Abstract
Wastewater discharge evidently increased bacterial diversity in the receiving waterbodies. The objective of this study was to evaluate the effectiveness of a constructed wetland in reducing fecal indicator bacteria (FIB) and antibiotic resistant genes (ARGs). We determined the prevalence and attenuation of fecal indicator bacteria including Escherichia coli and enterococci, along with ARGs, and human-associated Bacteroidales (HF183) markers by quantitative polymerase chain reaction (qPCR) method. Three types of water samples (inlet, intermediate, and outlet) from a constructed wetland were collected once a month from May to December in 2013. The overall reduction of E. coli was 50.0% based on culture method. According to the qPCR result, the overall removal rate of E. coli was only 6.7%. Enterococci were found in 62.5% of the wetland samples. HF183 genetic marker was detected in all final effluent samples with concentration ranging from 1.8 to 4.22 log10 gene copies (GC)/100 ml. Of the ARGs tested, erythromycin resistance genes (ermF) were detected in 79.2% of the wetland samples. The class 1 integrase (intI1) was detected in all water samples with concentration ranging from 0.83 to 5.54 log10 GC/100 ml. The overall removal rates of enterococci, HF183, intI1, and ermF were 84.0%, 66.6%, 67.2%, and 13.1%, respectively.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Wastewater is increasingly recognized as another option that can be treated and then used as a resource (Greenway 2005). One major issue with the use of this wastewater for potable reuse is the presence of both chemical and microbial pollutants that would adversely affect human health (Tran et al. 2015, 2016). The removal of pathogenic bacteria and antibiotic resistance genes (ARGs) from municipal wastewater treatment processes such as membrane bioreactor systems and conventional activated sludge systems, as well as more advanced Bardenpho treatment, has been well documented (Chen et al. 2010, 2016; Le et al. 2018; Xue et al. 2019). However, there are a few disadvantages of these technologies, such as expensive, not entirely feasible for application in rural areas, and unwanted chemical by-products (Guo et al. 2008; Wu et al. 2015; Waller and Bruland 2016; Lu et al. 2016). Compared with wastewater treatment technology, constructed wetlands, on the other hand, have been recognized as environmental-friendly and eco-technology due to their low-cost and easy operation (Faulwetter et al. 2009; Tu et al. 2014; Rachmadi et al. 2016; Chen et al. 2016), although it has disadvantages such as requires large area and the removal efficiency may be affected by environmental factors (Ramirez et al. 2005).
Constructed wetlands are artificial designed and constructed to simulate natural processes to treat domestic wastewaters. As wastewater passing through natural wetlands, it is filtered through sediments and finally replenishes the aquifer, providing clean water to the city and other municipal sources (Gelt 1997). Previous study suggested that constructed wetlands were able to remove organics, total carbon, and nutrients effectively (Tu et al. 2014). It was also reported that fecal indicator bacteria removal in constructed wetland was ranged from 97 to 99%, such as coliforms and Escherichia coli (Quiñónez-Dìaz et al. 2001; Wu et al. 2010). In addition to removal of bacteria, previous researches demonstrated that mesocosm-scale constructed wetlands are capable of removing 18 ARGs varied between 50 and 85.8% (Chen et al. 2010, 2016). However, these experiments were conducted by using mesocosm-scale constructed wetlands. In 2017, Yi et al. (2017) found several ARGs (intl1, sul1, sul2, and qnrA) from landfill leachate could be effectively removed by one hybrid constructed wetlands system. So far, there is limited data on removal of ARGs in treated wastewater by full-scale constructed wetlands system. More importantly, unlike other water treatment facilities, physicochemical and biological factors may affect fecal indicator bacteria removal in constructed wetlands, such as seasonal fluctuations, vegetation, sedimentation, and water composition (Kadlec and Wallace 2008; Wu et al. 2016). Increased water temperature, pH, and retention time are known to increase treatment efficiency, while increased turbidity is associated with decreased pathogen removal (Maiga 2017).
In the present study, quantitative polymerase chain reaction (qPCR) results can be used to determine the effectiveness of the wetlands in removing pollution from water. The concentration of human-associated Bacteroidales marker (HF183) (Bernhard and Field 2000; McLellan and Eren 2014) and genetic markers targeting fecal indicator bacteria (FIB) were determined in inlet, intermediate, and outlet water samples. QPCR were applied to understand the sources of fecal contamination by targeting host-associated molecular markers (Xue et al. 2017, 2018a, b, c; Tandukar et al. 2018). When compared to other culture-based methods, PCR-based methods are able to provide source information for a water sample that has been recently contaminated as they are obligate anaerobes that only are able to survive for short periods of time and have proven to be more successful at accurately detecting DNA and provided more specific and accurate readings (Aly et al. 2012; Kapoor et al. 2015; Xue et al. 2017, 2018c). Antibiotics are used in medicine and livestock animals for prophylactic, therapeutic, and growth-promoting purposes (Davies and Davies 2010; Hijosa-Valsero et al. 2011). Antibiotic residues can be discharged into environments though feces or urine, posing risks to not only human health but also the environment (Costanzo et al. 2005; Kotzerke et al. 2008; Liu et al. 2009; Underwood et al. 2011). The increase in antibiotic usage in humans and livestock leads to spread of occurrence of antibiotic-resistant bacteria (ARB) and ARGs in the environment (Pruden et al. 2006; Tao et al. 2010; Su et al. 2012), which may lead to source of resistant bacteria in humans. We are particularly interested in ARGs of intI1 and ermF that are responsible for resistance to class I integrase and erythromycin antibiotics, respectively. Class 1 integrons have been considered to be the key of worldwide antibiotic resistance problem, and intI1 gene has been known to be a proxy for horizontal gene transfer of ARGs (Gillings et al. 2008; Le et al. 2016). The erythromycin treatment failure has been noted in the past few years and the erythromycin resistance might because of the presence of the ermF genes in the bacterial genome (Chung et al. 2002; Xing et al. 2015).
In the present study, we evaluated the removal efficiency of FIB, HF183 genetic marker, as well as intI1 and erm(F) in surface flow of the constructed wetlands and explore the relationship between FIB and HF183 genetic marker concentrations. We compared various physicochemical water quality parameters such as temperature, pH, and turbidity to investigate whether those factors have any effect or correlation on the prevalence of the bacteria and ARGs. In addition, we analyzed the inlet, intermediate, and outlet samples to compare the efficiency rates among each step of the treatment processes.
Materials and methods
Description of the constructed wetlands studied
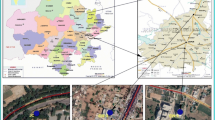
This study was conducted in the Sweetwater Wetlands located in the city of Tucson, Arizona. The Sweetwater Wetlands has been operated for about 20 years and is one of the most important functional, environmental, and educational components of Tucson’s reclaimed water system. It provides additional treatment of the treated wastewater from the Roger Road Water Reclamation Facility as well as backwash water from mixed media filters at the City of Tucson Water Reclamation Plant. This wetland is approximately 0.03 km2 in size with a retention time of 7 days (Vidales-Contreras et al. 2006; Rachmadi et al. 2016). Major vegetations include cattails, bulrushes, large cottonwood, and willow trees. The recovered wastewater is distributed for reuse in Tucson’s golf courses, parks, schools, and other large turf irrigation areas (Fig. 1) (https://www.tucsonaz.gov/water/about-sweetwater-wetlands-and-access).
Sample collection
In total, 24 surface water samples were collected from the Sweetwater Wetlands by using sterile 1-l plastic containers over a period of 8 months. The water samples were then kept on ice and delivered to Dr. Gerba’s laboratory at the University of Arizona within 6 h. The parameters of water temperature, dissolved oxygen, salinity, specific electrical conductivity, and turbidity were measured in situ by using the PCSTEst35 (Eutech Instruments, Singapore) through inlet, intermediate, and outlet samples.
DNA extraction from water samples
On the day of sample arrival, 500 ml from each sample was filtered through a 0.45 μm-pore-size membrane filter and 47-mm-diameter nitrocellulose membrane (Thermo Fisher Scientific, Waltham, MA) under vacuum. After filtration, sterile forceps were used to aseptically fold each of the membrane filters and placed in separate 50-ml conical tubes and stored at − 20 °C until the DNA extraction. Genomic DNA was extracted from membrane filters using the PowerSoil DNA Isolation Kit (Mo Bio Laboratories, Inc., Carlsbad, CA) according to the manufacturer’s instructions. Membrane filters were cut into small pieces with sterile scissors to maximize DNA extraction efficiency, and the extracted DNA was quantified with a NanoDrop ND-2000 UV spectrophotometer (Thermo Fisher Scientific, Walkham, MA). The DNA samples were stored at − 20 °C prior to use.
Quantitative real-time PCR
Quantitative PCR assays targeting E. coli, enterococci, human-associated Bacteroidales (HF183), and ARGs of erm(F) and intI1 were conducted (Table 1). For E. coli, enterococci, the PCR reaction mixture (20 μl) contained 1× PerfeCTa qPCR ToughMix (Quanta Biosciences, Beverly, MA), 0.5 μM of each primer, 0.5 μM of probe, and 2.5 μl of the template DNA. For HF183, erm(F), and intI1, the reaction mixture (15 μl) contained 1× SsoAdvanced Universal SYBR Green Supermix (Bio-Rad, Hercules, CA), 0.2 μM of each primer, and 2.5 μl of the template DNA. All PCR reactions were performed in duplicate using the Applied Biosystems StepOne™ Real-Time PCR system (Thermo Fisher Scientific). The amplification conditions consisted of a hold at 95 °C for 10 min, followed by 40 cycles of 95 °C 15 s, 60 °C 30s, and 72 °C 30s. A calibration curve with concentrations spanning the range from 10 to 106 gene copies per reaction with two replicates was generated. Duplicate no-template controls were included in each run. The calibration standard curves covering the range from 10 to 106 gene copies per reaction were prepared using serial dilutions of plasmid DNA or commercial genomic DNA purchased from ATCC (700926DQ for Escherichia coli and ATCC 29212Q-FZ for enterococci; Manassas, VA, USA). For SYBR Green-based qPCR assays (HF183), plasmid DNA obtained from Dr. Feng’s lab at Auburn University, Alabama. For ARGs SYBR Green-based qPCR assays, intI1 environmental E. coli isolate and erm(F) erythromycin-resistant environmental Staphylococcus spp. isolates were obtained from Dr. John Brook’s lab from USDA-ARS at Mississippi State. Sequencing results were confirmed by consulting the NCBI website using the nucleotide Basic Local Alignment Search Tool (BLAST). Plasmid DNA concentration was measured with a NanoDrop ND-2000 UV spectrophotometer (Thermo Fisher Scientific), and the gene copy numbers were calculated.
Data analyses
Statistical analyses were performed using SAS® 9.3 software. A Spearman’s test was used to identify relationships between the environmental parameters and the presence of E. coli, enterococci, HF183, and ARGs, if any. The coefficient correlation (r) value for each regression model was used to provide an indication of goodness of fit. An ANOVA test was used to identify any seasonal differences between site-specific data.
Results
Physicochemical and biological water quality parameters
Twenty-four water samples were collected from the Sweetwater Wetlands from the months of May to December in 2013. In our study, the correlation between physicochemical parameter and FIB as well as ARGs was studied. Turbidity of the water samples (Table S1) was highest in the inlet water, at a value of 30.2 NTU. E. coli was found in 100% of water samples from inlet, intermediate, and outlet locations. The inlet water samples had consistently high E. coli concentrations, with the highest concentration was found in November of 3.74 log10 of MPN/100 ml over the sampling period (Fig. S1). The geometric mean of E. coli concentration over the study period in outlet water samples was decreased by 50.0% compared with inlet water samples (Fig. 2).
Removal rate of FIB, HF183, and ARGs from inlet to intermediate and to outlet in Sweetwater constructed wetland. Positive value indicates the concentration of tested indicators reduced during the treatment. Negative value indicates the concentration of tested indicators increased during the treatment
Occurrence and reduction of E. coli, enterococci, and HF183 determined by qPCR
E. coli was found in 100% of the water samples during the study period with concentrations ranging from 0.68 Log10 GC/100 ml (intermediate sample of September) to 1.89 Log10 GC/100 ml (outlet sample of December) (Table 2). Except for December samples, the reduction of E. coli concentrations from inlet to intermediate samples ranged from 31.3% in May to 70.2% in October. Compared with intermediate water samples, the geometric mean of E. coli concentrations in outlet water samples increased by 65.8% (Fig. 2). E. coli concentrations in outlet water samples collected in September and December increased by 96.3% and 293.9%, respectively. The geometric mean of E. coli concentration over the study period in outlet water samples decreased by 6.7% compared with inlet water samples.
The enterococci markers were found in 62.5% (15/24) of water samples (Table 2), with concentrations ranging from 0.81 Log10 GC/100 ml (Intermediate sample of July) to 3.88 Log10 GC/100 ml GC/100 ml (inlet sample of May). From inlet to intermediate water samples in all months except for December, the removal rates of enterococci ranged from 70.0% in June to 99.0% in July. The overall concentration of enterococci in outlet water samples decreased by 84.0% compared with inlet water samples.
Human-associated Bacteroidales (HF183) was detected in 95.8% (23/24) of water samples, with concentrations ranging from 1.80 Log10 GC/100 ml (outlet water sample in Oct.) to 4.22 Log10 GC/100 ml (outlet sample of December). Except for June and September, the reduction rates of HF183 from inlet to intermediate water samples ranged from 63.0% (in August) to 91% (in July). The HF183 concentrations increased from inlet to intermediate in June and September, with increasing rate of 90% and 3%, respectively. Compared with intermediate water samples, the geometric mean of HF183 concentrations in outlet water samples increased by 37.2%. Except for a decrease of 64.7% from the inlet to intermediate sample in December, the geometric mean of HF183 in the outlet samples decreased by 66.6% when compared to inlet samples (Fig. 2).
Occurrence and reduction of ARGs determined by qPCR
In addition to FIB and HF183, the occurrence of ARGs, such as intI1 and erm(F), was also determined using qPCR method in the present study. All Sweetwater samples tested positive for intI1 marker. The concentrations of intI1 marker ranged from 0.83 Log10 GC/100 ml to 5.54 Log10 GC/100 ml. The removal rate of intI1 in intermediate samples ranged from 3.4% to 99.6% compared with inlet samples. The geometric mean of intI1 for outlet samples increased by 159.1% compared with intermediate samples. The overall removal rate of intI1 from inlet to outlet was 66.6%. The erm(F) marker was detected in 79.2% (19/24) of Sweetwater samples, with concentration ranged from 3.37 Log10 GC/100 ml (inlet water sample in July) to 2.28 Log10 GC/100 ml (inlet sample of Sept). From inlet to intermediate water samples, the reduction rate ranged from 5.7% in June to 83% in July. From inlet to outlet samples, the overall removal rate was 13.1%.
Discussion
It should be noticed that data on the water physicochemical and biological (E. coli IDEXX) parameters was recently published (Rachmadi et al. 2016). However, data included in the present study was to gain insight of removal efficiency and co-occurrence patterns between these parameters and ARGs. During the 8 months sampling period, E. coli was found in all water samples (24 samples) based on both IDEXX (culture method) and molecular qPCR (molecular method) results (Fig. S1; Table 2). Sweetwater Wetland water temperature ranged from 11.5 to 30.9 °C (Table S1), and a significant negative correlation (r = 0.64, p value < 0.01) was observed between E. coli (IDEXX) concentration and water temperature (Fig. S2). E. coli is likely to survive and regrow in aquatic environment (water and sediment) and can be affected by a number of environmental factors, such as water temperature, bacterial growth, sedimentation rate, and pollution sources (Solo-Gabriele et al. 2000; Karimi et al. 2014; Morató et al. 2014; Howitt et al. 2014; Nguyen et al. 2015). There are several reasons for this phenomenon: First, since we pooled E. coli results from all sampling sites together, it is possible that the reduction of E. coli by wetland treatment was higher in summer compared to winter due to different field condition in different seasons (more plants in summer). Previous study suggested that constructed wetlands with vegetation are more effective in removal of bacteria than without the vegetation (Maiga 2017); second, the E. coli concentration in Sweetwater Wetland can be affected by its water source, since the receiving water is treated wastewater. Previous studies have demonstrated that temperature is one of the most important factors driving the microbial community composition in wastewater treatment which will affect the bacterial removal rates (Ebrahimi et al. 2010; Wang et al. 2014; Lu et al. 2015; Meerbergen et al. 2016). In that case, higher temperature during summer increased the E. coli removal rates which is also observed in Fig. S1. In addition, pH has been studied extensively and was suggested affect the growth of bacteria in water, for example, microbes survive best at pH between 5.5 and 7.5 (McFeters and Stuart 1972; Blaustein et al. 2013). Previous study suggested that increased pH is linked to higher bacterial pathogen removal in constructed wetlands and that most bacteria are sensitive to high pH levels (Maiga 2017). The small change in pH among inlet, intermediate, and outlet samples has an insignificant effect on the occurrence of E. coli.
An interesting finding in the present study was that the concentration of E. coli, enterococci, and HF183 genetic markers decreased from inlet to intermediate and then increased back from intermediate to outlet (Fig. 2). The reason to explain this phenomenon was probably due to the fluctuation of turbidity during the treatment process. Water turbidity was found decreasing from inlet to intermediate, and then increased from intermediate to outlet (Table S1). Significant positive correlations between turbidity level and qPCR results for E. coli and HF183 were observed (Fig. 3). A weak positive correlation between turbidity and enterococci was also observed, even though it is not significant. The adsorption of DNA molecules on sediments have been studied extensively (Crecchio et al. 2005; Cai et al. 2006a, b; Yu et al. 2013; Xue and Feng 2018). It is reasonable to assume that the increase of water turbidity is a source of fecal indicator found in the water column. Increasing evidence suggests that sediments serve as a reservoir for pathogenic microorganisms of fecal origin (Wheeler Alm et al. 2003; Badgley et al. 2010). However, no significant correlation between turbidity and E. coli (IDEXX) was found, as qPCR is capable of detecting DNA from culturable cells, nonviable intact cells, viable but non-culturable (VBNC) cells, and extracellular free DNA (Levy-Booth et al. 2007; Rogers et al. 2011; Staley et al. 2012; Xue et al. 2017). Given the fact that qPCR-based MST methods target genetic materials rather than viable organisms, it is more likely that sorption characteristics of DNA molecules may be different compared with intact bacterial cells (Kim et al. 2011; Staley et al. 2012; Shelton et al. 2014).
Previous study evaluated the effectiveness of constructed wetland treatment on pathogen removal (Greenway 2005). This wetland showed a 95% removal rate of pathogens and fecal indicator organisms but also still contained fecal-coliform concentrations in the final effluent. Other constructed wetlands find that all pathogens are expected to be removed but are highly variable on the physicochemical factors (Chen et al. 2016; Le et al. 2016; Yi et al. 2017). The Sweetwater Wetland also had similar findings, where the overall fecal bacteria decreased, but was still prevalent in the final water sample. These studies of constructed wetlands all suggest a similar trend and efficacy of the wetland removal system: while the outlet samples still contain some pathogen bacteria, they are effective in reducing their concentrations. This could be due to extraneous influences, such as temperature, vegetation, sunlight or precipitation, or alternatively, an impact from high retention time to receive samples (Rachmadi et al. 2016). The correlation between environmental factors and bacteria removal efficiency have been studied. For instance, Karimi et al.’s research observed a negative correlation between the content of dissolved oxygen and microorganisms (Karimi et al. 2014). However, our study did not observe any correlation between physicochemical parameters and the removal effectiveness of the wetlands. It could possibly be due to the limited sample size in the present study. Future research is needed to look into the insight of the connection between these parameters and the performance of Sweetwater Wetland.
In addition, our results suggested that constructed wetlands are better at reducing enterococci concentrations than the other two indicators. In comparison, enterococci concentrations decreased by 84.0%, whereas HF183 genetic marker concentrations decreased by 66.6% and E. coli by 6.7%. The findings for HF183 are significant because this marker is an indicator of fecal pollution which may pose human health at risk due to the presence of enteric pathogens in fecal matter (Webster et al. 2013; Nayak et al. 2015). Regulation for E. coli and enterococci in drinking water is zero, so the prevalence of fecal indicator bacteria and HF183 in the final outlet water sample is not safe to be released to the public (EPA). As we mentioned previously, the reduction of E. coli can be affected by a number of environmental factors; we are not clear about why there was such a minimal decrease in E. coli compared to the other targets. Additionally, E. coli could have a slower decay rate than HF183 and enterococci and therefore have a higher prevalence than the other two indicators (Ahmed et al. 2016).
The removal of ARGs by constructed wetlands has been studied and results varied accordingly. The removal rates of ARGs in Chen et al.’s study (2016) ranged from 50.0 to 85.8%. In comparison, the overall removal efficiencies of ARGs in Yi et al.’s research varied significantly from − 145.6 to 98.9% depending on the type of gene (2017). It is noteworthy that even after wastewater treatment process, several ARGs were still found in final effluent at significant higher concentration (103 copies/ml) (Vaz-Moreira et al. 2014; Le et al. 2018). The removal of ARGs in Sweetwater Wetland is comparable to previous studies. Our results showed that Sweetwater Wetlands are less efficient in removing ermF than intl1. For example, the ermF concentrations were reduced by only 13.1% and intl1 markers had reached a 67.2% decrease. Although previous study suggested that the removal rates of ARGs in winter is higher than that in summer (Fang et al. 2017), we did not observe such pattern in our study due to the smaller sample size. However, we did observe the highest concentrations of both ARGs in the months of May and August, where water temperatures are higher (Table 2).
The molecular methods are more accurate than culture-based methods for detecting bacteria (Wade et al. 2010; Xue et al. 2018b) and have been essential to our study at providing information about prevalence of ARGs. Interestingly, we did not observe any correlation between qPCR and IDEXX results for E. coli. The difference between IDEXX and qPCR results reflects differences in natures of analyses as well as differences in persistence of live E. coli compared to DNA molecules in aquatic environment. As we mentioned before, qPCR is capable of detecting DNA from live cells, VBNC cells, and extracellular free DNA. The potential of E. coli regrow in aquatic environment may also alter the results. Previous studies have demonstrated that E. coli are likely to survive and regrow in sediment (Desmarais et al. 2002; Anderson et al. 2005). As the water level in Sweetwater Wetland is shallow, we therefore assume turbulences caused by water flow may resuspend sediments and subsequently release E. coli from sediment. Another study also suggests that when comparing E. coli concentrations using IDEXX and qPCR methods, qPCR is the more suitable method to use due to the potential loss of E. coli cells during sample processing while using the IDEXX method (Noble et al. 2010).
Conclusions
In our study, we investigated the occurrence of fecal indicator bacteria in the Sweetwater Wetlands, which allowed us to compare the bacterial resistance in a constructed wetland after wastewater treatment. Looking towards the qPCR results when compared with the inlet samples, the outlet samples all showed a decreased prevalence of bacteria. There is no correlation between temperature and FIB. Turbidity affected our qPCR results and had a positive correlation only to Enterococcus, HF183, and E. coli. Overall reduction rates for intl1 and erm(F) were 67.2% and 13.1%, respectively. E. coli rates decreased by 6.7% from inlet to outlet. Enterococcus and HF183 decreased by 84.0% and 66.6%, respectively. The prevalence of FIB even after treatment suggests that there could be additional factors to resilience, such as ARGs that should be further investigated to understand completely. Future study is needed to explore the pattern of removal efficiencies of ARGs and FIB throughout the entire year. The findings of our study, however, are promising and suggest that constructed wetlands are efficient in decreasing ARGs and fecal indicator bacteria in the water.
References
Ahmed W, Hughes B, Harwood V (2016) Current status of marker genes of Bacteroides and related taxa for identifying sewage pollution in environmental waters. Water 8:231. https://doi.org/10.3390/w8060231
Aly BH, Hamad MS, Mohey M, Amen S (2012) Polymerase chain reaction (PCR) versus bacterial culture in detection of organisms in otitis media with effusion (OME) in children. Indian J Otolaryngol Head Neck Surg 64:51–55. https://doi.org/10.1007/s12070-011-0161-6
Anderson KL, Whitlock JE, Harwood VJ (2005) Persistence and differential survival of fecal indicator bacteria in subtropical waters and sediments. Appl Environ Microbiol 71:3041–3048. https://doi.org/10.1128/AEM.71.6.3041-3048.2005
Badgley BD, Nayak BS, Harwood VJ (2010) The importance of sediment and submerged aquatic vegetation as potential habitats for persistent strains of enterococci in a subtropical watershed. Water Res 44:5857–5866. https://doi.org/10.1016/j.watres.2010.07.005
Bernhard AE, Field KG (2000) A PCR assay to discriminate human and ruminant feces on the basis of host differences in Bacteroides-Prevotella genes encoding 16S rRNA. Appl Environ Microbiol 66:4571–4574. https://doi.org/10.1128/AEM.66.10.4571-4574.2000
Blaustein RA, Pachepsky Y, Hill RL, Shelton DR, Whelan G (2013) Escherichia coli survival in waters: temperature dependence. Water Res 47:569–578. https://doi.org/10.1016/j.watres.2012.10.027
Cai P, Huang Q, Zhang X, Chen H (2006a) Adsorption of DNA on clay minerals and various colloidal particles from an Alfisol. Soil Biol Biochem 38:471–476. https://doi.org/10.1016/j.soilbio.2005.05.019
Cai P, Huang Q-Y, Zhang X-W (2006b) Interactions of DNA with clay minerals and soil colloidal particles and protection against degradation by DNase. Environ Sci Technol 40:2971–2976. https://doi.org/10.1021/es0522985
Chen J, Yu Z, Michel FC, Wittum T, Morrison M (2007) Development and Application of Real-Time PCR Assays for Quantification of erm Genes Conferring Resistance to Macrolides-Lincosamides-Streptogramin B in Livestock Manure and Manure Management Systems. Appl Environ Microbiol 73, 4407–4416. https://doi.org/10.1128/AEM.02799-06
Chen J, Michel FC, Sreevatsan S, Morrison M, Yu Z (2010) Occurrence and persistence of erythromycin resistance genes (erm) and tetracycline resistance genes (tet) in waste treatment systems on swine farms. Microb Ecol 60:479–486. https://doi.org/10.1007/s00248-010-9634-5
Chen J, Wei X-D, Liu Y-S, Ying GG, Liu SS, He LY, Su HC, Hu LX, Chen FR, Yang YQ (2016) Removal of antibiotics and antibiotic resistance genes from domestic sewage by constructed wetlands: optimization of wetland substrates and hydraulic loading. Sci Total Environ 565:240–248. https://doi.org/10.1016/j.scitotenv.2016.04.176
Chung WO, Gabany J, Persson GR, Roberts MC (2002) Distribution of erm(F) and tet(Q) genes in 4 oral bacterial species and genotypic variation between resistant and susceptible isolates. J Clin Periodontol 29:152–158. https://doi.org/10.1034/j.1600-051x.2002.290210.x
Costanzo SD, Murby J, Bates J (2005) Ecosystem response to antibiotics entering the aquatic environment. Mar Pollut Bull 51:218–223. https://doi.org/10.1016/j.marpolbul.2004.10.038
Crecchio C, Ruggiero P, Curci M, Colombo C, Palumbo G, Stotzky G (2005) Binding of DNA from on montmorillonite–humic acids–aluminum or iron hydroxypolymers. Soil Sci Soc Am J 69:834. https://doi.org/10.2136/sssaj2004.0166
Davies J, Davies D (2010) Origins and evolution of antibiotic resistance. Microbiol Mol Biol Rev 74:417–433. https://doi.org/10.1128/MMBR.00016-10
Desmarais TR, Solo-Gabriele HM, Palmer CJ (2002) Influence of soil on fecal indicator organisms in a tidally influenced subtropical environment. Appl Environ Microbiol 68:1165–1172. https://doi.org/10.1128/AEM.68.3.1165-1172.2002
Ebrahimi S, Gabus S, Rohrbach-Brandt E, Hosseini M, Rossi P, Maillard J, Holliger C (2010) Performance and microbial community composition dynamics of aerobic granular sludge from sequencing batch bubble column reactors operated at 20 degrees C, 30 degrees C, and 35 degrees C. Appl Microbiol Biotechnol 87:1555–1568. https://doi.org/10.1007/s00253-010-2621-4
Fang H, Zhang Q, Nie X, Chen B, Xiao Y, Zhou Q, Liao W, Liang X (2017) Occurrence and elimination of antibiotic resistance genes in a long-term operation integrated surface flow constructed wetland. Chemosphere 173:99–106. https://doi.org/10.1016/j.chemosphere.2017.01.027
Faulwetter JL, Gagnon V, Sundberg C, Chazarenc F, Burr MD, Brisson J, Camper AK, Stein OR (2009) Microbial processes influencing performance of treatment wetlands: a review. Ecol Eng 35:987–1004. https://doi.org/10.1016/j.ecoleng.2008.12.030
Frahm E, Obst U (2003) Application of the fluorogenic probe technique (TaqMan PCR) to the detection of Enterococcus spp. and Escherichia coli in water samples. J Microbiol Methods 52:123–131. https://doi.org/10.1016/S0167-7012(02)00150-1
Gelt J (1997) Constructed wetlands: using human ingenuity, Natural Processes to Treat Water, Build Habitat. https://wrrc.arizona.edu/publications/arroyo-newsletter/constructed-wetlands-using-human-ingenuity-natural-processes-treat-wa. Accessed 4 Jan 2019
Gillings M, Boucher Y, Labbate M, Holmes A, Krishnan S, Holley M, Stokes HW (2008) The evolution of class 1 integrons and the rise of antibiotic resistance. J Bacteriol 190:5095–5100. https://doi.org/10.1128/JB.00152-08
Greenway M (2005) The role of constructed wetlands in secondary effluent treatment and water reuse in subtropical and arid Australia. Ecol Eng 25:501–509. https://doi.org/10.1016/j.ecoleng.2005.07.008
Guo W, Vigneswaran S, Ngo H-H, Xing W, Goteti P (2008) Comparison of the performance of submerged membrane bioreactor (SMBR) and submerged membrane adsorption bioreactor (SMABR). Bioresour Technol 99:1012–1017. https://doi.org/10.1016/j.biortech.2007.03.012
Hijosa-Valsero M, Fink G, Schlüsener MP, Sidrach-Cardona R, Martín-Villacorta J, Ternes T, Bécares E (2011) Removal of antibiotics from urban wastewater by constructed wetland optimization. Chemosphere 83:713–719. https://doi.org/10.1016/j.chemosphere.2011.02.004
Howitt JA, Mondon J, Mitchell BD, Kidd T, Eshelman B (2014) Urban stormwater inputs to an adapted coastal wetland: role in water treatment and impacts on wetland biota. Sci Total Environ 485–486:534–544. https://doi.org/10.1016/j.scitotenv.2014.03.101
Kadlec RH, Wallace S (2008) Treatment wetlands, Second edn. CRC Press. https://doi.org/10.1201/9781420012514
Kapoor V, Pitkänen T, Ryu H, Elk M, Wendell D, Santo Domingo JW (2015) Distribution of human-specific Bacteroidales and fecal indicator bacteria in an urban watershed impacted by sewage pollution, determined using RNA- and DNA-based quantitative PCR assays. Appl Environ Microbiol 81:91–99. https://doi.org/10.1128/AEM.02446-14
Karimi B, Ehrampoush MH, Jabary H (2014) Indicator pathogens, organic matter and LAS detergent removal from wastewater by constructed subsurface wetlands. J Environ Health Sci Eng 12:52. https://doi.org/10.1186/2052-336X-12-52
Kim HB, Borewicz K, White BA, Singer RS, Sreevatsan S, Tu ZJ, Isaacson RE (2011) Longitudinal investigation of the age-related bacterial diversity in the feces of commercial pigs. Vet Microbiol 153:124–133. https://doi.org/10.1016/j.vetmic.2011.05.021
Kotzerke A, Sharma S, Schauss K, Heuer H, Thiele-Bruhn S, Smalla K, Wilke BM, Schloter M (2008) Alterations in soil microbial activity and N-transformation processes due to sulfadiazine loads in pig-manure. Environ Pollut 153:315–322. https://doi.org/10.1016/j.envpol.2007.08.020
Le T-H, Ng C, Chen H et al (2016) Occurrences and characterization of antibiotic-resistant bacteria and genetic determinants of hospital wastewater in a tropical country. Antimicrob Agents Chemother 60:7449–7456. https://doi.org/10.1128/AAC.01556-16
Le T-H, Ng C, Tran NH et al (2018) Removal of antibiotic residues, antibiotic resistant bacteria and antibiotic resistance genes in municipal wastewater by membrane bioreactor systems. Water Res 145:498–508. https://doi.org/10.1016/j.watres.2018.08.060
Levy-Booth DJ, Campbell RG, Gulden RH, Hart MM, Powell JR, Klironomos JN, Pauls KP, Swanton CJ, Trevors JT, Dunfield KE (2007) Cycling of extracellular DNA in the soil environment. Soil Biol Biochem 39:2977–2991. https://doi.org/10.1016/j.soilbio.2007.06.020
Liu F, Ying G-G, Tao R, Zhao JL, Yang JF, Zhao LF (2009) Effects of six selected antibiotics on plant growth and soil microbial and enzymatic activities. Environ Pollut 157:1636–1642. https://doi.org/10.1016/j.envpol.2008.12.021
Lu M-C, Chen YY, Chiou M-R, Chen MY, Fan HJ (2016) Occurrence and treatment efficiency of pharmaceuticals in landfill leachates. Waste Manag 55:257–264. https://doi.org/10.1016/j.wasman.2016.03.029
Lu X, Zhang X-X, Wang Z, Huang K, Wang Y, Liang W, Tan Y, Liu B, Tang J (2015) Bacterial pathogens and community composition in advanced sewage treatment systems revealed by metagenomics analysis based on high-throughput sequencing. PLoS One 10:e0125549. https://doi.org/10.1371/journal.pone.0125549
Ludwig W, Schleifer KH (2000) How quantitative is quantitative PCR with respect to cell counts? Syst Appl Microbiol 23:556–562
Maiga von S (2017) Constructed wetlands. In: Glob. Water Pathog. Proj. http://www.waterpathogens.org/book/constructed-wetlands. Accessed 4 Jan 2019
McFeters GA, Stuart DG (1972) Survival of coliform bacteria in natural waters: field and laboratory studies with membrane-filter chambers. Appl Microbiol 24:805–811
McLellan SL, Eren AM (2014) Discovering new indicators of fecal pollution. Trends Microbiol 22:697–706. https://doi.org/10.1016/j.tim.2014.08.002
Meerbergen K, Van Geel M, Waud M et al (2016) Assessing the composition of microbial communities in textile wastewater treatment plants in comparison with municipal wastewater treatment plants. Microbiology Open 6:e00413. https://doi.org/10.1002/mbo3.413
Morató J, Codony F, Sánchez O, Pérez LM, García J, Mas J (2014) Key design factors affecting microbial community composition and pathogenic organism removal in horizontal subsurface flow constructed wetlands. Sci Total Environ 481:81–89. https://doi.org/10.1016/j.scitotenv.2014.01.068
Nayak B, Weidhaas J, Harwood VJ (2015) LA35 poultry fecal marker persistence is correlated with that of indicators and pathogens in environmental waters. Appl Environ Microbiol AEM 81:00444–00415. https://doi.org/10.1128/AEM.00444-15
Nguyen MT, Jasper JT, Boehm AB, Nelson KL (2015) Sunlight inactivation of fecal indicator bacteria in open-water unit process treatment wetlands: modeling endogenous and exogenous inactivation rates. Water Res 83:282–292. https://doi.org/10.1016/j.watres.2015.06.043
Noble RT, Blackwood AD, Griffith JF, McGee CD, Weisberg SB (2010) Comparison of rapid quantitative PCR-based and conventional culture-based methods for enumeration of Enterococcus spp. and Escherichia coli in recreational waters. Appl Environ Microbiol 76:7437–7443. https://doi.org/10.1128/AEM.00651-10
Pruden A, Pei R, Storteboom H, Carlson KH (2006) Antibiotic resistance genes as emerging contaminants: studies in northern Colorado. Environ Sci Technol 40:7445–7450. https://doi.org/10.1021/es060413l
Quiñónez-Dìaz MJ, Karpiscak MM, Ellman ED, Gerba CP (2001) Removal of pathogenic and indicator microorganisms by a constructed wetland receiving untreated domestic wastewater. J Environ Sci Health Part A 36:1311–1320. https://doi.org/10.1081/ESE-100104880
Rachmadi AT, Kitajima M, Pepper IL, Gerba CP (2016) Enteric and indicator virus removal by surface flow wetlands. Sci Total Environ 542:976–982. https://doi.org/10.1016/j.scitotenv.2015.11.001
Ramirez E, Robles E, Bonilla P, Sainz G, Lopez M, de la Cerda JM, Warren A (2005) Occurrence of pathogenic free-living amoebae and bacterial indicators in a constructed wetland treating domestic wastewater from a single household. Eng Life Sci 5:253–258. https://doi.org/10.1002/elsc.200420071
Rogers SW, Donnelly M, Peed L, Kelty CA, Mondal S, Zhong Z, Shanks OC (2011) Decay of bacterial pathogens, fecal indicators, and real-time quantitative PCR genetic markers in manure-amended soils. Appl Environ Microbiol 77:4839–4848. https://doi.org/10.1128/AEM.02427-10
Shelton DR, Pachepsky YA, Kiefer LA, Blaustein RA, McCarty GW, Dao TH (2014) Response of coliform populations in streambed sediment and water column to changes in nutrient concentrations in water. Water Res 59:316–324. https://doi.org/10.1016/j.watres.2014.04.019
Solo-Gabriele HM, Wolfert MA, Desmarais TR, Palmer CJ (2000) Sources of Escherichia coli in a coastal subtropical environment. Appl Environ Microbiol 66:230–237. https://doi.org/10.1128/AEM.66.1.230-237.2000
Staley C, Reckhow KH, Lukasik J, Harwood VJ (2012) Assessment of sources of human pathogens and fecal contamination in a Florida freshwater lake. Water Res 46:5799–5812. https://doi.org/10.1016/j.watres.2012.08.012
Stokes HW, Nesbø CL, Holle M, Bahl MI, Gillings MR, Boucher Y (2006) Class 1 integrons potentially predating the association with tn402-like transposition genes are present in a sediment microbial community. J Bacteriol. 188, 5722–5730. https://doi.org/10.1128/JB.01950-05
Su H-C, Ying G-G, Tao R, Zhang RQ, Zhao JL, Liu YS (2012) Class 1 and 2 integrons, sul resistance genes and antibiotic resistance in Escherichia coli isolated from Dongjiang River, South China. Environ Pollut Barking Essex 1987 169:42–49. https://doi.org/10.1016/j.envpol.2012.05.007
Tandukar S, Sherchand JB, Xue J, Uprety S, Sherchan SP, Bhandari D, Malla B, Ghaju Shrestha R, Parajuli L, Poudel S, Dhital A, Haramoto E (2018) Prevalence and associated risk factors of Giardia duodenalis infection among school-going children in Nepal. Parasitol Res 117:287–293. https://doi.org/10.1007/s00436-017-5706-5
Tao R, Ying G-G, Su H-C, Zhou HW, Sidhu JPS (2010) Detection of antibiotic resistance and tetracycline resistance genes in Enterobacteriaceae isolated from the Pearl rivers in South China. Environ Pollut Barking Essex 1987 158:2101–2109. https://doi.org/10.1016/j.envpol.2010.03.004
Tran NH, Chen H, Reinhard M, Mao F, Gin KYH (2016) Occurrence and removal of multiple classes of antibiotics and antimicrobial agents in biological wastewater treatment processes. Water Res 104:461–472. https://doi.org/10.1016/j.watres.2016.08.040
Tran NH, Gin KY-H, Ngo HH (2015) Fecal pollution source tracking toolbox for identification, evaluation and characterization of fecal contamination in receiving urban surface waters and groundwater. Sci Total Environ 538:38–57. https://doi.org/10.1016/j.scitotenv.2015.07.155
Tu YT, Chiang PC, Yang J, Chen SH, Kao CM (2014) Application of a constructed wetland system for polluted stream remediation. J Hydrol 510:70–78. https://doi.org/10.1016/j.jhydrol.2013.12.015
Underwood JC, Harvey RW, Metge DW, Repert DA, Baumgartner LK, Smith RL, Roane TM, Barber LB (2011) Effects of the antimicrobial sulfamethoxazole on groundwater bacterial enrichment. Environ Sci Technol 45:3096–3101. https://doi.org/10.1021/es103605e
Vaz-Moreira I, Nunes OC, Manaia CM (2014) Bacterial diversity and antibiotic resistance in water habitats: searching the links with the human microbiome. FEMS Microbiol Rev 38:761–778. https://doi.org/10.1111/1574-6976.12062
Vidales-Contreras JA, Gerba CP, Karpiscak MM, Acuna-Askar K, Chaidez-Quiroz C (2006) Transport of coliphage PRD1 in a surface flow constructed wetland. Water Environ. Res. 78, 2253–2260
Wade TJ, Sams E, Brenner KP, Haugland R, Chern E, Beach M, Wymer L, Rankin CC, Love D, Li Q, Noble R, Dufour AP (2010) Rapidly measured indicators of recreational water quality and swimming-associated illness at marine beaches: a prospective cohort study. Environ Health 9:1
Waller VL, Bruland GL (2016) Fecal indicator bacteria dynamics in a surface flow constructed wetland in southwestern Illinois, USA. Wetlands 36:539–546. https://doi.org/10.1007/s13157-016-0763-6
Wang X, Xia Y, Wen X, Yang Y, Zhou J (2014) Microbial community functional structures in wastewater treatment plants as characterized by GeoChip. PLoS One 9:e93422. https://doi.org/10.1371/journal.pone.0093422
Webster LF, Graves DA, Eargle DA, Chestnut DE, Gooch JA, Fulton MH (2013) Assessment of animal impacts on bacterial water quality in a South Carolina, USA tidal creek system. Environ Monit Assess 185:7749–7756. https://doi.org/10.1007/s10661-013-3132-4
Wheeler Alm E, Burke J, Spain A (2003) Fecal indicator bacteria are abundant in wet sand at freshwater beaches. Water Res 37:3978–3982. https://doi.org/10.1016/S0043-1354(03)00301-4
Wu CY, Liu JK, Cheng SH, Surampalli DE, Chen CW, Kao CM (2010) Constructed wetland for water quality improvement: a case study from Taiwan. Water Sci Technol J Int Assoc Water Pollut Res 62:2408–2418. https://doi.org/10.2166/wst.2010.492
Wu H, Zhang J, Ngo HH, Guo W, Hu Z, Liang S, Fan J, Liu H (2015) A review on the sustainability of constructed wetlands for wastewater treatment: design and operation. Bioresour Technol 175:594–601. https://doi.org/10.1016/j.biortech.2014.10.068
Wu S, Carvalho PN, Müller JA, Manoj VR, Dong R (2016) Sanitation in constructed wetlands: a review on the removal of human pathogens and fecal indicators. Sci Total Environ 541:8–22. https://doi.org/10.1016/j.scitotenv.2015.09.047
Xing L, Yu H, Qi J, Jiang P, Sun B, Cui J, Ou C, Chang W, Hu Q (2015) ErmF and ereD are responsible for erythromycin resistance in riemerella anatipestifer. PLoS One 10:e0131078. https://doi.org/10.1371/journal.pone.0131078
Xue J, Caton K, Sherchan SP (2018a) Comparison of next-generation droplet digital PCR with quantitative PCR for enumeration of Naegleria fowleri in environmental water and clinical samples. Lett Appl Microbiol 67:322–328. https://doi.org/10.1111/lam.13051
Xue J, Feng Y (2018) Determination of adsorption and desorption of DNA molecules on freshwater and marine sediments. J Appl Microbiol 124:1480–1492. https://doi.org/10.1111/jam.13739
Xue J, Feng Y, Wijesinghe RU, Wood CW (2018b) Using Bacteroidales genetic markers to assess fecal pollution sources in coastal waters. Water Environ J 32:84–93. https://doi.org/10.1111/wej.12302
Xue J, Lamar FG, Zhang B, Lin S, Lamori JG, Sherchan SP (2017) Quantitative assessment of Naegleria fowleri and fecal indicator bacteria in brackish water of Lake Pontchartrain, Louisiana. Sci Total Environ 622–623:8–16. https://doi.org/10.1016/j.scitotenv.2017.11.308
Xue J, Lin S, Lamar FG, Lamori JG, Sherchan S (2018c) Assessment of fecal pollution in Lake Pontchartrain, Louisiana. Mar Pollut Bull 129:655–663. https://doi.org/10.1016/j.marpolbul.2017.10.053
Xue J, Schmitz BW, Caton K, Zhang B, Zabaleta J, Garai J, Taylor CM, Romanchishina T, Gerba CP, Pepper IL, Sherchan SP (2019) Assessing the spatial and temporal variability of bacterial communities in two Bardenpho wastewater treatment systems via Illumina MiSeq sequencing. Sci Total Environ 657:1543–1552. https://doi.org/10.1016/j.scitotenv.2018.12.141
Yi X, Tran NH, Yin T, He Y, Gin KYH (2017) Removal of selected PPCPs, EDCs, and antibiotic resistance genes in landfill leachate by a full-scale constructed wetlands system. Water Res 121:46–60. https://doi.org/10.1016/j.watres.2017.05.008
Yu WH, Li N, Tong DS, Zhou CH, Lin CX(C), Xu CY (2013) Adsorption of proteins and nucleic acids on clay minerals and their interactions: a review. Appl Clay Sci 80–81:443–452. https://doi.org/10.1016/j.clay.2013.06.003
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Responsible editor: Diane Purchase
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 24 kb)
Rights and permissions
About this article
Cite this article
Lamori, J.G., Xue, J., Rachmadi, A.T. et al. Removal of fecal indicator bacteria and antibiotic resistant genes in constructed wetlands. Environ Sci Pollut Res 26, 10188–10197 (2019). https://doi.org/10.1007/s11356-019-04468-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-019-04468-9