Abstract
The degradation of biodegradable plastics poses a significant environmental challenge and requires effective solutions. In this study, an esterase derived from a phyllosphere yeast Pseudozyma antarctica (PaE) enhanced the degradation and mineralization of poly(butylene succinate-co-adipate) (PBSA) film in soil. PaE was found to substitute for esterases from initial degraders and activate sequential esterase production from soil microbes. The PBSA film pretreated with PaE (PBSA-E) rapidly diminished and was mineralized in soil until day 55 with high CO2 production. Soil with PBSA-E maintained higher esterase activities with enhancement of microbial abundance, whereas soil with inactivated PaE-treated PBSA film (PBSA-inact E) showed gradual degradation and time-lagged esterase activity increases. The fungal genera Arthrobotrys and Tetracladium, as possible contributors to PBSA-film degradation, increased in abundance in soil with PBSA-inact E but were less abundant in soil with PBSA-E. The dominance of the fungal genus Fusarium and the bacterial genera Arthrobacter and Azotobacter in soil with PBSA-E further supported PBSA degradation. Our study highlights the potential of PaE in addressing concerns associated with biodegradable plastic persistence in agricultural and environmental contexts.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Agricultural mulch films made from biodegradable plastics (BPs) are plowed into soil after use, and eventually degraded to water and carbon dioxide (CO2) by soil microorganisms. Therefore, they are expected to be useful for post-use disposal in modern crop cultivation systems that benefit greatly from the use of plastic polymer-derived products, referred to as “Plasticulture” (Mormile et al. 2017). BPs are used in mulch films and can also be used in versatile products, such as nursery pots, bale wraps, strings, and rope (Guerrini et al. 2017).
However, the degradability of BPs depends on the environmental conditions. Several abiotic and biotic factors, such as humidity, temperature, organic matter content, and microbial activity, limited the biodegradation of BP in soil, with degradation occurring over several years (Li et al. 2014; Brodhagen et al. 2015; Martín-Closas et al. 2016; Sintim et al. 2020). Furthermore, manufacturers make BP mulches more durable to maintain their strength until harvest; this could slow down the degradation, and the remaining BP could accumulate in soil when the amount used exceeds the biodegradation rates. Accumulation of less biodegradable film fragments can adversely affect soil organisms as well as the agricultural and global environment by reducing water retention and soil permeability, as with non-degradable plastics (Miles et al. 2017).
It is thus necessary to develop plastics that are stable enough to maintain functionality during their use, which then biodegrade completely in an appropriate time scale. To address this need, we have been developing a new technique using a commercial BP-degrading enzyme to accelerate the degradation of biodegradable polyesters after use (Kitamoto et al. 2023). An esterase from the phyllosphere yeast Pseudozyma antarctia (PaE), which degraded various commercial BPs, was isolated, identified, and characterized (Kitamoto et al. 2011 and 2018, Shinozaki et al. 2013a and 2013b). PaE is an endoesterase that randomly degrades aliphatic and aliphatic–aromatic co-polyesters into oligomers and monomers with high activity compared to other known fungal BP-degrading enzymes (Sato et al. 2017; Kitamoto et al. 2023). When PaE was applied to commercial BP mulch films spread in a field, the strength of the film decreased the following day. After plowing, the size and total weight of the remaining fragments decreased further (Kitamoto et al. 2023). PaE pretreatment accelerated the reduction of residual solid commercial BP film, which was confirmed visually. Additionally, pretreatment promoted the recovery of soil fungal community structure, as depicted by electrophoresis profiles, from the impact of BP film addition in laboratory incubation experiments (Sameshima-Yamashita et al. 2019a).
To use PaE in the agricultural field, more information is needed regarding the fate of BP films cleaved by PaE, the role of PaE in the degradation of BP films, and the specific fungal and bacterial groups involved in this degradation process in soil. In this study, to address these knowledge gaps, we aimed to confirm the promotion of complete mineralization in PaE-pretreated BP films by measuring soil CO2 production, which is a direct indicator of biodegradation (Chinaglia et al. 2018; Francioni et al. 2022). We also aimed to investigate the degradation and mineralization processes by examining the changes in soil esterase activity and the abundance and structure of the soil microbial community.
Materials and methods
Preparation of soil and film samples
The soil used in this study (Andosol, loam, pH: 7.55, total carbon: 4.4%, weight ratio of organic carbon to total nitrogen: 11.9) was the same sample used by Yamamoto-Tamura et al. (2020), which was collected from the test field of the National Agricultural and Food Research Organization (36°01′22N, 140°06′52). The soil sample was sieved through a 2-mm mesh without air drying, and visible plant residues were carefully removed and stocked at 4 °C before use. In this investigation, a poly(butylene succinate-co-adipate) (PBSA) film was chosen due to its faster degradation in soil compared to other commonly used biodegradable polyesters for agricultural mulch films (Francioni et al. 2022; Tsuboi et al. 2022), making it suitable for the efficient evaluation of CO2 production. The PBSA film (provided by UNYCK Co., Ltd., Tokyo, Japan) was 15.0 ± 1.3 μm thick. The polyethylene film was also tested as a non-biodegradable plastic. The polyethylene film “Tenchi” (17 ± 2 μm thick) was purchased from Narutō Kasei K. K. (Tokyo, Japan). Both films were manually cut into 1 cm × 1 cm squares by the Heart Care Center Hitachinaka, Social Welfare Corporation Hamagiku-no-kai. The PBSA film pieces were stored at 4 °C after cutting to prevent unintended hydrolysis until they were used.
Enzyme production
Pseudozyma antarctica L1-S12 was cultivated in a jar fermentor (Sameshima-Yamashita et al. 2019b), and the culture filtrate was used to prepare the PaE solution. The strain was constructed as described in our previous study (Sameshima-Yamashita et al. 2019b), with slight modifications. The PaE gene expression cassette with a selective marker was amplified by polymerase chain reaction (PCR) and introduced into lysine auxotroph mutant GB-4(0)-L1 cells by electroporation, as described previously (Watanabe et al. 2016). The culture filtrate was diluted with sterilized water to 3.0 U and used as a PaE solution. Enzyme activity was evaluated based on the decreased turbidity of emulsified PBSA. As described previously (Shinozaki et al. 2013b), one unit (U) of PBSA degradation activity was defined as a 1 OD660 decrease per min in the reaction mixture (20 mM Tris–HCl, pH 6.8).
Experimental setup and PaE pretreatment of films
The four soil treatments used for further analysis over 167 days were as follows: (1) soil without any film or PaE treatment (control soil); (2) soil with non-degradable polyethylene film pretreated using PaE (soil with PE-E); (3) soil with PBSA film pretreated using PaE (soil with PBSA-E); and (4) soil with PBSA film pretreated using inactivated PaE (soil with PBSA-inact E). The PaE solution was autoclaved at 121 °C for 15 min to prepare the inactivated PaE solution (Ueda et al. 2015). Calcium carbonate (2% final concentration; SOFTON3200, Shiraishi Calcium Kaisha, Ltd., Osaka, Japan) was added to the PaE solution and the inactivated PaE solution to maintain a stable soil pH (Bache 1984). Although the optimum pH for the esterase activity of PaE is under alkaline conditions (Shinozaki et al. 2013b), the enzymatic degradation of PBSA produced oligomers and monomers with carboxylic acid ends, which lowered the pH of the reaction system. When the surface of a commercial biodegradable mulch film placed in the field is treated with PaE, the film decomposes more efficiently when combined with calcium carbonate (Kitamoto et al. 2023).
A total of 126 plastic film pieces (polyethylene 0.235 g or PBSA 0.3 g) were soaked in the PaE solution (24 μL/piece) in a 500 mL beaker for 3 min. Film pieces treated with inactivated PaE for 3 min were also prepared. Subsequently, 18 g dry-equivalent soil was added to each beaker containing the PaE-pretreated film pieces. The contents were thoroughly mixed and transferred to a plastic case with a lid (inner diameter: 50 mm, inner depth: 18 mm; Shiga Insect Promotion Company, Tokyo, Japan). For each treatment, triplicates were prepared for microbiological assays, and 3 − 5 replicates were prepared for CO2 measurement. The plastic cases were incubated at 25 °C in the dark, and the water content was adjusted and maintained at 60% (w/w) of the maximum water-holding capacity (MWHC) by weekly refilling of water decrements. In this study, we chose a small case containing 18 g of dry-equivalent soil to allow for a wider variety of treatments and replicates. This decision was informed by prior research on organic matter decomposition in soil, specifically referencing Wagai et al. (2013).
Soil samples were collected from three plastic cases for microbiological assays for each treatment on days 0, 1, 4, 7, 14, 19, 26, 55, 112, and 167 after the start of incubation. The sampled soil from each case was transferred to a 500 mL beaker and thoroughly mixed. Portions of the soil mixture were stored at − 20 °C for DNA analysis and at − 80 °C for esterase activity measurement until analysis. After sampling, the remaining soil from all three cases was combined and mixed well, and the visually discernible PBSA film pieces were picked up from a 40 g (wet weight) portion of the merged mixture, as shown in Fig. 1.
Evaluation of CO2 production from incubated soil
The CO2 measurement was conducted using the closed chamber method (Bekku et al. 1995) with a high-precision infrared gas analyzer (IRGA) (Li-7000, LiCor, Inc., Lincoln, NE, USA). The Li-7000 system consisted of closed chambers (diameter: 125 mm, height with lid: 120 mm) equipped with a butyl rubber septum and a pressure adjust bag attached to the lid. During each measurement, three plastic cases representing one replicate for each treatment were placed inside the chamber. The chamber was then hermetically sealed with water placed in an indent at the base of the chamber.
To measure CO2, after closing the chamber with a lid, a 0.5 mL headspace gas sample was taken from the chamber and injected into a N2 gas stream flow (500 mL min−1) connected to the Li-7000 system. The signal output of each injection was recorded by a CDS analyzer, and the area was calculated using CDS-Lite software ver. 5.0 (LAsoft, Ltd., Chiba, Japan). The CO2 concentration of the headspace gas was determined by comparing it to the area of standard CO2 gas (1.61%, air balance) using a linear regression relationship. The increase in CO2 concentration in each chamber was monitored at regular intervals (4 sampling points, 30 min apart) and expressed as a function of time (Bekku et al. 1995). The CO2 flux in each chamber was calculated using the following equation:
where P is the atmospheric pressure, Vc is the volume of the camber, Vp is the volume of the plastic case, R is the ideal gas constant, T is the absolute temperature, a is the CO2 slope, and W is the soil weight. CO2 emission rates for each treatment were measured on days 1, 4, and 7, and then on a weekly or monthly basis until the end of incubation. The CO2 production resulting from the degradation of films (PBSA-E and PBSA-inact E) was estimated by comparing the difference between the soil with films and the control (soil only).
Evaluation of soil esterase activity using p-nitrophenyl valerate (pNP-C5) and p-nitrophenyl laurate (pNP-C12)
To investigate the temporal variation in esterase activities for different acyl chain lengths in the soil, the activity of soil samples was determined by hydrolyzing p-nitrophenyl valerate (C5, Sigma-Aldrich, St Louis, MO, USA) and p-nitrophenyl laurate (C12, Tokyo Chemical Industry Co., Ltd., Tokyo, Japan) according to our previous study (Tsuboi et al. 2018) using Tris-maleate buffer, pH 6.0 (Sakai et al. 2002).
DNA extraction from soil samples
Total DNA was extracted from each soil sample (approx. 0.5 g) using a FastDNA SPIN Kit for Soil (Q-Biogene, Carlsbad, CA, USA) according to the manufacturer’s protocol, with a minor modification, as follows. In soil samples with buried PBSA films, DNA was extracted from the soil matrix and two film pieces until the films were observed. To enhance DNA recovery, a 10% casein solution (w/v) was added to the extraction buffer (Takada-Hoshino and Matsumoto 2004; Ikeda et al. 2008). The extracted DNA was further purified using a DNA Clean and Concentrator-25 kit (ZYMO RESEARCH, Irvine, CA, USA) according to the manufacturer’s protocol. The purified DNA samples were utilized for further molecular analyses to determine the microbial community structure and quantify fungi and bacteria.
Real-time quantitative PCR (qPCR)
Fungal and bacterial abundance in the soil samples was quantified using qPCR of the large subunit (LSU) rDNA copy numbers and 16S rDNA copy numbers, respectively. Each standard sample was generated using PCR amplicons from the Saccharomyces cerevisiae S288c genome using ITS1-LR3 primers (White et al. 1990; Vilgalys and Hester 1990) and the Escherichia coli DH5α genome using 63F-1492R primers (Marchesi et al. 1998; Lane 1991). The DNA concentration of the standards was determined using the Qubit BR dsDNA kit (Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s instructions. qPCR was conducted using SYBR Premix ExTaq II (TaKaRa BIO, Shiga, Japan) with the NL1F-LS2R primer pair for fungi and the 338f-518r primer pair for bacteria (Barnard et al. 2013) according to the manufacturer’s instructions on the StepOne Plus Real-Time System (Thermo Fisher Scientific).
Analysis of the fungal and bacterial communities
The fungal and bacterial communities were investigated with Miseq using a dual-index two-step approach. The ITS2 region for fungi and the 16S rDNA V4 region for bacteria were amplified using modified ITS3-Mix and ITS4-Mix primers (Keiblinger et al. 2018) and the 515f and 806r primers (Caporaso et al. 2012), respectively. These primers comprised an Illumina sequencing primer region. To improve sequencing quality (Lundberg et al. 2013), 6 mer Ns and 3–6 mer Ns were fused for fungal primers and bacterial primers, respectively, between the sequencing primer region and the target-specific region. The PCR mixture contained 25 μL of 2 × Q5 High-Fidelity Master Mix (New England Biolabs Japan, Tokyo, Japan), 1 μL (bacteria)/2.5 μL (fungi) of each primer (10 μM), 1 μL of the purified template DNA sample (1 ng/μL for bacteria and 25 ng/μL for fungi), and nuclease-free water added to a total volume of 50 μL. The thermal conditions for the first PCR are shown in Table S1 (Supplementary information). The first PCR products were purified, followed by the second PCR and purification of its products, in accordance with the Nextera library protocol from Illumina (2013).
Paired-end sequencing was conducted using Illumina Miseq, and the resulting demultiplexed fastq files corresponding to each sample were obtained directly. We removed adaptor sequences from the fastq files using Cutadapt (Martin 2014). Following this, low-quality regions in the sequences were trimmed using Trimmomatic (Bolger et al. 2014), applying the parameters “SLIDINGWINDOW:20:20 MINLEN:50”. Subsequently, forward and reverse reads were merged using fastq-join (Aronesty 2013). Data analyses, including assembly and quality filtering, were performed using QIIME ver. 1.9.1 (Caporaso et al. 2010). The chimera sequences were removed from the filtered sequences using VSEARCH (Rognes et al. 2016) based on the UCHIME algorithm (Edgar et al. 2011) using the UNITE database (https://doi.org/10.15156/BIO/587476) (Kõljalg et al. 2013) for the fungal ITS2 region sequences and the SILVA database (release 128) (Quast et al. 2013) for the bacterial 16S rDNA sequences. These sequences were then clustered into operational taxonomic units (OTUs) with a cut-off of 97% sequence similarity using QIIME. The taxonomic assignment of each OTU was carried out based on the UNITE database for fungal ITS2 region sequences and the SILVA database for bacterial 16S rRNA gene sequences. All sequences of the partial fungal ITS2 and bacterial 16S rDNA amplicon have been deposited into DDBJ/EMBL/GenBank databases under accession numbers DRR492222-DRR492332 and DRR491506-DRR491616, respectively.
Statistical analyses
Statistical differences in CO2 production, pNP-C5 and pNP-C12 hydrolytic activities, and copy numbers of fungal LSU rRNA and bacterial 16S rRNA genes among the four soil systems were calculated using a Tukey–Kramer test in “R” statistics software version 3.5.2 (R Development Core Team 2018).
PBSA film-degrading potential of fungal strains
Eight fungal strains, including seven of the genus Arthrobotrys (Arthrobotrys oligospora MAFF425031 and MAFF425032; A. superba MAFF236540; Arthrobotrys sp. MAFF243669, MAFF243670, MAFF243671, and MAFF243672) and one of the genus Tetracladium (Tetracladium setigerum MAFF425374) were obtained from the MAFF Genebank project culture collection at the Genetic Resources Center of the National Agriculture and Food Research Organization (NARO). These fungi were used to evaluate PBSA-degrading potential because the fungi belonging to these genera became dominant after burying the PBSA film in the soil in the present study. The degradation of emulsified PBSA (Bionolle® EM-301; Showa Denko K. K., Tokyo, Japan) on an agar plate and solid PBSA film (Bionolle® 3001 G, Showa Denko K. K., 20 ± 3 μm thick) was evaluated as described by Koitabashi et al. (2012) with slight modification as follows. In this study, potato dextrose agar medium (PDA, Nissui Co., Tokyo, Japan) was used for the preculture of the strains instead of the FMZ medium (a fungal minimal medium in the original method) to enhance the hyphae growth of the tested strains.
Results
Evaluation of CO2 production from soil and degradation of PBSA film pieces
Pieces of visible film were periodically recovered from the soil (Fig. 1). Only a few pieces of PBSA-E films could be collected from the day after they were buried in the soil, and none could be recovered after 55 days. Meanwhile, the PBSA-inact E films in the soil were partially degraded, but most remained over time (Fig. 1).
To evaluate PBSA film mineralization in the soil, the soil CO2 flux was measured in parallel (Fig. 2a and b). Intense CO2 production was observed for the first 7 days in all experiments. CO2 production was the highest among all treatments from day 1 to 55 (p < 0.05) in soil with PBSA-E (Fig. 2a), which then decreased and maintained values lower than those in the soil with PBSA-inact E but higher than those in the control soil (Fig. 2b). In soil with PBSA-inact E, CO2 production showed significantly higher values (p < 0.05) than those in control soil from days 4 to 167, except on days 7, 26, 132, and 146 (Fig. 2b). CO2 production in the soil with PE-E showed no significant differences from that in the control soil (Fig. 2b).
Temporal variations in CO2 production from the four soil treatments: the control soil and soils with PE pretreated using PaE (PE-E), PBSA pretreated using PaE (PBSA-E), and PBSA pretreated using inactivated PaE (PBSA-inact E). The whole view (a) and enlarged view from 0 to 0.08 mg CO2 g−1 dry soil day−1 in the CO2 flux (b) are shown. Means (n = 3) are presented with standard errors as black bars
Temporal variation in soil esterase activity hydrolyzing pNP-C5 and pNP-C12
In soil with PBSA-E, both pNP-C5 and pNP-C12 hydrolytic activities showed significantly higher or higher tendency values than those in the control soil and other treatments from day 1 to 112 after starting the incubation (Fig. 3a and b). The pNP-C5 hydrolytic activity retained higher values than 200 nmol g−1 min−1 from days 4 to 112 and then dropped (Fig. 3a), while the pNP-C12 hydrolytic activity of the soil gradually decreased (Fig. 3b). In soil with PBSA-inact E, the pNP-C5 and pNP-C12 hydrolytic activities showed increasing tendencies on days 56 and 26, respectively (Fig. 3a and b). The esterase activities of the two substrates were almost the same as those in the control soils for the first 19 days of incubation, showing a lack of PaE activity in the added solution. In soil with PE-E, because of PaE pretreatment, the hydrolytic activity of pNP-C5 and pNP-C12 was higher than that of the control on day 1. However, they rapidly decreased to the control level on day 55. In the control soil, the hydrolytic activities of pNP-C5 and pNP-C12 were constant during incubation (Fig. 3a and b).
Temporal variations in p-nitrophenyl valerate (pNP-C5) (a) and p-nitrophenyl laurate (pNP-C12) hydrolytic activity (b) based on the pNP production rate in the four different soil treatments: control soil and soils with PE pretreated using PaE (PE-E), PBSA pretreated using PaE (PBSA-E), and PBSA pretreated using inactivated PaE (PBSA-inact E). Means (n = 3) are presented with standard errors as black bars. Sampling was performed on days 0, 1, 4, 7, 14, 19, 26, 55, 112, and 167 after starting the incubation
Temporal variation in fungal and bacterial abundance
The temporal variations in fungal abundance based on LSU rDNA copy numbers were determined using qPCR (Fig. 4a). In soils with PBSA-E and PBSA-inact E, the fungal abundance increased by up to 10 times during the first 19 days, and was maintained at more than 108 copies until day 167, while that in control soil was mostly constant during incubation. The abundance increased more rapidly in soil with PBSA-E than in soil with PBSA-inact E. Fungal abundance in soil with PE-E showed values similar to those in the control soil, with no significant differences between them.
Temporal variations in fungal and bacterial abundance in four different soil treatments: control soil and soils with PE pretreated using PaE (PE-E), PBSA pretreated using PaE (PBSA-E), and PBSA pretreated using inactivated PaE (PBSA-inact E). Fungal abundance was measured based on LSU rRNA gene (a) and bacterial abundance was based on 16S rRNA gene copy numbers (b). Means (n = 3) are presented with standard errors as black bars. Sampling was performed on days 0, 1, 4, 7, 14, 19, 26, 55, 112, and 167 after starting the incubation
Temporal variations in bacterial abundance were measured based on 16S rDNA copy numbers (Fig. 4b). In soil with PBSA-E, the bacterial abundance gradually increased up to day 26 and then decreased, while that in soil with PBSA-inact E peaked at day 14 and quickly decreased. The bacterial abundance in the soil with PE-E was constant and showed no significant differences from that in the control soil.
Temporal variations in microbial community structures
In this study, the number of analyzed amplicons from the fungal ITS region and bacterial 16S rDNA ranged from 57,064 to 254,643 reads and from 11,489 to 135,730 reads, respectively (Tables S2 and S3 (Supplementary information)). Although the analyzed read numbers varied, the community structures were similar among the triplicate samples (Figs. S1 and S2 (Supplementary information)).
Composition of fungal communities
The temporal changes in the dominant fungal taxa in soil with PBSA-E and PBSA-inact E were different (Fig. 5a and 6 and Table S4 (Supplementary information)). The phylum Ascomycota, including Sordariomyctes, Orbiliomycetes, Leotiomycetes, Eurotiomycetes, and Dothideomycetes, was the most dominant, and variable phyla were observed in both soils throughout the experiment. In the control soil and the soil with PE-E, the fungal community structures on all sampling days were similar.
Temporal variations in the fungal (a) and bacterial community composition (b) in the four different treatments: the control soil and soils with PE pretreated using PaE (PE-E), PBSA pretreated using PaE (PBSA-E), and PBSA pretreated using inactivated PaE (PBSA-inact E). The relative abundance of each fungal and bacterial taxon is presented as the mean values of triplicates. The sampling days were 0, 1, 4, 7, 14, 19, 26, 55, 112, and 167 days after starting the incubation
Temporal variations of the dominant phylogenetic groups of fungal and bacterial communities in soils with PBSA pretreated using PaE (PBSA-E) and PBSA pretreated using inactivated PaE (PBSA-inact E). The percentages of each fungal and bacterial phylogenetic group are presented as the mean values of triplicate samples. Sampling was performed at 0, 1, 4, 7, 14, 19, 26, 55, 112, and 167 days after starting the incubation
In soil with PBSA-E, class Sordariomyctes rapidly increased in relative abundance from day 1 and dominated the community with 70.8% at day 7, maintaining a high proportion (over 39%) throughout the incubation period (Fig. 5a). In the Sordariomyctes class, two Fusarium-related OTUs (OTU433F and OTU474F) showed increased proportions for the first 7 days (Fig. 6). The ratio of classes Orbiliomycetes, Leotiomycetes, and Eurotiomycetes gradually increased from the start of incubation and peaked on days 19 (5.8%), 55 (9.8%), and 112 (21.3%), respectively (Fig. 5a). The representative OTUs in each class also showed similar tendencies, including the genus Arthrobotrys-related OTU449F in Orbiliomycetes, genus Tetracladium-related OTU542F in Leotiomycetes, and genus Exophiala-related OTU620F in Eurotiomycetes (Fig. 6).
Meanwhile, in soil with PBSA-inact E, the class Orbiliomycetes showed increased relative abundance from day 1 (1.4%) to day 26 (38.4%) and maintained dominance (31.7%) at day 55 (Fig. 5a). In the Orbiliomycetes class, the representative OTUs were related to the genus Arthrobotrys (Table S4 (Supplementary information)). OTU630F first increased to 12.2% on day 7, and OTU449F became dominant (over 20%) from day 14 to 55 (Fig. 6). After day 112, class Leotiomycetes, including Tetracladium-related OTU542F, showed an abundance of over 20% (Fig. 5a and 6). On day 167, the class Eurotiomycetes, including Exophiala-related OTU620F, comprised more than 5% (Fig. 6 and Table S4 (Supplementary information)). Compared to the soil with PBSA-E, OTU449F, OTU542F, and OTU620 reached a peak later in soil with PBSA-inact E, and the former two showed higher relative abundance, while the latter showed a lower relative abundance (Fig. 6).
Composition of bacterial communities
Similar to the fungal communities, the dominant bacterial taxa in soils with PBSA-E and PBSA-inact E showed temporal changes that differed from each other. Furthermore, the bacterial communities were relatively constant and similar in the control soil and in the soil with PE-E during incubation (Fig. 5b and 6 and Table S5 (Supplementary information)).
In the soil with PBSA-E, a relatively high proportion of Actinobacteria was found on day 1 (35.1%) and day 4 (32.0%) (Fig. 5b), and the genus Arthrobacter of the Actinobacteria-related OTU (OTU300601B) seemed likely to be more abundant (over 2%) from day 1 to 7 than that in the control soil (Fig. 6). The relative abundance of Gammaproteobacteria increased from day 1 (3.0%) to 14 (25.2%), and then comprised approximately 15% at day 55 (Fig. 5b). The genus Azotobacter in Gammaproteobacteria-related OTUs, especially OTU212542B and OTU267604B, was dominant from day 7 to 55 (Fig. 6).
In soil with PBSA-inact E, the temporal changes in dominant bacterial taxa were different from those in soil with PBSA-E. Relatively high proportions of Betaproteobacteria (about 16%) were observed at days 4 and 7 (Fig. 5b). Similar to the relatively gradual increase in Alphaproteobacteria toward the end of incubation (9.7–14.2%), the proportion of the genus Bradyrhizobium in subphylum Alphaproteobacteria-related OTU165401B gradually increased from 0.19% on day 1 to 0.71% on day 167.
PBSA film-degrading potential of fungal strains
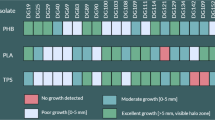
All tested fungal strains of Arthrobotrys and Tetracladium were able to form the clear zone on the emulsified PBSA on the agar plate (Fig. S3). In the case of the solid-state PBSA film, five of the Arthrobotrys strains, A. oligospora MAFF425031, Arthrobotrys sp. MAFF243669, MAFF243672, MAFF243670, and MAFF243671 decomposed the film on the agar plate (Fig. S4). The other two Arthrobotrys strains and T. setigerum MAFF425374 did not.
Discussion
In this study, we confirmed the effectiveness of pretreatment with PaE in promoting the degradation and mineralization of PBSA films to CO2 in cultivated soil. Our data indicated that PaE not only substitutes for esterases from initial degraders, but also stimulates esterase production by indigenous soil microbes. In addition, our next-generation sequence analyses elucidated the microbe genera that were involved in the sequential degradation of partially degraded PBSA films by PaE and new PBSA film.
In soil with PBSA-E, most of the PBSA pieces disappeared from day 1 after the start of incubation (Fig. 1) and were rapidly degraded to monomers and oligomers by PaE, as previously reported (Sato et al. 2017). Succinate and butanediol monomers showed fast mineralization (within a few hours) by microbial uptake and utilization (Nelson et al. 2022). In this study, the CO2 production peaked during the first 7 days, accompanied by enhanced fungal and bacterial abundance. These results indicate that PaE promoted PBSA degradation to CO2 or microbial biomass. The CO2 production rates then decreased and maintained higher values than those in the soil with PBSA-inact E until day 55, followed by a slower rate of release (Fig. 2a). This suggests that the organic carbon that soil microbes could assimilate was mostly consumed at this point. The addition of PaE with PE to soil showed little or undetectable effects on soil respiratory activity because PE was not degradable in soil, even after treatment with PaE, and the added PaE was negligible as a carbon source.
Our results indicated that this enhancement of PBSA degradation by PaE depended on the sequential cooperation of the added PaE and esterase-like enzymes from indigenous soil microbes, based on the following observations: (1) pNP-C5 and pNP-C12 hydrolytic activities showed high and similar values in soils with PBSA-E and PE-E one day after the start of incubation. (2) While these activities maintained higher values in soil with PBSA-E, they quickly decreased to the control level in soil with PE-E at day 55 (Fig. 3). (3) Fungal and bacterial abundance increased in soil with PBSA-E and showed higher values than in soils with other treatments during PBSA degradation. Shackle et al. (2006) advocated a “pump-priming mechanism” in which the supplemented enzyme can act to induce further enzyme production by microorganisms via increased degradation of the corresponding substrate. In the present study, PaE may also act as a driver of this mechanism underlying the enhanced degradation of PBSA films in soil.
In soils with PBSA-E, class Actinobacteria, genus Arthrobacter-related OTUs of bacteria, and class Sordariomycetes, genus Fusarium-related OTUs of fungi, were dominant for the first 7 days (Fig. 6). During this period, the data of CO2 production and soil esterase activities showed that the degradation and mineralization of PBSA proceeded intensely. The quick domination of these microorganisms may be attributed to their preference for nutrient-rich environments (Leplat et al. 2013; Bazhanov et al. 2017). One of the Arthrobacter strains was reported to possess an esterase gene (Nishizawa et al. 1995), and this genus was reported to be a potential degrader of PBSA in soils (Bandopadhyay et al. 2020). The genus Arthrobacter colonized biodegradable mulch films based on poly(butylene terephthalate-co-adipate) (PBAT) in soil, which is a biodegradable polyester (Bandopadhyay et al. 2020). Cutinase from F. solani can hydrolyze PBS films (Hu et al. 2016), and domination of F. solani in soils with buried PBSA films has been reported previously (Yamamoto-Tamura et al. 2020; Purahong et al. 2021; Tanunchai et al. 2021). These factors were considered to contribute to the degradation of not only PBSA degradation products by PaE, but also PBSA polymers. Bacterial genus Azotobacter-related OTUs then dominated and sustained high relative abundance during the incubation from day 14 to 55 (Fig. 6); in this period, CO2 showed a second peak, and pNP-C5 hydrolytic activity remained high. The bacterial genus Azotobacter has been found in soils containing buried PBSA films (Tanunchai et al. 2021, 2022). This genus is known as producers and degraders of polyhydroxyalkanoate, a biodegradable polyester (Page 1992) although its potential for degrading PBSA remains to be determined as far as we know. Additionally, it has been proposed that diazotrophic bacteria, including genus Azotobacter, can enhance PBSA degradation by increasing fungal abundance and activity, facilitated by nitrogen fixation (Tanunchai et al. 2022). As genus Azotobacter is a renowned diazotroph, it likely plays a crucial role in providing fixed nitrogen to soil with PBSA-E, where rapid degradation of PBSA films may deplete nitrogen. These suggests that genus Azotobacter might contribute to the degradation of PBSA-E in soil. However, further research is essential to elucidate the specific ecological roles of genus Azotobacter in soils containing PBSA-E.
The PBSA degradation without PaE required time for the induction and the increase of degraders in soil. Our results indicated that the fungal genera Arthrobotrys and Tetracladium played essential roles in the early stages of PBSA film degradation in soil. In this study, according to the increase in their relative abundance, pNP-C12 and pNP-C5 hydrolytic activity increased in PBSA-inact E. In the plate culture analyses, the genus Arthrobotrys had a solid PBSA film-degradation potential (Fig. S4a, d–g (Supplementary information)). This result suggests that the genus Arthrobotrys attacks the PBSA film initially. Arthrobotrys is reported to show a high abundance in the plastisphere of BP mulch films (Qi et al. 2022) and is well known as a nematode-capturing fungi (Nordbring-Hertz et al. 2011). The nematode epidermis is covered with a cuticle layer, whose structure is composed of ester compounds like BP (Chisholm and Xu 2012). The genus Tetracladium, which dominated, followed by the genus Arthorobotrys, seemed to play a role in the next step of PBSA degradation. Purahong et al. reported that Tetracladium spp. are important PBSA colonizers and potential PBSA decomposers (Purahong et al. 2021), whose genomes are enriched for esterase and pectate lyase domains (Anderson and Marvanová 2020). A strain belonging to the genus Tetracladium examined in this study showed a clear zone on the PBSA emulsion-containing culture plate (Fig. S3h (Supplementary information)), even though this strain did not degrade solid PBSA films. PBSA emulsions are small PBSA particles dispersed in water. One to two percent of strains in PBSA emulsion-degrading bacteria isolated from soil degrade solid biodegradable films (Kitamoto et al. 2011). In soil, after PBSA films are partially degraded into a form that disperses in water, PBSA polymer chains are depolymerized by more types of soil bacteria or fungi, such as Tetracladium. Because these genera were less abundant in soil with PBSA-E, PaE was considered to contribute to the degradation of PBSA films on behalf of these genera.
Conclusion
Enzymatic treatment with PaE enhanced the degradation and mineralization of biodegradable plastic in soil through the collaborative action of PaE and the soil microbial community. This approach offers promising potential to address agricultural and environmental concerns regarding the persistence of BPs in soil. Given the heterogeneity, complexity, and variability of soil, further experiments using different soil types or field trials should be performed in the future. This study identified candidate microbial degraders of PBSA film and observed their sequential succession during PBSA degradation in soil. Interestingly, these microbial groups appear to be commonly involved in the degradation of biodegradable polyesters, even across different geographical locations in various studies. Our findings open up new avenues for research, such as exploring the capabilities of these microbes in artificially synthesized and simplified microbial communities—known as the synthetic community (SynCom) approach (Vorholt et al. 2017). This approach can help clarify the direct contributions of these microbes to the degradation process. Strengthening our understanding of these mechanisms will be vital in developing more effective and targeted strategies for biodegradable plastic waste management.
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Anderson JL, Marvanová L (2020) Broad geographical and ecological diversity from similar genomic toolkits in the ascomycete genus Tetracladium. BioRxiv. https://doi.org/10.1101/2020.04.06.027920
Aronesty E (2013) Comparison of sequencing utility programs. Open Bioinforma J 7:1–8. https://doi.org/10.2174/1875036201307010001
Bache BW (1984) The role of calcium in buffering soils. Plant Cell Environ 7:391–395. https://doi.org/10.1111/j.1365-3040.1984.tb01428.x
Bandopadhyay S, Liquet Y, González JE, Henderson KB, Anunciado MB, Hayes DG, DeBruyn JM (2020) Soil microbial communities associated with biodegradable plastic mulch films. Front Microbiol 11:587074. https://doi.org/10.3389/fmicb.2020.587074
Barnard RL, Osborne CA, Firestone MK (2013) Responses of soil bacterial and fungal communities to extreme desiccation and rewetting. ISME J 7:2229–2241. https://doi.org/10.1038/ismej.2013.104
Bazhanov DP, Yang K, Li H, Li C, Li J, Chen X, Yang H (2017) Colonization of plant roots and enhanced atrazine degradation by a strain of Arthrobacter ureafaciens. Appl Microbiol Biotechnol 101:6809–6820. https://doi.org/10.1007/s00253-017-8405-3
Bekku Y, Koizumi H, Nakadai T, Iwaki H (1995) Measurement of soil respiration using closed chamber method: an IRGA technique. Ecol Res 10:369–373. https://doi.org/10.1007/BF02347863
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Brodhagen M, Peyron M, Miles C, Inglis DA (2015) Biodegradable plastic agricultural mulches and key features of microbial degradation. Appl Microbiol Biotechnol 99:1039–1056. https://doi.org/10.1007/s00253-014-6267-5
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N, Owens SM, Betley J, Fraser L, Bauer M, Gormley N, Gilbert JA, Smith G, Knight R (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624. https://doi.org/10.1038/ismej.2012.8
Chinaglia S, Tosin M, Degli-Innocenti F (2018) Biodegradation rate of biodegradable plastics at molecular level. Polym Degrad Stab 147:237–244. https://doi.org/10.1016/j.polymdegradstab.2017.12.011
Chisholm AD, Xu S (2012) The Caenorhabditis elegans epidermis as a model skin. II: differentiation and physiological roles. Wiley Interdiscip Rev Dev Biol 1:879–902. https://doi.org/10.1002/wdev.77
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Francioni M, Kishimoto-Mo AW, Tsuboi S, Hoshino YT (2022) Evaluation of the mulch films biodegradation in soil: a methodological review. Ital J Agron 17:1936. https://doi.org/10.4081/ija.2021.1936
Guerrini S, Borreani G, Voojis H (2017) Biodegradable materials in agriculture: case histories and perspectives. In: Malinconico M (ed) Soil degradable bioplastics for a sustainable modern agriculture. Springer, Berlin, Heidelberg, pp 35–65. https://doi.org/10.1007/978-3-662-54130-2_3
Hu X, Gao Z, Wang Z, Su T, Yang L, Li P (2016) Enzymatic degradation of poly(butylene succinate) by cutinase cloned from Fusarium solani. Polym Degrad Stab 134:211–219. https://doi.org/10.1016/j.polymdegradstab.2016.10.012
Ikeda S, Tsurumaru H, Wakai S, Noritake C, Fujishiro K, Akasaka M, Ando K (2008) Evaluation of the effects of different additives in improving the DNA extraction yield and quality from andosol. Microbes Environ 23:159–166. https://doi.org/10.1264/jsme2.23.159
Illumina (2013) 16S Metagenomic sequencing library: preparation preparing 16S ribosomal RNA gene amplicons for the Illumina MiSeq system. https://support.illumina.com/documents/documentation/chemistry_documentation/16s/16s-metagenomic-library-prep-guide-15044223-b.pdf. Accessed 1 Mar 2019
Keiblinger KM, Schneider M, Gorfer M, Paumann M, Deltedesco E, Berger H, Jöchlinger L, Mentler A, Zechmeister-Boltenstern S, Soja G, Zehetner F (2018) Assessment of Cu applications in two contrasting soils-effects on soil microbial activity and the fungal community structure. Ecotoxicology 27:217–233. https://doi.org/10.1007/s10646-017-1888-y
Kitamoto HK, Shinozaki Y, Cao XH, Morita T, Konishi M, Tago K, Kajiwara H, Koitabashi M, Yoshida S, Watanabe T, Sameshima-Yamashita Y, Nakajima-Kambe T, Tsushima S (2011) Phyllosphere yeasts rapidly break down biodegradable plastics. AMB Express 1:44. https://doi.org/10.1186/2191-0855-1-44
Kitamoto H, Yoshida S, Koitabashi M, Yamamoto-Tamura K, Ueda H, Yarimizu T, Sameshima-Yamashita Y (2018) Enzymatic degradation of poly-butylene succinate-co-adipate film in rice husks by yeast Pseudozyma antarctica in indoor conditions. J Biosci Bioeng 125:199–204. https://doi.org/10.1016/j.jbiosc.2017.08.017
Kitamoto H, Koitabashi M, Sameshima-Yamashita Y, Ueda H, Takeuchi A, Watanabe T, Sato S, Saika A, Fukuoka T (2023) Accelerated degradation of plastic products via yeast enzyme treatment. Sci Rep 13:2386. https://doi.org/10.1038/s41598-023-29414-1
Koitabashi M, Noguchi MT, Sameshima-Yamashita Y, Hiradate S, Suzuki K, Yoshida S, Watanabe T, Shinozaki Y, Tsushima S, Kitamoto HK (2012) Degradation of biodegradable plastic mulch films in soil environment by phylloplane fungi isolated from gramineous plants. AMB Express 2:40. https://doi.org/10.1186/2191-0855-2-40
Kõljalg U, Nilsson RH, Abarenkov K, Tedersoo L, Taylor AFS, Bahram M, Bates ST, Bruns TD, Bengtsson-Palme J, Callaghan TM, Douglas B, Drenkhan T, Eberhardt U, Dueñas M, Grebenc T, Griffith GW, Hartmann M, Kirk PM, Kohout P, Larsson E, Lindahl BD, Lücking R, Martín MP, Matheny PB, Nguyen NH, Niskanen T, Oja J, Peay KG, Peintner U, Peterson M, Põldmaa K, Saag L, Saar I, Schüßler A, Scott JA, Senés C, Smith ME, Suija A, Taylor DL, Telleria MT, Weiss M, Larsson K-H (2013) Towards a unified paradigm for sequence-based identification of fungi. Mol Ecol 22:5271–5277. https://doi.org/10.1111/mec.12481
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. John Wiley & Sons Ltd., New York, pp 115–175
Leplat J, Friberg H, Abid M, Steinberg C (2013) Survival of Fusarium graminearum, the causal agent of Fusarium head blight. A review. Agron Sustain Dev 33:97–111. https://doi.org/10.1007/s13593-012-0098-5
Li C, Moore-Kucera J, Miles C, Leonas K, Lee J, Corbin A, Inglis D (2014) Degradation of potentially biodegradable plastic mulch films at three diverse US locations. Agroecol Sustain Food Syst 38:861–889. https://doi.org/10.1080/21683565.2014.884515
Lundberg DS, Yourstone S, Mieczkowski P, Jones CD, Dangl JL (2013) Practical innovations for high-throughput amplicon sequencing. Nat Methods 10:999–1002. https://doi.org/10.1038/nmeth.2634
Marchesi JR, Sato T, Weightman AJ, Martin TA, Fry JC, Hiom SJ, Dymock D, Wade WG (1998) Design and evaluation of useful bacterium-specific PCR primers that amplify genes coding for bacterial 16S rRNA. Appl Environ Microbiol 64:795–799. https://doi.org/10.1128/AEM.64.2.795-799.1998
Martin M (2014) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17:10–12. https://doi.org/10.14806/ej.17.1.200
Martín-Closas L, Costa J, Cirujeda A, Aibar J, Zaragoza C, Pardo A, Suso ML, Moreno MM, Moreno C, Lahoz I, Mácua JI, Pelacho AM (2016) Above-soil and in-soil degradation of oxo- and bio-degradable mulches: a qualitative approach. Soil Res 54:225–236. https://doi.org/10.1071/SR15133
Miles C, DeVetter L, Ghimire S, Hayes DG (2017) Suitability of biodegradable plastic mulches for organic and sustainable agricultural production systems. Hortscience 52:10–15. https://doi.org/10.21273/HORTSCI11249-16
Mormile P, Stahl N, Malinconico M (2017) The world of plasticulture. In: Malinconico M (ed) Soil degradable bioplastics for a sustainable modern agriculture. Springer, Berlin, Heidelberg, pp 1–21. https://doi.org/10.1007/978-3-662-54130-2_1
Nelson TF, Baumgartner R, Jaggi M, Bernasconi SM, Battagliarin G, Sinkel C, Künkel A, Kohler HE, McNeill K, Sander M (2022) Biodegradation of poly(butylene succinate) in soil laboratory incubations assessed by stable carbon isotope labelling. Nat Commun 13:5691. https://doi.org/10.1038/s41467-022-33064-8
Nishizawa M, Shimizu M, Ohkawa H, Kanaoka M (1995) Stereoselective production of (+)-trans-chrysanthemic acid by a microbial esterase: cloning, nucleotide sequence, and overexpression of the esterase gene of Arthrobacter globiformis in Escherichia coli. Appl Environ Microbiol 61:3208–3215. https://doi.org/10.1128/aem.61.9.3208-3215.1995
Nordbring-Hertz B, Jansson H-B, Tunlid A (2011) Nematophagous fungi. In: eLS (ed). https://doi.org/10.1002/9780470015902.a0000374.pub3
Page WJ (1992) Production of polyhydroxyalkanoates by Azotobacter vinelandii UWD in beet molasses culture. FEMS Microbiol Lett 9:149–157. https://doi.org/10.1111/j.1574-6968.1992.tb05832.x
Purahong W, Wahdan SFM, Heinz D, Jariyavidyanont K, Sungkapreecha C, Tanunchai B, Sansupa C, Sadubsarn D, Alaneed R, Heintz-Buschart A, Schädler M, Geissler A, Kressler J, Buscot F (2021) Back to the future: decomposability of a biobased and biodegradable plastic in field soil environments and its microbiome under ambient and future climates. Environ Sci Technol 55:12337–12351. https://doi.org/10.1021/acs.est.1c02695
Qi Y, Ossowicki A, Yergeau É, Vigani G, Geissen V, Garbeva P (2022) Plastic mulch film residues in agriculture: impact on soil suppressiveness, plant growth, and microbial communities. FEMS Microbiol Ecol 98:1–10. https://doi.org/10.1093/femsec/fiac017
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596. https://doi.org/10.1093/nar/gks1219
R Development Core Team (2018) R: a language and environment for statistical computing. https://www.R-project.org/. Accessed 1 Mar 2019
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Sakai Y, Isokawa M, Masuda T, Yoshioka H, Hayatsu M, Hayano K (2002) Usefulness of soil p-nitrophenyl acetate esterase activity as a tool to monitor biodegradation of polybutylene succinate (PBS) in cultivated soil. Polym J 34:767–774. https://doi.org/10.1295/polymj.34.767
Sameshima-Yamashita Y, Ueda H, Koitabashi M, Kitamoto H (2019a) Pretreatment with an esterase from the yeast Pseudozyma antarctica accelerates biodegradation of plastic mulch film in soil under laboratory conditions. J Biosci Bioeng 127:93–98. https://doi.org/10.1016/j.jbiosc.2018.06.011
Sameshima-Yamashita Y, Watanabe T, Tanaka T, Tsuboi S, Yarimizu T, Morita T, Koike H, Suzuki K, Kitamoto H (2019b) Construction of a Pseudozyma antarctica strain without foreign DNA sequences (self-cloning strain) for high yield production of a biodegradable plastic-degrading enzyme. Biosci Biotechnol Biochem 83:1547–1556. https://doi.org/10.1080/09168451.2019.1571898
Sato S, Saika A, Shinozaki Y, Watanabe T, Suzuki K, Sameshima-Yamashita Y, Fukuoka T, Habe H, Morita T, Kitamoto H (2017) Degradation profiles of biodegradable plastic films by biodegradable plastic-degrading enzymes from the yeast Pseudozyma antarctica and the fungus Paraphoma sp. B47–9. Polym Degrad Stab 141:26–32. https://doi.org/10.1016/j.polymdegradstab.2017.05.007
Shackle V, Freeman C, Reynolds B (2006) Exogenous enzyme supplements to promote treatment efficiency in constructed wetlands. Sci Total Environ 361:18–24. https://doi.org/10.1016/j.scitotenv.2005.09.032
Shinozaki Y, Kikkawa Y, Sato S, Fukuoka T, Watanabe T, Yoshida S, Nakajima-Kambe T, Kitamoto HK (2013a) Enzymatic degradation of polyester films by a cutinase-like enzyme from Pseudozyma antarctica: surface plasmon resonance and atomic force microscopy study. Appl Microbiol Biotechnol 97:8591–8598. https://doi.org/10.1007/s00253-012-4673-0
Shinozaki Y, Morita T, Cao XH, Yoshida S, Koitabashi M, Watanabe T, Suzuki K, Sameshima-Yamashita Y, Nakajima-Kambe T, Fujii T, Kitamoto HK (2013b) Biodegradable plastic-degrading enzyme from Pseudozyma antarctica: cloning, sequencing, and characterization. Appl Microbiol Biotechnol 97:2951–2959. https://doi.org/10.1007/s00253-012-4188-8
Sintim HY, Bary AI, Hayes DG, Wadsworth LC, Anunciado MB, English ME, Bandopadhyay S, Schaeffer SM, DeBruyn JM, Miles CA, Reganold JP, Flury M (2020) In situ degradation of biodegradable plastic mulch films in compost and agricultural soils. Sci Tot Environ 727:138668. https://doi.org/10.1016/j.scitotenv.2020.138668
Takada-Hoshino Y, Matsumoto N (2004) An improved DNA extraction method using skim milk from soils that strongly adsorb DNA. Microbes Environ 19:13–19. https://doi.org/10.1264/jsme2.19.13
Tanunchai B, Juncheed K, Wahdan SFM, Guliyev V, Udovenko M, Lehnert AS, Alves EG, Glaser B, Noll M, Buscot F, Blagodatskaya E, Purahong W (2021) Analysis of microbial populations in plastic-soil systems after exposure to high poly(butylene succinate-co-adipate) load using high-resolution molecular technique. Environ Sci Eur 33:105. https://doi.org/10.1186/s12302-021-00528-5
Tanunchai B, Kalkhof S, Guliyev V, Wahdan SFM, Krstic D, Schädler M, Geissler A, Glaser B, Buscot F, Blagodatskaya E, Noll M, Purahong W (2022) Nitrogen fixing bacteria facilitate microbial biodegradation of a bio-based and biodegradable plastic in soils under ambient and future climatic conditions. Environ Sci Process Impacts 24:233–241. https://doi.org/10.1039/d1em00426c
Tsuboi S, Tanaka T, Yamamoto-Tamura K, Kitamoto H (2018) High-throughput method for the evaluation of esterase activity in soils. J Microbiol Met 146:22–24. https://doi.org/10.1016/j.mimet.2018.01.009
Tsuboi S, Yamamoto-Tamura K, Takada A, Yonemura S, Hoshino YT, Kitamoto H, Kishimoto-Mo AW (2022) Selection of p-nitrophenyl fatty acid substrate suitable for detecting changes in soil esterase activity associated with degradation of biodegradable polyester mulch films: a field trial. Ital J Agron 17:3. https://doi.org/10.4081/ija.2022.2040
Ueda H, Mitsuhara I, Tabata J, Kugimiya S, Watanabe T, Suzuki K, Yoshida S, Kitamoto H (2015) Extracellular esterases of phylloplane yeast Pseudozyma antarctica induce defect on cuticle layer structure and water-holding ability of plant leaves. Appl Microbiol Biotechnol 99:6405–6415. https://doi.org/10.1007/s00253-015-6523-3
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172:4238–4246. https://doi.org/10.1128/jb.172.8.4238-4246.1990
Vorholt JA, Vogel C, Carlström CI, Müller DB (2017) Establishing causality: opportunities of synthetic communities for plant microbiome research. Cell Host Microbe 22:142–155. https://doi.org/10.1016/j.chom.2017.07.004
Wagai R, Kishimoto-Mo AW, Yonemura S, Shirato Y, Hiradate S, Yagasaki Y (2013) Linking temperature sensitivity of soil organic matter decomposition to its molecular structure, accessibility, and microbial physiology. Glob Chang Biol 19:1114–1125. https://doi.org/10.1111/gcb.12112
Watanabe T, Morita T, Koike H, Yarimizu T, Shinozaki Y, Sameshima-Yamashita Y, Yoshida S, Koitabashi M, Kitamoto H (2016) High-level recombinant protein production by the basidiomycetous yeast Pseudozyma antarctica under a xylose-inducible xylanase promoter. Appl Microbiol Biotechnol 100:3207–3217. https://doi.org/10.1007/s00253-015-7232-7
White TJ, Bruns ST, Lee SF, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Yamamoto-Tamura K, Hoshino YT, Tsuboi S, Huang C, Kishimoto-Mo AW, Sameshima-Yamashita Y, Kitamoto H (2020) Fungal community dynamics during degradation of poly(butylene succinate-co-adipate) film in two cultivated soils in Japan. Biosci Biotechnol Biochem 84:1077–1087. https://doi.org/10.1080/09168451.2020.1713718
Acknowledgements
The authors thank Showa Denko K. K. and UNYCK for providing the PBSA emulsion and films, as well as the staff of the Institute for Agro-environmental Sciences on National Agriculture and Food Research Organization for the experimental preparation.
Funding
This work was supported by the Science and Technology Research Promotion Program for Agriculture, Forestry, Fisheries, and Food Industry (grant number: 25017AB), as well as the research program on the development of innovative technology grants from the Project (JPJ007097) of the Bio-oriented Technology Research Advancement Institution (BRAIN).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Ayaka W. Kishimoto-Mo, Yuko Takada Hoshino, and Kimiko Yamamoto-Tamura designed and conducted the incubation experiments. Shun Tsuboi collected the data from microbiological analyses and analyzed all the data. Yuka Sameshima-Yamashita and Hiroko Kitamoto conducted the preparation and treatment of PaE. Natsuki Omae, Tomotake Morita, and Hirohide Uenishi conducted the next generation sequencing and data analyses. The first draft of the manuscript was written by Shun Tsuboi, Yuko Takada Hoshino, and Kimiko Yamamoto-Tamura, and all authors commented on previous versions of the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tsuboi, S., Hoshino, Y.T., Yamamoto-Tamura, K. et al. Enhanced biodegradable polyester film degradation in soil by sequential cooperation of yeast-derived esterase and microbial community. Environ Sci Pollut Res 31, 13941–13953 (2024). https://doi.org/10.1007/s11356-024-31994-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-024-31994-y