Abstract
The aim of this study was to survey marine Actinomycetia (former class Actinobacteria) from nearshore sediments in Kaohsiung Harbor and deep-sea sediments in the Bashi Channel, and to screen them for their abilities to produce metabolites with antimicrobial and cytotoxic activities. A total of 811 actinomycetes were isolated from six marine sediment samples; 70 isolates (8.63%) showed antimicrobial activity using the agar plug method against at least one of two test strains of Bacillus subtilis and Staphylococcus aureus. Twenty-two isolates that displayed high antimicrobial activity were selected for 16S rDNA sequencing and seawater requirement assays. Phylogenetic analysis of the sequences indicated that these isolates were members of the Streptomycetaceae family (1 isolate) and five actinomycete genera: Streptomyces (15), Actinoalloteichus (2), Saccharomonospora (2), Saccharopolyspora (1), and Nocardiopsis (1). Significantly different actinomycete populations were found in the nearshore and deep-sea sediments. Fourteen isolates (63.6%) were found to be obligate marine bacteria, and 2 (9.1%) isolates showed better growth in the presence of seawater. Two isolates, BC01-N-GI05 and BC01-N-MA22, which exhibited antimicrobial activity against at least four test microorganisms as well as strong cytotoxicity (IC50 ≤ 1 μg/mL) against at least two cancer cell lines, were deemed good candidates for further natural product isolation and characterization of bioactive components. Our results suggest that marine sediments from Kaohsiung Harbor and the Bashi Channel could be promising sources of antibiotic-producing Actinomycetia.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Marine microorganisms, having been estimated to account for more than 70% of the world’s marine biomass, are essential in ocean nutrient recycling (Bar-On et al. 2018). They are also mostly under-explored resources with the potential to produce bioactive natural products that are absent in terrestrial microbes (Priyanka et al. 2019; Sharma et al. 2019). The class Actinomycetia (former class Actinobacteria) (Salam et al. 2020) is one of the most prolific supplier of bioactive secondary metabolites (Duncan et al. 2014; Gozari et al. 2019) and accounts for the production of more than 70% of all known drugs and 63% of all known bioactive prokaryotic metabolites (Bérdy 2012), such as antibacterial, antiviral, anticancer, and immunosuppressive agents (Manivasagan et al. 2014; Gao et al. 2021). It is also one of the largest classes of bacteria (Bull and Stach 2007), and is widely distributed in terrestrial, freshwater, and marine ecosystems (Kurtbӧke 2017) where it may function as organic matter and xenobiotic compound degraders (McCarthy and Williams 1992; Solyanikova and Golovleva 2015). Previous studies have shown that marine actinomycetes are not only an active part of marine microbes, but also form stable and persistent populations in different marine environments (Jensen et al. 2005; Das et al. 2006; Patin et al. 2017; Mahapatra et al. 2020). Though bioactive compounds continue to be screened from cultured terrestrial actinomycetes, it has been reported that the opportunity to find new lead compounds from terrestrial actinomycetes has been decreasing over the last three decades (Mincer et al. 2002; Donadio et al. 2010; Davies-Bolorunduro et al. 2021). In addition, many novel medically important compounds have been isolated from marine actinomycetes, including abyssomycin C, a polycyclic antibiotic (Riedlinger et al. 2004), lajollamycin, a lactam antibiotic (Manam et al. 2005), and salinosporamide A, a proteasome inhibitor (Feling et al. 2003). For this reason, cultivation of marine actinomycete taxa for the screening of novel bioactive compounds has become a major focus in marine drug discovery research.
In the past few years, the misuse and overuse of antibiotics in humans and animals has led to the rapid development of multidrug-resistant microbial pathogens, resulting in problems regarding the treatment of infectious diseases (Selvin et al. 2009). Microbial pathogen resistance to antibiotics has become a large global health concern of the twenty-first century (Riyanti et al. 2020). The current antibiotic development pipeline has been found to be insufficient to fight the steady increase in antibiotic-resistant pathogens (Gozari et al. 2019), and it has been predicted that by 2050, drug-resistant pathogens could kill 10 million people per year (de Kraker et al. 2016). In this light, new antibiotic compounds are urgently needed to meet this challenge.
Marine sediments, which cover nearly two-thirds of the Earth’s surface and more than 95% of which are 1000 m below sea level, are a wealthy and common source of diverse Actinomycetia (Bull and Stach 2007; Kamjam et al. 2017; Patin et al. 2017), many of which have been shown to produce novel bioactive metabolites (Olano et al. 2009; Duncan et al. 2014). In addition, diverse new rare (or uncommon) species from novel genera or high taxa of Actinomycetia have also been discovered from marine sediments (Maldonado et al. 2005; Jung et al. 2007; Subramani and Sipkema 2019). These actinomycetes have already adapted to their living environments; therefore, the metabolites they produce vary by locations (Hoshino et al. 2020; Stincone and Brandelli 2020), such as hydrothermal vents (Teske et al. 2002), heavily-polluted coastlines (Ouchene et al. 2022), the deep-sea floor (Jensen et al. 2005; Meena et al. 2019), and polar regions (Yuan et al. 2014). Among these marine environments, the deep sea is a unique, extreme, and difficult-to-access environment characterized by a unique high pressure, low temperature, and low availability of nutrients. The increasing numbers of novel metabolites from actinomycetes have rendered the bioprospecting of these microbes from deep-sea sediments a topic of great interest (Kamjam et al. 2017).
Kaohsiung Harbor, the largest international port and industrial city in Taiwan, is situated on the southwest coast of Taiwan at the intersection of the Taiwan strait and Bashi Channel. It has been suggested that highly-polluted environments, such as Kaohsiung Harbor, could be an interesting source of bioactive actinomycetes (Ouchene et al. 2022). Therefore, the objective of this work was to isolate marine actinomycetes from Kaohsiung Harbor’s nearshore sediments and the Bashi Channel’s deep-sea sediments, southwestern Taiwan, using a culture-based method, screen for antibiotic-producing potentials of these culturable actinomycetes, and finally assay their metabolites for cytotoxic activity.
Materials and methods
Marine sediment sample collection
All marine sediment samples were collected from southwestern Taiwan. Nearshore marine sediment samples KH01, KH02, and KH03 were collected at a depth of about 9-10 m using a sediment sampler grabber in the inner Kaohsiung Harbor, Taiwan (KH01: 22°31′31.38”N, 120°19′6.08″E; KH02: 22°31′40.65”N, 120°18′49.25″E; KH03: 21°32′31.56”N, 120°49′19.01″E). Deep-sea marine sediment samples BC01, BC02, and BC03 were collected using a core sampler at a depth of about 1500 m in the Bashi Channel (21°11′25.79”N, 119°27′43.44″E), with core depths of 0, 15, and 30 cm, respectively. After collection, samples were placed in labeled sterile polyethylene bags, stored at 4 °C in an ice box, brought to the laboratory, and processed within 24 hours.
Pretreatment of sediment samples
To inhibit the growth of non-actinomycetes, a pretreatment procedure was applied to the sediment samples to select spore-forming actinomycetes. Each marine sediment sample was divided aseptically into two groups, each containing about 5 g of marine sediments. The first group (F) of sediment samples was frozen in a − 20 °C refrigerator for five days, while the second group (N) of sediment samples did not undergo any pretreatment.
Isolation of Actinomycetia
For the isolation of actinomycetes, about 1 g of pretreated/non-pretreated sediment samples was serially diluted in sterile seawater and 0.1 mL of diluted samples was then spread evenly onto Petri dishes containing isolation media. Five different types of isolation media were used for the isolation of actinomycetes from sediment samples: (1) actinomycete isolation agar (AIA) (Hi media, India), (2) Gause modified agar Ι (GI) (Ivanitskaia et al. 1978; Chen et al. 2012), (3) Gause modified agar ΙI (GII) (Ivanitskaia et al. 1978), (4) marine agar (MA) (Hi media, India), and (5) soil agar (SA) (Suthindhiran and Kannabiran 2009). All five media were supplemented with nystatin (50 mg/L) and nalidixic acid (25 mg/L) to suppress the growth of fungi and Gram-negative bacteria, respectively. All the above media except MA were prepared with seawater. The isolation plates were incubated at 25 °C in the dark for two to six weeks. Individual colonies were repeatedly streaked onto fresh isolation medium to obtain pure cultures.
Microorganisms for antimicrobial testing
For the antimicrobial activity assay, Bacillus subtilis BCRC 1477 and Staphylococcus aureus ATCC 12600 were used as examples of Gram-positive test bacteria, while Escherichia coli ATCC 11775, Pseudomonas aeruginosa BCRC 10303, and Vibrio harveyi BCRC 13812 were selected as Gram-negative test bacteria, and Candida albicans BCRC 22903 was chosen as a test fungus. Most of the test microorganisms were purchased from the Bioresource Collection and Research Center (BCRC) of the Food Industry Research and Development Institute (FIRDI), Taiwan, except E. coli and S. aureus, which were a generous gift from Prof. J. K. Liu, Dept. of Biological Science, National Sun Yat-sen University, Taiwan.
P. aeruginosa and B. subtilis were cultured and maintained in nutrient broth (NB) and agar (NA) at 25 °C for 18 h. S. aureus and E. coli were cultured and maintained in Luria-Bertani broth (LB) and agar (LA) at 37 °C for 18 h. V. harveyi was cultured and maintained in marine broth (MB) and agar (MA) at 25 °C for 24 h. C. albicans was cultured and maintained in yeast malt broth (YM) and agar (YA) at 30 °C for 48 h.
Antimicrobial activities of actinomycete isolates
The agar plug method (Stern et al. 2006) was used for rapid in vitro evaluation and screening of the antimicrobial activities of the actinomycete isolates. Briefly, each isolate was grown for 10 days at 25 °C on M1 agar medium (1% starch, 0.4% yeast extract, 0.2% peptone, 80% seawater, and 1.6% agar). Agar plugs (8 mm in diameter) were excised from 10-day-old growths in Petri dishes. The agar plugs were placed onto new agar plates freshly inoculated with a suitable amount of the test microorganism (about 108 CFU/mL and 106 CFU/mL for the test bacteria and test fungus, respectively), and the plates were incubated for 24 h at temperature appropriate for the respective test microbe. Diameters of clear inhibition zones formed around the agar plug were then measured and recorded.
16S rRNA gene sequencing and construction of phylogenetic tree
Twenty-two actinomycete isolates that exhibited significant antimicrobial activity were identified based on sequencing of the 16S rRNA gene. The bacterial universal primer pairs 8F: 5’-GAGAGTTTGATCCTGGCTCAG-3′ and 1492R: 5’-CTACGGCTACCTTGTTACGA-3′ (Yuan et al. 2014) were used to amplify a partial 16S rDNA sequence. Actinomycetal genomic DNA was extracted using a FavorPrep Tissue Genomic DNA Extraction Mini kit (Favorgen, Taiwan) according to the manufacturer’s instructions. PCR amplification of partial 16S rDNA sequences was conducted following protocols described previously (Flemer et al. 2012). All the obtained sequences were then compared against the Ribosomal Data Project (RDP) (Cole et al. 2014) and NCBI NT databases (Benson et al. 2013) to identify the reference strains and closest relatives, respectively, using the nucleotide BLAST algorithm (Altschul et al. 1990). Actinomycete sequence identification was achieved using an RDP classifier with a confidence threshold value of 80% (Wang et al. 2007).
To carry out phylogenetic analysis, alignment of all the obtained and reference sequences was performed using the ClastalW algorithm (Thompson et al. 1994) in the MEGA XI platform (Tamura et al. 2021). A neighbor-joining tree was also created in the MEGA XI platform with a maximum composite likelihood model (Tamura et al. 2004) for evolutionary distance estimation. The robustness of the neighbor-joining tree was estimated through 1000 bootstrap replicates (Felsenstein 1985). The 16S rRNA sequence of B. subtilis DSM10 (AJ276351) was used as the outgroup.
Seawater requirement
The seawater requirement of each actinomycete isolates was determined using both M1 agar medium and M1 agar medium prepared using deionized water. Isolates that only grew on M1 agar were designated as obligate marine bacteria. Isolates that grew on both agar media, but grew better on M1 agar medium, were designated as having better growth with seawater.
Preparation of crude extracts
Each actinomycete isolate was fermented for 10 days in a 2-L Erlenmeyer flask containing 1 L M1 medium (Jensen et al. 2005) on a rotatory shaker (150 rpm, 25 °C). Following fermentation, the culture broth of each strain was harvested and exhaustively extracted three times with ethyl acetate (700 mL × 3). The organic layers were combined and evaporated to dryness under reduced pressure. The dry extracts were weighed and stored in the deep freezer. The extracts were used for the paper-disc diffusion antimicrobial assays and cytotoxic activity assays.
Antimicrobial activities of actinomycete crude extracts
The antimicrobial activities of crude metabolites against test microorganisms were also tested using the paper disc diffusion method according to Rios et al. (1988). In brief, a test bacterium or fungus with a final concentration of approximately 108 or 106 CFU/mL, respectively, was spread on the top of an agar plate. Sterilized filter paper discs (6 mm in diameter; Toyo Roshi, Japan) loaded with 50 μL of each extract at a concentration of 20 mg/mL were placed on the surface of the inoculated agar plates. The diameters of inhibition zones around each paper disc were measured after incubation in Petri dishes for 24 h under suitable conditions for the bacterium or fungus. An antibiotic mixture, 10 μg/disc chloramphenicol and 10 μg/disc nystatin, was used as positive control. No activity was observed for any of the test microbes in blank tests.
Detection of cytotoxic activity
Cytotoxic activity following treatment with crude extract was evaluated using a 3-(4, 5-dimethyl-thiazol-2-yl)-2, 5-diphenyltetra-zolium bromide (MTT) assay following previously-established procedures (Lu et al. 2009). Three cell lines, HL-60 (human promyelocytic leukemia), K-562 (human chronic myelogenous leukemia), and Molt-4 (human acute lymphoblastic leukemia) were used for the MTT assay; all were purchased from the BCRC of FIRDI. These cell lines were cultured in RPMI-1640 medium supplemented with 10% fetal bovine serum (FBS), 2 mM glutamine, 100 units/mL penicillin and 100 μg/mL streptomycin in a humidified incubator in the presence of 5% CO2 at 37 °C.
Nucleotide sequence accession numbers
The 16S rRNA gene sequences of the 22 isolates obtained in the present work are available in the NCBI GenBank database under accession numbers OP349179-OP349200.
Results
Isolation of Actinomycetia from marine sediments
In this study, 811 presumptive actinomycete isolates were recovered from six marine sediment samples using five different culture media. Of the 811 isolates, 101 (12.45%) were cultured from Kaohsiung Harbor sediments (sample KH01-KH03) and 710 (87.55%) were from Bashi Channel sediments (sample BC01-BC03) (Table 1). It is obvious that the number of actinomycetes isolated from deep-sea samples was much higher than the number recovered from nearshore samples.
Antimicrobial activities-primary and secondary screening
To quickly screen and identify antibiotic-producing actinomycete isolates, we carried out primary and secondary screening using the agar plug method. All 811 actinomycete isolates were first subjected to primary screening to examine their antimicrobial activities against two test bacteria B. subtilis and S. aureus (Table 1). Among them, 8.63% (70 isolates) possessed inhibitory activities against at least one of the test organisms (Table 1). The percentages of isolates obtained from the nearshore and deep sea sites possessed inhibitory activities against at least one of the test organisms were 11.88% (12) and 8.17% (58), respectively (Table 1). Of the 70 active isolates, 6.91% (n = 56) strains showed activity against S. aureus, 7.40% (n = 60) against B. subtilis, and 5.67% (n = 46) against both S. aureus and B. subtilis. There was no significant difference between the percentages of antibiotic-producing isolates obtained from the nearshore and deep sea (P = 0.4, Mann–Whitney U test).
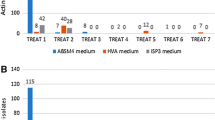
A secondary screening of 62 isolates that showed an inhibition zone >1 mm in diameter against either of the test strains in the primary screening was performed using the same detection method in order to detect their antimicrobial activities against all six test strains, B. subtilis, E. coli, P. aeruginosa, S. aureus, V. harveyi, and C. albicans. The results showed that of the 62 isolates (Figs. 1), all had activity against one or both Gram-positive bacterium (B. subtilis and/or S. aureus), 20.97% (n = 13) had activity against at least one Gram-negative bacterium (E. coli, V. harveyi, and/or P. aeruginosa), 20.97% (n = 13) had activity against both Gram-positive and Gram-negative bacteria (Fig. 1), and 3.23% (n = 2) had activity against the test fungus (C. albicans). A total of 80.65% (n = 50), 17.74% (n = 11), 9.68% (n = 6), and 1.62% (n = 1) isolates inhibited at least two, three, four, and five of the test strains, respectively (Fig. 1). It should be noted that the reason for all the isolates exhibiting antimicrobial activity against the Gram-positive bacteria was that we employed two Gram-positive bacteria as test bacteria in the primary screening.
The percentage of susceptibility of B. subtilis, S. aureus, E. coli, V. harveyi, P. aeruginosa, and C. albicans to the 62 isolates was 90.32% (n = 56), 85.48% (n = 53), 17.74% (n = 11), 8.06% (n = 5), 8.06% (n = 5), and 3.23% (n = 2) (Fig. 2), respectively, and the percentage of susceptibility to the 62 isolates with an inhibition zone diameter > 5 mm was 53.23%, 53.23%, 9.67%, 1.61%, 1.61% and 1.61%, respectively.
Identification of antibiotic-producing actinomycete isolates
Based on antimicrobial (strong and broad spectrum) activity in the secondary screening and location, 22 actinomycetes, 27.27% (n = 6) from nearshore and 72.73% (n = 16) from the deep sea, displayed high antimicrobial activity were selected for sequencing. A phylogenetic tree (Fig. 3) was constructed using the 16S rRNA gene sequences of these 22 potent isolates and reference strains downloaded from the RDP database using B. subtilis DSM10 as an outgroup. BLAST comparison of these sequences against the NCBI RefSeq database (Benson et al. 2013) is presented in Online Resource 1: Table S1. It should be emphasized that these isolates were neither a representative sample of culturable actinomycete bacteria nor a representative sample of culturable actinomycete bacteria with antimicrobial activities.
Neighbor-joining tree based on partial 16S rRNA gene sequences of 22 selected actinomycete strains isolated from marine sediments. Bootstrap values (expressed as percentages) greater than 60% are shown next to the branch points. The 16S rRNA sequence of Bacillus subtilis DSM10 was used as the outgroup. The scale bar corresponds to 0.02 substitutions per nucleotide position
In addition, we also ran the RDP classifier program (Wang et al. 2007) to identify the 22 potent isolates based on their 16S rRNA sequences. Among the 22 isolates, one isolate (BC03-N-GI66) could not be classified to the genus level, but was likely affiliated to the Streptomycetaceae family. The other 21 isolates were grouped into five genera: Streptomyces (15 isolates; 68.17%), Actinoalloteichus (2; 9.09%), Saccharomonospora (2; 9.09%), Saccharopolyspora (1; 4.55%), and Nocardiopsis (1; 4.55%).
Among the 22 isolates, Streptomycetaceae was the most commonly distributed and abundant family, accounting for 83.33% and 68.75% of the isolates (Fig. 4) from the nearshore and deep sea, respectively. Members of the three genera Actinoalloteichus, Saccharomonospora, and Saccharopolyspora were isolated only from deep-sea sediments, whereas the member of the genus Nocardiopsis was isolated only from nearshore sediments. These results suggested that there might be significant differences in the actinomycete population between nearshore and deep-sea sediments.
Antimicrobial activities of crude fermentation extracts
The 22 potent isolates were also selected for antimicrobial analysis with crude extracts. The culture extracts obtained from liquid cultures of these isolates were used to evaluate the antimicrobial potential by the paper disc method towards the same group of six test strains used in secondary screening. As shown in Table 2, these isolates truly can produce some organic compounds that showed broad-spectrum activity against the test microbes. Nineteen isolates (86.83%) could produce antibiotics that inhibited more than 4 test microbes, six isolates KH01-F-MA23 (Streptomyces NR_041423), KH01-F-MA24 (Streptomyces NR_041423), BC01-N-GI05 (Streptomyces NR_044035), BC01-N-GI24 (Streptomyces NR_119347), BC01-N-MA13 (Streptomyces NR_15202), and BC01-N-MA14 (Actinoalloteichus NR_043574) inhibited 5 test microbes, and 3 isolates KH01-N-SA02 (Streptomyces NR_041423), KH03-F-GII50 (Nocardiopsis NR_026340), and BC01-N-MA22 (Streptomyces NR_133874) inhibited all six test microbes. All isolates’ activity spectra covered both Gram-positive and Gram-negative bacteria, and 19 isolates (86.83%) inhibited both bacteria and the test fungus.
Seawater requirement assay
All the 22 isolates could grow on media prepared with seawater: 14 (63.6%) isolates were found to be obligate marine bacteria, and 2 (9.1%) isolates showed better growth in the presence of seawater (Table 2). These 16 isolates, 5 from nearshore and 9 from the deep sea, can be defined as marine-based bacteria (Jensen and Fenical 1995), and were considered to be indigenous marine bacteria or in a process of adaption to marine habitats. Interestingly, isolates from sample BC03, from the deep bottom of the Bashi Channel, had the lowest percentage (2 isolates, 33.3%) of marine-based bacteria.
Cytotoxic activity assay
The bioactive extracts of the 22 actinomycete strains were further subjected to cytotoxic activity determination using the MTT method. As shown in Table 3, 77.27% (n = 17) of strains displayed cytotoxic activity towards at least one target cell line, and 59.09% (n = 13) of strains displayed cytotoxic activity towards all three target cell lines; 31.81% (n = 7) of strains, including KH01-N-SA01 (Streptomyces NR_119346), BC01-N-GI05, BC01-N-MA19 (Streptomyces NR_044035), BC01-N-MA22, BC02-F-GII54 (Actinoalloteichus NR_04357), BC03-F-MA58 (Streptomyces NR_114820), and BC03-N-MA46 (Saccharopolyspora NR_024839), revealed strong (IC50 ≤ 1 μg/mL) cytotoxic activity toward at least two cell lines and may have great potential for antitumor treatment.
From Tables 2 and 3, we can conclude that the crude extracts of two isolates, BC01-N-GI05 and BC01-N-MA22, both from the deep bottom of the Bashi Channel, produced antibiotics that inhibited more than five test microbes and displayed activity towards all three target cell lines, two with strong activity, showing great potential for the development of antitumor antibiotic agents.
Discussion
In this study, actinomycete isolates from marine sediments of Kaohsiung Harbor and Bashi Channel were screened for metabolites with antimicrobial activity. Of a total of 811 actinomycete isolates, including 101 and 710 isolates from the nearshore and deep-sea environments, respectively, 12 (11.88%) and 58 (8.17%) were found to display antimicrobial activity towards at least one test organism based on primary screening (Table 1). Previous research has examined the antimicrobial activities of actinomycetes from costal marine sediments of Bejaia City, Algeria (Ouchene et al. 2022), northern Portugal (Ribeiro et al. 2020), and southwestern India (Chakraborty et al. 2015), and stated that respectively 9.7%, 11.5%, and 7.4% of the isolated actinomycetes possessed antimicrobial activity; these results were in line with our observations. In contrast, the percentages of actinomycetes possessing antimicrobial activity in our study was lower than the results of Özcan et al. (2013) (25.0%) as well as Ramesh and Mathivanan (2009) (53.4%), who studied the marine sediments from the coasts of the Turkey’s Anatolian Peninsula and India’s east coast on the Bay of Bengal, respectively. Generally, the percentage of actinomycetes isolated from marine sediments that possess antimicrobial activity differs broadly from study to study (Ramesh and Mathivanan 2009; Özcan et al. 2013; Chakraborty et al. 2015; Chen et al. 2021). However, it is important to understand that the percentage of isolates with antimicrobial activity can be affected by several parameters, such as culture media, fermentation conditions, number and species of test bacteria, and method used for assaying antimicrobial activity (Shnit-Orland and Kushmaro 2009).
Secondary screening (Figs. 1 and 2) shows that 62 isolates (100%) had activity against Gram-positive bacteria (B. subtilis and/or S. aureus) and only 13 isolates (20.97%) had activity against Gram-negative bacteria (E. coli, V. harveyi, and/or P. aeruginosa). Several previous studies reported that Gram-positive bacteria are more sensitive to antibiotic substances than Gram-negative bacteria (Khan et al. 2010; Kuo et al. 2021; Ouchene et al. 2022). For example, Khan et al. (2010) analyzed the antibacterial activities of 462 actinomycete strains isolated from 18 marine sponge samples, and found that strains active against Gram-positive bacteria outnumbered those active against Gram-negative bacteria by a ratio of 144 to 24. This is because the cell wall of Gram-positive bacteria is made predominantly of a thick layer of peptidoglycan, which can easily absorb foreign materials such as antibiotic substances, thus allowing them to kill the organism (Wang et al. 2020). Therefore, it is easy to establish that using more Gram-positive species as indicator bacteria may detect more antibiotic-producing isolates during the screening process.
Comparing the antimicrobial activities of the same isolates between different assay methods, the paper disc method (Table 2) generally displayed activity against significantly more kinds of Gram-negative bacteria and the test fungus than the agar plug method did (Fig. 1). This inconsistence has been detected previously (Abneuf et al. 2016; Rante et al. 2016). There are two possible reasons for this observation. First, a high concentration of actinomycete metabolites (crude extracts) used in the paper disc assay (Özcan et al. 2013). Second, the concentrations of these metabolites decreased as they diffuse through the agar in the agar-plug assay. Our results indicated that the paper disc method may be a more sensitive method by which to detect antimicrobial activity against Gram-negative bacteria and fungi than the agar plug method. In addition, it was also found that marine sediments from Kaohsiung Harbor and Bashi Channel both are a source of diverse actinomycetes with broad-spectrum antimicrobial activity.
Phylogenetic analysis (Fig. 3) of 22 selected isolates revealed that they belonged to five genera Streptomyces, Saccharomonospora, Actinoalloteichus, Saccharopolyspora, and Nocardiopsis. Blast analysis indicated that the range of sequence similarity for these isolates to their closest-related species was 98.3% to 100% (Online Resource 1: Table S1). Among them, isolate BC01-N-MA19 showed a 98.3% similaity to its closest-related strain Streptomyces xiamenensis MCCC 1A01550 which was originally isolated from mangrove sediments in Xiamen, China. As 98.65% was proposed by Kim et al. (2014) as the threshold of 16S rRNA gene sequence similarity for differentiating two bacterial species, isolate BC01-N-MA19 could be considered to belong to a new Streptomyces species. However, further studies are required to confirm its taxonomy.
Among the 22 selected isolates, Streptomyces (68.18%) was found to be the largest group represented (Fig. 3). In addition, most of the highly-potent actinomycetes isolated in this study were Streptomyces. For example, 7 of the 9 isolates that showed activity against at least 5 test microbes (Table 2) and 5 of the 7 isolates that displayed cytotoxic activity towards three target cell lines (Table 3), were all Streptomyces. These results suggested that antimicrobial and/or cytotoxic activity-oriented screening strategy reduce the diversity of isolated marine actinomycetes. Streptomyces species are economically important and the largest representative genus of Actinomycetia. The reason that Streptomyces spp. are high prolific natural product producers is probably because they have more biosynthetic genes (Arakawa 2018), such as PKS (polyketide synthases) and NRPS (non-ribosomal peptide synthetases), that are involved in natural product synthesis.
In the present study, it was found that of the 22 selected Actinomycetes, a high percentage (63.6%) of strains showed an obligatory requirement for seawater for growth (Table 2). Also, the percentage of seawater-requiring actinomycetes from Kaohsiung Harbor (66.7%) was about the same as that from Bashi Channel (62.2%). These seawater-requiring strains included members of genus Streptomyces (10 isolates, 62.5%), Nocardiopsis (1, 100%), Actinoalloteichus (2, 100%), and Saccharomonospora (1, 50%). Jensen et al. (2005) found that 59.1% of actinomycetes recovered from 227 marine sediments, 33 algae and 15 sponges collected around the island of Guam were seawater-requiring, a result in line with ours. The high percentage of seawater-requiring actinomycetes in this study suggests that these isolates may highly-adapted to the marine environment and could be novel ecotypes.
In conclusion, both nearshore and deep-sea sediments from Kaohsiung Harbor and Bashi Channel, respectively, are promising sources of bioactive marine Actinomycetia strains with the ability to produce new antimicrobial agents. In addition, selected active strains were found to exert cytotoxicity against cancer cell lines and broad-spectrum antimicrobial activity, and some of the strains were indigenous marine bacteria. Subsequent research into these bioactive actinomycetes will focus on compound identification and structure elucidation in order to unravel their biotechnological potential.
Abbreviations
- AIA:
-
Actinomycete isolation agar
- BCRC:
-
Bioresource Collection and Research Center
- FIRDI:
-
Food Industry Research and Development Institute
- GI:
-
Gause modified agar Ι
- GII:
-
Gause modified agar ΙI
- HL-60:
-
Human promyelocytic leukemia
- IC50 :
-
Half maximal inhibitory concentration
- K-562:
-
Human chronic myelogenous leukemia
- MA:
-
Marine agar
- Molt-4:
-
Human acute lymphoblastic leukemia
- RDP:
-
Ribosomal Data Project
- SA:
-
Soil agar
References
Abneuf MA, Khrishnan A, Aravena MG, Pang K-L, Convey P, Alias SA (2016) Antimicrobial activity of microfungi from maritime Antarctic soil. Czech Polar Rep 6:141–154. https://doi.org/10.5817/CPR2016-2-13
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Arakawa K (2018) Manipulation of metabolic pathways controlled by signaling molecules, inducers of antibiotic production, for genome mining in Streptomyces spp. Antonie Van Leeuwenhoek 111:743–751. https://doi.org/10.1073/pnas.1711842115
Bar-On YM, Phillips R, Milo R (2018) The biomass distribution on earth. Proc Natl Acad Sci U S A 115:6506–6511. https://doi.org/10.1073/pnas.1711842115
Benson DA, Cavanaugh M, Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW (2013) GenBank. Nucleic Acids Res 41:D36–D42. https://doi.org/10.1093/nar/gks1195
Bérdy J (2012) Thoughts and facts about antibiotics: where we are now and where we are heading. J Antibiot Res 65:385–395. https://doi.org/10.1038/ja.2012.27
Bull AT, Stach JEM (2007) Marine actinobacteria: new opportunities for natural product search and discovery. Trends Microbiol 15:491–499. https://doi.org/10.1016/j.tim.2007.10.004
Chakraborty RD, Chakraborty K, Thilakan B (2015) Isolation and characterization of antagonistic Streptomyces spp. from marine sediments along the southwest coast of India. IJMS 44:1742–1749
Chen YH, Kuo J, Sung PJ, Chang YC, Lu MC, Wong TY, Liu JK, Weng CF et al (2012) Isolation of marine bacteria with antimicrobial activities from cultured and field-collected soft corals. World J Microbiol Biotechnol 28:3269–3279. https://doi.org/10.1007/s11274-012-1138-7
Chen L, Wang Z, Du S, Wang G (2021) Antimicrobial activity and functional genes of Actinobacteria from coastal wetland. Curr Microbiol 78:3058–3067. https://doi.org/10.1007/s00284-021-02560-3
Cole JR, Wang Q, Fish JA, Chai B, McGarrell DM, Sun Y, Brown CT, Porras-Alfaro A et al (2014) Ribosomal database project: data and tools for high throughput rRNA analysis. Nucleic Acids Res 42:D633–D642. https://doi.org/10.1093/nar/gkt1244
Das S, Lyla PS, Khan SA (2006) Marine microbial diversity and ecology: importance and future perspectives. Curr Sci 90:1325–1335
Davies-Bolorunduro OF, Osuolale O, Saibu S, Adeleye IA, Aminah NS (2021) Bioprospecting marine actinomycetes for antileishmanial drugs: current perspectives and future prospects. Heliyon 7:e07710. https://doi.org/10.1016/j.heliyon.2021.e07710
de Kraker MEA, Stewardson AJ, Harbarth S (2016) Will 10 million people die a year due to antimicrobial resistance by 2050. PLoS Med 13:e1002184. https://doi.org/10.1371/journal.pmed.1002184
Donadio S, Maffioli S, Monciardini P, Sosio M, Jabes D (2010) Antibiotic discovery in the twenty-first century: current trends and future perspectives. J Antibiot Res 63:423–430. https://doi.org/10.1038/ja.2010.62
Duncan K, Haltli B, Gill KA, Kerr RG (2014) Bioprospecting from marine sediments of New Brunswick, Canada: exploring the relationship between total bacterial diversity and actinobacteria diversity. Mar Drugs 12:899–925. https://doi.org/10.3390/md12020899
Feling RH, Buchanan GO, Mincer TJ, Kauffman CA, Jensen PR, Fenical W (2003) Salinosporamide A: a highly cytotoxic proteasome inhibitor from a novel microbial source, a marine bacterium of the new genus Salinospora. Angew Chem Int Ed Engl 42:355–357. https://doi.org/10.1002/anie.200390115
Felsenstein J (1985) Confidence limits on phylogenies with a molecular clock. Syst Zool 34:152–161. https://doi.org/10.2307/2413323
Flemer B, Kennedy J, Margassery LM, Morrissey JP, O'Gara F, Dobson AD (2012) Diversity and antimicrobial activities of microbes from two Irish marine sponges, Suberites carnosus and Leucosolenia sp. J Appl Microbiol 112:289–301. https://doi.org/10.1111/j.1365-2672.2011.05211.x
Gao HM, Xie PF, Zhang XL, Yang Q (2021) Isolation, phylogenetic and Gephyromycin metabolites characterization of new exopolysaccharides-bearing Antarctic Actinobacterium from feces of emperor penguin. Mar Drugs 19. https://doi.org/10.3390/md19080458
Gozari M, Zaheri A, Jahromi ST, Gozari M, Karimzadeh R (2019) Screening and characterization of marine actinomycetes from the northern Oman Sea sediments for cytotoxic and antimicrobial activity. Int Microbiol 22:521–530. https://doi.org/10.1007/s10123-019-00083-3
Hoshino T, Doi H, Uramoto GI, Wörmer L, Adhikari RR, Xiao N, Morono Y, D'Hondt S et al (2020) Global diversity of microbial communities in marine sediment. Proc Natl Acad Sci U S A 117:27587–27597. https://doi.org/10.1073/pnas.1919139117
Ivanitskaia LP, Singal EM, Bibikova MV, Vostrov SN (1978) Directed isolation of Micromonospora generic cultures on a selective medium with gentamycin. Antibiotiki 23:690–692
Jensen PR, Fenical W (1995) The relative abundance and seawater requirements of gram-positive bacteria in near-shore tropical marine samples. Microb Ecol 29:249–257. https://doi.org/10.1007/bf00164888
Jensen PR, Gontang E, Mafnas C, Mincer TJ, Fenical W (2005) Culturable marine actinomycete diversity from tropical Pacific Ocean sediments. Environ Microbiol 7:1039–1048. https://doi.org/10.1111/j.1462-2920.2005.00785.x
Jung SY, Kim HS, Song JJ, Lee SG, Oh TK, Yoon JH (2007) Aestuariimicrobium kwangyangense gen. nov., sp. nov., an LL-diaminopimelic acid-containing bacterium isolated from tidal flat sediment. Int J Syst Evol Microbiol 57:2114–2118. https://doi.org/10.1099/ijs.0.64917-0
Kamjam M, Sivalingam P, Deng Z, Hong K (2017) Deep sea actinomycetes and their secondary metabolites. Front Microbiol 8:760. https://doi.org/10.3389/fmicb.2017.00760
Khan ST, Takagi M, Shin-ya K (2010) Diversity, salt requirement, and antibiotic production of Actinobacteria isolated from marine sponges. Actinomycetologica 24:18–23. https://doi.org/10.3209/saj.SAJ240101
Kim M, Oh HS, Park SC, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351. https://doi.org/10.1099/ijs.0.059774-0
Kuo J, Chang CF, Chi WC (2021) Isolation of endophytic fungi with antimicrobial activity from medicinal plant Zanthoxylum simulans Hance. Folia Microbiol 66:385–397. https://doi.org/10.1007/s12223-021-00854-4
Kurtbӧke Dİ (2017) Ecology and habitat distribution of Actinobacteria. In: Wink J, Mohammadipanah F, Hamedi J (eds) Biology and biotechnology of Actinobacteria. Springer International Publishing, Cham, pp 123–149. https://doi.org/10.1007/978-3-319-60339-1_6
Lu M-C, Du Y-C, Chuu J-J, Hwang S-L, Hsieh P-C, Hung C-S, Chang F-R, Wu Y-C (2009) Active extracts of wild fruiting bodies of Antrodia camphorata (EEAC) induce leukemia HL 60 cells apoptosis partially through histone hypoacetylation and synergistically promote anticancer effect of trichostatin A. Arch Toxicol 83:121–129. https://doi.org/10.1007/s00204-008-0337-3
Mahapatra GP, Raman S, Nayak S, Gouda S, Das G, Patra JK (2020) Metagenomics approaches in discovery and development of new bioactive compounds from marine actinomycetes. Curr Microbiol 77:645–656. https://doi.org/10.1007/s00284-019-01698-5
Maldonado LA, Fenical W, Jensen PR, Kauffman CA, Mincer TJ, Ward AC, Bull AT, Goodfellow M (2005) Salinispora arenicola gen. nov., sp. nov. and Salinispora tropica sp. nov., obligate marine actinomycetes belonging to the family Micromonosporaceae. Int J Syst Evol Microbiol 55:1759–1766. https://doi.org/10.1099/ijs.0.63625-0
Manam RR, Teisan S, White DJ, Nicholson B, Grodberg J, Neuteboom STC, Lam KS, Mosca DA et al (2005) Lajollamycin, a nitro-tetraene spiro-β-lactone-γ-lactam antibiotic from the marine actinomycete Streptomyces nodosus. J Nat Prod 68:240–243. https://doi.org/10.1021/np049725x
Manivasagan P, Venkatesan J, Sivakumar K, Kim SK (2014) Pharmaceutically active secondary metabolites of marine Actinobacteria. Microbiol Res 169:262–278. https://doi.org/10.1016/j.micres.2013.07.014
McCarthy AJ, Williams ST (1992) Actinomycetes as agents of biodegradation in the environment — a review. Gene 115:189–192. https://doi.org/10.1016/0378-1119(92)90558-7
Meena B, Anburajan L, Vinithkumar NV, Kirubagaran R, Dharani G (2019) Biodiversity and antibacterial potential of cultivable halophilic actinobacteria from the deep sea sediments of active volcanic Barren Island. Microb Pathog 132:129–136. https://doi.org/10.1016/j.micpath.2019.04.043
Mincer TJ, Jensen PR, Kauffman CA, Fenical W (2002) Widespread and persistent populations of a major new marine actinomycete taxon in ocean sediments. Appl Environ Microbiol 68:5005–5011. https://doi.org/10.1128/AEM.68.10.5005-5011.2002
Olano C, Méndez C, Salas JA (2009) Antitumor compounds from marine actinomycetes. Mar Drugs 7:210–248. https://doi.org/10.3390/md7020210
Ouchene R, Intertaglia L, Zaatout N, Kecha M, Suzuki MT (2022) Selective isolation, antimicrobial screening and phylogenetic diversity of marine actinomycetes derived from the Coast of Bejaia City (Algeria), a polluted and microbiologically unexplored environment. J Appl Microbiol 132:2870–2882. https://doi.org/10.1111/jam.15415
Özcan K, Aksoy SÇ, Kalkan O, Uzel A, Hames-Kocabas EE, Bedir E (2013) Diversity and antibiotic-producing potential of cultivable marine-derived actinomycetes from coastal sediments of Turkey. J Soils Sediments 13:1493–1501. https://doi.org/10.1007/s11368-013-0734-y
Patin NV, Schorn M, Aguinaldo K, Lincecum T, Moore BS, Jensen PR (2017) Effects of actinomycete secondary metabolites on sediment microbial communities. Appl Environ Microbiol 83:e02676–e02616. https://doi.org/10.1128/aem.02676-16
Priyanka S, Jayashree M, Shivani R, Anwesha S, Bhaskara Rao KV, I AE (2019) Characterisation and identification of antibacterial compound from marine actinobacteria: in vitro and in silico analysis. Journal of Infection and Public Health 12: 83-89. https://doi.org/10.1016/j.jiph.2018.09.005
Ramesh S, Mathivanan N (2009) Screening of marine actinomycetes isolated from the Bay of Bengal, India for antimicrobial activity and industrial enzymes. World J Microbiol Biotechnol 25:2103–2111. https://doi.org/10.1007/s11274-009-0113-4
Rante H, Alam G, Usmar U, Anwar R, Ali A (2016) Isolation of sponge bacterial symbionts from Kodingareng Keke Island-Makassar Indonesia which is potential as a producer of antimicrobial compounds. J Pure Appl Microbiol 16:737–743. https://doi.org/10.22207/JPAM.16.1.79
Ribeiro I, Girão M, Alexandrino DAM, Ribeiro T, Santos C, Pereira F, Mucha AP, Urbatzka R et al (2020) Diversity and bioactive potential of Actinobacteria isolated from a coastal marine sediment in Northern Portugal. Microorganisms 8:1691. https://doi.org/10.3390/microorganisms8111691
Riedlinger J, Reicke A, Zähner H, Krismer B, Bull AT, Maldonado LA, Ward AC, Goodfellow M et al (2004) Abyssomicins, inhibitors of the para-aminobenzoic acid pathway produced by the marine Verrucosispora strain AB-18-032. J Antibiot Res 57:271–279. https://doi.org/10.7164/antibiotics.57.271
Rios JL, Recio MC, Villar A (1988) Screening methods for natural products with antimicrobial activity: a review of the literature. J Ethnopharmacol 23:127–149. https://doi.org/10.1016/0378-8741(88)90001-3
Riyanti BW, Liu Y, Sharma A, Mihajlovic S, Hartwig C, Leis B, Rieuwpassa FJ et al (2020) Selection of sponge-associated bacteria with high potential for the production of antibacterial compounds. Sci Rep 10:19614. https://doi.org/10.1038/s41598-020-76256-2
Salam N, Jiao JY, Zhang XT, Li WJ (2020) Update on the classification of higher ranks in the phylum Actinobacteria. Int J Syst Evol Microbiol 70:1331–1355. https://doi.org/10.1099/ijsem.0.003920
Selvin J, Shanmughapriya S, Gandhimathi R, Seghal Kiran G, Rajeetha Ravji T, Natarajaseenivasan K, Hema TA (2009) Optimization and production of novel antimicrobial agents from sponge associated marine actinomycetes Nocardiopsis dassonvillei MAD08. Appl Microbiol Biotechnol 83:435–445. https://doi.org/10.1007/s00253-009-1878-y
Sharma S, Fulke AB, Chaubey A (2019) Bioprospection of marine actinomycetes: recent advances, challenges and future perspectives. Acta Oceanol Sin 38:1–17. https://doi.org/10.1007/s13131-018-1340-z
Shnit-Orland M, Kushmaro A (2009) Coral mucus-associated bacteria: a possible first line of defense. FEMS Microbiol Ecol 67:371–380. https://doi.org/10.1111/j.1574-6941.2008.00644.x
Solyanikova IP, Golovleva LA (2015) Physiological and biochemical properties of actinobacteria as the basis of their high biodegradative activity. Appl Biochem Microbiol 51:143–149. https://doi.org/10.1134/S0003683815020180
Stern NJ, Svetoch EA, Eruslanov BV, Perelygin VV, Mitsevich EV, Mitsevich IP, Pokhilenko VD, Levchuk VP et al (2006) Isolation of a Lactobacillus salivarius strain and purification of its bacteriocin, which is inhibitory to Campylobacter jejuni in the chicken gastrointestinal system. Antimicrob Agents Chemother 50:3111–3116. https://doi.org/10.1128/aac.00259-06
Stincone P, Brandelli A (2020) Marine bacteria as source of antimicrobial compounds. Crit Rev Biotechnol 40:306–319. https://doi.org/10.1080/07388551.2019.1710457
Subramani R, Sipkema D (2019) Marine rare actinomycetes: a promising source of structurally diverse and unique novel natural products. Mar Drugs 17:249. https://doi.org/10.3390/md17050249
Suthindhiran K, Kannabiran K (2009) Hemolytic activity of Streptomyces VITSDK1 spp. isolated from marine sediments in Southern India. J Med Mycol 19:77–86. https://doi.org/10.1016/j.mycmed.2009.01.001
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci U S A 101:11030–11035. https://doi.org/10.1073/pnas.0404206101
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Teske A, Hinrichs KU, Edgcomb V, de Vera GA, Kysela D, Sylva SP, Sogin ML, Jannasch HW (2002) Microbial diversity of hydrothermal sediments in the Guaymas Basin: evidence for anaerobic methanotrophic communities. Appl Environ Microbiol 68:1994–2007. https://doi.org/10.1128/aem.68.4.1994-2007.2002
Thompson JD, Higgins DG, Gibson TJ (1994) Clustal W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680. https://doi.org/10.1093/nar/22.22.4673
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. https://doi.org/10.1128/aem.00062-07
Wang W, Park KH, Lee J, Oh E, Park C, Kang E, Lee J, Kang H (2020) A new thiopeptide antibiotic, micrococcin P3, from a marine-derived strain of the bacterium Bacillus stratosphericus. Molecules 25:4383. https://doi.org/10.3390/molecules25194383
Yuan M, Yu Y, Li HR, Dong N, Zhang XH (2014) Phylogenetic diversity and biological activity of actinobacteria isolated from the Chukchi shelf marine sediments in the Arctic Ocean. Mar Drugs 12:1281–1297. https://doi.org/10.3390/md12031281
Acknowledgments
This work was supported by a grant from Taiwan’s Ministry of Science and Technology (MOST105-2611-M-291-007) and intramural funding from the National Museum of Marine Biology and Aquarium.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 16 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kuo, J., Chen, KT., Lu, MC. et al. Screening of marine Actinomycetia with bioactive metabolites from nearshore and deep sea marine sediments in southwestern Taiwan. Biologia 78, 2551–2562 (2023). https://doi.org/10.1007/s11756-023-01397-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-023-01397-4