plate in Fig. 4b were extracted by methanol and its molecular mass was determined by a direct MALDI-TOF mass spectrometer
Discussion
In this study, we identified a novel β-agarase AgaJ11 in an agar-degrading marine bacterium, G. joobiniege G7 [21]. This novel enzyme has a molecular mass of 35 kDa and is an agarase with optimum activity at pH 4.5 and temperature 40 °C (Fig. 2). The end product of agarose hydrolysis by AgaJ11 is NA2, NA4, and NA6 (Fig. 4).
AgaJ11 has a well-conserved domain with β-agarases belonging to the GH16 family. AgaJ11 has the amino acids that need to form tunnel-shaped active site cavities (Trp85, Trp165, Asp171, Glu174, Asp176, Glu179, Phe198, Arg200, Glu280). AgaJ11 also has a conserved active domain (EIDXXE) containing hydrophobic residues (Ile175, Val177) between two catalytic residues (Glu 174, 179), which is a common feature of the GH16 family.
AgaJ11 is an acidic β-agarase with the maximum enzymatic activity at pH 4.5 and retaining 70 % of its maximum enzymatic activity at pH 4. AgaJ11 is active only at a narrow pH range from 4 to 5, and less than 30 % of its enzymatic activity is retained at other pH conditions. Until now, most characterized β-agarases have the maximum enzymatic activity at a neutral pH range from 6 to 8 [1]. However, two recently discovered β-agarases have been found to work best in acidic conditions. AgaP from the marine bacterium Pseudoalteromonas sp. AG4 shows the maximum activity at pH 5.5 [32, 33]. AgaA from the agar-degrading Pseudoalteromonas sp. AG52 also has an optimal pH at 5.5 [34, 35]. Comparative sequence analyses using BLAST showed that AgaJ11 displayed 32 and 31 % identities with AgaP and AgaA, respectively. Among characterized agarases, AgaJ11 displayed the highest sequence identity (39 %) with a β-agarase DagA from S. coelicolor A3(2) [11]. However, the purified DagA exhibited the maximal agarase activity at pH 7 [11]. Therefore, these data suggest that AgaJ11 from G. joobiniege G7 is a novel GH16 β-agarase with the maximum enzymatic activity at a low pH.
Neoagarooligosaccharides produced by agarase have various biological activities such as moisturizing the skin [36], whitening of melanoma cells [36], antibacterial activity [32], and improving metabolic syndrome in mice (unpublished data). Therefore, AgaJ11 characterized in this study may be a useful enzyme with broad applications in food, pharmaceutical, and cosmetic industries.
References
Chi, W. J., Chang, Y. K., & Hong, S. K. (2012). Agar degradation by microorganisms and agar-degrading enzymes. Applied Microbiology and Biotechnology, 94, 917–930.
Duckworth, M., & Yaphe, W. (1972). The relationship between structures and biological properties of agars. In K. Nisizawa (Ed.), Proceedings of the 7th international seaweed symposium (pp. 15–22). New York: Halstead Press.
Morrice, L. M., McLean, M. W., Long, W. F., & Williamson, F. B. (1983). Porphyran primary structure. An investigation using β-agarase I from Pseudomonas atlantica and 13C-NMR spectroscopy. European Journal of Biochemistry, 133, 673–684.
Hehemann, J. H., Correc, G., Barbeyron, T., Helbert, W., Czjzek, M., & Michel, G. (2010). Transfer of carbohydrate-active enzymes from marine bacteria to Japanese gut microbiota. Nature, 464, 908–912.
Chi, W. J., Park da, Y., Seo, Y. B., Chang, Y. K., Lee, S. Y., & Hong, S. K. (2014). Cloning, expression, and biochemical characterization of a novel GH16 β-agarase AgaG1 from Alteromonas sp. GNUM-1. Applied Microbiology and Biotechnology, 98, 4545–4555.
Hsu, P. H., Wei, C. H., Lu, W. J., Shen, F., Pan, C. L., & Lin, H. T. (2015). Extracellular production of a novel endo-β-agarase AgaA from Pseudomonas vesicularis MA103 that cleaves agarose into neoagarotetraose and neoagarohexaose. International Journal of Molecular Sciences, 16, 5590–5603.
Liao, L., Xu, X. W., Jiang, X. W., Cao, Y., Yi, N., Huo, Y. Y., Wu, Y. H., Zhu, X. F., Zhang, X. Q., & Wu, M. (2011). Cloning, expression, and characterization of a new β-agarase from Vibrio sp. strain CN41. Applied and Environmental Microbiology, 77, 7077–7079.
Dong, J., Tamaru, Y., & Araki, T. (2007). A unique β-agarase, AgaA, from a marine bacterium, Vibrio sp. strain PO-303. Applied Microbiology and Biotechnology, 74, 1248–1255.
Park, Y., Chi, W. J., Park, J. S., Chang, Y. K., & Hong, S. K. (2015). Cloning, expression, and biochemical characterization of a GH16 β-agarase AgaH71 from Pseudoalteromonas hodoensis H7. Applied Biochemistry and Biotechnology, 175, 733–747.
Temuujin, U., Chi, W. J., Chang, Y. K., & Hong, S. K. (2012). Identification and biochemical characterization of Sco3487 from Streptomyces coelicolor A3(2), an exo- and endo-type β-agarase-producing neoagarobiose. Journal of Bacteriology, 194, 142–149.
Temuujin, U., Chi, W. J., Lee, S. Y., Chang, Y. K., & Hong, S. K. (2011). Overexpression and biochemical characterization of DagA from Streptomyces coelicolor A3(2): an endo-type β-agarase producing neoagarotetraose and neoagarohexaose. Applied Microbiology and Biotechnology, 92, 749–759.
Han, W., Cheng, Y., Wang, D., Wang, S., Liu, H., Gu, J., Wu, Z., & Li, F. (2016). Biochemical characteristics and substrate degradation pattern of a novel exo-type β-agarase from the polysaccharide-degrading marine bacterium Flammeovirga sp. strain MY04. Applied Microbiology and Biotechnology, 82, 4944–4954.
Hehemann, J. H., Correc, G., Thomas, F., Bernard, T., Barbeyron, T., Jam, M., Helbert, W., Michel, G., & Czjzek, M. (2012). Biochemical and structural characterization of the complex agarolytic enzyme system from the marine bacterium Zobellia galactanivorans. The Journal of Biological Chemistry, 287, 30571–30584.
Cui, F., Dong, S., Shi, X., Zhao, X., & Zhang, X. H. (2014). Overexpression and characterization of a novel thermostable β-agarase YM01-3, from marine bacterium Catenovulum agarivorans YM01(T). Marine Drugs, 12, 2731–2747.
Lee, D. G., Jeon, M. J., & Lee, S. H. (2012). Cloning, expression, and characterization of a glycoside hydrolase family 118 β-agarase from Agarivorans sp. JA-1. Journal of Microbiology and Biotechnology, 22, 1692–1697.
Li, G., Sun, M., Wu, J., Ye, M., Ge, X., Wei, W., Li, H., & Hu, F. (2015). Identification and biochemical characterization of a novel endo-type β-agarase AgaW from Cohnella sp. strain LGH. Applied Microbiology and Biotechnology, 99, 10019–10029.
Li, J., Sha, Y., Seswita-Zilda, D., Hu, Q., & He, P. (2014). Purification and characterization of thermostable agarase from Bacillus sp. BI-3, a thermophilic bacterium isolated from hot spring. Journal of Microbiology and Biotechnology, 24, 19–25.
Ohta, Y., Hatada, Y., Miyazaki, M., Nogi, Y., Ito, S., & Horikoshi, K. (2005). Purification and characterization of a novel α-agarase from a Thalassomonas sp. Current Microbiology, 50, 212–216.
Zhu, Y., Zhao, R., Xiao, A., Li, L., Jiang, Z., Chen, F., & Ni, H. (2016). Characterization of an alkaline β-agarase from Stenotrophomonas sp. NTa and the enzymatic hydrolysates. International Journal of Biological Macromolecules, 86, 525–534.
Tawara, M., Sakatoku, A., Tiodjio, R. E., Tanaka, D., & Nakamura, S. (2015). Cloning and characterization of a novel agarase from a newly isolated bacterium Simiduia sp. strain TM-2 able to degrade various seaweeds. Applied Biochemistry and Biotechnology, 177, 610–623.
Chi, W. J., Park, J. S., Kwak, M. J., Kim, J. F., Chang, Y. K., & Hong, S. K. (2013). Isolation and characterization of a novel agar-degrading marine bacterium, Gayadomonas joobiniege gen, nov, sp. nov., from the Southern Sea, Korea. Journal of Microbiology and Biotechnology, 23, 1509–1518.
Araki, C. (1959). Seaweed polysaccharides. In M. L. Wolfrom (Ed.), Carbohydrate chemistry of substances of biological interests (pp. 15–30). London: Pergamon Press.
Potin, P., Richard, C., Rochas, C., & Kloareg, B. (1993). Purification and characterization of the α-agarase from Alteromonas agarlyticus (Cataldi) comb. nov., strain GJ1B. European Journal of Biochemistry, 214, 599–607.
Ha, S. C., Lee, S., Lee, J., Kim, H. T., Ko, H. J., Kim, K. H., & Choi, I. G. (2011). Crystal structure of a key enzyme in the agarolytic pathway, α-neoagarobiose hydrolase from Saccharophagus degradans 2-40. Biochemical and Biophysical Research Communications, 412, 238–244.
Hehemann, J. H., Smyth, L., Yadav, A., Vocadlo, D. J., & Boraston, A. B. (2012). Analysis of keystone enzyme in agar hydrolysis provides insight into the degradation (of a polysaccharide from) red seaweeds. The Journal of Biological Chemistry, 287, 13985–13995.
Sugano, Y., Kodama, H., Terada, I., Yamazaki, Y., & Noma, M. (1994). Purification and characterization of a novel enzyme, α-neoagarooligosaccharide hydrolase (α-NAOS hydrolase), from a marine bacterium, Vibrio sp. strain JT0107. Journal of Bacteriology, 176, 6812–6818.
Frey, P. A. (1996). The Leloir pathway: a mechanistic imperative for three enzymes to change the stereochemical configuration of a single carbon in galactose. The FASEB Journal, 10, 461–470.
Yun, E. J., Lee, S., Kim, H. T., Pelton, J. G., Kim, S., Ko, H. J., Choi, I. G., & Kim, K. H. (2015). The novel catabolic pathway of 3, 6-anhydro-L-galactose, the main component of red macroalgae, in a marine bacterium. Environmental Microbiology, 17, 1677–1688.
Zor, T., & Selinger, Z. (1996). Linearization of the Bradford protein assay increases its sensitivity: theoretical and experimental studies. Analytical Biochemistry, 236, 302–308.
Segel, I. H. (1976). Enzyme kinetics. In Biochemical calculations. How to solve mathmatical problems in general biochemistry, 2nd edn (pp. 214–229). New York: Wiley.
Duckworth, M., & Yaphe, W. (1970). Thin-layer chromatographic analysis of enzymic hydrolysates of agar. Journal of Chromatography, 49, 482–487.
Fu, X. T., & Kim, S. M. (2010). Agarase: review of major sources, categories, purification method, enzyme characteristics and applications. Marine Drugs, 8, 200–218.
Oh, C., Nikapitiya, C., Lee, Y., Whang, I., Kim, S. J., Kang, D. H., & Lee, J. (2010). Cloning, purification and biochemical characterization of beta agarase from the marine bacterium Pseudoalteromonas sp. AG4. Journal of Industrial Microbiology & Biotechnology, 37, 483–494.
Yasuike, M., Nakamura, Y., Kai, W., Fujiwara, A., Fukui, Y., Satomi, M., & Sano, M. (2013). Draft genome sequence of Agarivorans albus strain MKT 106 T, an agarolytic marine bacterium. Genome Announcements, 1(4), e00367–e00313.
Oh, C., Nikapitiya, C., Lee, Y., Whang, I., Kang, D. H., Heo, S. J., Choi, Y. U., & Lee, J. (2010). Molecular cloning, characterization and enzymatic properties of a novel β-agarase from a marine isolate Psudoalteromonas sp. AG52. Brazilian Journal of Microbiology, 41, 876–889.
Kobayashi, R., Takisada, M., Suzuki, T., Kirimura, K., & Usami, S. (1997). Neoagarobiose as a novel moisturizer with whitening effect. Bioscience, Biotechnology, and Biochemistry, 61, 162–163.
Acknowledgments
This work was supported by the National Institute of Biological Resources (NIBR) funded by the Ministry of Environment (MOE) of the Republic of Korea (NIBR201629201) and the Advanced Biomass R&D Center (ABC) of Global Frontier Project funded by the Ministry of Science, ICT and Future Planning (ABC-2015M3A6A2065700).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Jung, S., Jeong, BC., Hong, SK. et al. Cloning, Expression, and Biochemical Characterization of a Novel Acidic GH16 β-Agarase, AgaJ11, from Gayadomonas joobiniege G7. Appl Biochem Biotechnol 181, 961–971 (2017). https://doi.org/10.1007/s12010-016-2262-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-016-2262-x