Abstract
Here, we report the comprehensive genome analysis of Enterobacter cloacae SBP-8 and its pan-genome comparison to other Enterobacter species. Genome comparison was performed by the BioRing Image Generator Tool, which predicted its genome similarity to other closest strains. Detailed annotations of the genome revealed different families of carbohydrate-degrading enzymes (CAZymes) that might be involved in the degradation of decaying plant materials with industrial significance for biomass research and the food industry. Moreover, the microbial secretome contains a number of proteins that generally interact with other microbes, environmental space, and the host. The secretion of proteins is performed by the secretory system present in the bacterial cell, however, only a few secretome proteins have been studied in limited bacterial strains. Therefore, we explored the secretome analysis of E. cloacae SBP-8 by nano LC-MS/MS and identified 776 proteins that provide insight into the role of these proteins in bacterial adaptation to different environments. The secretome proteins were classified into molecular, biological, and cellular functions using Blast2GO, whereas the subcellular localization was predicted by the PSORB tool. Many of the secretome proteins belonged to virulence proteins including Type VI secretion system (T6SS) proteins, chemotaxis & fimbriae formation proteins, outer membrane proteins (OMPs), and exotoxin secretion. Moreover, STRING analysis showed the interaction among the identified proteins. The detailed investigations of secretome proteins may provide an understanding of the possible pathogenicity of test organism E. cloacae SBP-8, and is also clinically important for the development of biomarkers and therapeutic approaches.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Enterobacter genus belongs to the gram-negative rod-shaped bacteria, facultative anaerobe, non-spore-forming, belonging to Enterobacteriaceae family. E. cloacae is mainly known as an opportunistic pathogen that causes various infections and has emerged as a nosocomial pathogen (Mezzatesta et al. 2012). Despite the presence of E. cloacae in diverse environments, their pathogenic mechanisms and features contributing to virulence, and antibiotic resistance are not understood yet. Due to the high prevalence of β-lactamases and carbapenemases, this organism is involved in nosocomial infections after Escherichia coli and Klebsiella pneumoniae (Jarlier and INVS 2014). WHO (World Health Organization) has declared the Enterobacter species among the top ten pathogens with multiple drug resistance (Rello et al. 2019). The other member, E. aerogenes also show virulence due to the horizontal acquisition of antibiotic resistance genes and mobile elements from other Enterobacteriaceae species, which are integrated into the genome as its own ancestral heritage (Diene et al. 2013).
Among the Enterobacteriaceae, E. cloacae are ubiquitous in terrestrial and aquatic environments, and are also commonly found in the intestinal microflora of humans and animals. This habitat diversity resulted in the genetic variety of E. cloacae (Mezzatesta et al. 2012). Despite their prominent role in virulence, previous studies (Bhise et al. 2017; de Zélicourt et al. 2018), demonstrated the role of Enterobacter sp. as a plant growth-promoting microorganism, and the genome sequence was analyzed to unravel the plant growth-promoting genes (Andrés-Barraoet al. 2017). In our previous study (Singh et al. 2017), we observed the various plant growth-promoting genes in SBP-8, however, detailed analysis of the genome was lacking. Therefore, in the present study, we investigated the presence of antibiotic resistance genes using the CARD (comprehensive antibiotic resistance database), virulence factors by the VFDB (virulence factor database), secondary metabolite production using antiSMASH, the presence of CAZymes family enzymes, metabolic pathway genes using the KEGG (kyoto encyclopedia of genes and genomes) database, and pan-genome analysis.
The extracellular metabolites secreted by Enterobacter species might also help the microorganism survive under stressful conditions. Genome analysis of E. cloacae ‘Ghats1’ unravel various genes related to chemotaxis, motility and hydrolytic enzymes, which supports the endophyte nature of the bacterium that colonizes and adapts to plants (Shastry et al. 2020). Genome analysis of E. cloacae ATCC identified multidrug efflux genes (37), antimicrobial resistance (7), β-lactamase (11), and operons (7) involved in heavy metal resistance (Ren et al. 2010). The genome analysis predicted CAZymes that have a prominent role in the production of biofuels, and various food products with prebiotic characteristics etc. (Linares-Pasten et al. 2014). In a previous study, we reported the preliminary genome analysis of test isolate SBP-8 (Singh et al. 2017), however, detailed genome investigation was lacking.
Genomic assessment only allows the theoretical prediction of genes, however, proteomic approaches are adequate to evaluate the protein profiles in particular conditions like culture growth phase, temperature, and growth medium etc., to compare the exact sets of proteins expressed in a particular condition (Chitlaru and Shafferman 2009). Moreover, it leads to a detailed characterization of secreted proteins involved in drug resistance and furthermore, these secreted proteins represent a valuable target as a future drug (Van Oudenhove and Devreese 2013; Ding et al. 2020). The secretome analysis of Bacillus velezensis LC1 demonstrated a large number of proteins involved in the degradation of cellulase, and hemicellulase etc. by using LC-MS/MS (liquid chromatography with tandem mass spectrometry) (Tang et al. 2019). The secreted proteins of Burkholderia cepacia such as proteases, cytotoxins, and hemolysins cause life-threatening infections in cystic fibrosis patients (Carvalho et al. 2007).
The genomic and comparative genomic analyses only provide information about the presence and absence of genes in a particular strain, however, the secretome analysis reflects the secreted proteins and their possible contribution to the bacterial lifestyle. The secreted proteins may also modulate the immune system of the host for virulence effects (Lee and Schneewind 2001), therefore, secretome analysis unravels the proteins involved in pathogenesis and virulence, if any (Wang and Wang 2019). The factors involved in virulence may not be predicted at the genomic level, therefore, we analyzed the extracellular proteome of E. cloacae SBP-8 using the LC-MS/MS approach. Overall, 776 proteins were identified in the culture supernatant, and further detailed analysis was performed.
Materials and methods
Genome analysis of SBP-8
In brief, the selected strain was isolated from the rhizospheric soil of Sorghum bicolor growing in the desert region of Rajasthan, India (28°18′N, and 74°58′E). De-novo genome sequencing was performed by the Illumina Paired-end (PE) sequencing platform and Illumina paired-end raw reads were quality-checked using FastQC. De-novo assembly of Illumina PE data was performed using the SPAdes assembler and assembled contigs were further scaffolded using the SSPACE program. The gene ontology of SBP-8 was performed against the Uniprot Bacterial Database using the Diamond Blast Program (Buchfink 2015), and pathway analysis was carried out using the KAAS database (Moriya et al. 2007). The gene sequences were used for metabolic pathway analysis by using BLAST comparisons against the manually curated KEGG (Kanehisa et al. 2004). The comparison and annotation of orthologous gene clusters were analyzed using Orthovenn2 using default parameters using protein sequences of E. cloacae SBP-8 (NZ_CP017413.1), E. mori LMG 25,706 (NZ_GL890780.1), E. cancerogenus ATCC 35,316 (LR593888.1), E. cloacae ATCC 13,047 (NC_014121.1), E. cloacae GS1 (AJXP01000001.1), and E. cloacae subsp. dissolvens SDM (CP003678.1) (Xu et al. 2019). The genomic island of SBP-8 was predicted by IslandViewer4 (Bertelli et al. 2017). These genomes were selected based on the closest similarity in the RAST data base. The circular genome comparison of SBP-8 was performed against the reference genomes of E. cloacae GS1, E. cloacae ATCC 13,047, E. cloacae subsp. dissolvens SDM, E. mori LMG 25,706, and E. cancerogenus ATCC 35,316 using the BRIG (Blast Ring Image Generator) Tool (Alikhan et al. 2011). The secondary metabolite biosynthetic gene clusters (BGCs) were identified by antiSMASH version 5.1.2 and BAGEL-4 and the cluster finder algorithm was used to detect BGC-like regions in the genome (Weber et al. 2015; Blin et al. 2019).
Pan-genome analysis
To identify the presence of core genes and shared genes among the selected genome, pan-genome analysis was performed using Roary 3.11.2 with default settings. The PROKKA 1.14.5 was used for the generation of GFF3 genome files for selected genomes including SBP-8 (Seemann 2014). The program SNP-sites 2.5.1 (https://github.com/sanger-patho gens/snp-sites) was used for the analysis of the maximum likelihood (ML) phylogenetic tree (Page et al. 2016). For identification of core genes in the tested genome, Phandango was used and Roary software was used to assess the proportions of the pan-genome.
AMRs and CAZyme analysis
To identify the presence of any AMR genes in the SBP-8 genome, CARD database was searched against the SBP-8 genome sequence using a homology-based approach (BLASTX) (Mcarthur et al. 2013). To reveal the presence of CAZymes such as glycoside hydrolases (GHs), polysaccharide lyases (PLs), carbohydrate esterases (CEs), polysaccharide lyases (PLs), glycosyl transferases (GTs) and carbohydrate-binding modules CBMs, the protein sequences of E. cloacae SBP-8 was annotated using the dbCAN2 server (Zhang et al. 2018) against the CAZy database (Buchfink et al. 2015a, b; Lombard et al. 2014).
Bacteria culturing for secretome study
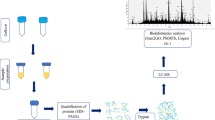
A loopful of bacterial colonies of E. cloacae SBP-8 harvested from an LB-agar plate were inoculated into 5 ml of LB-broth medium (Himedia, India), and incubated at 37 ℃ on a rotator shaker at 180 rpm. After overnight incubation (approximately 18 h), the broth cultures were centrifuged at 5,000 g for 20 min. To test the viability of bacteria, 100 µl of aliquots was spread on an LB-agar plate. The extracellular released proteins in the supernatants were separated and filtered through a 0.2 μm filter to remove any residual bacteria.
The extracellular proteins in the supernatant were extracted by TCA precipitation following the standard method (Deatherage Kaiser et al. 2015) with minor modifications. For the extraction, the ratio of TCA (100% w/v) to the supernatant was 1:4 and incubated at -20 ℃ for 30 min. Further, the extracted protein in the broth was recovered in pellet form by centrifugation at 12,000 g at 4 ℃ for 20 min. The obtained pellet was washed with 500 µL cold acetone two times and desalting of the protein sample was performed by diluting the protein sample to 500 µL in lysis buffer. Further, ultracentrifugation was done through a 3 K Ultra-0.5 centrifugal filter device at 14,000 g for 15 min at 4 ℃. The concentrated proteins from the columns were eluted in sterile collection tubes by another centrifugation at 2,000 g for 2 min at 4 ℃. A negative control containing only the broth medium was used in parallel.
Nano LC-MS/MS
Protein identification was performed using the nano LC-ESI-MS/MS method at the proteomics laboratory (VProteomics, India). The replicate protein samples from the secretome were pooled and tryptic digestion was performed with 400 ng of proteins (Sigma-Aldrich, USA). The digested peptides were applied to nanoLC-ESI-MS/MS (Shevchenko et al. 2006). The trapped peptides were desalted using 1% acetonitrile/0.5% formic acid as eluent. The eluted peptides were separated on a C18, 75 μm × 150 mm column using a 5 to 40% acetonitrile /0.1% formic acid gradient within 2 h. The MS spectra were recorded as per the manufacturer’s instrument settings for nanoLC-ESI-MS/MS analyses with a scan range of 300–1500 m/z (Advani et al. 2019). The identification of proteins was performed by the analysis of MS/MS spectra to the Mascot search engine and through a non-redundant protein database (Perkins et al. 1999).
Bioinformatics analysis
The signal peptides compelling the secretion of secretome proteins were predicted using SiganlP (http://www.cbs.dtu.dk/services/SignalP/) (Bendtsen et al. 2004), and PSORTb (http://www.psort.org/psortb/) tools (Yu et al. 2010). Proteins secreted by non-classical routes were determined by SecretomeP 2.0 http://www.cbs.dtu.dk/services/SecretomeP/) (Bendtsen et al. 2005a). The lipoproteins were searched using LipoP (http://www.cbs.dtu.dk/services/LipoP/) and PRED-LIPO (http://bioinformatics.biol.uoa.gr/PRED-LIPO/input.jsp) (Bagos et al. 2008). The functional annotation of the secretome proteins was performed using Gene Ontology (GO) tool and Blast2GO (Conesa and Gotz 2008; Conesa et al. 2005). The functional categorization of enzymatic proteins distributed in various metabolic pathways was predicted by the KEGG database. The presence of any virulence proteins was predicted by the Virulence Factor Data Base (Chen et al. 2005). The protein-protein interaction was analyzed by STRING (von Mering et al. 2005).
Results
Genome analysis
The genome sequence of SBP-8 is available with Genbank accession no. CP016906. The functional analysis of COGs (clusters of orthologous groups) showed a good number of genes involved in LysR-family transcriptional regulator (65), MFS family permeases (55), type I fimbriae protein (30), DNA-binding transcriptional regulator (PurR family 26), and OMR (21) etc. (Fig. 1a). KEGG analysis identified that the highest number of genes (175) belonged to ABC transporters, followed by the two-component system (146), purine metabolism (66), and quorum sensing (59) etc. (Fig. 1b). Using Island viewer, GI (genomic island) was identified in the E. cloacae SBP-8 genome which includes various proteases and survival proteins that assist the bacteria in survival under varying environments (Suppl. Figure 1).
a Kyoto Encyclopedia of Genes and Genomes (KEGG) were utilized for the retrieval of metabolic pathways of E. cloacae SBP-8 genes. KAAS server at KEGG database used for genes functional annotation by BLAST comparison with the manually curated database of KEGG- GENES; 1b The cluster of orthologus analysis in E. cloacae SBP-8 using the COGs database based on orthologous groups. The identified genes in SBP-8 were divided into several functional subcategories based on the COG annotation (http://www.ncbi.nlm.nih.gov/COG/)
Comparative genome analysis
The genomic features of all strains used for comparative analysis have been summarized in Table 1. The Orthovenn2 analysis demonstrated the presence of unique and shared proteins by the tested genome. It was found that six proteins were shared by all six species. Similarly, 127 proteins were shared by five species, 2206 shared by four species, 1551, and 456 shared among three, and two species, respectively (Fig. 2a). A total of 59 protein clusters were specific to only a single genome. Out of the 59 gene clusters, 16 belonged to E. cloacae ATCC 13,047, 14 to E. cloacae subsp. dissolvens SDM, 11 to E. cloacae SBP-8, 8 to E. cancerogenus ATCC 35,316, and 3 belongs to E. cloacae GS1. In the cell graph, the first pattern shows the gene clusters, the second pattern represents the cluster counts and the third pattern in terms of the stacker bar displayed at the right position illustrates the total protein counts (Fig. 2b). The pairwise heatmap generated by SBP-8 and other five tested strains highlight the overlapping gene clusters in a pairwise manner. Between SBP-8 and E. cloacae GS1, the lowest thresholds of overlapping gene clusters were recorded (Fig. 2c).
a OrthoVenn diagram showing the number of common and separate protein clusters in E. cloacae SBP-8 (NZ_CP017413.1), E. mori LMG 25,706 (NZ_GL890780.1), E. cancerogenus ATCC 35,316 (LR593888.1), E. cloacae ATCC 13,047 (NC_014121.1), E. cloacae GS1 (AJXP01000001.1), and E. cloacae subsp. dissolvens SDM (CP003678.1) b The occurrence table contains groups of gene clusters like cluster count and protein count. Row indicates the orthologous gene cluster for multiple species that summarized as a cell graph and column indicates different closely related bacterial species, c The pairwise protein sequence comparison for heatmap showing orthologous clusters between SBP-8 and other closely related strains
The circular genome comparison of E. cloacae SBP-8 was carried out against five reference genomes i.e. E. cloacae GS1, E. cloacae ATCC 13,047, E. cloacae subsp. dissolvens SDM, E. mori LMG 25,706, and E. cancerogenus ATCC 35,316 using the BRIG (v 0.95) tool which helps to determine the homologous regions between sets of genomes. The output image shows the highest homology with the reference genome of E. cloacae 13,047 (98.7% identity and 96% coverage), followed by E. cloacae subsp. dissolvens SDM (97.5% identity and 94% coverage), E. cloacae GS1 (96.8% identity and 93% coverage), E. mori LMG 25,706 (95.6% identity and 93.5% coverage), and E. cancerogenus ATCC 35,316 (95% identity and 92.5% coverage) (Suppl. Figure 2).
The pan-genome analysis was performed with nine closely related genomes of E. cloacae (i.e. E. cloacae 109, E. cloacae 3849, E. cloacae 764, E. cloacae A1137, E. cloacae STN0717, E. cloacae EN3600, E. cloacae dissolvens SDM, E. cloacae MY490, and E. cloacae GGT036 in the Roary tool (Fig. 3a). The matrix showed that the most closely related strains of SBP-8 belonged to E. cloacae MY490 and E. cloacae GGT036. The pan-genome showed that all strains contained 14,919 protein-encoding genes in their genome. Further analysis predicted that 1959 genes (21%) were core genomes, whereas 6587 genes (44%) were identified as shell genomes. A total of 6364 genes (40%) were identified as cloud genomes (Fig. 3b). The analysis showed that each strain contained 71 unique genes which correspond to approximately 2% of each genome (Fig. 3c).
Prediction of biosynthetic gene clusters (BGCs)
The anti-SMASH analysis identified genes clustering for non-ribosomal peptide synthesis (NRPS) like amonabactin (Region 1), thiopeptides like O-antigen (Region 2), and siderophores like aerobactin (Region 3) (Fig. 4). As per anti-SMASH, amonabactin is confined in the nucleotide position from approximately 1,238,311 to 1,282,171. The amonabactin gene cluster showed the highest similarity of 60% to the bacillibactin region of Bacillus velezensis FZB42 and 57% similarity to the amonabactin region of Aeromonas hydrophila ATCC 7966 (Fig. 4). The lowest similarity of 7% was observed for the malleobactin region of Burkholderia thailandensis E264. A 14% similarity of the thiopeptide gene cluster was observed for Pseudomonas aeruginosa (Fig. 4). It is localized in the region from 1,590,173 to 1,616,470. region 3 gene clusters belonging to aerobactin showed 100% similarity to Pantoea ananatis, 66% similarity to Xenorhabdus szentirmaii DSM 16,338 and localized in the region from 3,095,268 to 3,109,696 (Fig. 4). Using BAGEL-4, we identified the gene cluster for outer membrane lipoprotein, molybdenum-containing enzyme dimethyl sulfoxide reductase, metabolic active enzyme formate acetyltransferase, and phosphoserine aminotransferase etc. (Fig. 5; Suppl. Table 1)
AMRs and CAZy analysis
CARD analysis identified the antimicrobial resistance genes belonging to beta-lactam antibiotics, MFS-family of antibiotic efflux pumps, the ABC-antibiotic efflux pumps, the RND-family of antibiotic efflux pumps, and fosfomycin thiol transferase (Suppl. Figure 3). The identified AMR genes and their functional AMR- mechanism have been summarized in Suppl. Table 2. The CAZy analysis of E. cloacae SBP-8 identified 71 GHs, 31 GTs, 8 CEs, 4 AAs, 5 CBMs, and 1 PLs (Suppl. Table 3). Among the GHs, the higher subcategory was found to be GH13, and GH109, followed by GH23. Among GTS, the major subcategory was observed for GT2 followed by GT9 and GT4. Among CEs, the major subcategory was found to be CE1 and CE4. In AAs, an equal number of genes were recorded for each subcategory. Among CBMs, an equal number was noted for CBM48 and CBM 73. A sum of 1145 CAZyme-encoding sequences were observed among SBP-8, and closely related species. The number of CAZyme-encoding sequences was highest (n = 140) and lowest (n = 94) for E. cloacae ATCC 13,047 and E. cloacae GS1, respectively (Fig. 6).
Analysis of the secretome of E. cloacae SBP-8
By the analysis of the supernatant of the bacterial broth culture, we found 776 proteins in the supernatant by LC-MS/MS-based approach (Suppl. Figure 4). The detailed information about the secretome proteins has been provided in Suppl. File 2. The secretome was further categorized into three groups based on the signal peptide such as non-classical (63%), classical (31%), and lipoprotein (6%) (Fig. 7a). The secretome study of SBP-8 also reveals the localization of secretomes in the cytoplasmic, periplasmic, cytoplasmic membrane, outer membrane, and extracellular proteins. Among all secretome proteins, 64% were associated with the cytoplasmic, 10% with the periplasmic, 7% with the cytoplasmic membrane, 4% with the outer membrane, and 2% with extracellular proteins. A total of 13% of proteins were identified as unknown proteins (Fig. 7b).
a The categorization of E. cloacae SBP-8 secretome proteins into non-classical, classical and lipoproteins; b Subcellular localization of the identified secretome proteins. The FASTA protein sequences was analyzed by bioinformatics tool PSORTb for their subcellular localization such as extracellular, cytoplasmic, outer membrane, periplasmic, and cytoplasmic membrane
Gene ontology analysis
Gene ontology (GO) analysis of the secretome of E. cloacae SBP-8 was performed using the Blast2GO tool, which categorized the proteins into three groups “biological process”, “molecular function” and “cellular component”. In the biological process, 67% were associated with the metabolic process, 10% biosynthetic process, 6% regulation process, and 4% in localization and cellular response, respectively (Fig. 8a). In the ‘molecular functions’ 59% proteins are associated with binding proteins, 12% with catalytic activity, 8% with hydrolase activity and 7% with transferase activity (Fig. 8b). In the category of cellular component, 20% were associated to intracellular anatomical structure and membrane protein, 22% with cytoplasmic protein, 7% with cell periphery, 6% with periplasmic space and organelle protein, and 4% with extracellular, flagellar, and cell projection proteins, respectively (Fig. 8c).
Virulence proteins in E. cloacae SBP-8 secretome
The secretome study identified the different virulence proteins in E. cloacae SBP-8 using VFDB. Targeted Type VI secretion system (T6SS) proteins and their effector proteins like Hcp (Hemolysin co-regulated protein), VgrG (valine-glycine repeat protein), TssB, TssC, ClpB, VasK, VasI and VasK were found (Table 2). In the secretome, we observed the various exotoxin proteins related to hemolysin, structural toxin protein, putative aminomethyltransferase, ornithine carbamoyltransferase, precolibactin peptidase, and colibactin biosynthesis acyltransferase etc. (Table 3). Various antimicrobial resistance proteins belonging to acriflavine resistance, putative carbonic anhydrase, cyanide-forming glycine dehydrogenase, and fatty acid efflux system protein etc. were noted (Table 4). The various biofilm-forming proteins related to type 3 fimbrial major pilin (mrkA), biofilm-associated protein (bap), and biofilm-controlling response regulator (bfmR) etc. were observed (Table 5). The various flagellar proteins and the associated flagellar components such as flagellar hook protein (FlgE), flagellar hook associated protein (FlgK & FlgL), flagellar basal body rod protein (FlgG), and ATP-binding protein (FlhG) etc. were identified (Suppl. Table 4).
KEGG pathway analysis
The KEGG analytical tool was used for the prediction of the pathway associated with all the proteins in the secretome. The KEGG pathway analysis revealed the secretome proteins involved in the 90 different metabolic pathways. Out of which, the most abundant protein number of 25 was involved in purine metabolism. A total of 23 proteins were linked to glycolysis & pyruvate metabolism (11.47%), 20 to glycolysis & pyruvate metabolism (11.01%), and 19 to amino acid & pyrimidine metabolism etc. (Suppl. Figure 5a). Based on the abundance (8–13), proteins related to glyoxylate & galactose metabolism, drug metabolism, carbon fixation, citric acid cycle, fatty acid biosynthesis, and arginine & glutathione metabolism have been summarized in Suppl. Figure 5b.
Protein network analysis
The protein-protein network analysis among the chaperone proteins, transporter protein, outer membrane proteins, and flagellar proteins was performed using the STRING database. The PPI analysis of chaperone proteins showed a close interaction between chaperone proteins DnaK and ClpB. Similarly, a tight association was recorded among chaperone protein SurA and Skp (Suppl. Figure 6a). The observed STRING results showed a BamA (outer membrane protein assembly factor) was dominant, which showed the close interaction with BamC (outer membrane protein assembly factor), and BamB (outer membrane protein assembly factor) (Suppl. Figure 6b). Among flagellar proteins, close association was observed among flagellar hook-associated protein, flagellar hook protein FlgE, and flagellar basal-rod protein FlgG (Suppl. Figure 6d). The identified transporters proteins showed a varying level of interaction. The integrated protein-protein interaction was observed between arginine ABC transporter ATP-binding protein ArtP, ABC transporter arginine-binding protein 2, and arginine ABC transporter substrate-binding protein. Similarly, a close association was observed in-between aminotransferase, N-acetylglucosamine transporter subunit IIABC, and sucrose-specific transporter subunit IIBC (Suppl. Figure 6d).
Discussion
Genome sequencing has unraveled the understanding of the genetic potential of microorganisms and also provides valuable insight about the genetic adaptation of bacteria in diverse environments (Yi et al. 2018). KEGG pathway analysis showed a higher number of genes involved in transporter and two-component systems. The transporter genes play an important role in bacterial growth and survival, as they enable the bacterium to defend against endogenous and environmental stressors (Piepenbreier et al. 2017). Some of the transporters also possess secondary functions and their functions are attributed to sensory, and signaling systems. These functionalities equip the bacteria for decision-making processes that contribute to the bacterial cell adapting any changes in their surrounding environment or intracellular conditions (Schleif 2000). Transporters also enhance substrate utilization by modulating nutrient-sensing which results in enhanced intracellular substrate concentration which activates the genes related to increased rates of import and metabolism (Fritz et al. 2014). In the absence of transporters, substrates are not allowed to enter the cytoplasm of a cell (Megerle et al. 2008).
Bacterial two-component systems (TCSs) play essential roles in cell-cell communication, adaptation to changing environments, and pathogenesis in the case of a pathogenic bacterium. Their absence in humans and other mammals, represents the potential targets for designing, and developing new antibiotics with broader activity (Varughese 2002). Some of the TCSs regulate the gene clusters associated with cell growth, biofilm formation, and virulence activity (Gotoh et al. 2010; Schaefers et al. 2017). The relationship between TCSs and virulence in bacterial pathogens has helped in the development of suitable inhibitors aimed at signal transduction in bacteria. Recently, TCSs have opened a new avenue for antibacterial drug design and they impair the upstream regulatory functions related to pathogen physiology (Poole 2012). Thus, a detailed understanding of TCSs provides an alternate strategy for combating microbial infections (Gotoh et al. 2010).
The analysis of gene distribution within COG categories revealed the main functional gene cluster (> 50) belonging to the LysR family of DNA- binding transcriptional regulators and MFS family permeases. LysR family transcriptional regulators can function either as activators or repressors of gene expression (Schell 1993; Maddocks and Oyston 2008). These regulators control a wide variety of genes related to virulence (Doty et al. 1993), motility (Heroven and Dersch 2006), metabolism (Hartmann et al. 2013), and quorum sensing (O’Grady et al. 2011). MFS (major facilitator superfamily) family transporters comprise a functionally diverse type of transporter superfamily, and are widely expressed in many domains of life (Saier et al. 2014). These transporters are involved in the transport of monosaccharides, peptides, drugs, nucleotides, iron chelates, and many inorganic cations and anions (Guan and Kaback 2006; Newstead et al. 2011).
The genomic analysis identified the various genes associated with secondary metabolite production, predicted using BAGEL-4 and anti-SMASH. The genomes of Enterobacter sakazakii (Mullane et al. 2008), Enterobacter sp. CGMCC 5087 (Liu et al. 2018), and many other strains of Enterobacter (Mohite et al. 2022) also reported the gene clusters for non-ribosomal peptides, thiopeptides and siderophore productions. Members of amonabactin-producing bacteria play an essential role in iron acquisition from transferrin or lactroferrin, and in the establishment of pathogenesis (Balado et al. 2015; Esmaeel et al. 2016; Stintzi and Raymond 2000). The O-specific polysaccharides or O-antigens are major surface components of Gram-negative bacteria and are highly variable in structure. These are predominantly found in many pathogenic microorganisms (Rietschel et al. 1996), however, another study demonstrated their role in the interaction between Rhizobiaceae and plants (Kannenberg et al. 1998). Aerobactin is synthesized by the iucABCD-encoded gene products, and is a hydroxamate siderophore produced by many pathogenic microorganisms (Carbonetti and Williams 1984; Gross et al. 1984).
Pan-genome analysis indicated that every Enterobacter strain encodes a certain number of unique genes and the Enterobacter genus possesses an open pan-genome harboring genes that are not shared by other strains. As the genomes of more strains are sequenced, many new genes will continue to be identified (Xing et al. 2019). The pan-genome features enable the strains to survive in different environmental niches. The genome comparison showed its high resemblance to other E. cloacae genomes. The detailed genome analysis identified various genes for complex carbohydrate degradation. The identified GHs and GTs in the SBP-8 genome are well known for the hydrolysis of glycosidic bonds (Lopez-Mondejar et al. 2016), whereas AAs, PLs, and CBMs are involved in the degradation of biopolymers (Koeck et al. 2014).
The secretome analysis of E. cloacae SBP-8 by mass spectrometry provides a detailed study of various proteins involved in the interaction of bacteria with the host and their environment (Boekhorst et al. 2006). Bioinformatics-based analysis using tools like signalP, secretome P, and PRED LIPO, categorized the secretome proteins into three groups classical, non-classical, and lipoproteins. The lipoproteins are produced by many prokaryotes and further translocated through the Sec or Tat-dependent pathway across the membrane (Thompson et al. 2010). These peripherally anchored proteins play an important role in the virulence, physiology, and immune response in microbial pathogens. Furthermore, these are recognized as excellent vaccine targets (Nguyen and Gotz 2016). In M. tuberculosis, these lipoproteins impart an important role in virulence and evasion of the immune system (Su et al. 2016).
In bacterial pathogens, iron is involved in supporting the growth and virulence activity during the infection process. Due to a lack of free iron availability, bacterial pathogens utilize alternate strategies to acquire host-associated iron proteins (Brown and Holden 2002). In the secretome, we observed proteins related to iron acquisition, however, some of the identified proteins are also involved in virulence such as the FagC protein (Billington et al. 2002). We observed CiuA siderophore, a high-affinity iron uptake system localized in the operon ciuABCDE. Similarly, we observed the Fhu siderophore protein, an integral part of the ferric hydroxymate uptake system. Previous studies showed that in Listeria monocytogene, Fhu contributes to the ferric hydroxamate uptake from complex conjugated forms of iron (Jin et al. 2006; Xiao et al. 2011).
A number of chaperone proteins were identified in which DnaK (Hsp60) expresses on the cell surface that can be used as adhesins and virulence factors to release them into the extracellular milieu (Goulhen et al. 1998; Henderson et al. 2006). These chaperones also maintain protein disaggregation in the presence of a prolonged stress environment. The other observed chaperone protein e.g. elongation factor Tu facilitates protein acclimation under stress conditions. Additionally, elongation factor Tu interacts with unfolded/or denatured proteins similar to other chaperones that are involved in protein folding, and renaturation under stress conditions (Caldas et al. 1998, 2000).
Outer membrane proteins play an important role in the regulation or transportation of metabolites as well as small molecules between bacteria and the extracellular milieu, and play an important factor in the maintenance of drug resistance (Trias et al. 1989). Here, we identified 30 proteins as outer membrane proteins belonging to E. cloacae SBP-8, which include outer membrane protein (OmpA, OmpC, OmpW, OmpX), outer membrane protein assembly factor (BamA, BamB, BamC), outer membrane channel protein ToIC, and outer membrane lipoprotein Lpp which shows higher expression (Nirujogi et al. 2014). The OmpA provides linkage between the outer membrane and the underlying peptidoglycan layer. The Ompx outer membrane protein belongs to the highly conserved protein family, and plays an important role in the virulence and neutralization of the host defense mechanism (Heffernan et al. 1994). Bam stands for β-barrel assembly machinery complex, which acts for the insertion of β-barrel protein into the outer membrane. BamA is a 16-stranded β-barrel POTRA (five polypeptides transport-associated) domain that is present on the periplasm (Bakelar et al. 2016). Exotoxins are the diffusible proteins secreted from the pathogens to their external environment. The numbers of exotoxins investigated in E. cloacae SBP-8 are argK, clbG, clbP, cyaB, cylF, cylG, hlyA, hlyB, and rtxA, in which clbG is mainly involved in the synthesis of colibactin, and the development of meningitis in mice (Wang et al. 2021).
Adhesion factors in pathogenic bacteria play an important role in host-microbe interactions, colonization, persistence, and virulence (Hammerschmidt 2006; Navarre and Schneewind 1999). In the secretome of SBP-8, we observed the pili protein that plays an important role in cellular adhesion (Soares et al. 2013b). Flagellar proteins are associated with bacterial adhesion as well as invasion by providing motility to target cells and receptors, mainly FliC the flagellin protein involved in adhesion and invasion. FliC contains C- and A-terminal domains due to which it is involved in innate immunity (Haiko and Westerlund-wikström 2013). A number of flagellar proteins were investigated in this study and the enrichment of flagellar proteins or clusters in E. cloacae SBP-8 suggested a better understanding of pathogenesis during the extracellular transportation of the proteins. The flagellar functions are mainly studied for bacterial pathogens and virulence, due to the applications of flagellin and flagella in vaccine development and diagnosis (Gat et al. 2011). Due to the high immunogenic property of FliC, it can be used as a vaccine adjuvant with poorly immunogenic antigens, such as FliC or anti-FliC antibodies for the diagnosis of diseases like melioidosis and inflammatory bowel (Wajanarogana et al. 2013). Flagellar biosynthesis regulator FlhF, flagellin family FliC & FlaA, flagellar basal body protein FlgD & FlgG, flagellar hook protein FlgE/FlgK/FlgL, and ATP synthase Flil & PapX protein regulates flagellum synthesis to repress motility. These are the flagellum proteins that are involved in pathogenesis (Macnab 2004).
By using the KEGG database, nine pathways proteins were associated with antibiotic biosynthesis. The identified antibiotic biosynthesis pathways enzymes such as O-acetyltransferase, phosphorylase, kinase, reductase, phosphoribosyl-transferase, dehydrogenase, hexokinase, phosphatase, glucokinase, and penicillinase are essential for the antibiotic biosynthesis (Barnard-Britson et al. 2012; Liu et al. 2016). The most prominent pathways belonged to pyruvate metabolism, purine metabolism, and glycolysis/gluconeogenesis which help the bacteria in the utilization of nutrients (Cezairliyan and Ausubel 2017).
The KEGG enrichment analysis identified the most dominant pathways as the metabolic pathway, biosynthetic pathway, and degradation pathway in which the maximum numbers of proteins are involved. The other dominating pathways are purine metabolism, pyruvate metabolism, and amino acid metabolism in which nucleosidase, kinase, formyltransferase, cyclohydrolase, phosphorylase, reductase, carboxykinase, isomerase, and so many other enzymes are involved. Metabolic pathways are generally required by the bacteria during growth in the host (Tang et al. 1999). Many proteins associated with glycolytic pathways like 2,3 bisphosphoglycerate-phosphoglycerate mutase, enolase, pyruvate dehydrogenase, and acyl-transferase were observed. Many of the glycolytic enzymes can show multi-functional proteins rather than just basic components of the glycolytic pathway. A previous study demonstrated the secretion of glycolytic pathway enzymes under varying pH conditions (Giardina et al. 2014).
In the secretome, we noted conserved ATP-binding cassette transporter proteins like Opp-transporter that play diverse functions related to the virulence, signalling and nutrition of bacteria (Yu et al. 2010). Additionally, these Opp-proteins are involved in uptake of peptides from the external medium and contribute to infection (Danelishvili et al. 2014). In the secretome, we observed the RND-family of multidrug export proteins that contributes resistance against the microbiocidal peptides (Blodkamp et al. 2016). In the secretome, we also observed the export proteins related to ABC-type antimicrobial peptide transporters and the MATE family protein NorM.
In conclusion, in-depth genome analysis of SBP-8 revealed the presence of CAZymes, which have an important role in complex carbohydrate metabolism, and thereby, represent their significance in the industrial sector. A remarkably high proportion of genes were involved in transporters and various metabolic pathways which might play an important role in environmental adaptation and stress tolerance. Similarly, the presence of many BGCs in the SBP-8 genome contributes to its environmental suitability through different metabolic pathways. Additionally, the secretome analysis by nanoLC-MS/MS showed the various secreted proteins that might contribute to the possible virulence features of the environmental isolate E. cloacae SBP-8. These proteins and their related corresponding genes will be studied using a suitable model system to understand the molecular mechanism of pathogenicity in the future. We believe that detailed genome and secretome characterization will facilitate further exploitation of Enterobacter strains for biotechnology applications. Overall, in-depth genome characterization and secretome analysis of an environmental Enterobacter bacterium provide insight into metabolic potential, biotechnological applications, and possible pathogenicity.
Data availability
The data that support the findings of this study are available from the corresponding author, upon reasonable request.
References
Advani J, Verma R, Chatterjee O, Balaya RDA, Najar MA, Ravishankara N, Suresh S, Pachori PKD, Gupta UD, Pinto SM, Chauhan DSD, Tripathy SP, Gowda H, Prasad TSK (2019) Rise of clinical microbial proteogenomics: a multiomics approach to non-tuberculous mycobacterium—the case of Mycobacterium abscessus UC22. J Integr Biol 23:1–16. https://doi.org/10.1089/omi.2018.0116
Alikhan NF, Petty NK, Ben Zakour NL, Beatson SA (2011) BLAST Ring Image Generator (BRIG): simple prokaryote genome comparisons. BMC Genomics 12:402. https://doi.org/10.1186/1471-2164-12-402
Andrés-Barrao C, Lafi FF, Alam I, De Zélicourt A, Eida AA, Bokhari A, Alzubaidy H, Bajic VB, Hirt H, Saad MM (2017) Complete genome sequence analysis of Enterobacter sp. SA187, a plant multi-stress tolerance promoting endophytic bacterium. Front Microbiol 8:2023. https://doi.org/10.3389/fmicb.2017.02023
Bagos PG, Tsirigos KD, Liakopoulos TD, Hamodrakas SJ (2008) Prediction of lipoprotein signal peptides in Gram-positive bacteria with a hidden markov model. J Proteome Res 7(12):5082–5093. https://doi.org/10.1021/pr800162c
Bakelar J, Buchanan SK, Noinaj N (2016) The structure of the β-barrel assembly machinery complex. Science 351:180–186. https://doi.org/10.1126/science.aad3460
Balado M, Souto A, Vences A, Careaga VP, Valderrama K, Segade Y, Rodríguez J, Osorio CR, Jiménez C, Lemos ML (2015) Two catechol siderophores, acinetobactin and amonabactin, are simultaneously produced by Aeromonas salmonicida subsp. salmonicida sharing part of the biosynthetic pathway. ACS Chem Biol 10(12):2850–2860. https://doi.org/10.1021/acschembio.5b00624
Barnard Britson S, Chi X, Nonaka K, Spork AP, Tibrewal N, Goswami A, Pahari P, Ducho C, Rohr J, Van Lanen SG (2012) Amalgamation of nucleosides and amino acids in antibiotic biosynthesis: discovery of an L-threonine: uridine-5′-aldehyde transaldolase. J Am Chem Soc 134:18514–18517. https://doi.org/10.1021/ja308185q
Bendtsen JD, Nielsen H, von Heijne G, Brunak S (2004) Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 340:783–795. https://doi.org/10.1016/j.jmb.2004.05.028
Bendtsen JD, Kiemer L, Fausboll A, Brunak S (2005a) Nonclassical protein secretion in bacteria. BMC Microbiol 5:58. https://doi.org/10.1186/1471-2180-5-58
Bertelli C, Laird MR, Williams KP, Lau BY, Hoad G et al (2017) IslandViewer 4: expanded prediction of genomic islands for larger-scale datasets. Nucleic Acids Res 45:W30–W35. https://doi.org/10.1093/nar/gkx343
Bhise KK, Bhagwat PK, Dandge PB (2017) Plant growth-promoting characteristics of salt tolerant Enterobacter cloacae strain KBPD and its efficacy in amelioration of salt stress in Vigna radiata L. J. Plant Growth Regul 36(1):215–226. https://doi.org/10.1007/s00344-016-9631-0
Billington SJ, Esmay PA, Songer JG, Jost BH (2002) Identification and role in virulence of putative iron acquisition genes from Corynebacterium pseudotuberculosis. FEMS Microbiol Lett 208(1):41–45. https://doi.org/10.1111/j.1574-6968.2002.tb11058.x
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY, Medema MH, Weber T (2019) antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res 47:W81–W87. https://doi.org/10.1093/nar/gkz310
Blodkamp S, Kadlec K, Gutsmann T, Naim HY, von Köckritz-Blickwede M, Schwarz S (2016) In vitro activity of human and animal cathelicidins against livestock-associated methicillin-resistant Staphylococcus aureus. Vet Microbiol 194:107–111. https://doi.org/10.1016/j.vetmic.2015.09.018
Boekhorst J, Helmer Q, Kleerebezem M, Siezen RJ (2006) Comparative analysis of proteins with a mucus-binding domain found exclusively in lactic acid bacteria. Microbiology 152:273–280. https://doi.org/10.1099/mic.0.28415-0
Brown JS, Holden DW (2002) Iron acquisition by Gram-positive bacterial pathogens. Microbes Infect 4:1149–1156. https://doi.org/10.1016/S1286-4579(02)01640-4
Buchfink B, Xie C, Huson DH (2015a) Fast and sensitive protein alignment using DIAMOND. Nat Methods 12:59–60. https://doi.org/10.1038/nmeth.3176
Buchfink B, Xie C, Huson D (2015b) Fast and sensitive protein alignment using DIAMOND. Nat Methods 12:59–60. https://doi.org/10.1038/nmeth.3176
Caldas TD, El Yaagoubi A, Richarme G (1998) Chaperone properties of bacterial elongation factor EF-Tu. J Biol Chem 273:11478–11482. https://doi.org/10.1074/jbc.273.19.11478
Caldas T, Laalami S, Richarme G (2000) Chaperone properties of bacterial elongation factor EF-G and initiation factor IF2. J Biol Chem 275:855–860. https://doi.org/10.1074/jbc.275.2.855
Carbonetti NH, Williams PH (1984) A cluster of five genes specifying the aerobactin iron uptake system of plasmid ColV-K30. Infect Immun 46(1):7–12. https://doi.org/10.1128/iai.46.1.7-12.1984
Carvalho APDA, Ventura GMC, Pereira CB, Leão RS, Folescu TW, Higa L, Teixeira LM, Maciel, Plotkowski MC, Merquior VLC, Albano RM, Marques EA (2007) Burkholderia cenocepacia, B. multivorans, B. Ambifaria and B. Vietnamiensis isolates from cystic fibrosis patients have different profiles of exoenzyme production. Apmis 115:311–318. https://doi.org/10.1111/j.1600-0463.2007.apm_603.x
Cezairliyan B, Ausubel FM (2017) Investment in secreted enzymes during nutrient-limited growth is utility dependent. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.1708580114
Chen L, Yang J, Yu J, Yao Z, Sun L, Shen Y, Jin Qi (2005) VFDB: a reference database for bacterial virulence factors. Nucleic Acids Res 33:D325–D328. https://doi.org/10.1093/nar/gki008
Chitlaru T, Shafferman A (2009) Proteomic studies of Bacillus anthracis. Future Microbiol 4:983–998. https://doi.org/10.2217/fmb.09.73
Conesa A, Gotz S (2008) Blast2GO: a comprehensive suite for functional analysis in plant genomics. Int J Plant Genomics 1–12. https://doi.org/10.1155/2008/619832
Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18):3674–3676. https://doi.org/10.1093/bioinformatics/bti610
Danelishvili L, Babrak L, Rose SJ, Everman J, Bermudez LE (2014) Mycobacterium tuberculosis alters the metalloprotease activity of the COP9 signalosome. mBio 5:e01278–e01214. https://doi.org/10.1128/mbio.01278-14
de Zélicourt A, Synek L, Saad MM, Alzubaidy H, Jalal R, Xie Y, Andrés-Barrao C, Rolli E, Guerard F, Mariappan KG, Daur I (2018) Ethylene induced plant stress tolerance by Enterobacter sp. SA187 is mediated by 2-keto-4‐methylthiobutyric acid production. PLoS Genet 14(3):e1007273. https://doi.org/10.1371/journal.pgen.1007273
Deatherage Kaiser BL, Wunschel DS, Sydor MA, Warner MG, Wahl KL, Hutchison JR (2015) Improved proteomic analysis following trichloroacetic acid extraction of Bacillus anthracis spore proteins. J Microbiol Methods 118:18–24. https://doi.org/10.1016/j.mimet.2015.08.008
Diene SM, Merhej V, Henry M, El Filali A, Roux V, Robert C, Azza S, Gavory F, Barbe V, La Scola B, Raoult D, Rolain JM (2013) The rhizome of the multidrug-resistant Enterobacter aerogenes genome reveals how new killer bugs are created because of a sympatric lifestyle. Mol Biol Evol 30:369–383. https://doi.org/10.1093/molbev/mss236
Ding M, Tegel H, Sivertsson Å, Hober S, Snijder A, Ormö M, Strömstedt PE, Davies R, Holmberg Schiavone L (2020) Secretome-based screening in target discovery. SLAS Discov 25(6):535–551. https://doi.org/10.1177/2472555220917113
Doty SL, Chang M, Nester EW (1993) The chromosomal virulence gene, cheV, of Agrobacterium tumefaciens is regulated by a LysR family member. J Bacteriol 175:7880–7886. https://doi.org/10.1128/jb.175.24.7880-7886.1993
Esmaeel Q, Chevalier M, Gabrielle C, Leclere V (2016) Non ribosomal peptide synthetase with a unique iterative-alternative-optional mechanism catalyzes amonabactin synthesis in Aeromonas. Appl Microbiol Biotechnol 100:8453–8463. https://doi.org/10.1007/s00253-016-7773-4
Fritz G, Megerle JA, Westermayer SA, Brick D, Heermann R, Jung K, Radler JO, Gerland U (2014) Single cell kinetics of phenotypic switching in the arabinose utilization system of E. Coli. PLoS ONE 9:e89532. https://doi.org/10.1371/journal.pone.0089532
Gat O, Galen JE, Tennant S, Simon R, Blackwelder WC, Silverman DJ, Pasetti MF, Levine MM (2011) Cell-associated flagella enhance the protection conferred by mucosally-administered attenuated Salmonella Paratyphia vaccines. PLoS Negl Trop Dis 5(11):e1373. https://doi.org/10.1371/journal.pntd.0001373
Giardina S, Scilironi C, Michelotti A, Samuele A, Borella F, Daglia M, Marzatico F (2014) In vitro anti-inflammatory activity of selected oxalate degrading probiotic bacteria: potential applications in the prevention and treatment of hyperoxaluria. J Food Sci 79:384–390. https://doi.org/10.1111/1750-3841.12344
Gotoh Y, Eguchi Y, Watanabe T, Okamoto S, Doi A, Utsumi R (2010) Two-component signal transduction as potential drug targets in pathogenic bacteria. Curr Opin Microbiol 13:232–239. https://doi.org/10.1016/j.mib.2010.01.008
Goulhen F, Hafezi A, Uitto VJ, Hinoda D, Nakamura R, Grenier D, Mayrand D (1998) Subcellular localization and cytotoxic activity of the GroEL-like protein isolated from actinobactera ctinomycetemcomitans. Infect Immun 66:5207–5313. https://doi.org/10.1128/iai.66.11.5307-5313.1998
Gross R, Engelbrecht F, Braun V (1984) Genetic and biochemical characterization of the aerobactin synthesis operon on pColV. Mol Gen Genet 196(1):74–80. https://doi.org/10.1007/BF00334095
Guan L, Kaback HR (2006) Lessons from lactose permease. Annu Rev Biophys Biomol Struct 35:67–91. https://doi.org/10.1146/annurev.biophys.35.040405.102005
Haiko J, Westerlund-Wikström B (2013) The role of the bacterial flagellum in adhesion and virulence. Biology (Basel) 2:1242–1267. https://doi.org/10.3390/biology2041242
Hammerschmidt S (2006) Adherence molecules of pathogenic Pneumococci. Curr Opin Microbiol 9(1):12–20. https://doi.org/10.1016/j.mib.2005.11.001
Hartmann T, Zhang B, Baronian G, Schulthess B, Homerova D, Grubmüller S, Kutzner E, Gaupp R, Bertram R, Powers R, Eisenreich W, Kormanec J, Herrmann M, Molle V, Somerville GA, Bischoff M (2013) Catabolite control protein E (CcpE) is a LysR-type transcriptional regulator of tricarboxylic acid cycle activity in Staphylococcus aureus. J Biol Chem 288:36116–36128. https://doi.org/10.1074/jbc.M113.516302
Heffernan EJ, Wu L, Louie J, Okamoto S, Fierer J, Guiney DG (1994) Specificity of the complement resistance and cell association phenotypes encoded by the outer-membrane protein genes rck from Salmonella typhimurium and ail from Yersinia-Enterocolitica. Infect Immunol 62:5183–5186. https://doi.org/10.1128/iai.62.11.5183-5186.1994
Henderson B, Allan E, Coates AR (2006) Stress wars: the direct role of host and bacterial molecular chaperones in bacterial infection. Infect Immun 74(7):3693–3706. https://doi.org/10.1128/iai.01882-05
Heroven AK, Dersch P (2006) RovM, a novel LysR-type regulator of the virulence activator gene rovA, controls cell invasion, virulence and motility of Yersinia pseudotuberculosis. Mol Microbiol 62:1469–1483. https://doi.org/10.1111/j.1365-2958.2006.05458.x
Jarlier V (2014) INVS Surveillance of multidrug resistant bacteria in french healthcare facilities BMR-raisin network données 2012. Saint-Maurice: Institut de Veille Sanitaire. Available at: http://www.invs.sante.fr
Jin B, Newton SMC, Shao Y, Jiang X, Charbit A, Klebba PE (2006) Iron acquisition systems for ferric hydroxamates, haemin and haemoglobin in Listeria monocytogenes. Mol Microbiol 59:1185–1198. https://doi.org/10.1111/j.1365-2958.2005.05015.x
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32:277–280. https://doi.org/10.1093/nar/gkh063
Kannenberg EL, Reuhs BL, Forsberg LS, Carlson RW (1998) Lipopolysaccharides and K-antigens: their structures, biosynthesis and functions. In: Spaink HP, Kondorosi A, Hooykaas PJJ (eds) The Rhizobiaceae. Kluwer Academic, Dordrecht, pp 119–154. https://doi.org/10.1007/978-94-011-5060-6_7
Koeck AS, Loker F, Miglior DF, Kelton J, Jamrozik J, Schenkel FS (2014) Genetic relationships of clinical mastitis, cystic ovaries, and lameness with milk yield and somatic cell score in first-lactation Canadian holsteins. J Dairy Sci 97:5806–5813. https://doi.org/10.3168/jds.2013-7785
Lee VT, Schneewind O (2001) Protein secretion and the pathogenesis of bacterial infections. Genes Dev 15:1725–1752. https://doi.org/10.1101/gad.896801
Linares-Pasten J, Andersson M, Karlsson E (2014) Thermostable glycoside hydrolases in biorefinery technologies. Curr Biotechnol 3:26–44. https://doi.org/10.2174/22115501113026660041
Liu Z, Jeppesen PB, Gregersen S, Bach Larsen L, Hermansen K (2016) Chronic exposure to proline causes amino acid toxicity and impaired beta-cell function: studies in vitro. Rev Diabet Stud 13(1):66–78. https://doi.org/10.1900/RDS.2016.13.66
Liu C, Zhang K, Cao W, Zhang Ge, Chen G, Yang H, Wang Q, Liu H, Xian M, Haibo Zhang H (2018) Genome mining of 2-phenylethanol biosynthetic genes from Enterobacter sp. CGMCC 5087 and heterologous overproduction in Escherichia coli. Biotechnol Biofuels. https://doi.org/10.1186/s13068-018-1297-3
Lombard V, Golaconda Ramulu H, Drula E, Coutinho PM, Henrissat B (2014) The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res 42:490–495. https://doi.org/10.1093/nar/gkt1178
Lopez-Mondejar R, Zuhlke D, Becher D, Riedel K, Baldrian P (2016) Cellulose and hemicellulose decomposition by forest soil bacteria proceeds by the action of structurally variable enzymatic systems. Sci Rep 6:25279. https://doi.org/10.1038/srep25279
Macnab RM (2004) Type III flagellar protein export and flagellar assembly. Biochem Biophys Acta 1694:207–217. https://doi.org/10.1016/j.bbamcr.2004.04.005
Maddocks SE, Oyston PC (2008) Structure and function of the LysR-type transcriptional regulator (LTTR) family proteins. Microbiology 154:3609–3623. https://doi.org/10.1099/mic.0.2008/022772-0
McArthur AG, Waglechner N, Nizam F, Yan A, Azad MA, Baylay AJ, Bhullar K, Canova MJ, De Pascale G, Ejim L, Kalan L, King AM, Koteva K, Morar M, Mulvey MR, O’Brien JS, Pawlowski AC, Piddock LJ, Spanogiannopoulos P, Sutherland AD, Tang I, Taylor PL, Thaker M, Wang W, Yan M, Yu T, Wright GD (2013) The comprehensive antibiotic resistance database. Antimicrob Agents Chemother 57:3348–3357. https://doi.org/10.1128/aac.00419-13
Megerle JA, Fritz G, Gerland U, Jung K, Rädler JO (2008) Timing and dynamics of single cell gene expression in the arabinose utilization system. Biophys J 95:2103–2115. https://doi.org/10.1529/biophysj.107.127191
Mezzatesta ML, Gona F, Stefani S (2012) Enterobacter cloacae complex: clinical impact and emerging antibiotic resistance. Future Microbiol 7:887–902. https://doi.org/10.2217/fmb.12.61
Mohite OS, Lloyd CJ, Monk JM, Weber T, Palsson BO (2022) Pangenome analysis of Enterobacteria reveals richness of secondary metabolite gene clusters and their associated gene sets. Synth Syst Biotechnol 7(3):900–910. https://doi.org/10.1016/j.synbio.2022.04.011
Mullane N, O’Gaora P, Nally JE, Iversen C, Whyte P, Wall PG, Fanning S (2008) Molecular analysis of the Enterobacter sakazakii O-antigen gene locus. Appl Environ Microbiol 74(12):3783–3794. https://doi.org/10.1128/AEM.02302-07
Navarre WW, Schneewind O (1999) Surface proteins of gram-positive bacteria and mechanisms of their targeting to the cell wall envelope. Microbiol Mol Biol Rev 63:174–229. https://doi.org/10.1128/mmbr.63.1.174-229.1999
Newstead S, Drew D, Cameron AD, Postis VLG, Xia X, Fowler PW et al (2011) Crystal structure of a prokaryotic homologue of the mammalian oligopeptide-proton symporters, PepT1 and PepT2. EMBO J 30:417–426. https://doi.org/10.1038/emboj.2010.309
Nguyen MT, Götz F (2016) Lipoproteins of Gram-positive bacteria: key players in the immune response and virulence. Microbiol Mol Biol Rev 80:891–903. https://doi.org/10.1128/mmbr.00028-16
Nirujogi RS, Pawar H, Renuse S, Kumar P, Chavan S, Sathe G, Sharma J, Khobragade S, Pande J, Modak B, Prasad TS, Harsha HC, Patole MS, Pandey A (2014) Moving from unsequenced to sequenced genome: reanalysis of the proteome of Leishmania donovani. J Proteom 97:48–61. https://doi.org/10.1016/j.jprot.2013.04.021
Page AJ, Taylor B, Delaney AJ, Soares J, Seemann T, Keane JA, Harris SR (2016) SNP-sites: rapid efficient extraction of SNPs from multi-FASTA alignments. Microb Genom 292(4):e000056. https://doi.org/10.1099/mgen.0.000056
Perkins DN, Pappin DJ, Creasy DM, Cottrell JS (1999) Probability based protein identification by searching sequence databases using mass spectrometry data. Electrophor Int J 20(19991201):3551–3567. https://doi.org/10.1002/(SICI)15222683.)20:18%3C3551::AIDELPS3551%3E3.0.CO;2-2
Piepenbreier H, Fritz G, Gebhard S (2017) Transporters as information processors in bacterial signaling pathways. Mol Microbiol. https://doi.org/10.1111/mmi.13633
Poole K (2012) Bacterial stress responses as determinants of antimicrobial resistance. J Antimicrob Chemother 67:2069–2089. https://doi.org/10.1093/jac/dks196
Rello J, Kalwaje Eshwara V, Lagunes L, Alves J, Wunderink RG, Conway-Morris A, Nicolas Rojas J, Alp E, Zhang Z (2019) A global priority list of the TOpTEn resistant microorganisms (TOTEM) study at intensive care: a prioritization exercise based on multi-criteria decision analysis. Eur J Clin Microbiol Infect Dis 38:319–323. https://doi.org/10.1007/s10096-018-3428-y
Ren Y, Ren Y, Zhou Z, Guo X, Li Y, Feng Lu L, Wang L (2010) Complete genome sequence of Enterobacter cloacae subsp. cloacae type strain ATCC 13047. https://doi.org/10.1128/JB.00067-10. Genome Announ
Rietschel ET, Brade H, Holst O, Brade L, Müller-Loennies S, Mamat U, Zähringer U, Beckmann F, Seydel U, Brandenburg K, Ulmer AJ, Mattern T, Heine H, Schletter J, Loppnow H, Schönbeck U, Flad HD, Hauschildt S, Schade UF, Di Padova F, Kusumoto S, Schumann RR (1996) Bacterial endotoxin: chemical constitution, biological recognition, host response, and immunological detoxification. Curr Top Microbiol Immunol 216:39–81. https://doi.org/10.1007/978-3-642-80186-0_3
Saier MH Jr, Reddy VS, Tamang DG, Västermark (2014) The transporter classification database. Nucleic Acids Res 42:D251–D258. https://doi.org/10.1093/nar/gkt1097
Schaefers MM, Liao TL, Boisvert NM, Roux D, Yoder-Himes D, Priebe GP (2017) An oxygen-sensing two-component system in the Burkholderia cepacia complex regulates biofilm, intracellular invasion, and pathogenicity. PLOS Pathog 13:e1006116. https://doi.org/10.1371/journal.ppat.1006116
Schell MA (1993) Molecular biology of the LysR family of transcriptional regulators. Annu Rev Microbiol 47:597–626. https://doi.org/10.1146/annurev.mi.47.100193.003121
Schleif R (2000) Regulation of the L-arabinose operon of Escherichia coli. Trends Genet 16:559–565. https://doi.org/10.1016/S0168-9525(00)02153-3
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Shastry RP, Welch M, Rai VR, Ghate SD, Sandeep K, Rekha PD (2020) The whole-genome sequence analysis of Enterobacter cloacae strain Ghats1: insights into endophytic lifestyle-associated genomic adaptations. Arch Microbiol 202(6):1571–1579. https://doi.org/10.1007/s00203-020-01848-5
Shevchenko A, Tomas H, Havlis J, Olsen JV, Mann M (2006) In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat Protoc 1:2856–2860. https://doi.org/10.1038/nprot.2006.468
Singh RP, Nalwaya S, Jha PN (2017) The draft genome sequence of the plant growth promoting rhizospheric bacterium Enterobacter cloacae SBP-8. Genome Data 12:81–83. https://doi.org/10.1016/j.gdata.2017.03.006
Soares SC, Silva A, Trost E, Blom J, Ramos R, Carneiro A, Ali A, Santos AR, Pinto AC, Diniz C, Barbosa EG, Dorella FA, Aburjaile F, Rocha FS, Nascimento KK, Guimarães LC, Almeida S, Hassan SS, Bakhtiar SM, Pereira UP, Abreu VA, Schneider MP, Miyoshi A, Tauch A, Azevedo V (2013b) The pan-genome of the animal pathogen Corynebacterium pseudotuberculosis reveals differences in genome plasticity between the biovar ovis and equi strains. PLoS ONE 8:e53818. https://doi.org/10.1371/journal.pone.0053818. https://doi.org/10.1371/journal.pone.0053818
Stintzi A, Raymond KN (2000) Amonabactin-mediated iron acquisition from transferrin and lactoferrin by Aeromonas hydrophila: direct measurement of individual microscopic rate constants. JBIC 5:57–66. https://doi.org/10.1007/PL00010655
Tang H, Li Y, Zheng L, Lei L, Yang X, Luo C (2019) Comparative genomic and secretomic characterisation of endophytic Bacillus velezensis LC1 producing bioethanol from bamboo lignocellulose. PREPRINT (Version 1) res. https://doi.org/10.1007/s00203-021-02306-6. Square
Thompson BJ, Widdick DA, Hicks MG, Chandra G, Sutcliffe IC, Palmer T, Hutchings MI (2010) Investigating lipoprotein biogenesis and function in the model Gram-positive bacterium Streptomyces coelicolor. Mol Microbiol 77:943–957. https://doi.org/10.1111/j.1365-2958.2010.07261.x
Trias J, Dufresne J, Levesque RC, Nikaido H (1989) Decreased outer membrane permeability in imipenem-resistant mutants of Pseudomonas aeruginosa. Antimicrob Agents Chemother 33:1201–1206. https://doi.org/10.1128/aac.33.8.1202
Varughese KI (2002) Molecular recognition of bacterial phosphorelay proteins. Curr Opin Microbiol 5:142–148. https://doi.org/10.1016/S1369-5274(02)00305-3
von Mering C, Jensen LJ, Snel B, Hooper SD, Krupp M, Foglierini M, Jouffre N, Huynen MA, Bork P (2005)STRING: known and predicted protein-protein associations, integrated and transferred across organisms. Nucleic Acids Res 33(suppl_1):D433–D437. https://doi.org/10.1093/nar/gki005
Wajanarogana S, Nimnuch P, Thongmee A, Kritsiriwuthinan K (2013) Potential of recombinant flagellin fragment from Burkholderia thailandensis as an antigen for melioidosis antibody detection by indirect ELISA. Mol Cell Probes 27:98–102. https://doi.org/10.1016/j.mcp.2012.11.001
Xiao Q, Jiang X, Moore KJ, Shao Y, Pi H, Dubail I, Charbit A, Newton SM, Klebba PE (2011) Sortase independent and dependent systems for acquisition of haem and haemoglobin in Listeria monocytogenes. Mol Microbiol 80(6):1581–1597. https://doi.org/10.1111/j.1365-2958.2011.07667.x
Xing J, Li X, Sun Y, Zhao J, Miao S, Xiong Q, Zhang Y, Zhang G (2019) Comparative genomic and functional analysis of Akkermansia muciniphila and closely related species. Genes Genomics 41:1253–1264. https://doi.org/10.1007/s13258-019-00855-1
Xu L, Dong Z, Fang L, Luo Y, Wei Z, Guo H, Zhang G, Gu YQ, Coleman-Derr D, Xia Q, Wang Y (2019) OrthoVenn2: a web server for whole-genome comparison and annotation of orthologous clusters across multiple species. Nucleic Acids Res 47:W52–W58. https://doi.org/10.1093/nar/gkz333
Yu NY, Wagner JR, Laird MR, Gabor Melli G, Rey S, Lo R, Dao P, Sahinalp SC, Ester M, Foster LJ, Brinkman FSL (2010) PSORTb 3.0: improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics 26(13):1608–1615. https://doi.org/10.1093/bioinformatics/btq249
Acknowledgements
RPS acknowledge the Dept. of Biotechnology, Jaypee Institute of Information Technology for providing the necessary infrastructure.
Funding
The work was supported by the Ramalingaswami Re-entry Fellowship-2021 provided by Department of Biotechnology, Government of India.
Author information
Authors and Affiliations
Contributions
KK and PKS: analyzed the data. Y Ma and RPS: designed the work and wrote the original draft. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Ethical approval and consent to participate
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for publication
All authors accepted to publish the submitted manuscript.
Competing interests
The authors declare that they don’t have any financial or personal conflicts that might have an effect on the research reported in this study.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kumari, K., Sharma, P., Ma, Y. et al. Comparative genomics and secretome profiling of Enterobacter cloacae SBP-8. Biologia 79, 1957–1973 (2024). https://doi.org/10.1007/s11756-024-01664-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-024-01664-y