Abstract
Denitrification in river sediments plays a very important role in removing nitrogen in aquatic ecosystem. To gain insight into the key factors driving denitrification at large spatial scales, a total of 135 sediment samples were collected from Huaihe River and its branches located in the northern of Anhui province. Bacterial community composition and denitrifying functional genes (nirS, nirK, and nosZ) were measured by high-throughput sequencing and real-time PCR approaches. Potential denitrification rate (PDR) was measured by acetylene inhibition method, which varied from 0.01 to 15.69 μg N g−1 h−1. The sequencing results based on 16S rRNA gene found that the main denitrification bacterial taxa included Bacillus, Thiobacillus, Acinetobacter, Halomonas, Denitratisoma, Pseudomonas, Rhodanobacter, and Thauera. Therein, Thiobacillus might play key roles in the denitrification. Total nitrogen and N:P ratio were the only chemical factors related with all denitrification genes. Furthermore, nirS gene abundance could be more susceptible to environmental parameters compared with nirK and nosZ genes. Canonical correspondence analysis indicated that NO3−, NO2−, NH4+ and IP had the significant impacts on the nirS-encoding bacterial community and spatial distributions. There was a significantly positive correlation between Thiobacillus and nirS gene. We considered that higher numbers of nosZ appeared in nutrient rich sediments. More strikingly, PDR was positively correlated with the abundance of three functional genes. Random forest analysis showed that NH4+ was the most powerful predictor of PDR. These findings can yield practical and important reference for the bioremediation or evaluation of wetland systems.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Rivers play a substantial role in regional and global elemental biogeochemical cycles (Aufdenkampe et al. 2011), which can serve as a good sentinel of environmental changes in terrestrial and atmospheric processes (Williamson et al. 2008; Crump et al. 2009). However, nutrients originated from domestic sewage, farm drainage, industrial effluent, etc. not only lead to the deterioration of water quality in river systems, but also influence the function of river ecosystems. The nutrients in the aquatic ecosystems are foremost deposited in the sediment, harboring a lot of active and diverse microbia which affect the dynamic of nutrients (Cheng et al. 2014; Liu et al. 2014). For example, the denitrifiers dwelling in the river sediments can remove nitrate to alleviate eutrophication of river (Guo et al. 2013). The denitrifiers are widely distributed in wetland systems, including pools, rivers, lakes, and constructed wetlands (Huang et al. 2011; Ji et al. 2012;Yao et al. 2016).
Denitrification consists of many chemical steps, and each step is catalyzed by enzyme(s) from microorganisms. The reaction of reducing nitrite (NO2−) to nitric oxide (NO) catalyzed by nitrite reductase (encoded by nirS and nirK genes) is the central limiting step of denitrification (Kandeler et al. 2006). The terminal step of denitrificaton (N2O→N2) is catalyzed by nitrous oxide reductases (reducing nitrous oxide to dinitrogen, encoded by nosZ gene). Additionally, nirK, nirS, and nosZ are commonly used as the marker genes to detect the microbia involved in complete denitrification and predict denitrification-derived N2O emission (Myrtotsiknia, Paranychianakis, et al. 2015).
Denitrification can be achieved by phylogenetically unrelated denitrifier assemblages (Zhou et al. 2016), and there are huge differences in the physiology of denitrifying bacteria (Li et al. 2018). Hence, several previous studies have indicated that denitrifier community and denitrification rate in sediment/soil could be affected by environmental variables (Banerjee and Siciliano 2012; Myrtotsiknia, Paranychianakis, et al. 2015; Li et al. 2018). The utilization of molecular biological techniques, especially high-throughput sequencing and qPCR, greatly promoted studies on denitrifiers and denitrification in environmental samples (Banerjee and Siciliano 2012; Kozich et al. 2013). Simultaneously, several bacteria belonging to phylum Proteobacteria, including Thauera, Paracoccus, Hypomicrobium, and Comamonas can participate in denitrification process (Wang et al. 2014b). Bacteria having the potential of denitrification belong to diverse groups and possess various physiological traits and metabolic potential. Denitrifiers play a key role to sustain denitrification potential by altering environmental factors in river sediments (Bowles et al. 2012; Guo et al. 2013). The contents of ammonia, nitrate, total nitrogen, and total phosphorus might have important roles in shaping wetland denitrifier community structure (Li et al. 2018). Nitrate concentration is considered as the most important factor for controlling the denitrification process in river sediment (Piña-Ochoa, and lvarez-Cobelas, M. 2006). Meanwhile, the distribution of denitrifying functional genes nirS and nirK is environment-specific, and the relationship between nirS- and nirK-encoding denitrifiers is considered to be regulated by factors such as nutrient enrichment (Ji et al. 2012; Wang et al. 2014b). Furthermore, many studies also investigated the abundance and distribution of denitrifying genes in sediments and found that environmental factors, such as organic matter, dissolved inorganic nitrogen (DIN) (NO3−, NO2−, and NH4+), dissolved oxygen, and redox potential, significantly affected the distribution of denitrifying genes in sediments (Bowles et al. 2012; Myrtotsiknia, Paranychianakis, et al. 2015). Denitrifying genes have been also found to be correlated with potential denitrification rates (PDRs) (Čuhel et al. 2010; Semedo and Song 2020). Studies also indicated that the denitrification rates might been mediated by denitrifier community structure (Li et al. 2018; Xiong et al. 2017). Moreover, the sediment/soil ammonia, nitrate, nitrite, total phosphorus, and pH might exert a certain influence on denitrification rates (Xiong et al. 2017; Lisa et al. 2015). Detailed studies, therefore, are warranted to explore the relationship between denitrifier abundance and denitrification potential in the river sediments.
Huaihe River is located in plain area with the low flow velocity. An amount of nutrients including nitrogen, phosphorus, and organic matter from domestic sewage, farm drainage, industrial effluent, etc. enter into the main stream and its branches, resulting in deterioration of water quality. In the past 20 years, water pollution treatment in the Huaihe River basin has drawn great attentions of Chinese government and scientists. Although some effective measurements have been taken, the contents of nitrogen, phosphorus, and organic matter in sediments are still high. As two obligatory denitrification intermediates, NO and N2O elicit a harmful effect on biological metabolism and the natural environment. NO, a toxic substance, is recognized as an indirect greenhouse gas (Zhou et al. 2018). The reduction reaction of NO caused by microorganisms is the main source of N2O. And N2O is a powerful greenhouse gas (310 times stronger than carbon dioxide’s greenhouse effect), and it is also the main substance that destroys the ozone layer (Kuypers et al. 2018). In studies regarding sediment characteristics and denitrification potential, denitrifier community with functional gene abundance can be used to identify key factors to assess the function of river ecosystems, which improve river management and accelerate N loss in the form N2 from river sediment.
In this study, river sediment samples from the main stream and 22 branches located in Anhui province with different nutrient content were collected and a series of chemical and microbial analysis were carried out. The contents of organic matter, nitrogen, and phosphorus were measured. The microbial communities were investigated by high-throughput sequencing and qPCR. We explored two questions in the current study. We first explored whether differences in nutrient level are reflected in differences in their denitrifier communities and gene abundance. Second, we aimed to identify the key biological and environmental drivers that explain the observed changes in PDR associated with nitrogen removal.
Materials and methods
Site description
The Huaihe River (30° 55′–36°36′ N, 111°55′–120°45′ E) is located in the eastern China, between the Yangtze River and the Yellow River (Jiang 2011). The Huaihe basin (approximately 270,000 km2) is situated in a transition zone of northern-southern in china (Meng et al. 2014; He et al. 2015). The average precipitation rate in the basin is about 883mm annually. For this basin, the average annual temperature ranges 13.2–15.7°C; the annual evaporation ranges from 900 to 1500mm, and frost free period is about 200–240 days (He et al. 2015). In the basin, a complex interaction of meteorological and hydrological processes occurs which frequently trigger and exacerbate flood and drought events (Wang et al. 2014a; Zhang et al. 2015). Previous studies indicated that above 50% water resources of Huaihe River have been over-exploited (Jiang 2011). Agricultural cultivation and livestock production have been a long history in this basin. Textile, household appliances, steel, cement, and fertilizer are the major industries located along the main stream and branches of Huaihe River, which are running through the main economic areas in the middle-eastern of China (Tian et al. 2013). In recent decades, significant amount of nutrients such as nitrogen and phosphorous from farm drainage, domestic sewage, industrial effluent, etc. had entered into the main stream and its branches that are deposited in the river sediment, which led to severe degeneration of river ecosystems.
Sample collection and pretreatment
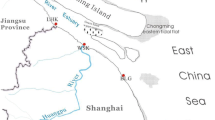
In this study, the main stream and the leftward branches located in the Anhui province were chosen as the investigated subject. The length of main stream of Huaihe River in Anhui province is more than 400km, and its leftward branches in Anhui province mainly includes Honghe River, Guhe River, Runhe River, Shayinghe River, Xifeihe River, Cihuai River, Xinhe River, Qianhe River, Guohe River, Beifeihe River, Xiehe-Huihe River, Tuohe River, Bianhe River, and Suihe River. A total of 135 sections from main stream and its branches were chosen to collect the sediment samples (Fig. 1 and Table S1). The sampling time was from July 2015 to September 2016.
Sketch map of sampling sites of rivers in northern Anhui province. Z Zhaohe River, Y Youhe River, XS Xinhe River, X Xiehe River, XF Xifeihe River, WJ Wujiahe River, W Guohe River, T Tuohe River, TA Tanghe River, S Shayinghe River, R Runhe River, Q Quanhe River, QI Qianhe River, L Suihe River, H Huihe River, HG Huaigan River, HO Honghe River, G Guhe River, CH Cihuai River, C Cihe River, XB Bianhe River, BT Beituohe River, BF Beihe River
Based on different river morphological characteristics, river junctions, nearby possible pollution sources and other factors, port, and wharves with relatively slow velocity and intensive human activities were selected as sampling sections. In each sampling section, 5 subsamples of surface sediment (depth: 0–10cm) were collected by Pedersen sampler and then mixed into a sample. Immediately after collection, sediments were placed into plastic bags and held on ice until return to the laboratory. Although the study was conducted on a wide range of areas, we were able to collect river sediment samples within 2–3 days of certain city. Hence, samples were held on ice approximately 2–3 days. Centralized collection prevented the effects of long time intervals on sediment samples as much as possible. One portion of each sample was used for physical and chemical analysis, and another was stored in the refrigerating cabinet with −20 °C and used for DNA extraction.
Chemical analysis
The pH meter (Mettler Toledo FE20, sedimentmass:H2Ovolume=1g:5ml) was used to assess pH. The organic matter (OM) was obtained from the subsequent loss of weight after continuously drying the sample in a muffle furnace at 550±5°C for 6 h (Parker 1983). The total nitrogen (TN) content was measured using kjeldahl determination device (Smart et al. 1983). The mixture (5 g samples and 20% KCl) was added to the triangle bottle and then filtered after being shaken for 0.5 h. Concentrations of NH4+-N (phenol-sodium hypochlorite spectrophotometry), NO3−-N (ultraviolet spectrophotometry) and NO2−-N (diazo coupled spectrophotometry) in sediment samples were determined using the filter by a UV-1800 spectrophotometer (Shimadzu, Kyoto, Japan). SMT (standard measurement and test) (Ruban et al. 2001) method was used to extract the total phosphorus and various forms of phosphorus. Total phosphorus, organic phosphorus, and inorganic phosphorus were measured by the molybdenum blue colorimetric method at 660 nm.
DNA extraction and high-throughput sequencing
Total DNA in sediment samples were extracted by using the PowerSoil® DNA isolation kit (Mo Bio Carlsbad, USA) according to the manufacturer’s protocol. Each extracted genomic DNA was stored at −20 °C before further analysis.
The 16S rRNA gene was amplified by PCR for multiplexed pyrosequencing with barcoded primers, and the V3-V4 region of the bacterial 16S rDNA was amplified by primers 338F: ACTCCTACGGGAGGCAGCA and 806R: GGACTACHVGGGTWTCTAAT. The resulting amplicons were sequenced by an Ilumina Miseq PE250 machine at Shanghai Majorbio Biopharm Technology Co., Ltd (Shanghai, China).
Real-time fluorescent quantitative PCR
Quantitative analyses on samples from the denitrification were performed on gene fragments including nirK, nirS, and nosZ clade I (Myrtotsiknia, Paranychianakis, et al. 2015). Nitrous oxide reductase (NosZ), the enzyme associated with terminal step in denitrification, however, is not always affiliated with denitrifying microorganisms. Phylogenetic analyses have shown that clade II nosZ is affiliated with a much broader diversity of microorganisms than those with clade I nosZ, the former including both non-denitrifiers and denitrifiers (Joanne et al. 2020). The primer sequence is listed in Table S2. Amplification was performed using TaqMan Universal PCR Master Mix (Bestar Biosystem, German), and the PCR program was based on the specifications. Real-time PCRs were carried out on a Stepone real-time PCR system (Applied Biosystems USA). Each PCR mixture (10 μL) contained 5uL of Bestar® SYBR qPCR Master Mix Ex TaqTM II (2×), 0.25 μL of each primer (concentration of 10 μM), 0.2 μL of ROX reference dye (50×), 3.3 μL of ddH2O, and 1μL of template DNA (Bestar Biosystem, German). The PCR temperature program was initiated with 2 min at 95°C, followed by 40 cycles of 10 s at 95°C and 30 s at the specific annealing temperature (Table S2), and 30 s at 72 °C. A melting curve for SYBR Green assay was generated to verify the specificity of amplification. All sediment DNA samples and negative controls were made in triplicate. After purifying PCR fragments of the respective denitrifying genes using M13 PCR from clones, standard curves were generated based on a serial dilution of known copies of PCR fragments. The R2 value of each standard curve was above 0.99.
Denitrification potential
Denitrification potential was measured in triplicate sediment slurries using the acetylene inhibition technique (Magalhães et al. 2005). A total of 30mL mixture (0.18g·L−1 glucose and 0.1g·L−1 KNO3) was used to incubate approximately 20g fresh sediment sample in 100mL sterile bottle. The bottle was flushed with nitrogen gas for 10 min brought to atmospheric pressure and add high amount of O2-free acetylene gas (C2H2) to block the final denitrification step of N2O to N2. Therefore, the potential denitrification rate (PDR) could calculate as the increment of N2O emissions. After culturing the sealed culture flask at 200 rpm and 25 °C for 6 h, 8 h, 12 h, and 24 h, the concentration of the produced gas N2O was measured on a gas chromatograph equipped with an electron capture detector (ECD). The temperatures of the detector were set at 360°C. The ECD used high-purity nitrogen as a carrier gas. The column temperature was maintained at 50 °C.
Statistical analysis
Table S3 presents the information of sequence numbers from raw to filtered sequences for each sample. The clean reads were defined as sequences >200 bp and <1000 bp in length, with an average quality score >25, with filtering the chimera sequence, without ambiguous base calls, and with at least an 80% match to a previously determined 16S rRNA gene sequence by QIIME 1.8.0. SILVA database (SILVA SSU and LSU databases 128) was used for QIIME. Denitrifying floras were selected through KEGG database: First, search the name of the genus in the KEGG database, and subsequently, the genome appeared. Click in to find its keywords and finally determine its function. If the function of the genus has not been determined in the database, the references were searched to verify whether the genus could participate in the denitrification process.
Canonical correspondence analysis (CCA) was performed with R statistical platform using the “cca” and “envfit” function of the vegan package to analyze the important environmental variables of explaining changes in the denitrification community. Pearson correlation analysis was used to further explore the link between environmental factors and denitrifier abundance. P < 0.05 represented statistically significant.
Before the analysis, the abundance data of functional genes was logarithmically transformed to approximate normality. All data including nutrients content, gene abundance and potential denitrification rates (PDRs) were processed by column normalization. Moreover, to eliminate the interference of autocorrelation between environmental variables, partial Mantel test was employed to investigate the influence of environmental factors on denitrifying gene abundance and potential denitrification rates. Meanwhile, the relationships between the functional genes and PDR were analyzed by partial Mantel test. The results obtained were demonstrated in the “R.” Finally, random forest was used to analyze the contribution of nutrients and gene abundance to PDR changes.
Results and discussion
Chemical properties of river sediments
Table 1 presents the chemical properties of river sediments. The organic matter (OM) content varied from 10.31 to 173.09 (g.kg−1). Similarly, ammonia nitrogen (NH4+-N) content varied from 2.87 to 304.46 (mg.kg−1), and nitrate (NO3−-N) content ranged from 0.10 to 157.48 (mg.kg−1). While total nitrogen (TN) content showed variation from 0.01 to 4.77 (g.kg−1), total phosphate (TP) content was in a range of 0.15 to 2.11 (g.kg−1).
Highly correlated correlations were found among nutrients (Fig. 2) which indicated the similar environmental sources. According to the standard values of the nutrient content in the sediment (Lange 1992), the evaluation standard values of total nitrogen and total phosphorus are 0.55 g/kg and 0.60 g/kg, respectively. The higher contents of nitrogen and phosphorus in branch sediments might be mainly attributed to the terrigenous input caused by agricultural production (non-point sources) and the domestic sewage and industrial wastewater discharge (point sources) (Wang et al. 2003; Wang et al. 2013). Higher nutrient contents in the sediments of the branches might be attributed to the lower water velocity and farmland irrigation (Wang et al. 2013; Jingqiu et al. 2015). In the densely populated area, the organic matter, nitrogen, and phosphorus in domestic sewage from villages and small towns often drained into rivers with surface runoff in the flood season and deposited in the river sediments (Wang et al. 2013).
Bacterial community structure and their abundance in river sediments
Bacteria living in or on the sediments at the bottom of the aquatic system play a substantial role in the organic pollutant degradation, biogeochemical cycles, and microbial food webs (Buesing and Gessner 2006).
The sequence reads were taxonomically assigned to 55 phyla in this study. Therein, the dominant bacteria (with an average relative abundance more than 1%) mainly assigned to 12 phyla (Table S4) including phyla Proteobacteria, Chloroflexi, Bacteroidetes, Acidobacteria, and Actinobacteria, indicating that these phyla were the most important in the bacterial communities of sediments. In these dominant taxa, Firmicutes, Nitrospirae, and Cyanobacteria displayed the largest variation among 135 samples (Table S4). In parallel, α-, β-, δ-, and γ-proteobacteria (mean relative abundance) accounted for 4.19%, 16.39%, 13.35%, and 13.66%, respectively. Previous studies found that β-proteobacteria, γ-proteobacteria, δ-proteobacteria, Bacteroidetes, and Chloroflexi were dominant divisions in sediments (Haller et al. 2011; Ligi et al. 2013;Nunoura et al. 2013). In addition, 33 denitrifying genera were found, in which the average relative abundance of Thiobacillus (Betaproteobacteria) and Halomonas (Gammaproteobacteria) reached 3.739% and 2.575%, respectively (Fig. 3). In these dominant bacteria, genera Bacillus, Halomonas, Pseudomonas, and Thiobacillus have been shown to be involved in denitrification (Falk et al. 2006; Huang et al. 2011; Han and Li 2016).
Quantities of denitrifying functional genes in river sediments
Functional genes nirK, nirS, and nosZ are common molecular markers for denitrifiers and widely distributed in sediments (Huang et al. 2011).
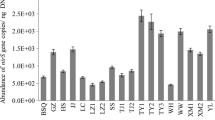
Table S5 presented the abundance of three functional genes (nirK, nirS, and nosZ) involved in denitrification in river sediment investigated. The abundances of nirK, nirS, and nosZ genes ranged from 2.17×106 to 9.00×108, 6.63×106 to 1.54×1010, and 3.3×105 to 1.73×109 gene copies per grams of dry sediment, respectively. In this study, nirS displayed higher abundance than nirK and nosZ genes in most sediment samples collected, which might indicate a favorable habitat distribution for nirS-type denitrifiers (Myrtotsiknia, Paranychianakis, et al. 2015). The differences between denitrifying gene abundance might imply some type of preferential cooperation or provide a proxy for the contribution of the functional groups on the corresponding process (Wang et al. 2014b; Myrtotsiknia, Paranychianakis, et al. 2015). Three functional genes were detected in all sediment samples, indicating the widespread presence of denitrifying bacteria in the river sediment of the mainstream and tributaries.
Figure 2 showed that TN and N:P were found to have significant effects on the abundance of three denitrifying genes (nirK, nirS, and nosZ). Compared to nirK and nosZ, nirS was significantly correlated with a higher number of environmental variables. Specifically, nirS gene was significantly affected by pH, OM, TN, and N:P (Mantel’s P < 0.01 and Mantel’s r > 0.15). Additionally, NO3− and C:N were also correlated to nirS gene (Mantel’s P < 0.05). However, the link between nirK gene and environmental factors was not strong (Mantel’s r < 0.15). Previous studies have shown that the critical role of nirK and nirS genes in nitrogen removal in estuarine and coastal ecosystems and their abundance in sediments were affected by environmental variables (Huang et al. 2011; Wang et al. 2014b). But compared with nirK and nosZ genes, nirS gene could be more susceptible to environmental factors (Fig. 2), which was similar to the result of Myrtotsiknia, Paranychianakis, et al. (2015).
Studies have approved that denitrifiers containing nirS and nirK genes in environments were mainly coming from different bacterial taxa including Alphaproteobacteria, Deltaproteobacteria, Gammaproteobacteria, Actinobacteria, Firmicutes, and Nitrospirae (Green et al. 2010; Xiong et al. 2014). Bacteria containing nirS genes included the genera Alcaligenes, Aromatoleum, Azoarcus, Bacillus, Brachymonas, Candidatus, Dechloromonas, Hahella, Halomonas, Herbaspirillum, Marinobacter, Paracoccus, Pseudomonas, Ralstonia, Rhodanobacter, Roseobacter, Thauera, Thiobacillus, and Thiomicrospira (Falk et al. 2006; Huang et al. 2011). Therein, genera Bacillus, Thiobacillus, Halomonas, Pseudomonas, and Rhodanobacter were widely distributed in sediment samples and displayed a higher relative abundance in this study area. In parallel, correlation analysis showed that Thiobacillus was positively correlated with nirS genes (Table 2). CCA showed that among the nutrient characteristics of Huaihe River sediment, NO3−, NO2−, NH4+, and IP had the significant impacts on the nirS-encoding bacterial community structure and spatial distributions (Fig. 4), and the nirK-encoding denitrifiers were influenced by NO3−, NH4+, and TP (Figure S1). Organic matter (OM) was the primary electron donor for the respiratory denitrifying bacteria (Burgin and Hamilton 2008), and the study found that the relative abundance of nirS-encoding denitrifier community and the quantity of nirS were both closely related to OM in the sediment. The variation percentages in other denitrifying communities were explained by various sediment chemical properties. Figure 4 also indicated that denitrifier abundance was also significantly affected by the content of phosphorus (TP and IP). TP was identified as a possible determinant of shaping denitrifier community structure in aquic brown soils in Northeast China and constructed wetland (Li et al. 2018; Yin et al. 2014). Due to the same terrestrial input, the strong correlation between P and N may partially account for this phenomenon. Another possible cause is the important role of P used as sources of energy in shaping the structure of bacterial communities (Burton and Johnston 2010; Xie et al. 2016). Nevertheless, the links between phosphorus and sediment denitrifiers remain unclear. The present study provided the evidence for the possible links of denitrifier abundance with sediment phosphorus.
Previous researches demonstrated that the abundance of nosZ genes was influenced by many environmental factors, such as sediment texture, carbon, total nitrogen, ammonia, nitrate, and C:N ratio (Huang et al. 2011; Myrtotsiknia, Paranychianakis, et al. 2015). Partial Mantel test presented that the abundance of nosZ gene was significantly affected by the contents of TN and N:P ratio in sediments, implying that the richer nitrogen nutrients can promote high gene abundance (Fig. 2). Laverman et al. (2010) considered that the higher numbers of nosZ were appeared in the nutrient rich sediments and possessed an efficiency of N2 production. Studies also found that the abundance of nosZ genes was stimulated by the nitrogen inputs and the accelerated N2O emissions (Myrtotsiknia, Paranychianakis, et al. 2015).
Bacterial nosZ is universal and abundant in sediments (Rusch and Gaidos 2013). In parallel, we observed similar impediments to functional-gene detection with commonly employed nosZ gene primers, which exclusively target genes from alpha-, beta-, and gamma-proteobacteria, suggesting that previous denitrifier community studies using nosZ analyses most likely do not elucidate the true diversity and abundance of denitrifying bacteria in the environment (Green et al. 2010). Phylogenetic analysis demonstrates that many bacteria containing nosZ genes are belonged to alpha-, beta-, and gamma-proteobacteria, such as genera Alkalilimnicola, Bradyrhizobium, Brucella, Cupriavidus, Hyphomicrobium, Marinobacter, Paracoccus, Pseudomonas, Ralstonia, Rhodopseudomonas, Roseobacter, Silicibacter, and Sinorhizobium (Henry et al. 2006; Magalhães and Hollibaugh 2008; Mills et al. 2008). This research showed that nosZ-type denitrifier community was mainly influenced by environmental factors such as NH4+, NO3−, TN, OM, and OP (Figure S1).
Denitrification potential of river sediments
In this study, denitrification potential rate (PDR) was expressed in N2O production rate which exhibited a range of 0.011 to 15.694 μgN·g−1·h−1, with a mean of 1.689 μgN·g−1·h−1 and a CV of 149.99%. Theoretically, PDR in sediments was influenced by the abundance of nirK, nirS, and nosZ genes as well as the environmental factors. Previous studies have demonstrated that PDR was significantly positively correlated with environmental factors such as ammonia, nitrate, nitrite, total phosphorus, and pH (Laverman et al. 2007; Kanyiginya et al. 2010). Figure 2 indicated that there was significant link between PDR and nutrients (OM, TN, NH4+-N, C:N, and N:P ratio) in this study. And it also highlighted the key contributions of NH4+-N and TN to the variations of PDR. A significant correlation (Mantel’s P < 0.01) between PDR and the abundance of functional genes was found (Fig. 5). Recent studies showed that the nosZ gene was correlated with PDR and hence could act as an index predicting PDR (Hallin et al. 2009; Petersen et al. 2012). Previous studies have demonstrated that both nirK and nirS gene abundance were correlated to PDR (Morales et al. 2010). Comparing to nirK and nosZ genes, the abundance of nirS genes had greater significant correlation coefficient with PDR (Fig. 5), suggesting that it was more suitable using nirS gene abundance to predict denitrification potential in this study. And the random forest analysis also showed high influence of nosZ to PDR. These assays are directly associated with the release of N2O or N2 and the consumption of NO3−, hence, with the activity of denitrifying microorganisms (Xiong et al. 2017;Lisa et al. 2015). Moreover, PDR has been confirmed to be related to changes in gene abundance (Semedo and Song 2020). Petersen et al. (2012) found that functional gene abundances of denitrifiers could predict the potential rate of denitrification. Several studies have also provided evidence for distinct cooccurrence patterns of the different types of denitrifiers between habitats and gene sharing among denitrifiers (Myrtotsiknia, Paranychianakis, et al. 2015). So we have concluded that all three genes may be more suitable to predict denitrification rates, since they respond to different environmental variables, but all of them correlate strongly with PDR. Nutrient level and gene abundance were used as candidate variables in random forest analysis to PDR (Table 3), altogether explaining 14.0% the PDR variations. Random forest (RF) means predictor importance of different indices as drivers for PDR. Analysis showed that NH4+ was the most important contributing factor driving the change of PDR (Table 3). To date, the links of denitrification rate with sediment physicochemical variables remain obscure. Studies have proved that denitrification enzyme activity in sediment was found to be possibly affected by nitrogen contents (Lan et al. 2015). The potential influences of NH4+-N, NO3−-N, and NO2−-N on denitrification rate were also found in sediments (Li et al. 2018). This result indicated that the concentration of NH4+ was more important than the abundance of denitrifying genes to explain the variation of PDR. The driving force of NH4+ on changes in PDR is probably due to the interdependence of nitrifying and denitrifying bacteria on substrates (NH4+ and NO3−) (Myrtotsiknia, Paranychianakis, et al. 2015). Recently, the simultaneous nitrification-denitrification process has also been widely reported (Jia et al. 2020). The importance of NH4+ in this study could also suggest that nitrification-denitrification and/or nitrifier denitrification may play an important role in driving PDR (Jia et al. 2020; Xiong et al. 2017). In summary, this study indicated that variations in PDR were related to changes in nitrogen nutrient concentration and denitrifier abundance.
Further analysis (Table 2) found that there was a significantly positive correlation between PDR and abundance of genera Thiobacillus, which indicated that bacteria from genera Thiobacillus played an important role in the denitrification and might contribute to the conversion/removal of nitrogen pollutants in river sediments. However, a significantly negative correlation was also found between the PDR and Halomonas abundance (r = −0.193 and P= 0.025), which speculate Halomonas may restrain the efficiency of N2 production. The results were instructive for possible bioremediation or evaluation of river sediments. For example, Thiobacillus may contribute to the conversion/removal of nitrogen pollutants in river sediments by facilitating denitrification.
Conclusions
We have concluded from our study that genera Thiobacillus played an important role in the denitrification of the river sediments. TN and N:P ratio were correlated with the abundance of all three denitrifying genes. Additionally, the abundance of nirS gene could be more easily influenced by environmental parameters, and NO3−, NO2−, NH4+, and IP had the significant impacts on the nirS-encoding bacterial community structure and spatial distributions. Quantitative response relationships between denitrification potential and functional genes were established, and PDR was positively correlated with denitrifying genes. Sediment nitrogen nutrient level and denitrifying gene abundance were shown to be tightly linked to PDR. Moreover, NH4+ was the key contributing factor in explaining variations in PDR. There could be a significant difference between community composition and activity throughout the depths sampled in this study as the redox conditions could change drastically, which required further study. Additionally, the physiology and ecological roles of functional genes (including other genes of nitrogen metabolism) and denitrifiers in denitrification mechanism and nitrogen oxide degradation in the environment should be of great interest for future studies.
References
Aufdenkampe AK, Mayorga E, Raymond PA, Melack JM, Doney SC, Alin SR, Aalto RE, Yoo K (2011) Riverine coupling of biogeochemical cycles between land, oceans, and atmosphere. Front Ecol Environ 9:53–60
Banerjee S, Siciliano SD (2012) Spatially tripartite interactions of denitrifiers in arctic ecosystems: activities, functional groups and soil resources. Environ Microbiol 14:2601–2613
Bowles MW, Nigro LM, Teske AP, Joye SB (2012) Denitrification and environmental factors influencing nitrate removal in Guaymas Basin hydrothermally altered sediments. Front Microbiol 3:377
Buesing N, Gessner MO (2006) Benthic bacterial and fungal productivity and carbon turnover in a freshwater marsh. Appl Environ Microbiol 72:596–605
Burgin AJ, Hamilton SK (2008) Have we overemphasized the role of denitrification in aquatic ecosystems? A review of nitrate removal pathways. Front Ecol Environ 5:89–96
Burton GA, Johnston EL (2010) Assessing contaminated sediments in the context of multiple stressors. Environ Toxicol Chem 29:2625–2643
Cheng W, Zhang J, Wang Z, Wang M, Xie S (2014) Bacterial communities in sediments of a drinking water reservoir. Ann Microbiol 64:875–878
Crump BC, Peterson BJ, Raymond PA, Amon RMW, Rinehart A, Mcclelland JW, Holmes RM (2009) Circumpolar synchrony in big river bacterioplankton. Proc Natl Acad Sci U S A 106:21208–21212
Čuhel J, Šimek M, Laughlin RJ, Bru D, Chèneby D, Watson CJ, Philippot L (2010) Insights into the effect of soil pH on N2O and N-2 emissions and denitrifier community size and activity. Applied & Environmental Microbiology Aem 76:1870–1878
Falk S, Hannig M, Gliesche C, Wardenga R, Ster MK, Jürgens K, Braker G (2006) nirS-containing denitrifier communities in the water column and sediment of the Baltic Sea. Biogeosciences 3:697–727
Green SJ, Prakash O, Gihring TM, Akob DM, Jasrotia P, Jardine PM, Watson DB, Brown SD, Palumbo AV, Kostka JE (2010) Denitrifying bacteria isolated from terrestrial subsurface sediments exposed to mixed-waste contamination. Appl Environ Microbiol 76:3244–3254
Guo GX, Deng H, Qiao M, Yao HY, Zhu YG (2013) Effect of long-term wastewater irrigation on potential denitrification and denitrifying communities in soils at the watershed scale. Environ Sci Technol 47:3105–3113
Haller L, Tonolla M, Zopfi J, Peduzzi R, Wildi W, Poté J (2011) Composition of bacterial and archaeal communities in freshwater sediments with different contamination levels (Lake Geneva, Switzerland). Water Res 45:1213–1228
Hallin S, Jones CM, Schloter M, Philippot L (2009) Relationship between N-cycling communities and ecosystem functioning in a 50-year-old fertilization experiment. ISME J 3:597–605
Han H, Li Z (2016) Effects of macrophyte-associated nitrogen cycling bacteria on ANAMMOX and denitrification in river sediments in the Taihu Lake region of China. Ecol Eng 93:82–90
He Y, Ye J, Yang X (2015) Analysis of the spatio-temporal patterns of dry and wet conditions in the Huai River Basin using the standardized precipitation index. Atmos Res 166:120–128
Henry S, Bru D, Stres B, Hallet S, Philippot L (2006) Quantitative detection of the nosZ gene, encoding nitrous oxide reductase, and comparison of the abundances of 16S rRNA, narG, nirK, and nosZ genes in soils. Appl Environ Microbiol 72:5181–5189
Huang S, Chen C, Wu Q, Zhang R (2011) Distribution of typical denitrifying functional genes and diversity of the nirS-encoding bacterial community related to environmental characteristics of river sediments. Biogeosci Discuss 8:5251–5280
Ji G, Wang R, Zhi W, Liu X, Kong Y, Tan Y (2012) Distribution patterns of denitrification functional genes and microbial floras in multimedia constructed wetlands. Ecol Eng 44:179–188
Jia Y, Zhou M, Chen Y, Hu Y, Luo J (2020) Insight into short-cut of simultaneous nitrification and denitrification process in moving bed biofilm reactor: effects of carbon to nitrogen ratio. Chem Eng J 400:125905
Jiang Y (2011) GIS stream network analysis for Huaihe River basin of China. Procedia Environ Sci 10:1553–1558
Jingqiu P, Changyuan T, Xianfang S (2015) Distributions of phosphorus fractions in the sediments of a river–lake system: a case study in Huai River catchment area, China. Water Sci Technol 72:824–834
Joanne C, Lynn C, Alexander K, Wendy H, Robert A (2020) Hierarchical detection of diverse Clade II (atypical) nosZ genes using new primer sets for classical- and multiplex PCR array applications. J Microbiol Methods 172:105908
Kandeler E, Deiglmayr K, Tscherko D, Bru D, Philippot L (2006) Abundance of narG, nirS, nirK, and nosZ genes of denitrifying bacteria during primary successions of a glacier foreland. Appl Environ Microbiol 72:5957–5962
Kanyiginya V, Kansiime F, Kimwaga R, Mashauri DA (2010) Assessment of nutrient retention by Natete wetland Kampala, Uganda. Physics & Chemistry of the Earth Parts A/b/c 35:657–664
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol 79:5112–5120
Kuypers M, Marchant H, Kartal B (2018) The microbial nitrogen-cycling network. Nat Rev Microbiol 16:263–276
Lan Y, Cui B, Han Z, Li X, Li F, Zhang Y (2015) Spatial distribution and environmental determinants of denitrification enzyme activity in reed-dominated raised fields. Chin Geogra Sci 25:438–450
Lange GJD (1992) Distribution of exchangeable, fixed, organic and total nitrogen in interbedded turbiditic/pelagic sediments of the madeira abyssal plain, eastern north atlantic. Mar Geol 109:95–114
Laverman AM, Meile C, Cappellen PV, Wieringa EBA (2007) Vertical distribution of denitrification in an estuarine sediment: integrating sediment flowthrough reactor experiments and microprofiling via reactive transport modeling. Appl Environ Microbiol 73:40–47
Laverman AM, Garnier JA, Mounier EM, Rooseamsaleg CL (2010) Nitrous oxide production kinetics during nitrate reduction in river sediments. Water Res 44:1753–1764
Li BX, Chen JF, Wu Z, Wu SF, Xie SG, Liu Y (2018) Seasonal and spatial dynamics of denitrification rate and denitrifier community in constructed wetland treating polluted river water. Int Biodeterior Biodegradation 126:143–151
Ligi T, Oopkaup K, Truu M, Preem JK, Nõlvak H, Mitsch WJ, Mander, Truu J (2013) Characterization of bacterial communities in soil and sediment of a created riverine wetland complex using high-throughput 16S rRNA amplicon sequencing. Ecol Eng 72:56–66
Lisa JA, Song B, Tobias CR, Hines DE (2015) Genetic and biogeochemical investigation of sedimentary nitrogen cycling communities responding to tidal and seasonal dynamics in cape fear river estuary. Estuar Coast Shelf Sci 167:A313–A323
Liu Y, Zhang J, Zhao L, Zhang X, Xie S (2014) Spatial distribution of bacterial communities in high-altitude freshwater wetland sediment. Limnology 15:249–256
Magalhães C, Hollibaugh JT (2008) Dynamics of nitrous oxide reductase genes (nosZ) in intertidal rocky biofilms and sediments of the Douro River estuary (Portugal), and their relation to N-biogeochemistry. Microb Ecol 55:259–269
Magalhães CM, Joye SB, Moreira RM, Wiebe WJ, Bordalo AA (2005) Effect of salinity and inorganic nitrogen concentrations on nitrification and denitrification rates in intertidal sediments and rocky biofilms of the Douro River estuary, Portugal. Water Res 39:1783–1794
Meng J, Wang T, Wang P, Giesy JP, Lu Y (2014) Perfluoroalkyl substances and organochlorine pesticides in sediments from Huaihe watershed in China. J Environ Sci 26:2198–2206
Mills HJ, Hunter E, Humphrys M, Kerkhof L, Mcguinness L, Huettel M, Kostka JE (2008) Characterization of nitrifying, denitrifying, and overall bacterial communities in permeable marine sediments of the northeastern Gulf of Mexico. Appl Environ Microbiol 74:4440–4453
Morales SE, Cosart T, Holben WE (2010) Bacterial gene abundances as indicators of greenhouse gas emission in soils. ISME J 4:799–808
Myrtotsiknia, Paranychianakis NV, Varouchakis EA, Nikolaidis NP. (2015). Environmental drivers of the distribution of nitrogen functional genes at a watershed scale. FEMS Microbiology Ecology 91
Nunoura T, Nishizawa M, Kikuchi T, Tsubouchi T, Hirai M, Koide O, Miyazaki J, Hirayama H, Koba K, Takai K (2013) Molecular biological and isotopic biogeochemical prognoses of the nitrification-driven dynamic microbial nitrogen cycle in hadopelagic sediments. Environ Microbiol 15:3087–3107
Parker JG (1983) A comparison of methods used for the measurement of organic matter in marine sediment. Chem Ecol 1:201–209
Petersen DG, Blazewicz SJ, Firestone M, Herman DJ, Turetsky M, Waldrop M (2012) Abundance of microbial genes associated with nitrogen cycling as indices of biogeochemical process rates across a vegetation gradient in Alaska. Environ Microbiol 14:993–1008
Piña-Ochoa E, lvarez-Cobelas, M. (2006) Denitrification in aquatic environments: a cross-system analysis. Biogeochemistry 81:111–130
Ruban V, López-Sánchez JF, Pardo P, Rauret G, Muntau H, Quevauviller P (2001) Harmonized protocol and certified reference material for the determination of extractable contents of phosphorus in freshwater sediments--a synthesis of recent works. Anal Bioanal Chem 370:224–228
Rusch A, Gaidos E (2013) Nitrogen-cycling bacteria and archaea in the carbonate sediment of a coral reef. Geobiology 11:472–484
Semedo M, Song B (2020) From genes to nitrogen removal: determining the impacts of poultry industry wastewater on tidal creek denitrification. Environ Sci Technol 54:146–157
Smart MM, Rada RG, Donnermeyer GN (1983) Determination of total nitrogen in sediments and plants using persulfate digestion. An evaluation and comparison with the kjeldahl procedure. Water Res 17:1207–1211
Tian D, Zheng W, Wei X, Sun X, Liu L, Chen X, Zhang H, Zhou Y, Chen H, Zhang H (2013) Dissolved microcystins in surface and ground waters in regions with high cancer incidence in the Huai River Basin of China. Chemosphere 91:1064–1071
Wang C, Wang Y, Mo Z, Wang Z (2003) Ecotoxicological examination of sediment extracts of Huaihe River, China by in vitro bioassays. Bull Environ Contam Toxicol 71:0782–0790
Wang P, Wang C, Zhao L, Schnoebelen DJ, Qian J, Hou J (2013) Nutrient speciation and distribution between surface water and sediment in the middle reach of the Huai River, China. J Environ Eng 139:226–234
Wang JZ, Chen TH, Zhu CZ, Peng SC (2014a) Trace organic pollutants in sediments from Huaihe River, China: evaluation of sources and ecological risk. J Hydrol 512:463–469
Wang L, Zheng B, Nan B, Hu P (2014b) Diversity of bacterial community and detection of nirS- and nirK-encoding denitrifying bacteria in sandy intertidal sediments along Laizhou Bay of Bohai Sea, China. Mar Pollut Bull 88:215–223
Williamson CE, Dodds W, Kratz TK, Palmer MA (2008) Lakes and streams as sentinels of environmental change in terrestrial atmospheric processes. Front Ecol Environ 6:247–254
Xie Y, Wang J, Wu Y, Ren C, Song C, Yang J, Yu H, Giesy JP, Zhang X (2016) Using in situ bacterial communities to monitor contaminants in river sediments. Environ Pollut 212:348–357
Xiong J, Ye X, Wang K, Chen H, Hu C, Zhu J, Zhang D (2014) Biogeography of the sediment bacterial community responds to a nitrogen pollution gradient in the East China Sea. Appl Environ Microbiol 80:1919–1925
Xiong Z, Guo L, Zhang Q, Liu G, Liu W (2017) Edaphic conditions regulate denitrification directly and indirectly by altering denitrifier abundance in wetlands along the Han River, China. Environ Sci Technol 51:5483–5491
Yao X, Zhang L, Zhang Y, Xu H, Jiang X (2016) Denitrification occurring on suspended sediment in a large, shallow, subtropical lake (Poyang Lake, China) ☆. Environ Pollut 219:501–511
Yin C, Fan FL, Song A, Li ZJ, Yu WT, Liang YC (2014) Different denitrification potential of aquic brown soil in Northeast China under inorganic and organic fertilization accompanied by distinct changes of nirS- and nirK-denitrifying bacterial community. Eur J Soil Biol 65:47–56
Zhang W, Pan S, Cao L, Cai X, Zhang K, Xu Y, Xu W (2015) Changes in extreme climate events in eastern China during 1960–2013: a case study of the Huaihe River Basin. Quat Int 380:22–34
Zhou SL, Huang TL, Zhang CH, Fang KK, Xia C, Bai SY, Zeng MZ, Qiu XP (2016) Illumina MiSeq sequencing reveals the community composition of NirS-Type and NirK-type denitrifiers in Zhoucun reservoir - a large shallow eutrophic reservoir in northern China. RSC Adv 6:91517–91528
Zhou H, Zhao Y, Yang H, Zhu L, Cai B, Luo S, Cao J, Wei Z (2018) Transformation of organic nitrogen fractions with different molecular weights during different organic wastes composting. Bioresour Technol 262:221–228
Acknowledgements
We appreciate many graduate students involved in analyzing the chemical properties and collecting soil samples who are not listed as co-authors.
Availability of data and materials
All data generated or analyzed during this study are included in this published article.
Funding
This study was funded by Major Science and Technology Program for Water Pollution Control and Treatment (2012ZX07204-004) and Wetlands Ecological Security Collaborative Innovation Center, Anhui University.
Author information
Authors and Affiliations
Contributions
MZZ was a major contributor in analyzing the data and writing the manuscript. UD provided language help. QYS originally designed the study. PXC carried out the chemical experiments of sediments, and XHW provided help in collecting samples. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhang, M., Daraz, U., Sun, Q. et al. Denitrifier abundance and community composition linked to denitrification potential in river sediments. Environ Sci Pollut Res 28, 51928–51939 (2021). https://doi.org/10.1007/s11356-021-14348-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-021-14348-w