Abstract
Sucrose is the principal form of photosynthesis products, and long-distance transport of sucrose requires sucrose transporters (SUTs) to perform loading and unloading functions. SUTs play an important role in plant growth, development and reproduction. In this study, five unique sucrose transporter (SbSUT) genes that contain full-length cDNA sequences were cloned from sweet sorghum, and these SbSUT genes were clustered into four different clades: SUT1, SUT3, SUT4 and SUT5. Heterologous expression of SbSUTs in yeast demonstrated that they were functional sucrose transporters. Tissue-specific expression profiles showed that sorghum SUT genes had different tissue-specific expression patterns, suggesting that sorghum SUT genes may play an important role in plant growth and developmental processes. After defoliation, expression patterns of SbSUT1, SbSUT2 and SbSUT4 were different in leaf sheaths, leaves and roots. Taken together, the results indicate that the above mentioned five unique sucrose transporter genes may play important roles in performing sucrose loading and unloading functions and that they exhibit different expression in response to leaf blade removal.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In most plant species, sucrose is the main form of carbohydrate that is delivered from photosynthetic source organs to heterotrophic sink organs, where sugars are used or stored (Kühn and Grof 2010). Sweet sorghum (Sorghum bicolor), as a promising energy crop, is known for its high sugar content in the culms and is a highly productive species that can produce biomass ranging from 58.3 to 80.5 tons of fresh stems per hectare in semiarid zones (Wang and Liu 2009). Plants possess a family of sucrose transporters (SUTs) involved in long-distance transport of sucrose in the phloem (Daie 1989), with SUTs performing loading and unloading functions (Reinders et al. 2006; Scofield et al. 2007; Slewinski and Braun 2010). Unlike the dicot SUTs, monocotyledonous SUT1 proteins are highly specific for sucrose. Sivitz et al. (2007) analyzed the substrate specificity for four characterized SUTs (AtSUC2, AtSUC9, ShSUT1 and HvSUT1). The results indicated that dicot type I SUTs (AtSUC2 and AtSUC9) had low substrate specificity, while monocot type II SUTs (ShSUT1 and HvSUT1) were more selective and did not transport most β-linked glucosides. ShSUT1 was previously shown to be expressed in maturing stems and plays an important role in the accumulation of sucrose in sugarcane stalks (Reinders et al. 2006). Because sorghum is agriculturally important, SUTs from cereals have been the focus of increasing research interest.
Plant SUT proteins were previously divided into three phylogenetic groups: type I was composed exclusively of dicot sequences, whereas type II and type III contained both monocot and dicot proteins (Aoki et al. 2003; Lalonde et al. 2004). Later, the type II subfamily was split into two groups, dividing the SUTs into four distinct classes (Sauer 2007). Then, the SUTs were described as five distinct clades: SUT1–SUT5. SUT1 and SUT5 (formerly type II) consist entirely of monocot SUTs, SUT2 (formerly type I) contains only dicot SUTs, and both SUT3 (formerly type II) and SUT4 (formerly type III) contain monocot and dicot SUTs (Braun and Slewinski 2009; Doidy et al. 2012). The majority of sucrose transporters have been characterized to date as energy-dependent sucrose-H+ symporters belonging to the major facilitator superfamily (MFS) with 12 transmembrane domains (Carpaneto et al. 2005; Sauer 2007).
Defoliation processes will inevitably cause carbon reallocation in plants, and carbon redistribution is an important factor affecting plant growth (Callow et al. 2005; Wang et al. 2012). Leaf sheaths function as sink organs under normal growth conditions. After defoliation, leaf sheaths change from sink tissues to source tissues and can accumulate up to 70 % fructans stored in vegetative regions (Volenec 1986; Housley and Volenec 1988). Defoliation in Lolium perenne led to a significant increase in the expression of LpSUT1 in leaf sheaths, with the increased LpSUT1 transcript levels possibly contributing to rapid refoliation (Berthier et al. 2009). In our previous study, LcSUT1 was up-regulated in leaf sheaths after defoliation through sucrose starvation instead of wound signaling (Su et al. 2013). However, the function of the other SUT genes following defoliation was not elucidated.
To advance the understanding of sink–source tissue relationships after defoliation, we isolated the sucrose transporters from sorghum and characterized their expression patterns in response to defoliation. Our data will yield new information about regulation mechanisms of plant sucrose transport and accumulation, which could help us improve crop yield by informing genetic engineering strategies.
Materials and methods
Plant materials and defoliation treatment
Sweet sorghum (S. bicolor cv. Rio) seeds were germinated on plates for 2 days, after which the seedlings were transferred into pots 8 cm in diameter and filled with vermiculite. Seedlings were grown in a greenhouse with a 16 h light/8 h dark photoperiod and 25/18 °C (day and night, respectively) temperature and were fertilized with Hoagland solution (Hoagland and Arnon 1950) every 3 days. Defoliation treatment was performed 10 days after sowing when the nutrients of seeds were exhausted (Su et al. 2011), and plants were defoliated at 5 cm above ground level at 10:00 a.m. (Berthier et al. 2009). The leaf sheaths, elongating leaves and roots were harvested after 0, 2, 6, and 12 h of regrowth, frozen in liquid nitrogen, stored at −80 °C and used for analyzing the effect of defoliation on Suc transport (three-replicate analysis). For the tissue-specific analysis, blades, leaf veins, source leaves, sheaths, phloem, pulps, inflorescences, and roots were harvested at the flowering stage.
Identification of SbSUTs genes
To isolate the full-length sequence of five sucrose transporters from sweet sorghum, we designed gene-specific primers for SbSUTs based on the conserved region for monocot (such as Zea mays and Saccharum hybridum) SUTs and then screened the sorghum genome database (JGI, http://www.jgi.doe.gov/).
To obtain the full-length cDNA of SbSUT genes from sweet sorghum, all organs were harvested at the flowering stage. Total RNA was isolated using the Trizol Kit (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions, and genomic DNA was removed via digestion with RNase-free DNase I (TaKaRa Biotechnology Co., Ltd., Japan). The cDNA first strand was synthesized using Prime Script RT-PCR kit (TaKaRa), following the manufacturer’s protocol. For amplification of SbSUTs genes, five pairs of primers were designed based on the putative SbSUT transcript sequences and are listed in Table 1. The amplification conditions were as follows: 95 °C for 5 min, followed by 35 cycles of 95 °C for 30 s, 58 °C for 30 s, 72 °C for 2 min, and a final extension at 72 °C for 10 min. All the PCR products were cloned into the pMD19-T vector and sequenced at Sangon Biotech (Shanghai Co., Ltd.).
Domain prediction and phylogenetic tree construction
The identity search for nucleotides and translated amino acids was carried out using the National Center for Biotechnology Information (NCBI) BLAST network server (http://www.ncbi.nlm.gov/BLAST). Transmembrane domain predictions were achieved using the TopPred2 database (http://mobyle.pasteur.fr/cgi-bin/portal.py?form=toppred#forms::toppred). The phylogenetic tree was generated by MEGA 4.1 (Tamura et al. 2007) using the protein sequence of SbSUTs and 33 other confirmed sucrose transporter sequences from publicly accessible databases.
Heterologous expression in yeast
For heterologous expression in yeast, cDNA harboring the full-length ORF was generated by RT-PCR using the following primers: SbSUT1 (5′-GGAATTCATGGCTCGCGGCGACGGC-3′, EcoR I site underlined, and 5′-GCGTCGACTCAGTGGCCGCCCGCGCT-3′, Sal I site underlined), SbSUT2 (5′-GGACTAGTATGGACGCCGGCACCGGG-3′, Spe I site underlined, and 5′-GCGTCGACGCCAAATCCATGGAAACCGG-3′, Sal I site underlined), SbSUT3 (5′-CGGAATTCATGGCTGCTGATGGCAC-3′, EcoR I site underlined, and 5′-GCGTCGACATGGCCTCCTCCCATGG-3′, Sal I site underlined), SbSUT4 (5′-GGAATTCATGCCGCCGCGCACGGCT-3′, EcoR I site underlined, and 5′-GCGTCGACTTATCGGTGCGTGCCAC-3′, Sal I site underlined) and SbSUT5 (5′-GGACTAGTATGGACGGTGGTGACGGC-3′, Spe I site underlined, and 5′-CGGAATTCGTGGCCGCCGCCCGCCAT-3′, EcoR I site underlined), and then ligated into the sites of the yeast shuttle vector pDR196 with the Saccharomyces PMA1 promoter. Competent yeast SUSY7/ura3 cells were prepared and transformed according the method of Weise et al. (2000).
Semi-quantitative RT-PCR and qRT-PCR analysis
The primers of semi-quantitative RT-PCR and qRT-PCR are listed in Table 1. Actin was used as the internal reference gene for assessing expression levels in sweet sorghum. For semi-quantitative RT-PCR, the following conditions were used: 5 min at 94 °C followed by 28 cycles (for SbSUT1), 30 cycles (SbSUT2, SbSUT3 and SbSUT4) or 35 cycles (SbSUT5) of 95 °C for 30 s, 55 °C for 30 s and 72 °C for 30 s while the Actin gene used 20 cycles.
For qRT-PCR, the cDNA was diluted 1:100 with easy dilution (Takara, Japan), and 2 μl was used as a template for qRT-PCR analysis. qRT-PCR was performed on the MX3000P (Stratagene) using SYBR® Premix Ex Taq™ (Takara, Japan) according to the manufactures’ instructions. The qRT-PCR program was performed as follows: 30 s at 95 °C, followed by 35 cycles of 95 °C for 15 s and 60 °C for 30 s. Fluorescence data were collected at each polymerization step. The relative expression levels of the SbSUT genes were calculated using the method of 2−∆∆CT described by Livaka and Schmittgen (2001). All of the qRT-PCR results were confirmed by three independent reactions.
Results
Isolation and sequence analysis of sucrose transporter proteins (SbSUTs) in sweet sorghum
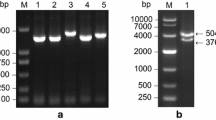
Five sucrose transporter cDNAs (Sb01g045720, Sb04g038030, Sb01g022430, Sb08g023310 and Sb04g023860) were isolated from sweet sorghum by reverse transcription polymerase chain reaction (RT-PCR) and designated as SbSUT1–SbSUT5. The sequences of SbSUT1 and SbSUT4 were deposited in the NCBI database with the accession numbers GU126430 and GU045300, respectively. The predicted amino acid sequences were characterized in Table 2. Deduced proteins included 12 transmembrane spanning regions, which are universal in plant sucrose transporters (Carpaneto et al. 2005). To understand the evolutionary relationship between plant SUTs, phylogenetic analyses were conducted using the MEGA 4.1 tool. As presented in the tree, SbSUT proteins belonged to four SUT clades and shared strong similarity with the identified functional sucrose transporters of Z. mays (Fig. 1).
Phylogenetic relationships between SbSUTs and other species’ SUTs. The transporters presented are the following: Arabidopsis thaliana—AtSUC1, At1g71880; AtSUC2, At1g22710; AtSUC3, At2g02860; AtSUC5, At1g71890; AtSUC9, At5g06170; Bambusa oldhamii—BoSUT5, AAY43226. Hordeum vulgare—HvSUT1, CAB75882; HvSUT2, CAB75881; Leymus chinensis—LcSUT1, HM769942; Lycopersicum esculentum—LeSUT1, CAA57726; LeSUT2, AAG12987; Lotus japonicus—LjSUT4, CAD61275; Oryza sativa—OsSUT1, AAF90181; OsSUT2, BAC67163; OsSUT3, BAB68368; OsSUT4, BAC67164; OsSUT5, BAC67165. Plantago major—PmSUC1, CAI59556; PmSUC2, X75764; PmSUC3, CAD58887; Pisum sativum—PsSUF4, A3DSX1; Sorghum Bicolor—SbSUT1, GU126430, SbSUT4, GU045300; Solanum tuberosum—StSUT1, CAA48915; StSUT4, AAG25923; Saccharum hybridum—ShSUT1, AAV41028; Triticum aestivum—TaSUT1A, AAM13408; TaSUT1B, AAM13409; TaSUT1D, AAM13410; Z. mays—ZmSUT1, BAA83501; ZmSUT2, AAS91375; ZmSUT3, ACF86653; ZmSUT4, AAT51689; ZmSUT5, ACF85284; ZmSUT6, ACF85673
Functional analysis of SbSUTs in yeast
To test whether SbSUT proteins were functional sucrose transporters, the full open reading frame of SbSUTs cDNAs were cloned into the yeast expression vector pDR196 and transformed into a modified version of the yeast strain SUSY7/ura3 (Riesmeier et al. 1992). This strain was unable to utilize exogenous sucrose, but if transformed with a functional SUT, it can import sucrose and hydrolyze it internally (Barker et al. 2000). Yeast transformed with pDR196-StSUT1 (Solanum tuberosum sucrose transporter 1) as a positive control (Riesmeier et al. 1994) grew better on media containing sucrose as the sole carbon source than yeast transformed with the empty pDR196 vector (negative control). A clear complementation experiment using the expression of SbSUTs in yeast strains showed that SbSUTs can import sucrose in media for SUSY7/ura3 growth, demonstrating that these five SbSUT genes were functional sucrose transporters (Fig. 2a, b).
Expression of SbSUTs in the yeast strain SUSY7/ura3. a Expression of SUTs in the yeast strain SUSY7/ura3 allows growth on glucose as the sole carbon source. The numbers 1, 2, 3, 4, and 5 represent transformation of the following constructs: PDR196-SbSUT1, PDR196-SbSUT2, PDR196-SbSUT3, PDR196-SbSUT4, and PDR196-SbSUT5, respectively. The number 6 represents transformation of PDR196-StSUT1 as a positive control, 7 represents transformation of PDR196 as a negative control, and 8 represents the blank control. b Expression of SbSUTs in the yeast strain SUSY7/ura3 allows growth on sucrose as the sole carbon source. The numbers 1, 2, 3, 4, and 5 represent transformation of the following constructs: PDR196-SbSUT1, PDR196-SbSUT2, PDR196-SbSUT3, PDR196-SbSUT4, and PDR196-SbSUT, respectively. The number 6 represents transformation of PDR196-StSUT1 as a positive control, and 7 represents transformation PDR196 as a negative control
Expression patterns of SbSUTs in different organs
Semi-quantitative RT-PCR was performed to determine the expression pattern of SbSUT genes in different tissues. The SbSUT1 gene was highly expressed in leaf sheath, phloem and pulp, and SbSUT2 showed higher expression levels in leaf sheaths and inflorescences. The expression of SbSUT4 was higher in leaf veins, pulp and inflorescences than in other organs, while SbSUT3 expression was basically the same in various organs. Interestingly, the expression level of the SbSUT5 transcript was higher in the pulp than in other organs, which suggested SbSUT5 may play important roles in sucrose storage in the pulp (Fig. 3).
The tissue-specific expression levels of SbSUTs. The semi-quantitative RT-PCR was used for analysis of the gene expression levels in different organs. The SbSUT1 transcript was amplified with 28 cycles, SbSUT2, SbSUT3 and SbSUT4 were amplified with 30 cycles, and SbSUT5 was amplified with 35 cycles and the Actin gene with 20 cycles
Relative expression levels of SbSUTs in young seedlings
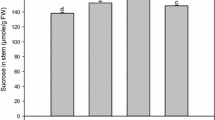
By 10 days after sowing, the nutrients in seeds are depleted and sweet sorghum seedling growth depends entirely on photosynthesis (Su et al. 2011). We compared the relative expression levels of five SbSUTs in young seedlings using qRT-PCR, finding that the expression levels of SbSUT3 and SbSUT5 were lower than that of SbSUT1, SbSUT2 and SbSUT4 (Fig. 4).
Different expression patterns of SbSUT1, SbSUT2 and SbSUT4 in three organs after defoliation
The leaf blade functions as a source tissue that performs photosynthesis and fixes carbon. Upon leaf blade removal, carbon becomes reallocated in whole plants. To find different functions of SbSUTs in response to defoliation, we detected the expression levels of SbSUT1, SbSUT2 and SbSUT4 after defoliation by qRT-PCR. Our results showed that the expression patterns of SbSUT1, SbSUT2, and SbSUT4 were different after defoliation. The SbSUT1 gene was significantly up-regulated in leaf sheaths after defoliation (Fig. 5a), while the expression level of SbSUT4 was not obviously changed (Fig. 5c). In contrast, SbSUT4 was significantly up-regulated in leaves and roots (Fig. 5c). The expression levels of SbSUT2 were increased in leaf sheaths and decreased in leaves and roots (Fig. 5b).
Relative expression levels of SbSUTs after defoliation. The relative expression levels of SbSUTs in leaves, leaf sheaths and roots were detected by qRT-PCR. a The transcript levels of SbSUT1 at 0 h was used as the control. b The transcript levels of SbSUT2 at 0 h was used as the control. c The transcript levels of SbSUT4 at 0 h was used as the control
Discussion
Nine sucrose transporter genes have been described in Arabidopsis (Kühn and Grof 2010), and five in the rice genome (Aoki et al. 2003). The monocots maize and Brachypodium are separated into five groups, with the fifth group made up exclusively of functionally uncharacterized monocot transporters (Braun and Slewinski 2009). We have cloned five SbSUT genes from sorghum (Table 2). Phylogenetic comparison of the members of the SUT family among sorghum, Z. mays, wheat, rice, and Arabidopsis showed that SbSUT proteins belong to four SUT clades. SbSUT1 and SbSUT3 belong to clade 1 and cluster with SUT isoforms from other grass species (e.g., rice, maize), while SbSUT2, SbSUT4 and SbSUT5 belong to clades 3, 4 and 5, respectively. SbSUTs share strong similarity with the identified functional sucrose transporters of Z. mays (Fig. 1). Experiments in yeast (Saccharomyces cerevisiae) have demonstrated biochemical activity for many SUTs (Chandran et al. 2003; Zhou et al. 2007; Payyavula et al. 2011). Based on this method, heterologous expression of SbSUTs in yeast proved that they were functional sucrose transporters (Fig. 2).
Previous studies have also described tissue-specific expression of sucrose transporter genes in numerous organs of different plant species. Examples include the following: OsSUT1 expression is observed in leaf blades, leaf sheaths, panicles, and germinating seeds, except in roots (Hirose et al. 1997; Chen et al. 2010). AtSUC1 is expressed in pollen tubes, funiculi, placenta and anther connective tissue, and the expression in the connective tissue suggests that AtSUC1 may play a role in the control of anther dehiscence (Stadler et al. 1999). AtSUC3 expression has been detected in numerous sink cells and tissues, such as guard cells, trichomes, germinating pollen, root tips, the developing seed coat, and stipules (Meyer et al. 2004). In maize plants, SUT1 mutants produced tassels with partially barren branches and limited spikelet development, and SUT1 mutant plants displayed strongly inhibited reproductive development as well as vegetative growth defects (Slewinski et al. 2009). When the expression of SbSUTs was compared among eight different sweet sorghum tissues, it was detectable in each tissue examined. This result suggests that SbSUTs may play different roles in plant growth, development and reproduction. SbSUT1 expression differed markedly among different tissues, with highest expression in the leaf sheath, phloem and pulp. SbSUT2 showed higher expression levels in leaf sheaths and inflorescences, whereas the expression of SbSUT4 was higher in leaf veins, pulp and inflorescences (Fig. 3). These results imply that preferential roles in sucrose loading into these organs exist. Additionally, SbSUT3 expression was basically the same in various organs, while the SbSUT5 transcript level was higher in pulp than in other organs, suggesting that SbSUT5 may play important roles in sucrose storage in the pulp (Fig. 3).
Relative expression levels of the SbSUT genes in seedlings showed that expression levels of SbSUT3 and SbSUT5 are lower than that of SbSUT1, SbSUT2 and SbSUT4 (Fig. 4) and that the expression patterns of SbSUT1, SbSUT2 and SbSUT4 in response to defoliation were detectable. Meyer et al. (2004) found that expression levels of AtSUC3, which belonged to the SUT1 group, were strongly induced upon wounding in Arabidopsis tissue. In Lolium perenne, the increased level of LpSUT1 after defoliation might represent one of the mechanisms contributing to rapid refoliation (Berthier et al. 2009). Combined with our defoliation treatment results of LcSUT1 (Su et al. 2013), we could infer that the up-regulation of SbSUT1 in leaf sheaths after defoliation may be the result of sucrose starvation. Furthermore, the SbSUT2 gene may be responsible for the loading of sucrose in leaf sheaths and the SbSUT4 gene may be in charge of the unloading of sucrose from the growing organs after defoliation (Fig. 5). In summary, the five SbSUTs were functional sucrose transporters, and SbSUT1, SbSUT2 and SbSUT4 have different functions in response to defoliation. These findings will help to elucidate the function of sucrose transporter genes on sucrose loading and unloading.
Author contribution
The study was conceived and designed by Liu G.S. and Chen S.Y. The experiments were carried out by Su M. and Li X.X., and the plant materials were prepared by Li X.F., Cheng L.Q. and Qi D.M. Li X.X. contributed to the manuscript preparation. All authors read and approved the final manuscript.
Abbreviations
- SUT:
-
Sucrose transporter
- MFS:
-
Major facilitator superfamily
- cDNA:
-
DNA complementary to RNA
- ORF:
-
Open reading frame
- DNase:
-
Deoxyribonuclease
- RT-PCR:
-
Reverse transcriptase PCR
- qRT-PCR:
-
Quantitative RT-PCR
References
Aoki N, Hirose T, Scofield GN, Whitfeld PR, Furbank RT (2003) The sucrose transporter gene family in rice. Plant Cell Physiol 44:223–232
Barker L, Kühn C, Weise A, Schulz A, Gebhardt C, Hirner B, Hellmanna H, Schulze W, Ward JM, Frommer WB (2000) SUT2, a putative sucrose sensor in sieve elements. Plant Cell 12:1153–1164
Berthier A, Desclos M, Amiard V, Morvan-Bertrand A, Demmig-Adams B, William WA, Turgeon R, Prud’homme M, Noiraud-Romy N (2009) Activation of sucrose transport in defoliated Lolium perenne L.: an example of apoplastic phloem loading plasticity. Plant Cell Physiol 50:1329–1344
Braun DM, Slewinski TL (2009) Genetic control of carbon partitioning in grasses: roles of sucrose transporters and Tie-dyed loci in phloem loading. Plant Physiol 149:71–81
Callow MN, Michell P, Baker JE, Cocks PS, Houg GM (2005) Response of herbage regrowth and water-soluble carbohydrate concentration of ryegrass species to defoliation practices when grown in a Mediterranean environment. Grass Forage Sci 60:59–67
Carpaneto A, Geiger D, Bamberg E, Sauer N, Fromm J, Hedrich R (2005) Phloem-localized, proton-coupled sucrose carrier ZmSUT1 mediates sucrose efflux under the control of the sucrose gradient and the proton motive force. J Biol Chem 280:21437–21443
Chandran D, Reinders A, Ward JM (2003) Substrate specificity of the Arabidopsis thaliana sucrose transporter AtSUC2. J Biol Chem 278:44320–44325
Chen JY, Liu SL, Siao W, Wang SJ (2010) Hormone and sugar effects on rice sucrose transporter OsSUT1 expression in germinating embryos. Acta Physiol Plant 32:749–756
Daie J (1989) Phloem loading of sucrose: update and opportunities in molecular biology. Plant Mol Biol Rep 7:106–115
Doidy J, Tuinen DV, Lamotte O, Corneillat M, Alcaraz G, Wipf D (2012) The Medicago truncatula sucrose transporter family: characterization and implication of key member in carbon partitioning towards arbuscular mycorrhizal fungi. Mol Plant 5:1346–1358
Hirose T, Imaizumi N, Scofield GN, Furbank RT, Ohsugi R (1997) cDNA cloning and tissue specific expression of a gene for sucrose transporter from rice (Oryza sativa L.). Plant Cell Physiol 38:1389–1396
Hoagland DR, Arnon DI (1950) The water-culture method for growing plants without soil. California Agric Exp Stn Circ 347(1950):536–537
Housley TL, Volenec JJ (1988) Fructan content and synthesis in leaf tissue of Festuca arundinacea. Plant Physiol 86:1247–1251
Kühn C, Grof C (2010) Sucrose transporters of higher plants. Curr Opin Plant Biol 13:288–298
Lalonde S, Wipf D, Frommer WB (2004) Transport mechanisms for organic forms of carbon and nitrogen between source and sink. Annu Rev Plant Biol 55:341–372
Livaka KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Meyer S, Lauterbach C, Niedermeier M, Barth I, Sjolund RD, Sauer N (2004) Wounding enhances expression of AtSUC3, a sucrose transporter from arabidopsis sieve elements and sink tissues. Plant Physiol 134:684–693
Payyavula RS, Tay KHC, Tsai C-J, Harding SA (2011) The sucrose transporter family in Populus: the importance of a tonoplast PtaSUT4 to biomass and carbon partitioning. Plant J 65:757–770
Reinders A, Sivitz AB, Hsi A, Grof CP, Perroux JM, Ward JM (2006) Sugarcane ShSUT1: analysis of sucrose transport activity and inhibition by sucralose. Plant Cell Environ 29:1871–1880
Riesmeier JW, Willmitzer L, Frommer WB (1992) Isolation and characterization of a sucrose carrier cDNA from spinach by functional expression in yeast. EMBO J 11:4705–4713
Riesmeier JW, Willmitzer L, Frommer WB (1994) Evidence for an essential role of the sucrose transporter in phloem loading and assimilate partitioning. EMBO J 13:1–7
Sauer N (2007) Molecular physiology of higher plant sucrose transporters. FEBS Lett 581:2309–2317
Scofield GN, Hirose T, Aoki N, Furbank RT (2007) Involvement of the sucrose transporter, OsSUT1, in the long-distance pathway for assimilate transport in rice. J Exp Bot 58:3155–3169
Sivitz AB, Reinders A, Johnson ME, Krentz AD, Grof CP, Perroux JM, Ward JM (2007) Arabidopsis sucrose transporter AtSUC9. High-affinity transport activity, intragenic control of expression, and early flowering mutant phenotype. Plant Physiol 143:188–198
Slewinski TL, Braun DM (2010) Current perspectives on the regulation of whole-plant carbohydrate partitioning. Plant Sci 178:341–349
Slewinski TL, Meeley R, Braun DM (2009) Sucrose transporter 1 functions in phloem loading in maize leaves. J Exp Bot 60:881–892
Stadler R, Truernit E, Gahrtz M, Sauer N (1999) The AtSUC1 sucrose carrier may represent the osmotic driving force for anther dehiscence and pollen tube growth in Arabidopsis. Plant J 19:269–278
Su M, Li XF, Ma XY, Peng XJ, Zhao AG, Cheng LQ, Chen SY, Liu GS (2011) Cloning two P5CS genes from bioenergy sorghum and their expression profiles under abiotic stresses and MeJA treatment. Plant Sci 181:652–659
Su M, Li XX, Li XF, Cheng LQ, Qi DM, Chen SY, Liu GS (2013) Molecular characterization and defoliation-induced expression of a sucrose transporter LcSUT1 gene in sheep grass (Leymus chinesis). Plant Mol Biol Rep 31:1184–1191
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Volenec JJ (1986) Nonstructural carbohydrates in stem base components of tall fescue during regrowth. Crop Sci 26:122–127
Wang F, Liu CZ (2009) Development of an economic refining strategy of sweet sorghum in the Inner Mongolia region of China. Energy Fuels 23:4137–4142
Wang XL, Liu D, Li ZQ (2012) Effects of the coordination mechanism between roots and leaves induced by root-breaking and exogenous cytokinin spraying on the grazing tolerance of ryegrass. J Plant Res 125:407–416
Weise A, Barker L, Kühn C, Lalonde S, Buschmann H, Frommer WB et al (2000) A new subfamily of sucrose transporters, SUT4, with low affinity/high capacity localized in enucleate sieve elements of plants. Plant Cell 12:1345–1355
Zhou Y, Qu H, Dibley KE, Offler CE, Patrick JW (2007) A suite of sucrose transporters expressed in coats of developing legume seeds includes novel pH-independent facilitators. Plant J 49:750–764
Acknowledgments
We thank Dr. W.B. Frommer for his kind supply of the yeast strain SUSY7/ura3, the vector pDR196 and PDR196-StSUT1. This work was supported by National Basic Research Program of China (‘‘973”, 2014CB138704), the National Natural Science Foundation of China (31070302), the Project of Ningxia Agricultural Comprehensive Development Office (NNTK-2013-03 (1)), the Knowledge Innovation Program of the Chinese Academy of Sciences (KSCX2-EW-B-05), and National High Technology Research and Development Program of China (“863”, 2011AA100209).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by Y. Wang.
Rights and permissions
About this article
Cite this article
Li, X., Su, M., Li, X. et al. Molecular characterization and expression patterns of sucrose transport-related genes in sweet sorghum under defoliation. Acta Physiol Plant 36, 1251–1259 (2014). https://doi.org/10.1007/s11738-014-1505-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11738-014-1505-0