Abstract
Five SUT genes, namely SoSUT1, SoSUT2, SoSUT3, SoSUT4, and SoSUT5, were cloned, and their expressions in roots, stems, leaves, inflorescence, and buds at physiological mature stage were analyzed by qRT-PCR. Major findings of the study were: (1) the molecular mass of deduced sugarcane sucrose transporter proteins was between 53.44 and 61.80 kDa, and the pIs were between 5.94 and 10.68; (2) all sugarcane sucrose transporter proteins had 12 typical transmembrane domains; (3) there were two DNA sequences encoding the SoSUT3 gene with six exons and five introns; (4) while SoSUT2, SoSUT4, and SoSUT5 belonged to Clades SUT2, SUT4, and SUT5, other SoSUT proteins belonged to Clade SUT3; (5) real-time PCR analyses results showed SoSUT1, SoSUT4, and SoSUT5 had highly abundant expression in the inflorescence, and SoSUT2 and SoSUT3 highly expressed in the buds and inflorescence at physiological maturity. These data may indicate that different SUT genes may be transcribed during the sucrose accumulation process in sugarcane stems and that the expressions of SoSUTs may play an important role in sugarcane inflorescence development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In sugarcane, biosynthesis of sucrose occurs in the leaf tissue, where considerable amounts of sucrose are measurable in vacuoles or plastids (Schneider and Keller 2009). Sucrose is the predominant form of assimilated carbon being transported through the phloem from the source to sink organs. Sucrose transportation involves two organs, i.e., the loading of sucrose in phloem, mostly in the leaf veins, and the storage of sucrose in parenchyma of the stem. Sucrose accumulation begins with the translocation of sucrose through the phloem sieve elements to the stem internodes (Braun et al. 2014). Sucrose accumulation is accompanied by continuous cleavage and synthesis of sucrose in the parenchyma cells of the stem (McCormick et al. 2009), where sucrose metabolism is catalyzed by several enzymes, including sucrose phosphate synthase (SPS), sucrose synthase (SS), soluble acid invertase (SAI), neutral invertase (NI), and cell wall invertase (CIN) (Sturm 1999). A previous study showed that sucrose content is positively correlated with SPS activity, but negatively correlated with SAI and NI activities (Pan et al. 2009). In most plants studied, sugar transporters is responsible for sucrose transportation across membrane, such as sucrose loading and/or unloading in phloem, sucrose importation into sink tissues, and sucrose supply for organ development process (Shiratake 2007; Geiger 2011). Sugar transporters belong to the major facilitator superfamily (MFS), which is one of the largest groups of secondary transporters that are ubiquitously conserved. The MFS proteins selectively transport a wide spectrum of substrates across biomembranes and play a pivotal role in multiple physiological processes (Reddy et al. 2012; Yan 2013).
A sucrose transporter (ShSUT1) was identified by Rae et al. (2005) in sugarcane leaves and ElSayed et al. (2010) in sugarcane stem. The expression of SUT1 transporter in yeast mutants showed that SUT1 acted as a sucrose–proton symporter. In sugarcane, ShSUT1 was localized only in the layer of cells surrounding the vascular bundle sheath, but was absent in phloem itself (Rae et al. 2005). Sugarcane is capable of storing large amounts of sucrose in its stem (Sachdeva et al. 2005). The sucrose transport and accumulation in sugarcane internodes are very complicated processes, and how SUTs function in these processes remains unclear.
In order to understand which SUT proteins play a more important role in sucrose transportation and accumulation in sugarcane, we cloned and molecularly characterized five SUT genes in this study. The expression patterns of these sugarcane SUTs in the roots, stem, leaves, inflorescence, and buds at physiological mature stage were also analyzed. Finally, how these five sugarcane SUTs relate to those from other plant genera is also discussed.
Materials and Methods
Plant Material
Sugarcane variety GT28 was planted on January 20, 2012, in a field plot at College of Agriculture, Guangxi University (Nanning, China). On December 22, 2012, maturing internodes were sampled, while root, stem, leaf, inflorescence, and bud samples were taken at physiological mature stage on January 22, 2013. Six stalks were collected. The internode immediately subtended by the fully expanded leaf was designated as internode 1, with internodes below given sequentially increasing numbers (Rae et al. 2005). The internodes were cut and chopped into small pieces with a sharp knife. Internodes with the same sequential number were mixed together, immediately frozen in liquid nitrogen, and stored in a − 80 °C freezer until use.
RNA Extraction and cDNA Synthesis
Total RNAs were extracted using Trizol-A+ (Tiangen, Beijing, China) according to the manufacturer’s instructions. All RNA samples were treated with RNase-free DNase to eliminate DNA contamination. First-strand cDNA was synthesized from 1 μg total RNA using MLV reverse transcriptase of TaKaRa (Otsu, Japan). The cDNA was quantified in a spectrophotometer and diluted to a concentration of 50 ng/µL for quantitative real-time PCR (qRT-PCR) analysis. Total leaf DNA was extracted using a Nuclean Plant Gene DNA Kit (Tiangen, Beijing, China).
Amplification of Sugarcane SUTs Gene
SoSUT1 and SoSUT2 genes were cloned in our laboratory previously using RT-PCR and RACE-PCR with a length of 1678 and 2132 bp, respectively (unpublished). Oligonucleotide primers were designed based on maize and sorghum SUT3, SUT4, and SUT5 nucleotide sequences, using vector NTI Advance 11.0., to amplify the open reading frames (ORF) of sugarcane SoSUT3, SUT4, and SoSUT5. The primer sequences for the coding OFR of the five SoSUT genes are shown in Table 1. The PCR thermal cycling program was 5 min at 94 °C, followed by 35 cycles of 50 s at 94 °C, 30 s annealing at 56–61 °C, 2 min at 72 °C, and a final extension for 10 min at 72 °C. Ethidium bromide-stained PCR products were visualized through 1% agarose gel electrophoresis and photographed under UV light.
The ORFs of SoSUTs were amplified from leaf DNA following a thermal cycle program of 1 min at 94 °C, 30 cycles of (10 s at 98 °C, 5 min at 68 °C), and 10 min at 72 °C. Upon separation through 1% agarose gel electrophoresis, the ethidium bromide-stained PCR products were purified using the Gel Extraction Kit of OMEGA (Norcross, USA) before being cloned into the pMD18-T vector using the TA cloning kit (TaKaRa, Japan).
Bioinformatic Analyses
The molecular weights, isoelectric point (pI) values, and amino acid sequences of the SUT proteins were predicted using ExPASy (http://expasy.org/tools). Structure and function of the SUTs were analyzed by SMART (http://smart.embl-heidelberg.de). Protein trans-membrane structure was analyzed using TMPRED Server v2.0 (http://www.ch.embnet.org/cgi-bin/TMPRED_form_parser). Amino acid sequences of SUTs from both dicotyledonous and monocotyledonous plants were identified by searching public databases available at Phytozome (www.phytozome.net). A phylogenetic dendrogram of plant SUTs was made based on the deduced amino acid sequences using MEGA 6.0 (http://www.megasoftware.net) (Tamura et al. 2013). The neighbor-joining method was used for constructing the phylogenetic tree based on bootstrap analysis for 100 replications.
Extraction of Sucrose and Total Sugar
For each sample, 2.5 g of frozen internode tissue without peel was ground to fine powder in liquid nitrogen in chilled mortar and pestle. The tissue powder was added to a 50-mL centrifuge tube with 10 mL 80% ethanol and heated for 30 min in an 80 °C water bath. After centrifugation at 13,000×g for 15 min, the supernatant was poured into a clean flask, and the sediment was re-extracted twice in 80% ethanol as described above. All the supernatant was amalgamated before incubation in a 90 °C water bath. When the supernatant was volatilized to about 2 mL, distillated water was added to bring total volume to 10 mL. Impurities were removed by going through a 0.44-µm filter, and the supernatant was measured for total sugar and sucrose content. All the prepared supernatant samples were stored at 4 °C for further measurements of sucrose and total sugar.
Sucrose content was determined using high-performance liquid chromatography (HPLC). The chromatographic separation was achieved on an YMC-Pack NH2 carbohydrate column (250 × 4.6 mm, 5 μm), which was held at a constant temperature of 40 °C, using a mobile phase composed of acetonitrile and water (80:20, v/v) at a flow rate of 1.0 mL/min. The prototype of chromatographic grade is 30 mg/mL of sucrose. The content of total sugar was determined by the colorimetric method of Miller (Miller 1959). The data were presented using the mean of three replicates and were analyzed by SPSS v.15 software (SPSS Inc. Chicago, IL, USA).
Real-Time PCR
qRT-PCR primers were designed based on sorghum genomic DNA and sugarcane EST sequences using vector NTI Advance 11.0. The primer sequence and expected size of the amplified products are listed in Table 1. qRT-PCR was performed using SYBR Premix Ex Tap™ of TaKaRa (Japan) on Real-Time PCR Detection System (Bio-Rad, USA). Total reaction volume was 20 μL, containing 12.5 μL of SYBR Premix Ex Tap™, 2 μL of cDNA (50 ng/μL), 1 μL of each 10 μM primer, and 4.5 μL ddH2O. Thermal cycling program was set at 95 °C for 30 s, followed by 40 cycles of 95 °C for 5 s, 60 °C for 20 s. Melting curve analysis was conducted for each reaction to confirm reaction specificity. All cDNA samples were analyzed in triplicate. Data were analyzed by the instrument software automatically and converted into a Microsoft Excel format. The relative quantification analysis was done by the \( 2^{{ - \Delta \Delta C_{\text{t}} }} \) method of Livak and Schmittgen (2001). The glyceraldehyde-3-phosphate dehydrogenase gene (GAPDH: EF189713) was used as reference, which is expressed at constant levels in a broad range of sugarcane tissues (Iskandar et al. 2004).
Results
SoSUT cDNA Cloning and Sequence Analysis
The full-length coding sequences of five sugarcane SUT genes were successfully cloned through RT-PCR using a mixed template of leaf, stem, root, and inflorescence cDNAs. The amplified ORFs of SoSUT1, SoSUT2, SoSUT3, SoSUT4, and SoSUT5 products were identified through 1% agarose gel electrophoresis (Fig. 1a). Sequencing of cloned ORFs showed a size of 1626, 1527, 1749, 1509, and 1605 bp, respectively. The SoSUT2 ORF encoded a putative protein of 582 amino acids, with 5′ and 3′ untranslated regions (UTR) of 25 and 358 bp, respectively. The predicted molecular mass of SoSUT2 protein was approximately 61.80 kDa versus 53–57 kDa for other SUT2 proteins. SoSUT2 has an acidic pI < 6, while other SUTs have alkaline pI > 6.5 (Table 2). The SoSUT1 sequence shared 99% homology with the ShSUT1 (AY780256) with a 3′ UTR of 124 bp, while SoSUT4 shared up to 99% homology with ShSUT4 (GQ485583). These five sugarcane SUT proteins were predicted to have 12 trans-membrane helices, a typical feature of sucrose transporters of MFS. The nucleotide sequences of these five SoSUTs were registered with the NCBI database as KF808327, KF808330, KM403103, GQ485583, and KF808328, respectively (Table 2).
Cloning of sucrose transporter (SoSUT) genes in sugarcane. a M, DL2000 DNA ladder; and Lanes 1, 2, 3, 4 and 5, PCR-amplified DNA products of SoSUT1, SoSUT3, SoSUT2, SoSUT4, and SoSUT5, respectively. b M, DL10000 DNA ladder; and Lane 1, PCR amplified DNA products of SoSUT3 with sizes (3760 and 5040 bp) were indicated
When sugarcane genomic DNA was used as the template, the SoSUT3 ORF primers amplified two PCR products of 3760 and 5040 bp, respectively (Fig. 1b). These two PCR products shared 92% sequence homology, each composed of six exons and five introns. The exons were located at positions 1–162, 1972–2037, 2154–2252, 2379–2443, 2518–3434, and 3543–3760 in one sequence and of 1–162, 3188–3253, 3434–3532, 3659–3723, 3798–4714, and 4823–5040 in another sequence, respectively. These two sequences have been registered in NCBI database as KM403104 and KM434771.
Multiple Sequence Alignment and Evolutionary Relationships of SoSUTs
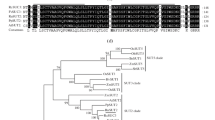
Analysis of the deduced amino acid sequences revealed that the five SoSUTs proteins contained 12 trans-membrane domains with both N- and C-termini at the cytoplasmic side. A conserved histidine residue (H), which is responsible for sucrose binding in transportation processes (Lu and Bush 1998), was present in all the five putative SUT peptides in the extracellular loop between the first and second membrane spanning helices (Fig. 2). These SUT peptides shared an amino acid sequence identity of 53–69% (Table 2), for example, 69% between SoSUT1 and SoSUT3, 53% between SoSUT1 and SoSUT4, and 56–64% among SoSUT1, SoSUT2 and SoSUT5.
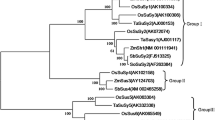
The SUT family is further divided into five subfamilies or clades based on homology, that is, SUT1, SUT2, SUT3, SUT4, and SUT5 (Fig. 3). Although phylogenetically separated from the dicot SUT1s, the SUT3s and SUT5s are only present in the monocot-specific branches. In sugarcane, SoSUT1 and SoSUT3 belonged to Clade SUT3, while SoSUT5 belonged to Clade SUT5, and SoSUT2 and SoSUT4 belonged to Clades SUT2 and SUT4, respectively. Both SUT2 and SUT4 were monocot- and dicot-specific subclades.
A phylogenetic tree of the sucrose transporter gene family of five plant species (At, Arabidopsis thaliana; Zm, Zea mays; So, Saccharum officinarum; Sb, Sorghum bicolor; Gm, Glycine max). Gene registration numbers of Arabidopsis thaliana and rice SUTs are taken from Shiratake (2007), and maize, sorghum and soybean SUTs are taken from Phytozome (www.phytozome.net). The unrooted N–J tree was constructed using MEGA 6.0 for sucrose transporter genes
Expression Patterns of Sugarcane SUT Family
At physiological mature stage, most of the SUT genes expressed in roots, stem, leaves, buds, and inflorescence tissues, but with an organ-specific regulation under normal growth conditions (Fig. 4). SoSUT1, SoSUT4, and SoSUT5 expressed the highest relative abundance in inflorescence. More SoSUT2 and SoSUT3 transcripts were detected in buds and inflorescence than in roots and stems. SoSUT5 transcripts were more abundant in inflorescence than in buds and stems, but were not detectable in leaves.
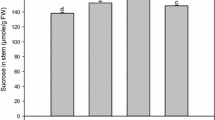
Initially, the sucrose and total sugar contents showed increasing from internodes + 1 to + 31 at maturity stage (Fig. 5). The difference of sucrose content between internodes + 16 and + 21 was significant by t test at the 0.05 level, while the differences of total sugar content among internodes + 16, + 21, + 26 and + 31 were not significant. In sugarcane internodes, a positive correlation coefficient of 0.975 was found between sucrose content and total sugar content (Table 3).
Expression analysis of the five SoSUT genes by qRT-PCR in different internodes showed that SoSUT2 and SoSUT5 had a similar gene expression pattern of increasing gradually from internodes + 16 to + 31 and reaching the maximum in internode + 31 (Fig. 6). SoSUT3 expression reached the maximum level in internode + 21 from its minimum level in internode + 1. However, SoSUT4 was highly expressed in internode + 1, but the least expression was found in internode + 6. In sugarcane internodes, the average sucrose content was positively correlated with total sugar (correlation coefficient value of 0.975) and with the abundance of gene transcripts of SoSUT1 (0.432), SoSUT2 (0.535), SoSUT3 (0.496), and SoSUT5 (0.369), but negatively correlated with SoSUT4 expression (− 0.560). The expression of SoSUT2 gene was positively correlated with SoSUT5 expression with a correlation coefficient of 0.977, which was very significant at the 0.01 level (Table 3).
Discussion
Due to its highly complex polyploid genome and the lack of a classical linkage map, whole-genome sequence is currently unavailable for sugarcane. Researchers have to rely on publically available genome sequence of its close relative sorghum, for comparative genomics studies in sugarcane (Jannoo et al. 2007; Wang et al. 2010). Since its first isolation from spinach (Riesmeier et al. 1993), many plant sucrose transporter (SUT) genes have been cloned and sequenced. For example, nine, five, and six SUT genes were identified in Arabidopsis, Oryza sativa (Shiratake 2007), and Theobroma cacao (Zhang et al. 2013), respectively. Phylogenetic analysis revealed that the plant SUT family can be divided into five subfamilies or distinct clades: SUT1 (dicot-specific), SUT3 and SUT5 (monocot-specific), and SUT2 and SUT4 (monocot and dicot). Members of the high-affinity dicot SUT1 clade, which exhibit apparent Km values between 0.07 and 2.0 mM sucrose phloem loading and long-distance transport, are expressed in the plasma membranes of sieve elements (SE) (Barker et al. 2000), companion cells (CC) or vegetative tissues (Aoki et al. 2004; Fan et al. 2009). The SUT2 and SUT4 transporters, which encompass both monocot and dicot members, demonstrate a lower affinity for sucrose that reported as Km values of 4–20 mM. The SUT2 transporters have been localized to SE plasma membranes in tomato, plantain (Barth et al. 2003) and Arabidopsis (Meyer et al. 2004). The members of the SUT4 clade catalyze sucrose uptake with low affinity, which have been identified in the chloroplast (Rolland et al. 2003), the tonoplast (Endler et al. 2006), and the plasma membrane (Chincinska et al. 2008). The SUT3 have been localized to the SE and CC plasma membrane, which has low affinity for sucrose, and exhibit apparent Km values of 2–8 mM sucrose (Reinders et al. 2006; Scofield et al. 2007). However, the SUT5 members have not been characterized yet. It is obvious that the subcellular localization of sucrose transporters is crucial for their functionality. Analysis of the SUT phylogenetic tree will provide new knowledge of the SUTs and its evolution in plants.
Analysis of the SoSUT expression in different tissues by RT-PCR in this study showed that the potential function of gene members could be predicted based on the gene expression profiles of the source and sink tissues in sugarcane. SoSUT1, SoSUT4, and SoSUT5 showed the highest relative expressions in inflorescence, and SoSUT2 and SoSUT3 transcripts were detected at higher levels in buds and inflorescence than in stems and roots. The result indicated that the expressions of SoSUTs may play an important role in sugarcane inflorescence development and seed formation. Carpaneto et al. (2005) reported that ZmSUT1 is capable of mediating both the sucrose uptake into the phloem in mature leaves (source) and the sucrose desorption from the phloem vessels into heterotrophic tissues (sink) of maize. Despite the genetic complexity, the identification of flowering genes in sugarcane will be beneficial for research into identifying potential targets to induce flowering on demand and understanding genetic variations between cultivars (Glassop et al. 2014).
Sucrose storage in sugarcane internodes is a developmentally regulated process, during which the parenchyma cells start to accumulate sucrose after a period of growth and expansion. Sucrose accumulation in storage tissues is accompanied by a continuous cycle of synthesis and cleavage of sucrose (McCormick et al. 2009). However, the role of these SUTs in sucrose accumulation in sugarcane is still need to be explored. This study is limited to one genotype GT28, further research is warranted to involve more sugarcane genotypes with a few contrasting traits that are associated with total sugar and sucrose yields. In addition, the expression profiles of these SoSUTs during entire growth need to be explored.
In a previous research, sugarcane ShSUT1 expression in maturing stems played an important role in the accumulation of sucrose in this tissue (Reinders et al. 2006). In rice, the OsSUT1 protein was found in the mature phloem of all the vegetative tissues involved in the long-distance assimilation transport pathway during grain filling. The OsSUT1 transcript was expressed in the uppermost internode of the rice plant (internode-1) (Scofield et al. 2007). In potato, the OsSUT5Z transgenic lines, the average tuber yield per plant was 1.9-fold higher than the controls. In average, more than 10 additional tubes were produced per plant without significant increase in average single tuber weight (Sun et al. 2011).
Sucrose transporter genes are located at key positions along the sucrose metabolic pathways, so there must be an effective regulatory mechanism to directly control the switch of sucrose transporters, and thereby to control the sucrose loading or unloading. All the evidence shows that sucrose transporters regulate the distribution of photosynthetic products through a variety of macromolecular events in plants (Braun and Slewinski 2009; Ma et al. 2009). Expression of SUT genes in the valuable sink organs has been improved through breeding to influence seed/tuber growth rate and yield (Sun et al. 2011; Li et al. 2011). In sugarcane internodes, sucrose content was positively correlated with the expression levels of SoSUT1, SoSUT2, SoSUT3 and SoSUT5, but was negatively correlated with the expression level of SoSUT4. This implies that different sucrose transporter genes may transcribe during the sucrose accumulation process in sugarcane stem.
Conclusions
Five sugarcane sucrose transporter (SUT) genes, namely SoSUT1, SoSUT2, SoSUT3, SoSUT4, and SoSUT5, were cloned, and their expressions in roots, stems, leaves, inflorescence, and buds at mature stage were analyzed. All sugarcane sucrose transporter proteins had 12 typical trans-membrane domains, deduced molecular mass of 53.44–61.80 kDa, and pI values of 5.94–10.68. There were two DNA sequences encoding the SoSUT3 gene with six exons and five introns. SoSUT2, SoSUT4 and SoSUT5 belonged to Clades SUT2 and SUT4, SUT5, and other SoSUT proteins belonged to Clade SUT3. Real-time PCR analyses results showed highly abundant expressions of SoSUT1, SoSUT4 and SoSUT5 in the inflorescence and of SoSUT2 and SoSUT3 in both buds and inflorescence at physiological mature stage. These results indicate that different sucrose transporter genes may be transcribed during the sucrose accumulation process in sugarcane stems and that the expressions of SoSUTs may play an important role in sugarcane inflorescence development and seed formation.
References
Aoki, N., G.N. Scofield, X.D. Wang, J.W. Patrick, C.E. Offler, and R.T. Furbank. 2004. Expression and localisation analysis of the wheat sucrose transporter TaSUT1 in vegetative tissues. Planta 219: 176–184.
Barker, L., C. Kühn, A. Weise, A. Schulz, C. Gebhardt, B. Hirner, H. Hellmann, W. Schulze, J.M. Ward, and W.B. Frommer. 2000. SUT2, a putative sucrose sensor in sieve elements. Plant Cell 12: 1153–1164.
Barth, I., S. Meyer, and N. Sauer. 2003. PmSUC3: characterization of a SUT2/SUC3-type sucrose transporter from Plantago major. Plant Cell 15: 1375–1385.
Braun, D.M., and T.L. Slewinski. 2009. Genetic control of carbon partitioning in grasses: roles of sucrose transporters and tiedyed loci in phloem loading. Plant Physiology 149: 71–81.
Braun, D.M., L. Wang, and Y.L. Ruan. 2014. Understanding and manipulating sucrose phloem loading, unloading, metabolism, and signaling to enhance crop yield and food security. Journal of Experimental Botany 65 (7): 1713–1735.
Carpaneto, A., D. Geiger, E. Bamberg, N. Sauer, J. Fromm, and R. Hdrich. 2005. Phloem-localized, proton-coupled sucrose carrier ZmSUT1 mediates sucrose efflux under the control of the sucrose gradient and the proton motive force. Journal of Biological Chemistry 280: 21437–21443.
Chincinska, I.A., J. Liesche, U. Krügel, J. Michalska, P. Geigenberger, B. Grimm, and C. Kühn. 2008. Sucrose transporter StSUT4 from potato affects flowering, tuberization, and shade avoidance response. Plant Physiology 146: 515–528.
ElSayed, A.I., M.F. Ramadan, and E. Komor. 2010. Expression of sucrose transporter (ShSUT1) in a Hawaiian sugarcane cultivar infected with sugarcane yellow leaf virus (SCYLV). Physiological and Molecular Plant Pathology 75: 56–63.
Endler, A., S. Meyer, S. Schelbert, T. SchneiderT, W. Weschke, S.W. Peters, F. Keller, S. Baginsky, E. Martinoia, and U.G. Schmidt. 2006. Identification of a vacuolar sucrose transporter in barley and Arabidopsis mesophyll cells by a tonoplast proteomic approach. Plant Physiology 141: 196–207.
Fan, R.C., C.C. Peng, Y.H. Xu, X.F. Wang, Y. Li, Y. Shang, S.Y. Du, R. Zhao, X.Y. Zhang, L.Y. Zhang, and D.P. Zhang. 2009. Apple sucrose transporter SUTl and sorbitol transporter SOT6 interact with cytochrome b5 to regulate their affinity for substrate sugars. Plant Physiology 150: 1880–1901.
Geiger, D. 2011. Plant sucrose transporters from a biophysical point of view. Molecular Plant 4: 395–406.
Glassop, D., A.L. Rae, and G.D. Bonnett. 2014. Sugarcane flowering genes and pathways in relation to vegetative regression. Sugar Tech 16: 235–240.
Iskandar, H.M., R.S. Simpson, R.E. Casu, G.D. Bonnett, D.J. Maclean, and J.M. Manners. 2004. Comparison of reference genes for quantitative real-time PCR analysis of gene expression in sugarcane. Plant Molecular Biology Reporter 22: 325–337.
Jannoo, N., L. Grivet, N. Chantret, O. Garsmeur, J.C. Glaszmann, P. Arruda, and A. D’Hont. 2007. Orthologous comparison in a gene-rich region among grasses reveals stability in the sugarcane polyploidy genome. Plant Journal 50: 574–585.
Li, F., C. Ma, X. Wang, C. Gao, J. Zhang, Y. Wang, N. Cong, X. Li, J. Wen, B. Yi, J. Shen, J. Tu, and T. Fu. 2011. Characterization of sucrose transporter alleles and their association with seed yield-related traits in Brassica napus L. BMC Plant Biology 11: 168–181.
Livak, K.J., and T.D. Schmittgen. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 25: 402–408.
Lu, J.M.Y., and D.R. Bush. 1998. His-65 in the proton-sucrose symporter is an essential amino acid whose modification with site-directed mutagenesis increases transport activity. Proceedings of the National Academy of Sciences of the USA 95: 9025–9030.
Ma, Y., T.L. Slewinski, R.F. Baker, and D.M. Braun. 2009. Tie-dyed 1 encodes a novel, phloem-expressed transmembrane protein that functions in carbohydrate partitioning. Plant Physiology 149: 181–194.
McCormick, A.J., D.A. Watt, and M.D. Cramer. 2009. Supply and demand: sink regulation of sugar accumulation in sugarcane. Journal of Experimental Botany 60: 357–364.
Meyer, S., C. Lauterbach, M. Niedermeier, I. Barth, R.D. Sjolund, and N. Sauer. 2004. Wounding enhances expression of AtSUC3, a sucrose transporter from Arabidopsis sieve elements and sink tissues. Plant Physiology 134: 684–693.
Miller, G.L. 1959. Use of dinitrosalicylic acid reagent for determination of reducing sugar. Analytical Chemistry 31: 426–428.
Pan, Y.Q., H.L. Luo, and Y.R. Li. 2009. Soluble acid invertase and sucrose phosphate synthase: key enzymes in regulating sucrose accumulation in sugarcane stalk. Sugar Tech 11: 28–33.
Rae, A.L., J.M. Perroux, and C.P. Grof. 2005. Sucrose partitioning between vascular bundles and storage parenchyma in the sugarcane stem: a potential role for the ShSUT1 sucrose transporter. Planta 220: 817–825.
Reddy, V.S., M.A. Shlykov, R. Castillo, E.I. Jr, and M.H.S. Sun. 2012. The major facilitator superfamily (MFS) revisited. FEBS Journal 279: 2022–2035.
Reinders, A., A.B. Sivitz, A. His, C.P.L. Grof, J.M. Perroux, and J.M. Ward. 2006. Sugarcane ShSUT1: analysis of sucrose transport activity and inhibition by sucralose. Plant, Cell and Environment 29: 1871–1880.
Riesmeier, J.W., B. Hirner, and W.B. Frommer. 1993. Potato sucrose transporter expression in minor veins indicates a role in phloem loading. Plant Cell Online 5: 1591–1598.
Rolland, N., M. Ferro, D. Seigneurin-Berny, J. Garin, R. Douce, and J. Joyard. 2003. Proteomics of chloroplast envelope membranes. Photosynthesis Research 78: 205–230.
Sachdeva, M., A.P.S. Mann, and S.K. Batta. 2005. Regulation of sucrose storage by amino acid arginine in sugarcane cell suspension cultures. Acta Physiologiae Plantarum 27: 37–43.
Schneider, T., and F. Keller. 2009. Raffinose in chloroplasts is synthesized in the cytosol and transported across the chloroplast envelope. Plant and Cell Physiology 50: 2174–2182.
Scofield, G.N., T. Hirose, N. Aoki, and R.T. Furbank. 2007. Involvement of the sucrose transporter, OsSUT1, in the long-distance pathway for assimilate transport in rice. Journal of Experimental Botany 58: 3155–3169.
Shiratake, K. 2007. Genetics of sucrose transporter in plants. Genes, Genomes and Genomics 1: 73–80.
Sturm, A. 1999. Invertases: primary structures, functions and roles in plant development and sucrose partitioning. Plant Physiology 121: 1–7.
Sun, A.J., Y. Dai, X.S. Zhang, C.M. Li, K. Meng, H.L. Xu, X.L. Wei, G.F. Xiao, P.B.F. Ouwerkerk, M. Wang, and M. Zhu. 2011. A transgenic study on affecting potato tuber yield by expressing the rice sucrose transporter genes OsSUT5Z and OsSUT2M. Journal of Integrative Plant Biology 53: 586–595.
Tamura, K., G. Stecher, D. Peterson, A. Filipski, and S. Kumar. 2013. MEGA6: molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution 30: 2725–2729.
Wang, J.P., B. Roe, S. Macmil, Q.Y. Yu, J. Murray, H.B. Tang, C.X. Chen, F. Najar, G. Wiley, J. Bowers, M.A.V. Sluys, D.S. Rokhsar, M.E. Hudson, S.P. Moose, A.H. Paterson, and R. Ming. 2010. Microcollinearity between autopolyploid sugarcane and diploid sorghum genomes. BMC Genomics 2010 (11): 261. https://doi.org/10.1186/1471-2164-11-261.
Yan, N. 2013. Structural advances for the major facilitator superfamily (MFS) transporters. Trends in Biochemical Sciences 38: 151–159.
Zhang, H.P., S.J. Zhang, G.H. Qin, L.F. Wang, T. Wu, K.J. Qi, and S.L. Zhang. 2013. Molecular cloning and expression analysis of a gene for sucrose transporter from pear (Pyrus bretschneideri Rehd.) fruit. Plant Physiology and Biochemistry 73: 63–69.
Funding
This study was supported in part by the grants from 863 Program project (2013AA102604), Guangxi Natural Science Foundation projects (2011GXNSFF018002, 2013NXNSFAA019073), International Scientific Cooperation Program project (2013DFA31600), Guangxi Funds for Bagui Scholars and Distinguished Experts (2013-3), and Project of Guangxi Sugarcane Innovation team of National Modern Agriculture Industry Technology System (gjnytxgxcxtd-03).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Niu, JQ., Huang, JL., Phan, TT. et al. Molecular Cloning and Expressional Analysis of Five Sucrose Transporter (SUT) Genes in Sugarcane. Sugar Tech 21, 47–54 (2019). https://doi.org/10.1007/s12355-018-0623-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12355-018-0623-1