Abstract
Substrates like sand or gravels and aquatic nutrient concentrations of rivers are highly heterogeneous, influencing the abundance of functional genes in epilithic biofilms where nitrification–denitrification processes take place. To analyze how the relative abundance of nitrifying/denitrifying genes and the associated microbes changes with the physical properties of substrates and aquatic concentrations of nutrients, this paper utilized metagenomics to comprehensively characterize these functional genes (i.e., amoA, hao, and nxrB involved in nitrification, and napA, narG, nirS, norB, and nosZ associated with denitrification) from epilithic biofilms collected along the Shitingjiang River in Southwest China and further obtained the relative abundance of major nitrifiers and denitrifiers. The results show that substrate size most significantly affects the relative abundance of hao and norB by altering the hydrodynamic conditions. In sampling sites with high heterogeneity in substrate size distribution, the relative abundance of most denitrifying genes is also higher. The carbon–nitrogen ratio negatively correlates with the relative abundance of all the nitrifying genes, while ammonium, total inorganic carbon, and total organic carbon concentrations positively affect the relative abundance of amoA and nxrB. As to the relative abundance of nitrifiers and denitrifiers, mainly belonging to phyla Proteobacteria and Actinobacteria, substrate heterogeneity and the aquatic concentrations of nutrients have greater influences than substrate size. Also, the substrate heterogeneity exerted positive influence on functional species of Pseudogemmobacter bohemicus and Paracoccus zhejiangensis. Considering the genes’ functions and the dominant species linked to denitrification, nitrous oxide is more likely to occur in rivers with higher heterogeneity and larger substrates.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Microbes are the major drivers of nitrification and denitrification processes which produce nitrate (NO3−), nitrite (NO2−), nitric oxide (NO), dinitrogen gas (N2), and nitrous oxide (N2O), resulting in geochemical and ecological consequences such as eutrophication and global warming (Magalhães et al. 2005). In stream ecosystems, complex microbial assemblages attaching to river substrates like sand or gravels are called epilithic biofilms (Lyautey et al. 2003), which are hot spots of nitrification and denitrification process by taking and transferring nutrients and providing aerobic-anaerobic microsites (Besemer 2015; Hanrahan et al. 2018).
Epilithic biofilms are sensitive to anthropogenic activities and geological changes in river ecosystems (Besemer 2015; Lear et al. 2008; Magalhães et al. 2005). As a result, the abundance and composition of nitrifier and denitrifier communities in epilithic biofilms also change with the environment. For example, denitrifier community structure shifts with the variation in nutrient load, especially the nitrogen load (Lyautey et al. 2013); higher abundance of Proteobacteria was observed as a response to higher nutrient concentrations (Qu et al. 2017); ammonia-oxidizing bacteria dominate the epilithic biofilms when ammonium concentration is high (Liu et al. 2018a); and water warming changes the abundance of denitrifiers to modulate dissolved nitrogen removal indirectly (Boulêtreau et al. 2014; Lear et al. 2008).
Due to the close attachment between nitrifier/denitrifier communities and environmental changes, the abundance and composition of these micro-organisms in epilithic biofilms may act as indicators of nitrification and denitrification processes in rivers. However, the functions of microbial communities are often decoupled from their taxonomic composition. Therefore, the role of microbial functional genes in nitrogen cycle should be emphasized (Wang et al. 2022). In recent years, metagenomics enables the identification of all known nitrogen-cycling genes changing with the environment. A series of studies based on metagenomics have shown that the hydrological and physicochemical factors exert profound impacts on the abundance of functional genes. For instance, Ren et al. (2017) found that glaciated area proportion, runoff proportion, and distance to glaciers affect the relative abundance of nitrifying and denitrifying genes more significantly than physicochemical factors including temperature, dissolved oxygen, ammonium, and nitrate in glacial-fed streams; Palacin-Lizarbe et al. (2019) revealed that the abundance of nirS was higher than nirK in more productive lakes; Vila-Costa et al. (2014) showed that amoA abundance was strongly correlated with nutrient concentrations such as ammonium and nitrate. In addition to nutrient concentrations in river water, the riverbed substrates with different physical properties, as the main attachment places for epilithic biofilms, could also affect the nitrifying and denitrifying genes. However, this factor has not been sufficiently investigated. In addition, it remains to be seen the relative importance between the two factors, i.e., substrate properties and aquatic nutrient concentrations.
In this study, we try to explore how the nitrifying/denitrifying genes (i.e., amoA, hao, nxrB, napA, narG, nirS, norB, and nosZ) in epilithic biofilms correlate with the size and size distribution (heterogeneity) of substrates and aquatic nutrient concentrations in the Shitingjiang River located in the southwest of China and experiencing dramatic natural and anthropogenic changes (Fan et al. 2016; Li et al. 2022). In addition, bacteria potentially carrying the focused genes were also identified. Given that certain genes, such as hao and norB, are involved in the formation of nitrous oxide, a much stronger greenhouse gas than carbon dioxide (Wang et al. 2013), our study also analyzed the nitrogen metabolic pathways and the enzyme in groups with different substrate sizes, heterogeneity, and nutrient concentrations and discussed potential nitrous oxide emissions in rivers associated with substrate physical properties which could be changed by natural incision of riverbed and water conservancy projects.
Materials and methods
Site description
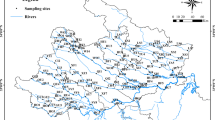
The Shitingjiang River (31° 10′–31° 17′ N, 104° 01′–104° 14′ E) is a 115 km-long tributary of the Mianyuanhe River (Fig. 1), originating from the Longmen Mountain and flowing to the Chengdu Plain in the southwest of China. The watershed area takes up ~ 1600 km2 with an average slope of ~ 3.3‰ (Fan et al. 2016). Shitingjiang River is highly heterogeneous in substrates, with particle size ranging from < 2 to > 256 mm, and the heterogeneity of substrates has been further increased since the Wenchuan earthquake (in 2008) which mobilized coarse particles. Commercial sediment extraction and constructions of water conservancy facilities also affect the heterogeneity of substrates. High nutrient concentrations are observed in the river due to fertilizer abuse and industrial wastewater discharge (Li et al. 2022).
Sketch map of the sampling sites in Shitingjiang river. (a) Location of the studied area in Sichuan Province in the southwest of China. (b) Tributaries of the Tuojiang River, including the Mianyuan River and the Shitingjiang River. (c) Locations of the ten sampling sites along the Shitingjiang River
Sample collection and pretreatment
The sampling was carried out from July 2 to July 10 in 2022 within a 25.8-km reach of the Shitingjiang River. At each sampling site, we randomly sampled submerged substrates from the river at a depth between 10 and 30 cm and divided the grain size into four groups, i.e., < 2 mm (D1), 2–40 mm (D2), 40–100 mm (D3), and 100–200 mm (D4). Then, epilithic biofilms were removed by scrapers from the upper surface of each substrate and put into pre-sterilized polyethylene tubes. For epipsammic biofilms (i.e., biofilm from particles smaller than 2 mm), the bead-beating procedure is an effective method. However, due to the constraints of time and environment for in situ monitoring, a rubber-tipped dropper was used to collect epipsammic biofilms after consulting with Majorbio Bio-Pharm Technology Co., Ltd (Shanghai, China). The procedure was as follows: (1) find the < 2-mm particles that were adjacent to the places where we got substrates of the larger-sized groups. To keep the epipsammic samples in consistent with the epilithic ones, only the epipsammic biofilms on the upper side of the < 2-mm particles were taken. Note that < 2-mm particles usually form a small continuous area, and the epipsammic biofilms growing after the particle deposits are also continuous. (2) A rubber-tipped dropper was used to suck up the epipsammic biofilms on the surface of the < 2-mm particles. (3) By extruding the water that had been sucked in with the epipsammic biofilms, the sample was placed in a pre-sterilized polyethylene tube. Note that only a very small amount of biofilms can be sucked up with every release, so Steps (2) and (3) needed to be repeated many times until 10 ml of sample was collected at one site.
The water samples were also collected from 0.2 m underwater using pre-sterilized bottles. Then, all the water and biofilm samples were immediately stored in thermos cabinet with ice bag and frozen at − 20 °C within 10 h to prevent the degradation of genes (Silva et al. 2021) for physiochemical and metagenomics analysis.
Meanwhile, the Wolman sampling method was used to estimate the grain size ranging from 2 to 256 mm on the bed of the stream. The Wolman method includes three steps: (1) establishing a grided area using step length (0.3–0.6 m) at the sampling site, (2) selecting one gravel randomly in each grid along the designed route and measure the length of its central axis, and (3) repeating the previous step until 100 gravels are measured (Bunte and Abt 2001; Galia et al. 2017; Wolman 1954).
Chemical analyses
Physicochemical properties of the overlying water samples were characterized including nitrate (NO3−), nitrite (NO2−), ammonium (NH+), total nitrogen (TN), total phosphorus (TP), chemical oxygen demand (COD), total inorganic carbon (TIC), total carbon (TC), and total organic carbon (TOC). The NO3− and NO2− concentrations were determined by UV spectrophotometer (SP-722E, Spectrum Shanghai, China). NO3− concentration was determined with potassium nitrate and hydrochloric acid at the peak absorption of 220 nm and 275 nm. The determination of NO2− concentration was conducted using N-1-naphthalene ethylenediamine based on National Standards (GB 7493–87) (Zhang et al. 2022). And the NH+ concentration was measured using the Nessler Reagent Spectrophotometry method (Zhang et al. 2022). The determinations of TN and TP were conducted using alkaline potassium persulfate digestion UV spectrophotometry (Hao 2014) and ammonium molybdate spectrophotometric method (Li et al. 2012), respectively. TOC, TIC, and TC concentrations were analyzed on a vario TOC SELECT (elementar-Analysensysteme GmbH, Germany) (Qu et al. 2017). The dichromate method was used for the COD measurement (Anderson et al. 2007).
DNA extraction and metagenomic sequencing
Genomic DNA from epilithic biofilm samples of Shitingjiang River was extracted using E.Z.N.A® Soil DNA Kit (Omega Bio-tek, Norcross, GA, USA) and then detected by 1% agarose gel electrophoresis. Analyses of DNA purity and concentration were performed using NanoDrop 2000 spectrophotometer and TBS-380, respectively. Paired-end library was constructed using NEXTflex Rapid DNA-Seq (Bioo Scientific, Austin, TX, USA) based on random fragments of extracted DNA with an average size of about 400 bp (Covaris M220, Gene Company Limited, China), and then paired-end sequencing was performed on Illumina NovaSeq 6000 (Illumina Inc., NovaSeq Reagent Kits) at Majorbio Bio-Pharm Technology Co., Ltd. (Shanghai, China) according to the manufacturer’s instructions (www.illumina.com). Paired-end reads were trimmed and improved (length < 50 bp or with a quality value < 20 or having N base) by fastp (http://github.com/OpenGene/fastp, version 0.20.0) (Chen et al. 2018) on the Majorbio Cloud Platform (www.majorbio.com). Metagenomic data were assembled using MEGAHIT (http://github.com/voutcn/megahit, version 1.1.2) and filter out contigs with a length < 300 bp (Li et al. 2015). ORFs (Open Reading Frames) which are essential to discover specific protein-encoding genes were predicted using Prodigal (Hyatt et al. 2010). A non-redundant gene catalog was obtained using CD-HIT (http://www.bioinformatics.org/cd-hit/, version 4.6.1) (Fu et al. 2012) based on predicted ORFs with 90% identity and 90% coverage. The relative abundance of genes were calculated by aligning high-quality reads (95% identity) to non-redundant gene catalogs using SOAPaligner (http://soap.genomics.org.cn/, version 2.21) (Li et al. 2008).
Genes and microbial taxonomy annotation
The genes involved in nitrification (i.e., amoA, hao, nxrB) and denitrification (i.e., napA, narG, nirS, norB, and nosZ) were identified against the Kyoto Encyclopedia of Genes and Genomes (KEGG) database via Diamond (http://www.genome.jp/keeg/, version 94.2) (Fig. 2). Microbial composition assigned with aforementioned genes was characterized against NR database (https://ftp.ncbi.nlm.nih.gov/blast/db/FASTA/, nr_20200604) with an e-value cutoff of 1e−5 using Diamond (http://www.diamondsearch.org/index.php, version 0.8.35) (Buchfink et al. 2015).
Statistical methods
The data were analyzed on the online platform of Majorbio Cloud Platform (www.majorbio.com) and SPSS (version 26.0). The Skewness and Kurtosis tests were taken to test whether the relative abundance of functional genes followed normal distribution (SPSS version 26.0). Then significant difference (P < 0.05) among different size and heterogeneity of gravels was assessed using Kruskal–Wallis test (SPSS version 26.0) and Paired-Samples T test (SPSS version 26.0) (Xu et al. 2017). Significance of the relative abundance of genes involved in nitrification and denitrification among different substrate heterogeneity was calculated by permutational multivariate analysis of variance (PERMANOVA) (vegan package 2.4 in R 3.3.2) (P < 0.05). Analysis of unique and shared nitrifiers/denitrifiers of epilithic biofilms among groups with different substrate sizes were conducted using the VennDiagram package (R 3.3.2). The microbial community dissimilarities of nitrifiers/denitrifiers were examined using non-metric multidimensional scaling based on a Bray–Curtis similarity matrix (Kobayashi et al. 2009; Qu et al. 2017). After calculating the first axis of the lengths of gradient (based on the analysis result of Detrended Correspondence Analysis) (Griffith et al. 2001), we chose redundancy analysis (RDA) to reveal the correlation of nitrifying/denitrifying genes and their associated microbes with aquatic nutrients using Bray–Curtis distance matrix (vegan package 2.4 in R 3.3.2) (Qu et al. 2017; Ren et al. 2017). The relative abundance of functional gene proportions among different heterogeneity groups was visualized using Circos (Circos 0.67–7). Spearman analysis was conducted to compare the importance of aquatic nutrient concentrations and substrate size heterogeneity to the relative abundance of nitrifying/denitrifying genes and their associated microbes. Only the significant correlations were taken into consideration (P < 0.05).
Results and discussion
At one site, we collected four samples from the four substrate groups (i.e., D1, D2, D3, and D4), respectively, and we got a total of 40 samples at the ten sites. Each sample contained 10-ml biofilms.
Relative abundance of nitrifying/denitrifying genes and the associated microbes among different substrate size groups
The relative abundance of most nitrifying and denitrifying genes that followed the normal distribution shows significant difference among substrate size groups (P < 0.05 in the t test) (Table 1 and Table S1). In the case of hao, its relative abundance was the highest in Group D1, followed by Group D4 (Fig. 3). In contrast, the highest relative abundance of norB was found in the group with the largest size (D4), followed by Group D2 (Fig. 3). Besides, the presence of napA significantly varied among Groups D2, D3, and D4, while nxrB and nirS mostly differed between Groups D2 and D4. Groups D4 and D3 were also distinguished in the relative abundance of nosZ (Fig. 3).
Relative abundance of the nitrifying genes (amoA, hao, nxrB) and the denitrifying genes (napA, narG, nirS, norB, and nosZ) in groups with different substrate sizes. D1, substrate size < 2 mm; D2, substrate size within 2–40 mm; D3, substrate size within 40–100 mm; D4, substrate size within 100–200 mm. *P < 0.05; **P < 0.01; ***P < 0.001
By assigning all the concerned genes to specific bacteria, we found nitrifiers and denitrifiers mainly belonged to phyla Proteobacteria, Actinobacteria, Verrucomicrobia, and Planctomycetes (Fig. 4). In contrast to the functional genes, there was no significant difference in the relative abundance of bacteria at phylum level among the four substrate size groups (P > 0.05) (Table S3). Furthermore, we assigned all the concerned genes to the species level of nitrifiers and denitrifiers. The results showed that groups with different substrate size shared 465 species (74.6–77.8% of the total species number) for nitrifiers and 467 species for denitrifiers (74.4–80.4% of the total) (Fig. S1). The dominant species (i.e., top four species in relative abundance) among four substrate groups showed insignificant difference (Actinobacteria bacterium, Pseudogemmobacter bohemicus, Paracoccus zhejiangensis, Gemmobacter aestuarii for nitrifiers, and Actinobacteria bacterium, Pseudogemmobacter bohemicus, Paracoccus zhejiangensis, Rhodobacter veldkampii for denitrifiers, P > 0.05). The average relative abundance of Actinobacteria bacterium was 7.27%, 4.52%, 6.26%, and 7.72% in Group D1, D2, D3, and D4, respectively (Fig. 4c, d), while that of Pseudogemmobacter bohemicus varied from 4.31 to 5.50%, with the highest relative abundance found in Group D2 (Fig. 4c, d). The highest the relative abundance of Paracoccus zhejiangensis was in Group D2 while lowest was found in Group D3 (Fig. 4c, d). The highest relative abundance of Gemmobacter aestuarii as a nitrifier was observed in Group D2 (Fig. 4c). The lowest relative abundance of Rhodobacter valdkampii as a denitrifier was found in Group D4 (Fig. 4d).
Relative abundance of (a) nitrifiers at the phylum level, (b) denitrifiers at the phylum level, (c) nitrifiers at the species level, and (d) denitrifiers at the species level in groups with different substrate sizes. D1, substrate size < 2 mm; D2, substrate size within 2–40 mm; D3, substrate size within 40–100 mm; D4, substrate size within 100–200 mm
Although Aquabacterium pictum is the species with low relative abundance, the nitrifying and denitrifying genes in this species exhibited significant variation among the four substrate size groups (P < 0.05). The highest mean relative abundance of this species was found in Group D3 (1.33%), followed by Group D2 (1.06%), D4 (0.81%), and D1 (0.55%). The highest relative abundance of Verrucomicrobia bacterium as a nitrifier was observed in Group D1 (1.18%). The lowest relative abundance of Thermomonas spHDW16 as a denitrifier was found in Group D1 (0.22%).
The relationship between the relative abundance of genes and the composition of corresponding bacteria among different substrate size groups was also identified. For example, the substrate size group (i.e., Group D1) with the highest relative abundance of hao had the most various taxonomic compositions at both the genus level (including 50.27% of Nitrosomonas, 36.64% of Nitrospira, and 13.08% of Candidatus Brocadia, associated with genes showing significant difference among size groups) and the species level (including 39.22% of Nitrosomonas sp. ST-bin4, 20.97% of Nitrosomonas sp. Is79A3, and 26.19% of Nitrosomonas sp. Nm141, also associated with genes exhibiting significant difference among size groups). In contrast, in Group D3 where the relative abundance of hao was the lowest, the bacteria assigned from hao mainly belonged to genus Nitrosomonas (Fig. S2) (Merbt et al. 2015; Mußmann et al. 2013; Smith and Oerther 2006). In the case of norB, the highest relative abundance co-occurred the large proportions of genus Tabrizicola and species Tabrizicola sp. DJC in Group D4 (Fig. S2) (Lin et al. 2022; Wu et al. 2020).
Relative abundance of nitrifying/denitrifying genes and the associated microbes among different size heterogeneity groups
The ratio of d84 to d50 (di representing the substrate size larger than the ith percent of substrates in the river) was used to measure the heterogeneity in the distribution of substrate size (Cardinale et al. 2002). Here, we calculate d84/d50 (~ 3.4) of site A1 located in the mountain exit as the standard for median heterogeneity (MH) because the influences of human activities and earthquakes there are very small. Therefore, sampling sites with d84/d50 smaller or larger than 3.4 were identified as low heterogeneity (LH) or high heterogeneity (HH) (Table 2).
The relative abundance of most denitrifying genes (except gene norB) presented significant difference among heterogeneity groups using Kruskal–Wallis H test (P < 0.05) (Fig. S3). Their relative abundance was usually the highest in HH sampling sites. In the case of nosZ, however, the relative abundance was lower in HH sites than in the LH sites (Fig. 5a), while there was no significant difference in the relative abundance of nitrifying genes.
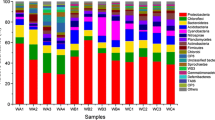
The nitrifiers/denitrifiers taxonomic community also distinguished between HH and LH sampling sites at the phylum level (identified by assigning all the concerned genes, see Fig. S4), as indicated by the results of non-metric multidimensional scaling (NMDS). In terms of relative abundance, the permutational multivariate analysis of variance (PERMANOVA) test based on the Bray–Curtis distance also identified significant difference (P < 0.05) between HH and LH sites in the nitrifiers, which is unlike the nitrifying genes, and in the denitrifiers, which is in agreement with the denitrifying genes. For instance, being the largest phyla of both nitrifiers and denitrifiers, Proteobacteria phylum have higher relative abundance in HH sampling sites (82.0% for HH and 76.9% for LH, P < 0.001), while Actinobacteria phylum have lower relative abundance in HH sampling sites (8.5% for the HH and 11.9% for the LH, P < 0.001) (Fig. S5).
Furthermore, the dominant species (identified as the top four species in relative abundance after assigning genes with significant difference among heterogeneity groups) and their relative abundance of nitrifiers and denitrifiers were analyzed in sampling sites with different heterogeneity. It showed that three species, namely Actinobacteria bacterium, Pseudogemmobacter bohemicus, and Paracoccus zhejiangensis, were dominated in both nitrification and denitrification processes (Fig. 5b), with relative abundance differing significantly between HH and LH sampling sites (Table S4). Hereinto, the relative abundance of species Actinobacteria bacterium was higher in LH sites, while that of species Pseudogemmobacter bohemicus and Paracoccus zhejiangensis was higher in HH sites (Fig. 5b). The higher relative abundance of Gemmobacter aestuarii as a nitrifier was observed in LH sites (Fig. 5b). The lower relative abundance of Rhodobacter valdkampii as a denitrifier was found in LH sites (Fig. 5b).
The co-occurrence of most denitrifying genes and the corresponding denitrifiers was found in different heterogeneity groups. Higher relative abundance of napA co-occurred with higher affiliated genus, Pseudogemmobacter and Paracoccus, in HH sampling sites. Similarly, higher relative abundance of norB and nirS also co-occurred genus Tabrizicola in HH sampling sites (Fig. 6).
Impact of aquatic nutrient concentrations on the relative abundance of nitrifying/denitrifying genes and the associated microbes
The results of redundancy analysis (RDA) showed that the nutrient concentrations were tightly correlated with the relative abundance of nitrifying genes but poorly correlated with that of the denitrifying genes. For example, ammonium (NH4+), total inorganic carbon (TIC), and total organic carbon (TOC) concentrations were positively correlated with the relative abundance of amoA and nxrB, while TIC and TOC concentrations negatively correlated with the relative abundance of hao (P < 0.05). Besides, the relative abundance of all the concerned nitrifying genes was negatively dependent on the total carbon and total nitrogen ratio (CN ratio).
When assigning genes to the bacteria, we found that dominant phyla Proteobacteria, Actinobacteria, Verrucomicrobia, and Planctomycetes were all shared for nitrifiers and denitrifiers in different sampling sites. Moreover, the cluster analysis of the relative abundance of nitrifiers and denitrifiers at phylum level in different sampling sites was visualized using the dendrograms based on Euclidean distance (Fig. 7).
For nitrifiers, sampling sites including B1, B3, E2, and F3 were grouped together; A1, C1, C4, D2, and E2 were clustered in Group 2; and A3 site was in Group 3. In the case of denitrifiers, sites including A1, C1, and A3 were clustered in Group 1; B1, B3, C4, E1, and D2 sites were grouped together; and E2 and F3 sites were in Group 3 (Fig. 8a). However, we found that the relative abundance of nitrifiers and denitrifiers at phylum level in different sampling sites differed significantly (P < 0.05) (Table S5).
a Hierarchical clustering analysis based on the Euclidean distance of the relative abundance of nitrifers/denitrifiers generated from the biofilm samples collected from 10 sampling sites. b, c The RDA plot revealing the association of nitrifying genes/nitrifiers at phylum level and denitrifying genes/denitrifiers at phylum level, respectively, in relation to nutrients concentrations. The length of an arrow represents the degree of correlation between nutrient concentrations and community distribution. The angle between two arrows represents the correlation between nutrients concentrations and nitrifying/denitrifying genes and associated microbes. Acute angle, positive correlation; obtuse angle, negative correlation; right angle, no correlation
Although the relative abundance of bacteria at different sampling sites were distinguished at the phyla level, the heterogeneous aquatic nutrient concentration characteristics can only explain part of the changes in the relative abundance of nitrifiers and denitrifiers, with the total contribution of the first two RDA axes accounted for 30.87% and 27.01% of the variance, respectively. However, TIC concentration and CN ratio were relatively the most influential factors for the relative abundance of nitrifiers and denitrifers (P < 0.05). The relative abundance of Actinobacteria was positively correlated to TIC concentrations and CN ratio, while for Proteobacteria, its relative abundance had negative correlation with CN ratio and positive correlation with TIC concentration (Fig. 8b, c).
Discussion
Reasons for the impact of substrate size
The impact of substrate size on the relative abundance of nitrifying and denitrifying genes can potentially be interpreted by three mechanisms. First, substrate size determines the mobility of the substrates, adjusts the hydrodynamic conditions, and changes availability of the light and nutrients (Arnon et al. 2013), thus affecting the relative abundance of functional genes of the biofilms attached to substrates of different size groups (Fig. 3). Second, the substrate size not only controls biofilm attachment area but also affects the porosity, permeability, and interspace between substrates indirectly (Santmire and Leff 2007). Greater porosity allows higher ingress of fresh water containing oxygen and nutrients through the sediments in some instances, which is important for microbe/substrate reactions and removal of microbial by-products. Third, Suarez et al. (2019) found that the biofilm thickness, closely related to substrate size (Ahmad et al. 2017), played a decisive role in river ecosystem functioning and microbial community compositions. However, we did not measure the thickness of biofilm samples in this study because of the lack of microscopic equipment for mm-level measurement, and we encourage further investigation on the impact of biofilm thickness. However, Santmire and Leff (2007) argued that the effect of substrate size on the bacterial community and abundance is not always evident in field surveys due to the changing environment surround sampling sites. Romaní and Sabater (2001) also found that epilithic biofilms may be less sensitive to the changes of physicochemical parameters in aquatic environments when the composition of autotrophic organisms is complicated and abundant. This explains why some genes in our study showed little difference among size groups nor did the taxonomic compositions or the relative abundance of most nitrifiers and denitrifiers showed significant difference among substrate size groups (Table S3).
Reasons for the impact of substrate heterogeneity in size distribution
Physical habitat heterogeneity of stream ecosystem influences microbial diversity, composition, transportation of active substance, and biogeochemical cycle (Lear et al. 2008; Singer et al. 2010; Keil et al. 2011; Cardinale et al. 2002). In particular, higher heterogeneity of substrate in size distribution reduces the space between particles because smaller particles fill voids created by larger particles. Therefore, the overall porosity is also reduced, prohibiting the transfer of nutrients and oxygen (Singer et al. 2010; Wilson and Dodds 2009). Besides, higher heterogeneity also enhances geomorphological complexity, consequently increases water residence time, and promotes nitrogen removal (Hanrahan et al. 2018). These factors all exert impacts on biofilm microbial communities and finally the ecological process (Besemer 2015; Cardinale et al. 2002). Our results confirmed the role of substrate heterogeneity in affecting the relative abundance by presenting the co-occurrence of high relative abundance of denitrifying genes, microbes, and high heterogeneity (Fig. 5a). For instance, higher relative abundance of species including Pseudogemmobacter bohemicus and Paracoccus zhejiangensis was observed in substrate with high heterogeneity (Fig. 5b). These two distinguished dominant species are significantly more efficient in denitrification under anaerobic conditions (Cydzik-Kwiatkowska. 2015; Liu et al. 2018b; Qu et al. 2016), even though they are capable of aerobic denitrification. In contrast, higher relative abundance of Actinobacteria bacterium was found in sites with low heterogeneity because Actinobacteria bacterium prefers aerobic conditions (Zhang et al. 2023). Hanrahan et al. (2018) and Wang et al. (2021) also suggested that nitrogen removal rate is accelerated as the increase of substrate heterogeneity in size distribution. Considering the crucial role of these two species in total nitrogen removal, we could reasonably deduce that substrate heterogeneity may affect the existence of Pseudogemmobacter bohemicus and Paracoccus zhejiangensis.
Relative importance of aquatic nutrient concentrations and substrate heterogeneity
The Spearman’ correlation analyses showed that aquatic nutrient concentrations were significant (P < 0.05) in affecting the relative abundance of nitrifying genes, while the substrate heterogeneity was insignificant. For the relative abundance of denitrifying genes, nutrient concentrations and substrate heterogeneity were both important for narG (P < 0.05); only NH4+ and TIC concentrations had significant correlations with nosZ (P < 0.01); substrate heterogeneity was the major factor for the relative abundance of nirS (Table S6). In the case of the nitrifiers and denitrifiers, aquatic nutrient concentrations were significantly related to the relative abundance of phyla Verrucomicrobia and Planctomycetes, while the substrate heterogeneity was insignificant (Table S7). In contrast, TOC concentrations and substrate heterogeneity were both important for the relative abundance of Proteobacteria (P < 0.05).
Nitrification/denitrification pathways and potential nitrous oxide (N2O) emission in streams
Using the KEGG pathway database, a nitrogen metabolic network was constructed. We identified eight enzymes that associated with nitrification (i.e., amoAB, hao, nxrAB) and denitrification (i.e., napAB, narGHI, nirSK, norBC, nosZ). Furthermore, we analyzed the relative abundance of the enzymes that differed significantly in groups with different substrate size, heterogeneity, and nutrient concentrations (P < 0.05). For enzymes including nxrAB, narGHI, and nosZ, substrate size, heterogeneity, and aquatic nutrient concentrations all exerted great impact on their relative abundance (Fig. 9). The higher relative abundances of these three enzymes were observed in group with the smallest substrate size (D1) and in HH sampling sites. Reasons for this phenomenon might attributed to that high heterogeneity in size distribution would result in varied microenvironments, thereby affecting enzymes involved in nitrogen cycle (Wilson and Dodds 2009). As to nitrification process, NH4+ concentration posed positive effects on three annotated enzymes (amoABC, hao, nxrAB), and other physicochemical properties like CN ratio, NO3−N, and TOC also showed different influence in the relative abundance of enzymes involved in nitrification. Also, heterogeneity and substrate size played a significant role in affecting the abundance of nitrifying genes, especially nxrAB. Meanwhile, in denitrification process, aquatic nutrient concentrations were also significantly correlated to relative abundance of denitrifying genes, like CN ratio to narGHI, and NH4+, NO3−, and TIC to nosZ. In addition, relative abundances of napAB, narGHI, nirSK, norBC, and nosZ were impacted significantly by the substrate size and/or heterogeneity (Fig. 9).
Nitrogen metabolic pathways (nitrification and denitrification process) and enzyme analysis in groups with different substrate sizes, heterogeneity, and nutrient concentrations. The color of each histogram corresponds with the color of each gene label. All the relative abundance of functional enzymes differs significantly (P < 0.05). LH, low heterogeneity; HH, high heterogeneity
N2O is an important greenhouse gas with ~ 296 times warming potential of carbon dioxide. By providing microsites for nitrification and denitrification processes, epilithic biofilms are the main N2O source in river ecosystems (Lear et al. 2008; Magalhães et al. 2005; Sanli et al. 2015; Vila-Costa et al. 2014). Denitrifying genes of norB and nosZ are closely related to N2O emission in the process of reducing nitric oxide to N2O (i.e., producing N2O, linked to norB) and converting nitrous oxide to nitrogen (i.e., consuming N2O, linked nosZ) (Magalhães et al. 2008). In this study, the higher relative abundance of norB and lower relative abundance of nosZ was observed in epilithic biofilms attached to substrates with bigger size and higher heterogeneity (Fig. 3 and Fig. 5a), implying the higher N2O emission in streams with such characteristics. This implication could also be confirmed by the varied relative abundances of norBC, nosZ, and narGHI, affected by substrate size and heterogeneity, illustrated in Fig. 9. Considering that substrate size and heterogeneity can be altered by river regulation works and stream habitat restoration projects (Hasselquist et al. 2018; Morley et al. 2008; Stout et al. 2017; Wells et al. 2008), possibility of N2O emission could be increased after these human projects which generally coarsen substrates in streams (Morley et al. 2008).
Limitations of this study
There are several limitations due to the constraints of measuring equipment and in situ sampling. First, previous studies showed that the chemical properties of particle also affect the microbial community compositions and functions (Qin et al. 2017; Welz et al. 2018). In our study, however, we only considered the impact of particle size while ignoring the impact of chemical properties. In the future, chemical properties of substrates should be investigated using XRF to analyze the comprehensive effects on functional gene abundance and microbial community compositions.
Second, we only performed a once-off sampling mainly because the nutrient concentrations of the Shitingjiang River, such as total nitrogen and total phosphorus, did not differ significantly within one season (i.e., dry season, wet season, or mean-flow season, Li et al. (2022)). In addition, the primary focus of our study is the spatial heterogeneity not the temporal variation of the nutrients concentrations. However, the nutrient concentrations can vary over time in natural rivers, which can significantly impact on the abundance of functional genes and microbial community composition, which should be taken into account in the future.
Third, we only collected the overlying water samples to analyze the nutrient concentrations (Richards et al. 2020) for the consistency of adopting aquatic environment of microbes attached to biofilms. However, for the biofilms that attached to < 2-mm particles, pore water can be a more direct and accurate reflection of the environments of microbes (Drummond et al. 2017). We encourage pore water sampling for better investigation of the correlation with aquatic nutrient concentrations.
Conclusion
Correlations of the relative abundance of nitrifying/denitrifying genes and their associated microbes in epilithic biofilms with substrate properties and aquatic nutrient concentrations were analyzed in the Shitingjiang River. We found that nitrifiers/denitrifiers were influenced by substrate heterogeneity and aquatic nutrient concentrations rather than substrate size. For nitrifying genes, aquatic nutrient concentrations were the main factors that affected their relative abundance, while for the denitrifying genes, substrate size and heterogeneity both exerted great impacts, especially for those genes and dominant species connected with N2O generation. Since nitrogen removal in the form of N2O tends to take place in more heterogeneous streams with larger substrates, river restoration and regulation projects which usually lead to coarser and more heterogenous substrates possibly increase N2O emission. Therefore, the relative abundance of key functional genes related to N2O production and consumption should be monitored after river projects in the future.
Data availability
Raw reads were deposited to NCBI Sequence Read Archive database (Accession Number: SRR 24182856, SRR24182855, SRR24182854, SRR24182853, SRR24182852, SRR24182851, SRR24182850, SRR24182849, SRR24182848, SRR24182847). And sequence data have been deposited in the NCBI Short Read Archive database (Accession Number: SRP432823).
References
Ahmad M, Liu S, Mahmood N et al (2017) Effects of porous carrier size on biofilm development, microbial distribution and nitrogen removal in microaerobic bioreactors. Biores Technol 234:360–369. https://doi.org/10.1016/j.biortech.2017.03.076
Anderson JE, Mueller SA, Kim BR (2007) Incomplete oxidation of ethylenediaminetetraacetic acid in chemical oxygen demand analysis. Water Environ Res 79:1043–1049. https://doi.org/10.2175/106143007X184104
Arnon S, Yanuka K, Nejidat A (2013) Impact of overlying water velocity on ammonium uptake by benthic biofilms. Hydrol Process 27(4):570–578. https://doi.org/10.1002/hyp.9239
Besemer K (2015) Biodiversity, community structure and function of biofilms in stream ecosystems. Res Microbiol 166:774–781. https://doi.org/10.1016/j.resmic.2015.05.006
Boulêtreau S, Lyautey E, Dubois S, Compin A, Delattre C, Touron-Bodilis A, Mastrorillo S, Garabetian F (2014) Warming-induced changes in denitrifier community structure modulate the ability of phototrophic river biofilms to denitrify. Sci Total Environ 466–467:856–863. https://doi.org/10.1016/j.scitotenv.2013.07.121
Buchfink B, Xie C, Huson DH (2015) Fast and sensitive protein alignment using DIAMOND. Nat Methods 12:59–60. https://doi.org/10.1038/nmeth.3176
Bunte K, Abt SR (2001) Sampling surface and subsurface particle-size distributions in wadable gravel-and cobble-bed streams for analyses in sediment transport, hydraulics, and streambed monitoring (No. RMRS-GTR-74). U.S. Department of Agriculture, Forest Service, Rocky Mountain Research Station, Ft. Collins, CO. https://doi.org/10.2737/RMRS-GTR-74
Cardinale BJ, Palmer MA, Swan CM, Brooks S, Poff NL (2002) The influence of substrate heterogeneity on biofilm metabolism in a stream ecosystem. Ecology 83:412–422. https://doi.org/10.1890/0012-9658(2002)083
Chen S, Zhou Y, Chen Y, Gu J (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890. https://doi.org/10.1093/bioinformatics/bty560
Cydzik-Kwiatkowska A (2015) Bacterial structure of aerobic granules is determined by aeration mode and nitrogen load in the reactor cycle. Bioresour Technol 181:312–320. https://doi.org/10.1016/j.biortech.2015.01.101
Drummond JD, Larsen LG, González-Pinzón R, Packman AI, Harvey JW (2017) Fine particle retention within stream storage areas at base flow and in response to a storm event. Water Resour Res 53:5690–5705. https://doi.org/10.1002/2016WR020202
Fan N, Nie R, Wang Q, Liu X (2016) Dramatic undercutting of piedmont rivers after the 2008 Wenchuan Ms 8.0 Earthquake. Sci Rep 6:37108. https://doi.org/10.1038/srep37108
Fu L, Niu B, Zhu Z, Wu S, Li W (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28:3150–3152. https://doi.org/10.1093/bioinformatics/bts565
Galia T, Škarpich V, Gajdošová K, Krpec P (2017) Variability of Wolman pebble samples in gravel/cobble bed streams. Acta Sci Pol Form Circumiectus 1:237–246
Griffith MB, Kaufmann PR, Herlihy AT, Hill BH (2001) Analysis of macroinvertebrate assemblages in relation to environmental gradients in rocky mountain streams. Ecol Appl 11:489–505. https://doi.org/10.1890/1051-0761(2001)011[0489:AOMAIR]2.0.CO;2
Hanrahan BR, Tank JL, Aubeneau AF, Bolster D (2018) Substrate-specific biofilms control nutrient uptake in experimental streams. Freshw Sci 37:456–471. https://doi.org/10.1086/699004
Hao DL (2014) Influence factors of alkaline potassium persulfate digestion uv spectrophotometry for determination of total nitrogen. China Water & Waste Water 1000–4602. https://doi.org/10.19853/j.zgjsps.1000-4602.2014.12.038
Hasselquist E, Polvi L, Kahlert M, Nilsson C, Sandberg L, McKie B (2018) Contrasting responses among aquatic organism groups to changes in geomorphic complexity along a gradient of stream habitat restoration: implications for restoration planning and assessment. Water 10:1465. https://doi.org/10.3390/w10101465
Hyatt D, Chen GL, LoCascio PF, Land ML, Larimer FW, Hauser LJ (2010) Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform 11:119. https://doi.org/10.1186/1471-2105-11-119
Keil D, Meyer A, Berner D, Poll C, Schützenmeister A, Piepho HP, Vlasenko A, Philippot L, Schloter M, Kandeler E, Marhan S (2011) Influence of land-use intensity on the spatial distribution of N-cycling microorganisms in grassland soils: spatial distribution of N-cycling soil microorganisms. FEMS Microbiol Ecol 77:95–106. https://doi.org/10.1111/j.1574-6941.2011.01091.x
Kobayashi Y, Kim C, Yoshimizu C, Kohzu A, Tayasu I, Nagata T (2009) Longitudinal changes in bacterial community composition in river epilithic biofilms: influence of nutrients and organic matter. Aquat Microb Ecol 54:135–152. https://doi.org/10.3354/ame01258
Lear G, Anderson MJ, Smith JP, Boxen K, Lewis GD (2008) Spatial and temporal heterogeneity of the bacterial communities in stream epilithic biofilms: heterogeneity of bacteria in stream biofilms. FEMS Microbiol Ecol 65:463–473. https://doi.org/10.1111/j.1574-6941.2008.00548.x
Li R, Li Y, Kristiansen K, Wang J (2008) SOAP: short oligonucleotide alignment program. Bioinformatics 24:713–714. https://doi.org/10.1093/bioinformatics/btn025
Li D, Liu CM, Luo R, Sadakane K, Lam TW (2015) MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31:1674–1676. https://doi.org/10.1093/bioinformatics/btv033
Li T, Zhou P, Ding Y, Tang Q, Zhou S, Liu Y (2022) Distribution characteristics and source analysis of nitrogen and phosphorus in different rivers in two water period: a case study of Pi River and Shiting River in the Upper Reaches of Tuo River in China. Int J Environ Res Public Health 19:12433. https://doi.org/10.3390/ijerph191912433
Li H, Zhang X, Yu L, Wang L, Chen S (2012) The determination of phosphate in seawater by reverse flow injection spectrophotometry. Rev Anal Chem 31. https://doi.org/10.1515/revac-2011-0024
Lin S-Y, Hameed A, Tsai C-F et al (2022) Description of Pseudogemmobacter faecipullorum sp. nov., isolated from poultry manure. FEMS Microbiol Lett 369:fnac112. https://doi.org/10.1093/femsle/fnac112
Liu Y, Liu J, Yao P, Ge T, Qiao Y, Zhao M, Zhang X-H (2018a) Distribution patterns of ammonia-oxidizing archaea and bacteria in sediments of the eastern China marginal seas. Syst Appl Microbiol 41:658–668. https://doi.org/10.1016/j.syapm.2018.08.008
Liu X, Shu Z, Sun D, Dang Y, Holmes DE (2018b) Heterotrophic nitrifiers dominate reactors treating incineration leachate with high free ammonia concentrations. ACS Sustain Chem Eng 6:15040–15049. https://doi.org/10.1021/acssuschemeng.8b03512
Lyautey E, Teissier S, Charcosset J, Rols J, Garabétian F (2003) Bacterial diversity of epilithic biofilm assemblages of an anthropised river section, assessed by DGGE analysis of a 16S rDNA fragment. Aquat Microb Ecol 33:217–224. https://doi.org/10.3354/ame033217
Lyautey E, Hallin S, Teissier S, Iribar A, Compin A, Philippot L, Garabetian F (2013) Abundance, activity and structure of denitrifier communities in phototrophic river biofilms (River Garonne, France). Hydrobiologia 716:177–187. https://doi.org/10.1007/s10750-013-1561-2
Magalhães CM, Joye SB, Moreira RM, Wiebe WJ, Bordalo AA (2005) Effect of salinity and inorganic nitrogen concentrations on nitrification and denitrification rates in intertidal sediments and rocky biofilms of the Douro River estuary, Portugal. Water Res 39:1783–1794. https://doi.org/10.1016/j.watres.2005.03.008
Magalhães C, Bano N, Wiebe WJ, Bordalo AA, Hollibaugh JT (2008) Dynamics of nitrous oxide reductase genes (nosZ) in intertidal rocky biofilms and sediments of the Douro River Estuary (Portugal), and their relation to N-biogeochemistry. Microb Ecol 55:259–269. https://doi.org/10.1007/s00248-007-9273-7
Merbt SN, Auguet J-C, Blesa A, Martí E, Casamayor EO (2015) Wastewater treatment plant effluents change abundance and composition of ammonia-oxidizing microorganisms in Mediterranean urban stream biofilms. Microb Ecol 69:66–74. https://doi.org/10.1007/s00248-014-0464-8
Morley SA, Duda JJ, Coe HJ, Kloehn KK, McHenry ML (2008) Benthic invertebrates and periphyton in the Elwha River basin: current conditions and predicted response to dam removal. Northwest Sci 82:179–196. https://doi.org/10.3955/0029-344X-82.S.I.179
Mußmann M, Ribot M, Von Schiller D, Merbt SN, Augspurger C, Karwautz C, Winkel M, Battin TJ, Martí E, Daims H (2013) Colonization of freshwater biofilms by nitrifying bacteria from activated sludge. FEMS Microbiol Ecol 85:104–115. https://doi.org/10.1111/1574-6941.12103
Novič M, Pihlar B, Dular M (1988) Use of flow injection analysis based on iodometry for automation of dissolved oxygen (Winkler method) and chemical oxygen demand (dichromate method) determinations. Fresenius z Für Anal Chem 332:750–755. https://doi.org/10.1007/BF01129769
Palacin-Lizarbe Carlos, Camarero L, Hallin S et al (2019) The DNRA-denitrification dichotomy differentiates nitrogen transformation pathways in mountain Lake Benthic habitats. Front Microbiol 10. https://doi.org/10.3389/fmicb.2019.01229
Qin K, Struewing I, Domingo J, Lytle D, Lu J (2017) Opportunistic pathogens and microbial communities and their associations with sediment physical parameters in drinking water storage tank sediments. Pathogens 6:54. https://doi.org/10.3390/pathogens6040054
Qu Z, Bakken LR, Molstad L, Frostegård Å, Bergaust LL (2016) Transcriptional and metabolic regulation of denitrification in Paracoccus denitrificans allows low but significant activity of nitrous oxide reductase under oxic conditions. Environ Microbiol 18:2951–2963. https://doi.org/10.1111/1462-2920.13128
Qu X, Ren Z, Zhang H, Zhang M, Zhang Y, Liu X, Peng W (2017) Influences of anthropogenic land use on microbial community structure and functional potentials of stream benthic biofilms. Sci Rep 7:15117. https://doi.org/10.1038/s41598-017-15624-x
Ren Z, Gao H, Elser JJ, Zhao Q (2017) Microbial functional genes elucidate environmental drivers of biofilm metabolism in glacier-fed streams. Sci Rep 7:12668. https://doi.org/10.1038/s41598-017-13086-9
Richards J, Tibby J, Barr C, Goonan P (2020) Effect of substrate type on diatom-based water quality assessments in the Mount Lofty Ranges, South Australia. Hydrobiologia 847:3077–3090. https://doi.org/10.1007/s10750-020-04316-9
Roig B, Pouly F, Gonzalez C, Thomas O (2001) Alternative method for the measurement of ammonium nitrogen in wastewater. Anal Chim Acta 437:145–149. https://doi.org/10.1016/S0003-2670(01)00944-8
Romaní AM, Sabater S (2001) Structure and activity of rock and sand biofilms in a mediterranean stream. Ecology 82:3232–3245. https://doi.org/10.1890/0012-9658(2001)082[3232:SAAORA]2.0.CO;2
Sanli K, Bengtsson-Palme J, Nilsson RH, Kristiansson E, Alm Rosenblad M, Blanck H, Eriksson KM (2015) Metagenomic sequencing of marine periphyton: taxonomic and functional insights into biofilm communities. Front Microbiol 6. https://doi.org/10.3389/fmicb.2015.01192
Santmire JA, Leff LG (2007) The effect of sediment grain size on bacterial communities in streams. J North Am Benthol Soc 26:601–610. https://doi.org/10.1899/06-130.1
Silva LCF, Lopes DRG, Lima HS, Quartaroli L, De Sousa MP, Waldow VDA, Akamine RN, De Paula SO, Silva CCD (2021) Comparison of methods for preservation of activated sludge samples for high-throughput nucleic acid sequencing and bacterial diversity analysis. Int Biodeterior Biodegrad 157:105139. https://doi.org/10.1016/j.ibiod.2020.105139
Singer G, Besemer K, Schmitt-Kopplin P, Hödl I, Battin TJ (2010) Physical heterogeneity increases biofilm resource use and its molecular diversity in stream mesocosms. PLoS One 5:e9988. https://doi.org/10.1371/journal.pone.0009988
Smith RC, Oerther DB (2006) Microbial community development in a laboratory-scale nitrifying activated sludge system with input from a side-stream bioreactor treating digester supernatant. Water Sci Technol 54:209–216. https://doi.org/10.2166/wst.2006.389
Stout TL, Majerova M, Neilson BT (2017) Impacts of beaver dams on channel hydraulics and substrate characteristics in a mountain stream. Ecohydrology 10:e1767. https://doi.org/10.1002/eco.1767
Suarez C, Piculell M, Modin O, Langenheder S, Persson F, Hermansson M et al (2019) Thickness determines microbial community structure and function in nitrifying biofilms via deterministic assembly. Sci Rep 9(1):5110. https://doi.org/10.1038/s41598-019-41542-1
Vila-Costa M, Bartrons M, Catalan J, Casamayor EO (2014) Nitrogen-cycling genes in epilithic biofilms of oligotrophic high-altitude lakes (Central Pyrenees, Spain). Microb Ecol 68:60–69. https://doi.org/10.1007/s00248-014-0417-2
Wang C, Zhu G, Wang Y, Wang S, Yin C (2013) Nitrous oxide reductase gene (nosZ) and N2O reduction along the littoral gradient of a eutrophic freshwater lake. J Environ Sci 25:44–52. https://doi.org/10.1016/S1001-0742(12)60005-9
Wang P, Li J, Luo X, Ahmad M, Duan L, Yin L, Fang B, Li S, Yang Y, Jiang L, Li W (2022) Biogeographical distributions of nitrogen-cycling functional genes in a subtropical estuary. Funct Ecol 36:187–201. https://doi.org/10.1111/1365-2435.13949
Wang W, Hu M, Shu X, et al (2021) Microbiome of permeable sandy substrate in headwater river is shaped by water chemistry rather than grain size and heterogeneity. Sci Total Environ 780:146552. https://doi.org/10.1016/j.scitotenv.2021.146552
Wells AJ, Balster NJ, VanWychen S, Harrington J (2008) Differences in belowground heterogeneity within a restoration of a dewatered reservoir in Southwestern Wisconsin. Restor Ecol 16:678–688. https://doi.org/10.1111/j.1526-100X.2008.00487.x
Welz PJ, Mbasha W, Smith I, Holtman G, Terblanche G, Le Roes-Hill M, Haldenwang R (2018) The influence of grain physicochemistry and biomass on hydraulic conductivity in sand-filled treatment wetlands. Ecol Eng 116:21–30. https://doi.org/10.1016/j.ecoleng.2018.02.017
Wilson KC, Dodds WK (2009) Centimeter-scale stream substratum heterogeneity and metabolic rates. Hydrobiologia 623:53–62. https://doi.org/10.1007/s10750-008-9647-y
Wolman MG (1954) A method of sampling coarse river-bed material. Trans Am Geophys Union 35:951. https://doi.org/10.1029/TR035i006p00951
Wu K, Yau Y, Sze ETP (2020) Application of anaerobic bacterial ammonification pretreatment to microalgal food waste leachate cultivation and biofuel production. Mar Pollut Bull 153:111007. https://doi.org/10.1016/j.marpolbul.2020.111007
Xu M, Fralick D, Zheng JZ, Wang B, Tu XM, Feng C (2017) The differences and similarities between two-sample t-test and paired t-test 29
Zhang X, Deng J, Yang C, Wang Z, Liu Y (2022) Selective reduction of nitrite to nitrogen by polyaniline-carbon nanotubes composite at neutral pH. Environ Res 214:114203. https://doi.org/10.1016/j.envres.2022.114203
Zhang H, Yang W, Ma B, Liu X, Huang T, Niu L, Zhao K, Yang Y, Li H (2023) Aerobic denitrifying using actinobacterial consortium: novel denitrifying microbe and its application. Sci Total Environ 859:160236. https://doi.org/10.1016/j.scitotenv.2022.160236
Acknowledgements
Financial support from the National Natural Science Foundation of China (Grant No. 52079094 and 52079095) is acknowledged. The authors are also grateful to Tang Liu for valuable suggestions, and Songyi Li, Min Liu, Xiaobing Meng, Yushuang Yang, and Guinan Liu for the help in field sampling.
Author information
Authors and Affiliations
Contributions
CL: data curation, formal analysis, writing—original draft, samples collecting, and visualization; YY: project administration, writing—original draft, samples collecting, funding acquisition, and supervision; SZ: project administration, samples collecting, and funding acquisition; XL: formal analysis and writing—review and editing; LP: writing—review and editing and samples collecting; ZY: writing—review and editing, supervision, and resources.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, C., Yue, Y., Zheng, S. et al. Impacts of substrate properties and aquatic nutrient concentrations on the relative abundance of nitrifying/denitrifying genes and the associated microbes in epilithic biofilms. Environ Sci Pollut Res 30, 120930–120944 (2023). https://doi.org/10.1007/s11356-023-30818-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-023-30818-9