Abstract
We investigated the parameter identification of a multi-scale physiological model of skeletal muscle, based on Huxley’s formulation. We focused particularly on the knee joint controlled by quadriceps muscles under electrical stimulation (ES) in subjects with a complete spinal cord injury. A noninvasive and in vivo identification protocol was thus applied through surface stimulation in nine subjects and through neural stimulation in one ES-implanted subject. The identification protocol included initial identification steps, which are adaptations of existing identification techniques to estimate most of the parameters of our model. Then we applied an original and safer identification protocol in dynamic conditions, which required resolution of a nonlinear programming (NLP) problem to identify the serial element stiffness of quadriceps. Each identification step and cross validation of the estimated model in dynamic condition were evaluated through a quadratic error criterion. The results highlighted good accuracy, the efficiency of the identification protocol and the ability of the estimated model to predict the subject-specific behavior of the musculoskeletal system. From the comparison of parameter values between subjects, we discussed and explored the inter-subject variability of parameters in order to select parameters that have to be identified in each patient.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Electrical stimulation (ES) can be used to induce the contraction of muscular fibers for rehabilitation purposes. It can be applied on the muscle surface (epimysial), through the motor nerve (neural) or spinal cord (epidural) [19]. Electrodes, in the first two cases can be either implanted or over the skin.
However, the stimulation patterns still need to be empirically tuned to obtain the desired functional effect. These trial sessions may be long and the results could be biased by the increased fatigue that occurs during the sessions. Indeed, the stimulation parameters are often not optimal regarding the individual subject’s characteristics and are very difficult to set in an objective manner. Therefore, an accurate muscle model may provide a simulation for analyzing the system behavior and could be used to optimize the ES tuning.
Muscle models could be based on black box approaches [15, 33, 39] or on physiological constitutive laws [24, 25, 45, 46]. Because of the difference in mass, geometry and dynamic characteristics between subjects and muscles, both types of models require an estimation of parameters specific to each subject and muscle. Based on a black box model, the authors in [15, 33, 39] introduced parameter identification methods through a model linearization in isometric conditions [15], or by using a known model structure, which allowed the authors to apply well-established identification methods in isometric and non-isometric conditions [33, 39]. However, the identification results were not clearly quantified and the validation was performed in either isometric conditions or with very specific and predefined input. Moreover, the models were not physiologically meaningful since the parameter values were meaningless and did not contain any information to gain insight into the basic underlying physiological/biomechanical processes.
On the other hand, physiological models have also been used and the identification of their parameters has been investigated in [12, 23, 43] through invasive protocols on isolated animal muscles. In the case of human applications, a noninvasive condition is usually required, where some internal states such as muscle lengths are not easily measurable, so numerical algorithms have to be adapted in a non-trivial way. The feasibility in isometric conditions was investigated to identify the parameters of a part of the model [5, 8, 27] or of the whole model [6]. In this latter work, the identification accuracy, quantified through normalized percentage errors, is between 8.49 and 32.2 %. In isometric conditions, the model is often simplified, limited and cannot describe the dynamic behavior of the musculoskeletal system. Under dynamic conditions, the model and the identification procedure become more difficult to handle, due to the influence of force–velocity and force–length phenomena.
In [7, 14, 16, 34], the authors identified the model parameters under dynamic conditions with a normalized root mean square errors of between 16 and 30 % in [16] and lower than 17 % in [14]. In [7], the identification accuracy was lower than 15 % and was quantified through the sum of absolute errors divided by the sum of the measurements, i.e. so-called fractional percent error. In these works the active muscle model was validated in dynamic conditions, while the identification procedure was based on isometric measurements [14, 34] or on both dynamic and isometric measurements [7, 16]. Hill-type muscle models were used in all of these cited works. Such models are based on a macroscopic and phenomenological formulation, and do not reflect enough nonlinear muscle properties. Besides, the authors in [32] evaluated the accuracy of such models and showed that Hill-model errors increase during movement for low stimulation rates (between 10 and 20 Hz), which are most relevant to normal movement conditions. The authors also highlighted the increase in Hill-model errors with increasing movement amplitude. Therefore, to overcome these limitations, it is necessary to use a model based on Huxley’s formulation [29], which takes microscopic phenomena into account, and includes more nonlinear dynamics that are most relevant in the representation of muscle behavior. Moreover, the identification of all parameters makes the identification protocol very complex and the selection of parameters to identify still remains very delicate. Indeed, as far as we know, no previous studies have determined the importance of each parameter in the model customization, by exploring their inter-subject variability.
In this work, we applied a new microscopic and physiological muscle model based on the Huxley formulation and previously developed in [29]. This model, which has so far only been used on isolated rabbit muscle in isometric conditions [23], was investigated here in human subjects with complete spinal cord injury (SCI) under dynamic conditions. Thus, we established a whole experimental protocol for parameter identification that we applied on knee joints, actuated by the electrically stimulated quadriceps muscle. The whole identification protocol was applied in isometric and dynamic conditions on each side in seven subjects with complete SCI. First, we applied an initial identification protocol including four steps in order to estimate the parameters which are common to Hill-type models and our model. Therefore, these methods are adaptations from existing identification methods applied on Hill-type models. We thus introduced an original identification protocol using dynamic measurements of the leg and based on the resolution of a nonlinear programming (NLP) problem to identify the stiffness of the serial element.
Our main aim is to facilitate clinical application of this identification procedure by physiotherapists. Therefore, we focused on two main points to reach this goal. The first involved simplification of the identification procedure by defining the relevant parameters to identify. We therefore discuss the inter-subject variability in the identification results to define the most subject-specific parameters. The second point is to establish and apply a safer and lighter identification protocol than the existing ones [8, 30], to estimate the stiffness of the serial element (SE) of muscle.
Thereafter, we introduce the new musculoskeletal model and describe the experimental setup and then we present the identification protocol and detail the NLP-based identification protocol. In Sect. 3, we present the experimental results obtained on human subjects. Finally, the results are discussed in Sect. 4.
2 Methods
2.1 Neuro-musculo-skeletal modeling
The musculoskeletal system of the knee joint controlled with the electrically stimulated quadriceps muscle is henceforth called the quadriceps–shank system. Its model consists of two parts:
-
model of electrically stimulated quadriceps muscle,
-
model of the controlled knee joint.
2.1.1 Electrically stimulated muscle model
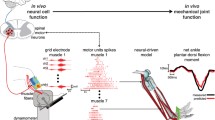
The muscle model, which was developed in a previous work [29], highlights a multi-scale aspect of muscle as a combination of macroscopic [24] and microscopic [25] model properties by describing its dynamics from the fiber to the whole muscle levels, as detailed in Appendix. It consists of two parts, as shown in Fig. 1a:
2.1.1.1 The activation model
It describes the generation of action potential and the initialization of contraction from the stimulation input. It includes two subparts:
-
The fiber recruitment model describes the spatial summation of activated fibers. It represents the relation between the electrical current applied on the nerve or the motor point and the rate of the activated fibers α (see Eq. 10).
-
The calcium dynamics model represents the electrochemical phenomena responsible for the force triggering within one fiber [20, 37]. This model, controlled by the stimulation frequency, generates a chemical pulse train u (Eq. 11), that introduces the temporal summation of forces.
2.1.1.2 The mechanical muscle model
It represents the contractile properties of the muscle–tendon structure. It is a Hill-type lumped element model [44] adapted for FES application. Figure 1b illustrates this model, which includes a contractile element (CE) in series with an elastic element (SE) whose stiffness is k s, and a viscoelastic parallel element (PE). With non-isolated muscles, PE is moved to the joint as a part of passive effects [16], as it cannot be easily evaluated alone. Therefore, the active force F, developed by the whole muscle–tendon structure, is equal to the active force of the contractile element F c (Fig. 1b). The specific model of CE, which describes the contraction under FES, is based on Huxley’s sliding filament theory [25] (see Appendix). It is represented by the dynamic equations of the force F c and stiffness K c of the contractile element, as presented in [29].
where, F cm and K cm correspond to maximal isometric force and stiffness, respectively. u, \(\Uppi_{\rm c}\) and U c are described by Eqs. (11) and (12) (see Appendix). Fl c is the CE force–length relationship, which relates the maximal isometric force to the strain of CE, \(\varepsilon_{\rm c}= \frac{L_{\rm c}-L_{\rm c0}}{L_{\rm c0}},\) with L c being the CE length and L c0 the rest length [21, 35]:
where b is the so-called shape parameter, which describes the overlapping level of filaments in sarcomeres.
Physiological models of electrically stimulated muscle: a overview of the muscle model. The stimulated muscle applies a force on the skeletal system. The joint position has a feedback on the muscle length and then on the contractile element length L c; b mechanical model of muscle includes the CE element, controlled by the recruitment rate α and the chemical pulse train u, the SE element, whose stiffness is k s, and the PE element; c the biomechanical model of knee joint. The stimulated quadriceps controls the knee joint extension while the gravity torque T g performs the flexion. The passive muscles include the passive part of muscle around the joint. The pulley radius r q is the moment arm of the quadriceps force and L f is the femur length
The mechanical interaction between CE and SE allows us to formulate the deformation velocity of the contractile element [29]:
Where:
and \(\varepsilon=\frac{L-L_0}{L_0}\) is the strain of the muscle–tendon structure, with L being the muscle−tendon length and L 0 its rest length. Then L = L c + L s , with L s being the SE length.
2.1.2 Dynamic model of a controlled knee joint
The knee joint is modeled in the sagittal plane as one degree of freedom θ, as illustrated in Fig. 1c. It is controlled by the quadriceps torque through a constant moment arm r q . The full knee extension is at 0° and the rest position is at θ 0 depending on the subject. The shank–foot group is considered as a single rigid body since we ensure that the foot motion according to the leg is very small during the experiment.
In this work, only the quadriceps muscle group was stimulated for the knee extension. Flexion was performed by the gravity torque T g due to the shank–foot group weight. The geometrical formulation of quadriceps muscle–tendon length is:
where L ext is the quadriceps length at the maximal extension (i.e. θ = 0°).
The dynamics of the knee joint motion around the rest position θ 0 is given by the following second-order nonlinear equation [42]:
where, θ r = θ 0 − θ is the knee joint angle w.r.t. the rest position and positive in the counterclockwise direction. \(T_{\rm q}=F \cdot r_{\rm q}\) is the active quadriceps torque. B and J are, respectively, the viscosity coefficient and the shank inertia around the center of rotation O. \(T_{\rm g}(\theta_{\rm r})=MgL_{\rm og}\sin(\theta_{\rm r})\) is the gravitational torque of the leg around the knee, where g is the gravity constant, M the mass of the leg and L og its center of mass w.r.t. the rotation point O. T e is an elastic torque that is usually considered to be highly nonlinear [13, 16].
The identification is complicated by the nonlinearity and difficulty of separating the gravity and elasticity torques in the measurements. Therefore, we considered the sum of the gravity and the elasticity torques as a single static torque (T s = T g + T e), as proposed in several previous works [7, 28, 40, 42]. From our measurements, which are discussed further (Sects. 2.3.1, 2.3.2), the static torque was modeled as:
where K is the parameter to identify.
2.2 Experimental setup
Experiments were conducted in the PROPARA rehabilitation center (Montpellier, France) with ten male patients who had a complete spinal cord lesion with an ASIA A score. An agreement from the local ethical committee and an informed and signed consent from the patients were obtained. The details of their clinical assessment are given in Table 1.
Subject 8 was excluded from the study because of muscle weaknesses and a high level of spasms. The first experiments, carried out on subjects 1 and 2, enabled us to solve some technical problems and adjust the protocol. Therefore, the results obtained on these subjects were incomplete and discarded. The subjects were seated on a chair with their hip flexed at approximately 90° and their thigh and back held against the seat. Therefore, since the flexion of the hip was fixed, the rectus femoris, which is biarticular muscle, is acting as monoarticular muscle on the knee joint within the quadriceps group. In this study, the right and left quadriceps–shank were separately considered. However, the choice of the first quadriceps–shank (right or left) tested was randomized.
The quadriceps muscle group was stimulated, with the PROSTIM stimulator (MXM-Sophia Antipolis, France and DEMAR, Montpellier, France), through surface electrodes, except for subject 10, for whom neural stimulation was performed using an implanted FES system [18]. The two surface electrodes of stimulation (10 cm × 5 cm, Cefar Medical, Lund, Sweden) were placed, one at the top of the rectus femoris and the other at the bottom of the vastus medialis, in order to stimulate the whole quadriceps muscle group. The stimulation frequency was set at 20 Hz for all subjects during all sessions. It was chosen to induce smooth contractions with the lowest fatigue effect possible according to [4]. The stimulation amplitude (current intensity) was fixed for all sessions for each subject. This stimulation amplitude was obtained during the first session by setting the pulse width at 300 μs, and increasing the amplitude until reaching level 3 on the Medical Research Council scale (MRC). The muscular force of quadriceps was controlled by the pulse width modulation, which was limited to PWmax = 420 μs for the safety of the patients.
In isometric conditions, the muscle torques were acquired using the BIODEX dynamometer (Biodex 3, Shirley Corporation, NY, USA), as presented in Fig. 2a and recorded on a PC through the Biopac MP100 acquisition device (Biopac Systems, CA, USA) and the isolation units INISO-Biopac, as presented in Fig. 2b. In dynamic conditions, the knee joint angles were acquired using an externally mounted electrogoniometer (Biometrics Ltd., VA, USA), as presented in Fig. 2c, and recorded on a PC through its acquisition system (DataLINK) as illustrated in Fig. 2d.
The knee joint angles and joint torques were sampled, respectively, at 100 Hz and 2 KHz, and then low-pass filtered at 30 Hz using the fourth-order Butterworth filter as in [16, 42].
2.3 Identification of physiological parameters
The physiological parameters of the model vary between subjects. Therefore, these subject-specific parameters had to be identified. The complexity of the current model increased the complexity of the identification procedure. Moreover, the parameter effects were hard to distinguish in one identification step in dynamic conditions. Therefore, we divided the identification protocol into several steps, with different measurement conditions, where the effect of each parameter was separated from the others.
The parameters to identify are pooled in two categories: the first one includes parameters common to Hill-type models, which were estimated in an initial identification phase, and the second are parameters specific to the current model (see Appendix), which were estimated in the second identification phase.
2.3.1 Initial identification phase
For this identification phase, we improved and adapted existing protocols and methodologies to the corresponding context of subjects with SCI. It includes four successive steps:
2.3.1.1 Estimation of geometrical parameters
In this step, we used a fifth-order polynomial equation, formulated in [22], to estimate the normalized muscle–tendon lengths for 10 regularly spaced knee joint positions, between full extension and 90° flexion. Then we combined this with the anthropometrical estimation of shank length [26] to estimate the values of L ext and r q (Eq. 5) using a linear least square method. Therefore, the geometrical parameters L ext and r q were estimated from the subject’s height without any limb measurement.
2.3.1.2 Identification of joint dynamics parameters
The knee joint dynamics parameters (Eq. 6) are J, B and K. Two measurement sets were performed without muscle activation (T q = 0).
-
The first test was conducted in static conditions (i.e. \(\dot{\theta}_{\rm r}=0\)). Thus for different knee joint angles, from the rest position to full extension with a step of 5°, the passive torques originating from the joint elasticity and gravity effects were measured. From the measurements (Fig. 4a), the relationship between the torques and knee joint angles was modeled by Eq. (7). This equation is linear with respect to parameter K, which was identified through a linear least square method.
-
In the second test, the passive pendulum test was applied [ 7, 13, 16, 28, 40]. By considering small displacements of the leg, the approximation sin(θ r) ≈ θ r was used. In the absence of muscular activity and considering the static torque model, Eq. (6) was linearized [28, 42]. The inertia J and the viscosity coefficient B were identified based on the damping ratio \(\zeta\) and the natural frequency ω n of the free response of a second-order linear system, such as in [28].
Identification of the quadriceps muscle stiffness parameter: a the identification principle based on the resolution of NLP problem; b profile of the input stimulation pulse width (subjects 1–9); c simulated knee joint angles with the whole estimated model and measured ones on the left leg of subject 3
Electromyography (EMG), through surface electrodes (Controle Graphique, Brie-Compte, France), was used to verify the absence of muscular activity.
2.3.1.3 Identification of the force–length relationship
The force–length relationship was identified in isometric conditions as in [12, 34, 41]. However, unlike these previous works, we considered the force–length relationship of the CE independently from the SE elastic properties. We assumed in this step that the SE was highly stiff compared to the CE [5, 12, 36, 38]. We applied a reduced stimulation in order to ensure lower CE stiffness compared to the SE stiffness. This was enough because the force–length relationship is independent of the muscle activation level [46]. Furthermore, we defined the rest length L c0 of CE as the length where the isometric force was maximal [46]. The quadriceps was stimulated with a pulse width (PW = 300 μs), and then the steady-state muscular active torques were measured (Fig. 2a, b) for seven knee joint angles (90°, 80°, 70°, 60°, 50°, 40°, 30°). The measured forces were filtered, as detailed in Sect. 2.2, calculated as the average of two trials and then normalized to the maximal obtained force.
For each knee joint position, the CE length L c was estimated by subtracting the tendon length L s from the muscle–tendon length L. For the current identification only, L s was assumed as a constant due to its high stiffness compared to the muscle. The muscle–tendon length L was calculated using Eq. (5) and the constant tendon length L s was obtained based on the proportion between the muscle and the tendon in the rest position [10], which corresponds to the position of the maximal isometric force. This approximation in static conditions, used only for the force–length relationship, was not used in the dynamic simulation thereafter. The inversion of Eq. (2) provides a linear function with respect to the parameters b and L c0. A linear least square method was thus applied to identify these parameters.
2.3.1.4 Identification of recruitment function parameters
Steady-state measurements were performed in isometric conditions to identify the recruitment function parameters, as done in [9, 11, 14, 27, 37, 43]. The knee joint was fixed at the position at which the torque was maximum. The quadriceps muscle was stimulated with 13 stimulation trains (duration 1.5 s), separated by a 4 s rest time. Pulse widths were fixed during each train, but increased between trains from 0 to maximum PWmax = 420 μs with a constant step. Isometric quadriceps torques were measured (Fig. 2a, b), filtered (Sect. 2.2) and averaged during the steady-state phase. The recruitment rates α were obtained by normalizing the torques to their maximum obtained at PWmax. A nonlinear least square method was applied to identify the parameters c 1, c 2 and c 3 of Eq. (10).
The maximal isometric force F cm, which corresponds to the saturation area, was thus deduced from the identified recruitment function. Finally, the maximal stiffness K cm chosen was proportional to the maximal isometric force F cm, according to the physiological model [29]. However, for a few subjects, K cm was increased to avoid oscillations that appeared in the dynamic response.
2.3.2 NLP-based identification of SE stiffness
In this second phase, the identification concerned the stiffness properties of the muscle–tendon structure. Since the maximal stiffness K cm was estimated previously (Sect. 2.3.1–2.3.4), we focused particularly on the identification of SE stiffness k s.
The muscle mechanical parameters, such as k s , are seldom specified in existing muscle models. Indeed, the series-elastic element and the contractile element are usually lumped in the same muscle–tendon structure and its stiffness is not considered [5, 7, 12, 17, 36, 38, 43].
As far as we know, very few works have focused on estimation of the stiffness in the muscle–tendon structure. In [23], the stiffness of series-elastic elements was considered and identified from the passive force–length relationship on rabbit. However, this passive force–length relationship is impossible to obtain in an experimental non-isolated muscle context. In [8, 30], a separate estimation of muscle and tendon stiffness was obtained from a linear measured compliance. This compliance has been measured on muscles from cat [30] and healthy human subjects [8] under strong and tetanic stimulation, during rapid muscle stretching under isometric conditions. However, these experimental identification protocols are very risky and inappropriate for SCI subjects due to their fragility and the very low muscle fatigue resistance.
The muscle model used here (Sect. 2.1.1) is more sensitive to parameter k s in dynamic conditions (i.e. \(\dot{\varepsilon} \neq 0\)) rather than in isometric conditions, since it is multiplied by \(\dot{\varepsilon}\) in Eq. (3). In a qualitative sense, generally the stiffness is relevant to the dynamic behavior. However, as far as we know, the dynamic conditions have never been considered for identification of stiffness in a muscle–tendon structure such as k s, nor for validation of the whole identified model, as presented in the current work.
Therefore, we defined an original identification protocol that is safer than existing ones [8, 30] using nonlinear optimization of the stiffness k s of SE elements along with the experimental data. This optimization is based on resolution of a nonlinear programming problem (NLP), which considers dynamic measurements of the shank motion without any mechanical constraint and the simulation of shank motion, using the model with parameters identified during the initial phase (Sect. 2.3.1). The quadriceps was stimulated in dynamic conditions, while the knee joint motion was recorded through the electrogoniometer (Fig. 2c, d). The input stimulation pulse width profile is presented in Fig. 3b. It was chosen to excite the internal dynamics of CE contraction. The same profile was applied for subject 10, but the intensity was modulated instead of the pulse width.
The use of an NLP-based identification method was due the nonlinearity of the model. It involved optimizing parameter k s under box constraints, which minimizes the quadratic criterion of output angular errors [θ r(t) − θ s(t)] (see Fig. 3a). It is formulated as follows:
where θ r represents the measured knee joint angles and θ s are the simulated ones using the whole identified model, including all of its initially identified parameters and the currently optimized parameter k s. t end is the movement duration used for the identification.
The algorithm of this constrained nonlinear optimization and its implementation are summarized belowFootnote 1 For this recursive identification, parameter k s was initialized with a value taken from literature [8].
3 Results
3.1 Performances of identification steps
The identification performances at all steps and the cross-validation results for all subjects are reported in Table 2. When the measured forces are too weak to be reliable, the results are represented by nonsignificant force (Nsf).
We selected an error criterion, called normalized root mean square deviation (NRMSD) to evaluate the performance of the identification method and the model fidelity:
where X m and X s are two vectors of the measured and simulated data and N is the number of samples within the considered duration.
For example, the identification results of the left quadriceps–shank of subject 3 are presented in Fig. 4.
In Fig. 4a, the simulated static torque closely fits the measured one, with a NRMSD of about 3.87 %. The results obtained for all subjects highlight a good static torque model accuracy and identification performance, with the NRMSD criteria being lower than 5 % (Table 2). Figure 4b highlights the close match of the identified knee joint dynamics model with the measured passive pendulum oscillations of the leg, as shown by a low NRMSD (3.26 %). However, we noticed small oscillations in the simulation around the rest position. These oscillations, due to the lack of coulombic friction in the model, are meaningless considering the lack of significant error. The results obtained in all subjects highlight the good identification performance, with NRMSDs lower than 5.1 % (Table 2).
The simulated force–length relationship (Eq. 2), as reproduced from the identified model, closely matches the measured one, as presented in Fig. 4c and confirmed by an NRMSD of under 4 %. For all subjects, most of these initial identification results illustrate good performance, with NRMSDs lower than 19 %. However, the results obtained for the right legs of subjects 5, 7 and 9 indicate lower performance, with an NRMSD ranging from 35 to 38 %. This was due to a reduced number (7 knee joint angle positions) of measurements so as to reduce a risk of muscle fatigue, so it could not compensate for the outlier data. It was also due to the small force levels that are not easy to simulate and can introduce a higher percentage of errors even if the absolute error is not very significant.
In Fig. 4d, the measured recruitment curve closely corresponds to the sigmoid model (NRMSD about 1.96 %). The results for all subjects showed good performance, confirmed by NRMSD to be lower than 6.2 %. In subject 10, who had a neural stimulation implanted system controlled through intensity modulation [18], a similar recruitment function was used, where the modulated parameter is the stimulation amplitude I.
The results of the NLP-based identification of parameter k s indicated close agreement between the measured knee joint angle trajectory and the simulated one, as presented in Fig. 3c and confirmed by a low NRMSD of about 4.77 %. The overall NLP-based identification demonstrated satisfactory results to a certain extent considering the difficult accessibility to this internal stiffness parameter. Their NRMSDs were lower than 19.9 %, except for the right leg of subject 5, where forces were nonsignificant, and the right leg of subject 4, where forces during this phase suddenly became high compared to previous measurements, indicating a variation in the muscle response to the stimulation over time.
3.2 Cross validations and model prediction performances
The cross validation of the subject-specific model was performed in dynamic conditions using a stimulation input that had not been applied during the two identification phases. The stimulation pulse width profile is presented in Fig. 5a. This estimated model includes all of the parameters identified at each step, and the parameter obtained from the NLP-based identification step as well.
Cross-validation results in dynamic condition: a profile of the input stimulation pulse width (subjects 1–9); b measured knee joint angle trajectory on the left leg of subject 3 and the two predicted ones, before and after stiffness parameter identification; c two knee joint angle error trajectories between each simulation and the measurement
To show the importance of k s identification in the final subject-specific model, two joint angle trajectory simulations were performed before and after k s identification and compared with the measurements on the left quadriceps–shank of subject 3, as presented in Fig. 5b. The two error trajectories between each simulation and the measurement are presented in Fig. 5c. The results highlight the importance of the NLP-based identification of k s and good final-model prediction of the knee dynamic behavior, since the NRMSD was 21.1 % before k s identification and became 5.17 % after. The final-model cross-validation results obtained in other subjects showed a quite good dynamic behavior prediction, with an NRMSD of under 19.7 %, except for the right leg of subject 5 and the left leg of subject 10, because of the weakness of their obtained forces, and for the right leg of subject 4, which still had a high NRMSD (≈50.5 %) due to abruptly high force measurements, which led to a poor NLP-based identification performance.
4 Discussion
The results of the NLP-based method highlight the feasibility of the new identification protocol for all subjects, with an NRMSD average of about is about 10.94 % (Table 2), except those who present small muscle contraction levels or a clear time-variant muscle response. With our method, these results demonstrate the identifiability of the internal stiffness parameter, which is significant with respect to the dynamic behavior and often excluded from experimental identifications, and set at literature values. The comparison between the two models (Fig. 5), without and with stiffness parameter identification, highlights the importance of identifying this parameter for improving the accuracy of the cross validation in dynamic conditions. The reason is that an identification in dynamical condition under an active condition of the muscle was required to improve the cross validation, since the initial identification steps did not consider this condition. The NLP-based identification protocol allows for safer clinical experiments because the dynamic measurements are performed without any physical constraint. In addition, this identification was able to compensate for the inaccuracy of the force–length identification results for the right legs of subjects 7 and 9, since it involved the whole model with the previously identified parameters. Indeed, the NLP-based identification results for these two cases were improved, with an NRMSD of about 8.63 and 10.5 %, while their force–length identification NRMSDs were about 35 and 38 %, respectively (see Table 2). However, this good NLP-based identification performance was not obtained for the right leg of subject 4 (NRMSD about 47.6 %), despite the good performances in its initial identification steps (3.13, 3.8, 10.4 and 2.44 %, see Table 2). This poor performance could be due to major variation problems with respect to the muscle response originating from the muscle fatigue or the surface electrode condition over time. It also shows the limits of NLP-based identification of stiffness parameter to deal with the time-varying parameters identified in the initial phase. NLP-based identification can be applied to identify other parameters as well, as long as the dynamic behavior is sensitive to them and every other parameters are previously estimated, as we tested it in [1] to estimate the recruitment function. On the other hand, some parameters such as k s should be identified only in dynamic conditions, as achieved here, due to their insignificant effects in isometric conditions.
The cross-validation results present satisfactory results and highlight the ability of the tuned model to predict the system dynamic behavior for all admissible subjects, with a low NRMSD average (≈12 %, see Table 2). These results indicate that the presented method together with the use of a complex Huxley-based muscle model could be very efficient for predicting the behavior in dynamic conditions.
The identified parameter values for all subjects are summarized in Table 3. To investigate the inter-subject variability and then the importance of each parameter in the identification protocol, we had to calculate the average of each parameter, also as summarized in Table 3. Each parameter average for our model, that is common to Hill-type models and was previously identified in [7, 15, 16, 35, 42] as summarized in Table 3, was used also for a comparison with literature values and discussion about its physiological relevance.
The geometrical parameters L ext and r q were estimated from the whole body height without any measurement (Sect. 2.3.1.1). Thus, the estimation performance was not quantified and the inter-subject variability study was thus meaningless. Furthermore, the average L ext value, which was around 42.2 cm, seemed realistic according to thigh lengths, which were roughly measured on each subject. In addition, the average value of the moment arm r q, which was 4.97 cm ± 4.88 %, is included in the range of values estimated in [3], i.e. between 4 and 6 cm.
The average K (10.7 N m/rad ± 23.8 %) was lower but close to values reported in [42] (14.3 ± 3.0 and 15.1 ± 3.9 N m/rad) and in [7] (12 and 14 N m/rad). The average J (0.3 kg m2 ± 22.9 %) was close to values estimated in [42] (0.32 ± 0.09 and 0.36 ± 0.13 kg m2), in [15] (0.35 ± 0.05 kg m2) and [7] (0.25 and 0.29 kg m2), and in the same range as the values estimated in [16] (0.41 ± 0.129 kg m2) and [28] (0.43 ± 0.18 kg m2).
The average viscosity coefficient B value (0.24 Nms/rad ± 35.4 %) was close to the values estimated in [15] (0.28 ± 0.016 Nm s/rad) and [16] (0.18 ± 0.05 Nm s/rad). It was, however, less close to the values obtained in [7] (0.15 and 0.08 Nm s/rad) and [42] (0.86 ± 0.69 and 0.58 ± 0.38 Nm s/rad). Finally, most values obtained are in the same order of magnitude of the ones reported in previous works. The variation is the expression of subject variability, quite high on patients with SCI. The average b value, which was around 0.42 ± 16.1 %, was very close to those introduced in [35], i.e. 0.4 for rectus femoris, and 0.45 for vasti. In addition, the average L c0 (about 9.22 cm) was quite close to the optimal fiber length values, cited in [10], of rectus femoris (8.4 cm), vastus medialis (8.9 cm), vastus intermedius (8.7 cm) and vastus lateralis (8.4 cm), which make up the quadriceps muscle group.
Average plateau level c 1 was 1.09 ± 11.9 %. It is realistic as it is close to 1, which represents the maximal recruitment rate. However, when this parameter was above 1, this showed that the identified recruitment function could theoretically exceed the maximal recruitment, since the real plateau was never reached, for safety reasons, and the maximal authorized stimulation was considered as 100 % of the recruitment. The average inflexion point c 3 value (0.62 ± 23.2 %) was close to 0.5, which represents the recruitment function where the half of stimulation level activates the half of total fibers. This means that the MRC index is meaningful to adjust a quasi-optimal range of PW variations through intensity adjustment (vice versa for subject 10).
In subject 10, who had a neural stimulation implanted system [18], the obtained parameters c 1, c 2 and c 3 were comparable to those of other subjects (Table 3) and were included in the statistical calculation. This showed that controlling the muscle force through the pulse width PW or the amplitude I of stimulation was similar.
From the identified parameter results, we calculated their relative standard deviations (RSD), which are included in Table 3. The RSD corresponds to the SD normalized to the average. It was used to quantify the inter-subject variability of each parameter in order to evaluate which parameters were more subject-specific and required more attention during the identification protocol.
The RSD of K and J, which were about 23.9 and 22.9 % respectively, highlighted a medium inter-subject variability, unlike the viscosity coefficient B, which presented a larger inter-subject variability, with a RSD of around 35.4 %. The variabilities in the joint elasticity K and viscosity J parameters were qualitatively realistic and expected because they matched the physical reality of the patient characteristics (see Table 1). Indeed, these variabilities matched the variability in body weight of the subjects (21.8 %), since K and J are mainly related to the body weight.
The RSD of L c0, which was equal to 3.77 %, highlighted a very low inter-subject variability. In addition, the shape parameter b represented a small inter-subject variability, with an RSD of about 16.1 %. The recruitment function identification results highlighted a low inter-subject variability of c 1 (with RSD = 11.9 %), a medium inter-subject variability of c 3 (with RSD = 23.2 %) and a very high inter-subject variability of c 2 (with RSD = 83.4 %). Moreover, parameter c 2, which represents the slope of the sigmoid recruitment function, is very important for FES control accuracy. Indeed, the bigger this slope is, the more difficult the accurate torque control is. Therefore, special attention should be given to the identification protocol for this parameter, which is highly specific to subjects and important in the model.
Values of parameters F cm, K cm represented very high inter-subject variability, as confirmed by their RSDs, which were respectively about 44.8 and 57.8 %. This high variability was expectable since the muscle strength varies with the patient age, the post-injury time (Table 1) and the recovery exercise rate in patients with SCI. This confirms the importance of identifying the maximal isometric force to personalize the model for each subject.
The few poor NLP identification results only concerned the right legs of subjects 4 and 5 (see Table 2), and were due to the muscle weakness and high time-variations rather than to the identification methodology. The k s values for these subjects were thus not included in the statistical computation of average and RSD. The high RSD value of parameter k s (61.2 %) and the marked difference between its minimal value (1 N/mm) and maximal value (12 N/mm), underlined a very high inter-subject variability and indicated the importance of identifying this parameter to obtain better accuracy with the subject-specific model. This confirmed the importance of identifying the SE stiffness k s, e.g. via our NLP-based identification method.
Besides, the identification results obtained in subject 10 (FES-implanted) were comparable to the results obtained in other subjects under surface stimulation. This also highlighted the feasibility of our identification strategy for applications involving neural stimulation with FES-implanted system.
Considering separately each quadriceps–leg of each subject, the cross-validation results, summarized in Table 2, highlighted 10 valid tests under surface stimulation and one valid test under implanted neural stimulation. This number of valid tests is higher than in the previous identification works, which were 9 in [42], 6 in [7], 5 in [8, 16], 4 in [34], 3 in [6] and 2 in [14]. The present work presents preliminary conclusions on a significant number of trials that confirm the relevance of the approach. However, our long-term goal is to make this protocol a clinical routine to extend the set of patient and then increase the statistical relevance.
5 Conclusion
In the current work, a whole parameter identification protocol was established and applied on a multi-scale model based on Huxley’s formulation. The experimental protocol was applied in vivo and noninvasively on 10 human subjects with a complete SCI. It was carried out on both knee joints controlled by their related electrically stimulated quadriceps muscles. Nine subjects were stimulated through surface electrodes, whereas one subject was stimulated through an implanted neural stimulator.
The whole identification protocol required isometric and dynamic measurements in passive and active conditions divided into several steps. In the initial identification steps, the geometrical parameters, knee joint mechanical parameters, force–length relationship and the recruitment function were estimated using adaptations of existing identification protocols applied in previous works on the Hill-type model. Least mean square methods were applied for these initial identification steps. In the last identification step, an original and safer identification protocol was applied to estimate the serial element stiffness of the quadriceps muscle. It is original since this parameter has seldom been considered and estimated in previous works, and it is safer as it considers dynamic measurements of shank motion without any mechanical constraint. This identification consists of a nonlinear optimization of the stiffness parameter based on the resolution of a nonlinear programming problem (NLP).
The results of each identification step and cross validation of the whole estimated model were evaluated through a quadratic error criterion between the simulated and measured data. It highlighted the good accuracy of each identification step and cross validation with a normalized RMS error of less than 20 %, except for a very few subjects, where the muscles were weak, quickly fatigued or presented a strong time-varying behavior. Cross validation in all subjects presented satisfactory results with an average normalized RMS error of about 12 %.
Our identification protocol should be generalized to other muscle–joint systems. However, practical issues related to the experimental position of the patients, the relevant choice of stimulated muscles and the stimulation selectivity are still open questions. Possible applications of the current results concern the synthesis of optimal electrical stimulation [2], the feedback control of the ES [14] and physiological interpretation or medical diagnosis based on model parameter monitoring.
Notes
This algorithm is based on trust-region method, which uses an approximation of the objective function with a simpler one in the neighborhood around the current variable, called the trust region [31]. For the implementation, Matlab optimization tools were used through the “fmincon” function.
References
Benoussaad M, Hayashibe M, Fattal C, Poignet P, Guiraud D (2009) Identification and validation of FES physiological musculoskeletal model in paraplegic subjects. In: 31st annual international conference of the IEEE engineering in medicine and biology society, pp 6538–6541
Benoussaad M, Poignet P, Guiraud D (2008) Optimal functional electrical stimulation patterns synthesis for knee joint control. In: Intelligent robots and systems IROS. IEEE/RSJ international conference on Nice pp 2386–2391. doi:10.1109/IROS.2008.4651112
Bonnefoy A, Doriot N, Senk M, Dohin B, Pradon D, Chze L (2007) A non-invasive protocol to determine the personalized moment arms of knee and ankle muscles. J Biomech 40:1776–1785
Chae J, Kilgore K, Triolo R, Creasey G, DiMarco A (2004) Functional neuromuscular stimulation. In: DeLisa J, Gans D (eds) Rehabilitation medicine: principles and practice, 4th edn. Philadelphia, J. B. Lippincott Company, pp 1405–1425
Chang YW, Su FC, Wu HW, An K N (1999) Optimum length of muscle contraction. Clin Biomech (Bristol, Avon) 14:537–542
Chia TL, Chow PC, Chizeck HJ (1991) Recursive parameter identification of constrained systems: an application to electrically stimulated muscle. IEEE Trans Biomed Eng 38:429–442. doi:10.1109/10.81562
Chizeck HJ, Chang S, Stein RB, Scheiner A, Ferencz DC (1999) Identification of electrically stimulated quadriceps muscles in paraplegic subjects. IEEE Trans Biomed Eng 46:51–61
Cook C, McDonagh M (1996) Measurement of muscle and tendon stiffness in man. Eur J Appl Physiol Occup Physiol 72:380–382
Crago PE, Peckham PH, Thrope GB (1980) Modulation of muscle force by recruitment during intramuscular stimulation. IEEE Trans Biomed Eng 27:679–684. doi:10.1109/TBME.1980.326592
Delp SL (1990) Surgery simulation: a computer graphics system to analyze and design musculoskeletal reconstructions of the lower limb. PhD thesis, Department of Mechanical Engeneering, Stanford University
Durfee WK, MacLean KE (1989) Methods for estimating isometric recruitment curves of electrically stimulated muscle. IEEE Trans Biomed Eng 36:654–667. doi:10.1109/10.32097
Durfee WK, Palmer KI (1994) Estimation of force-activation, force–length, and force–velocity properties in isolated, electrically stimulated muscle. IEEE Trans Biomed Eng 41:205–216. doi:10.1109/10.284939
Ferrarin M, Iacuone P, Mingrino A, Frigo C, Pedotti A (1996) A dynamic model of electrically activated knee muscles in healthy and paraplegics. chap. Neuroprosthetics from Basic Research to Clinical Applications A. Pedotti, M. Ferrarin, R. Riener, and J. Quintern, Eds. Berlin, Germany: Springer-Verlag pp. 81–90
Ferrarin M, Palazzo F, Riener R, Quintern J (2001) Model-based control of fes-induced single joint movements. IEEE Trans Rehabil Eng 9:245–257
Ferrarin M, Pedotti A (2000) The relationship between electrical stimulus and joint torque: a dynamic model. IEEE Trans Rehabil Eng 8:342–352
Franken DH, Veltink PP, Tijsmans IR, Nijmeijer DH, Boom PH (1993) Identification of passive knee joint and shank dynamics in paraplegics using quadriceps stimulation. IEEE Trans Rehabil Eng 1:154–164
Franken DH, Veltink PP, Tijsmans IR, Nijmeijer G, Boom PH (1995) Identification of quadriceps-shank dynamics using randomized interpulse interval stimulation. IEEE Trans Rehabil Eng 3:182–192
Guiraud D, Stieglitz T, Koch KP, Divoux JL, Rabischong P (2006) An implantable neuroprosthesis for standing and walking in paraplegia: 5-year patient follow-up. J Neural Eng 3:268–275
Harkema S, Gerasimenko Y, Hodes J, Burdick J, Angeli C, Chen Y, Ferreira C, Willhite A, Rejc E, Grossman RG, Edgerton VR (2011) Effect of epidural stimulation of the lumbosacral spinal cord on voluntary movement, standing, and assisted stepping after motor complete paraplegia: a case study. Lancet 377:1938–1947
Hatze H (1978) A general myocybernetic control model of skeletal muscle. Biol Cybern 28:143–157
Hatze H (1981) Myocybernetic Control Models of Skeletal Muscle. South Africa: University of South Africa
Hawkins D, Hull M (1990) A method for determining lower extremity muscle-tendon lengths during flexion/extension movements. J Biomech 23:487–494
Hayashibe M, Poignet P, Guiraud D, Makssoud HE (2008) Nonlinear identification of skeletal muscle dynamics with sigma-point kalman filter for model-based fes. In: 2008 IEEE international conference on robotics and automation. Pasadena, CA, USA, pp 2049–2054
Hill A (1938) The heat of shortening and the dynamic constants of muscle. R Soc Lond Proc Ser B 126:136–195
Huxley AF (1957) Muscle structure and theories of contraction. Prog Biophys Biophys Chem 7:255–318
de Leva P (1996) Adjustments to Zatsiorsky–Seluyanov as segment inertia parameters. J Biomech 29:1223–1230
Levy M, Mizrahi J, Susak Z (1990) Recruitment, force and fatigue characteristics of quadriceps muscles of paraplegics isometrically activated by surface functional electrical stimulation. J Biomed Eng 12:150–156. doi:10.1016/0141-5425(90)90136-B
Lin DC, Rymer WZ (1991) A quantitative analysis of pendular motion of the lower leg in spastic human subjects. IEEE Trans Biomed Eng 38:906–918. doi:10.1109/10.83611
Makssoud HE, Guiraud D, Poignet P, Hayashibe M, Wieber PB, Yoshida K, Azevedo-Coste C (2011) Multiscale modeling of skeletal muscle properties and experimental validations in isometric conditions. Biol Cybern 105:121–138. doi:10.1007/s00422-011-0445-7
Morgan DL (1977) Separation of active and passive components of short-range stiffness of muscle. Am J Physiol Cell Physiol 232:C45–C49
Nocedal J, Wright SJ (2006) Numerical optimization, 2nd edn. Springer Series in Operations Research and Financial Engineering, Springer, New York. doi:10.1007/978-0-387-40065-5
Perreault EJ, Heckman CJ, Sandercock TG (2003) Hill muscle model errors during movement are greatest within the physiologically relevant range of motor unit firing rates. J Biomech 36:211–218
Previdi F (2002) Identification of black-box nonlinear models for lower limb movement control using functional electrical stimulation. Control Eng Pract 10:91–99
Riener R, Ferrarin M, Pavan EE, Frigo CA (2000) Patient-driven control of FES-supported standing up and sittingdown: experimental results. IEEE Trans Rehabil Eng 8:523–529
Riener R, Fuhr T (1998) Patient-driven control of FES-supported standing up: a simulation study. IEEE Trans Rehabil Eng 6:113–124. doi:10.1109/86.895956
Riener R, Quintern J (1997) A physiologically based model of muscle activation verified by electrical stimulation. Bioelectrochemistry Bioenerg 43:257–264. doi:10.1016/S0302-4598(96)05191-4
Riener R, Quintern J, Psaier E, Schmidt G (1996) Physiological based multi-input model of muscle activation. Neuroprosthetics: from basic research to clinical applications. Springer, Berlin 12:95–114
Sapio VD, Warren J, Khatib O, Delp S (2005) Simulating the task-level control of human motion: amethodology and framework for implementation. Vis Comput 21:289–302
Schauer T, Negard NO, Previdi F, Hunt K, Fraser M, Ferchland E, Raisch J (2004) Online identification and nonlinear control of the electrically stimulated quadriceps muscle. Control Eng Pract 13:1207–1219
Scheiner A, Stein RB, Ferencz DC, Chizeck HJ (1993) Improved models for the lower leg in paraplegics. In: Engineering in medicine and biology society, the 15th annual international conference of the IEEE, San Diego, CA, USA, pp 1151–1152
Shue G, Crago P E, Chizeck HJ (1995) Muscle-joint models incorporating activation dynamics, moment-angle, and moment-velocity properties. IEEE Trans Biomed Eng 42:212–223. doi:10.1109/10.341834
Stein RB, Zehr EP, Lebiedowska MK, Popović DB, Scheiner A, Chizeck HJ (1996) Estimating mechanical parameters of leg segments in individuals with and without physical disabilities. IEEE Trans Rehabil Eng 4:201–211
Veltink PH, Chizeck HJ, Crago PE, El-Bialy A (1992) Nonlinear joint angle control for artificially stimulated muscle. IEEE Trans Biomed Eng 39:368–380. doi:10.1109/10.126609
Winters J (1990) Hill-based muscle models: a systems engineering perspective, chap 5. In: Winters JM, Woo SY (eds) Multiple muscle systems: biomechanics and movement organization. Springer, New York, pp 69–93
Zahalak GI (1981) A distribution-moment approximation for kinetic theories of muscular contraction. Math Biosci 55:89–114. doi:10.1016/0025-5564(81)90014-6
Zajac FE (1989) Muscle and tendon: properties, models, scaling, and application to biomechanics and motor control. Crit Rev Biomed Eng 17:359–411
Acknowledgments
The authors would like to thank the patients for their participation and patience, and also Maria Papaiordanidou, Robin Passama and Patrick Benoit for their help during the experiments.
Author information
Authors and Affiliations
Corresponding author
Appendix: Multi-scale muscle model
Appendix: Multi-scale muscle model
The appendix focuses on the contractile element for which a multi-scale approach was used to state, from the microscopic scale using Huxley’s formulation to a macroscopic and controlled muscle model that fully fulfills the well-established muscle law. At the microscopic scale, the authors in [29] started from Huxley’s formulation, which describes the distribution of the fraction of actin–myosin pairs. The key point is the choice of rate functions of attachment f and detachment g of the cross-bridges that allow integration together with a meaningful f and g definitions. Let us begin with the inputs of the model. We designed a model with two separate inputs, i.e. the recruitment rate α and the activation u. The first one is linked to the number of fired motor units deduced from the level of stimulation intensity and pulse width. It is a static relationship described by a sigmoid function [27] and modulated here by the pulse width PW of the electrical stimulus:
where c 1, c 2 and c 3 are parameters which represent the plateau level, maximum slope and inflexion point, respectively. The u function is linked to the calcium dynamics and expresses the rate of the creation or breakage of bridges. It is composed of three phases which are the muscle contraction and relaxation, and the transition between both states. It is defined by the following function [29]:
where \(\Uppi_{\rm c}\) is a function defined as follows:
where t r is the relative time position from the beginning of the transition phase. U c represents the rate of the actin–myosin cycle although U r represents the rate when mainly breakage occurs during the calcium restoring phase in the sarcoplasmic reticulum.
For all subjects, the same values τ c = 30 ms and τ r = 20 ms were obtained through curve fitting of isometric muscle forces under stimulation twitches. The values of U c and U r were fixed for all subject as well to 30 s−1 and 10 s−1, respectively.
From the activation function above and when stimulus is under way, f (although f is always zero) can be defined as follows:
where ξ is the relative position of the myosin head and the closest actin binding site. We consider g to be split in two parts, one linked to the chemical process from the cell at a rate u(t) that depends on the contraction phase and the other one linked to the relative shortening speed that may increase the detachment rate accordingly. It gives (compared to [29] the parameter a = 1):
Our definition is thus linked to the microscopic mechanisms involved within the muscle cell, but another assumption is now needed to change scale and to assess the macroscopic behavior. We do not compute the distribution n of the actin–myosin pairs fraction or express a specific distribution to solve the problem. Instead, we approximate the g function during the contraction phase as follows:
when f ≠ 0 and \(\xi \in [0,1].\) It can be shown that a constant can be added to this formulation without any mathematical change if the definition of g has to be adjusted. This is in fact equivalent to changing the U c and k 0 values (see below). In any case it is consistent with Huxley’s theory, which expresses that g is lower within than outside of this interval. Then we can write the modified g function:
such that f + g no longer depends on the variable ξ. The dynamics of the first two moments of the distribution n is then possible. The detailed computation is available in [29], and the main steps are:
with y linked to both the relative elongation of cross-bridges and relative position of thin and thick filaments, y 0 is the initial position of the spring, k 0 is the scaled stiffness at the sarcomere scale. The dynamics of these moments become:
Due to the wise definition of f and g, f + g does not depend on y, so integration is possible and we obtain:
fl is the force length relationship at the sarcomere scale, which represents the relation between the maximum number of actin–myosin pairs that can bind and the relative length of the contractile element. Then considering that all sarcomeres are identical, going to the fiber scale involves multiplying by a scale factor. And then going to the muscle scale involves scaling of the maximum available force and stiffness. This leads to Eq. (1) with macroscopic parameters which can be obtained by a combination of the microscopic parameters.
Rights and permissions
About this article
Cite this article
Benoussaad, M., Poignet, P., Hayashibe, M. et al. Experimental parameter identification of a multi-scale musculoskeletal model controlled by electrical stimulation: application to patients with spinal cord injury. Med Biol Eng Comput 51, 617–631 (2013). https://doi.org/10.1007/s11517-013-1032-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-013-1032-y