Abstract
The current industrial and human activities scenario has accelerated the widespread use of endocrine-disrupting compounds (EDCs), which can be found in everyday products, including plastic containers, bottles, toys, cosmetics, etc., but can pose a severe risk to human health and the environment. In this regard, fungal bioremediation appears as a green and cost-effective approach to removing pollutants from water resources. Besides, immobilizing fungal cells onto nanofibrous membranes appears as an innovative strategy to improve remediation performance by allowing the adsorption and degradation to occur simultaneously. Herein, we developed a novel nanostructured bioremediation platform based on polyacrylonitrile nanofibrous membrane (PAN NFM) as supporting material for immobilizing an endophytic fungus to remove bisphenol A (BPA), a typical EDC. The endophytic strain was isolated from Handroanthus impetiginosus leaves and identified as Phanerochaete sp. H2 by molecular methods. The successful assembly of fungus onto the PAN NFM surface was confirmed by scanning electron microscopy (SEM). Compared with free fungus cells, the PAN@H2 NFM displayed a high BPA removal efficiency (above 85%) at an initial concentration of 5 ppm, suggesting synergistic removal by simultaneous adsorption and biotransformation. Moreover, the biotransformation pathway was investigated, and the chemical structures of fungal metabolites of BPA were identified by ultra-high performance liquid chromatography - high-resolution mass (UHPLC-HRMS) analysis. In general, our results suggest that by combining the advantages of enzymatic activity and nanofibrous structure, the novel platform has the potential to be applied in the bioremediation of varied EDCs or even other pollutants found in water resources.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The awareness of the adverse effects that endocrine-disrupting compounds (EDCs) can pose on human health and the environment has increased in recent years (Magi et al. 2010; Mondal et al. 2021; Naveira et al. 2021; Alabi et al. 2021). Bisphenol A (BPA, 4,4-isopropylidenediphenol), a typical EDC, is commonly used to produce a wide variety of everyday materials, including food and beverage containers, kitchen utensils, children’s toys, and reusable plastic bottles (Torres-García et al. 2022). Several studies have reported that BPA can block the action of natural hormones, impairing normal growth, metabolism, and reproduction (Naveira et al. 2021). The chemical properties of BPA, such as low water solubility, low octanol-water partition coefficient, high soil-water portioning coefficient, and long half-life in sediments and soils, contribute to its bioaccumulation and biomagnification along the trophic chains, thereby causing long-term adverse effects to human health and the environment (Mansilha et al. 2013; Shi et al. 2018; Liao and Kannan 2019). Until now, efficient methods for removing low-concentration BPA from wastewater are scarce (Zielinska et al. 2019; Oliveira et al. 2020; Dhangar and Kumar 2020). For instance, conventional physical and chemical remediation technologies are usually high-cost and generate undesired byproducts (Sharma et al. 2018). Therefore, efficient, reliable, green, and economical approaches for BPA removal from water resources are in high demand.
Biological wastewater treatment that takes advantage of the natural role of organisms to transform or alter (through metabolic or enzymatic action) the structure of hazardous contaminants has proven to be a sustainable and cost-effective way to remediate EDCs (Roccuzzo et al. 2021; Zhuo and Fan 2021). Filamentous fungi have been used to biotransform several organic compounds by their intra- and extracellular enzymes, or through oxidizing radicals production (Chen et al. 2023). Therefore, many recalcitrant compounds can be transformed, even those with low solubility (Purohit et al. 2018; El-Gendi et al. 2021). More recently, processes based on the use of fungi in BPA-contaminated areas have been successfully reported (Delius et al. 2022; Wang et al. 2022).
Despite the advantages of fungal bioremediation, the use of free fungi cells faces challenges since mechanical disturbances may disrupt mycelia growth, and the mycelia development during the water treatment may also lead to operational issues such as clogging, nutrient addition, foaming, biomass aging, and microbial contamination (Mir-Tutusaus et al. 2018; Ahn et al. 2020; George et al. 2022). In this regard, fungal immobilization on supports provides a potential tool to overcome such drawbacks, as it prevents mycelial dispersion, enables easier solid-liquid separation, and improves the fungi enzymatic activity, which optimize the ability of fungi in any bioremediation process (Beltrán-Flores et al. 2022; Alam et al. 2023) and even heavy metal pollution (Chen et al. 2022).
An ideal support for fungal immobilization should be stable and provide a good surface for the fungus attachment and growth, along with large surface area/porosity, which can potentially benefit the cells immobilization and the mass transfer between media and entrapped cells (Rodríguez Couto 2009; Yang et al. 2022). In this scenario, electrospun nanofibrous membranes (NFMs) stand out as flexible and free-standing substrates for microbial cell immobilization due to their appealing properties such as large specific surface area, high porosity, large liquid permeability, cost-effectiveness, and easily adjustable properties (Balusamy et al. 2019; Mercante et al. 2021). Moreover, the fungal immobilization onto NFMs may further enhance the removal of recalcitrant compounds from contaminated areas by associating the adsorption potential of NFMs with the enzymatic degradation ability of fungi. Electrospun NFMs have been successfully applied as support for drug delivery, filtration, enzyme immobilization, and biosensors, among other applications. However, their use as support for fungal immobilization to enhance EDCs degradation has not been fully explored yet.
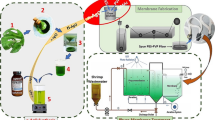
Herein, we demonstrate the feasibility of using an endophytic fungus immobilized on NFM as a nanohybrid platform for BPA bioremoval, as illustrated in Scheme 1. Specifically, polyacrylonitrile (PAN) electrospun NFM was used as supporting material to immobilize the endophytic fungus Phanerochaete sp. H2 isolated from Handroanthus impetiginosus leaves. The performance of the PAN@H2 NFM for BPA removal was characterized, and UHPLC-HRMS analysis allowed the chemical characterization of the BPA fungal metabolites. Our results contribute to elucidating the biotransformation pathway of BPA by endophytic fungus strain.
Materials and methods
Achievement and molecular identification of endophytic fungus
The endophytic fungus was recovered, as previously described by our group (do Nascimento et al. 2020), from Handroanthus impetiginosus (Mart. ex DC.) Mattos leaves, which were collected in Alfenas, Minas Gerais, Brazil (S21°18’49.15”, W45°57’28.53”) and identified by Dr. Lúcia G. Lohmann (Botanical Department of the Bioscience Institute of the University of São Paulo). The study with the isolated fungus was registered in the Brazilian System for the Management of Genetic Heritage and Associated Traditional Knowledge (SisGen) under code A6F76F0.
The endophytic fungus with promising features in the bioremediation screening was identified by sequencing the ITS (Internal Transcribed Spacer) region. Glass microspheres (425–600 μm in diameter, Sigma) were used for the extraction of DNA of the endophytic strain through physical lysis of the mycelium (Aamir et al. 2015). The ITS1-5.8 S-ITS2 region was amplified using ITS1 (5′-CCG TAG GTG AAC CTG CGG − 3′) and ITS4 primers (5′-TCC TCC GCT TAT TGA TAT GC-3′) (White et al. 1990). The amplification primers using the Big Dye Kit (Life Technologies) in the ABI 3500 Genetic Analyzer XL system carried out DNA fragments sequencing. Amplification of the D1/D2 domain of the LSU rRNA gene was achieved using the primers ITS1-F (TCCGTAGGTGAACCTGCGG) and NL-4 (5′-TCCTCCGCTTATTGATATGC-3′).
The editor BioEdit software assembled the forward and reverse sequences of fungus in contigs (Hall 1999). The BLAST program (National Center for Biotechnology Information) compared the sequences obtained with reference sequences from GenBank (Altschul et al. 1990). GenBank database received our sequence under the accession number MK737061. Based on the identity score, the closest reference sequence of endophytic fungus was obtained and used for further phylogeny analysis. CLUSTALX function aligned the sequences (Thompson et al. 1997), and the phylogenetic tree was constructed using the MEGA (Molecular Evolutionary Genetics Analysis) 4.0 software (Tamura et al. 2007).
The neighbor-Joining algorithm was used to establish the evolutionary relationship, and distances were calculated with the Kimura 2-parameter model (Kimura 1980). The statistical support of nodes was estimated by bootstrap analysis with 1000 replications (Felsenstein 1985). The analysis involved 32 nucleotide sequences. All positions containing gaps and missing data were eliminated. There was a total of 380 positions in the final dataset. The Maximum Composite Likelihood method computed the evolutionary distances. The rate variation among sites was modeled with a gamma distribution (shape parameter = 1). The obtained sequence was documented on the GenBank database under the accession number MK737061.
Polyacrylonitrile nanofibrous membrane preparation and fungus immobilization
Polyacrylonitrile (PAN, Mw = 120,000 g mol− 1) and N,N-dimethylformamide (DMF, anhydrous, 99.8%) were purchased from Sigma-Aldrich.
PAN electrospun nanofiber was obtained according to previously reported (Facure et al. 2022). The electrospinning solution was prepared by dissolving PAN (10% w/v) in DMF and stirring for 6 h at room temperature. The nanofibrous membrane was obtained using an electrospinning apparatus at an applied voltage of 12 kV, a feed rate of 0.5 mL h− 1, and a working distance of 12 cm. The nanofibrous membrane was collected in an aluminum foil, from which it could be easily removed after the completion of the electrospinning process.
To prepare the PAN@H2 membranes, 15.6 mg of PAN NFM was added to a Petri dish at the same time as the endophytic strain, which was cultured in Petri dishes containing potato dextrose agar (PDA, Kasvi, Curitiba, Brazil) at 28ºC for seven days. The successful immobilization of fungus onto PAN NFM was confirmed by scanning electron microscopy analysis (SEM, JOEL JSM-6510).
Biotransformation procedures
H2 free cells and PAN@H2 NFM were used in the biotransformation of BPA (Sigma Aldrich, ≥ 99%). 100 mL-Erlenmeyer flasks containing 50 mL of medium consisting of 0.18% glucose (Synth, São Paulo, Brazil), 0.06% peptone (Merck, Darmstadt, Germany), and 0.04% yeast extract (Acumedia, Baltimore, USA), pH 6.0, received five disks (5 mm) of H2 or PAN@H2 and BPA as a solution in dimethylsulfoxide (Synth, São Paulo, Brazil). Three controls were used with the following compositions: (i) culture medium, tetrahydrofuran, and fungus, with no substrate (BPA); (ii) culture medium and substrate but no fungus; and (iii) culture medium only.
The biotransformation assays were carried out at 28 °C using three different BPA concentrations: 5, 10, and 20 ppm. The absorbance of the supernatant withdrawn at different time intervals (0, 4, 12, 16, 20, and 24 h) was measured at the maximum absorbance wavelength for the BPA (λmax = 266 nm) using a Quimis Model Q780U. The experiments were carried out in triplicate and the results were reported as average with standard deviation. The percentage of biotransformation was calculated from the difference between initial and final values using the following equation:
where A0 is the absorbance at 0 h, and At is the absorbance at t hour.
Metabolite identification by UHPLC-HRMS
The monitoring of the biotransformation of BPA into its derivatives, as well as the comparison of the extracts of biotransformation and controls, were initially analyzed by Thin-layer chromatography (TLC). Previously, H2 and PAN@H2 were separated by filtration, and the fermentation broths were extracted with ethyl acetate (Synth, São Paulo, Brazil). Crude extracts of biotransformation and control experiments were achieved after solvent evaporation and dissolved in methanol for their analysis by TLC (Merck, Darmstadt, Germany). The mobile phase consisted of hexane/ethyl acetate 40:60 (v/v), and ultraviolet radiation (254 nm) was employed to observe the plates.
The crude extracts were also analyzed by ultra-high performance liquid chromatography - high-resolution mass (UHPLC-HRMS). UHPLC-HRMS apparatus contained an electrospray ionization (ESI) source and an Orbitrap technology analyzer. The flow rate was 400 µL.min-1, and the gradient elution system was 5 to 100% methanol (HPLC grade Tedia, Rio de Janeiro, Brazil) in water for over 30 min. A C18 column (ACE 150 mm×4.6 mm×3 μm) was used at a spectrometer operating at both positive and negative modes. The column temperature was set at 30ºC. The following parameters were used: scanning range of 120–1200 m/z to full MS, ESI MS resolution of 70.000 with lock mass, microbeam of 1, and maximum injection time of 250 ms. The parameters of the ESI ionization source were as follows: gas flow rate of 30 L/min; auxiliary gas flow rate of 10 L/min; positive voltage spray mode of 3.6 kV; negative voltage spray mode of 3.2 kV; and Slens level of 55. Nitrogen gas was used as a nebulizer in the collision cell. The mass spectra were obtained and processed using Xcalibur software (Thermo Fisher Scientific). The BPA metabolites were identified based on the accurate masses and analysis of their fragmentation patterns, shown in the MS/MS spectra.
Results
Endophytic fungi isolated from Handroanthus impetiginosus leaves (do Nascimento et al. 2020) were assayed for their abilities in the degradation of BPA. The endophytic fungus (H2) that displayed the best results in the biotransformation screening was identified through molecular techniques. The phylogenetic tree generated by the Neighbor-Joining approach based on ITS sequences of H2 and related species is shown in Figure S1 (Online Resource). The endophyte H2 clustered with sequences of Phanerochaete species, and it was identified as Phanerochaete sp. H2. Additionally, the identification of the strain was confirmed by analysis of its morphology.
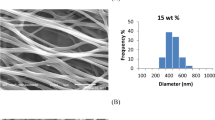
After identifying the selected endophytic strain, a polyacrylonitrile electrospun nanofiber (PAN NFM) produced by electrospinning was employed as support for Phanerochaete sp. H2 immobilization. The formation of the fungus biofilm onto PAN NFM was analyzed by scanning electron microscopy (SEM), whose images are shown in Fig. 1. The PAN NFM (Fig. 1A) presented a bead-free and smooth fibrous nature with an average diameter of 202 ± 32 nm. Figure 1B reveals the biofilm’s formation after the fungus growth (PAN@H2 NFM).
The potential of PAN@H2 NFM was then evaluated for bioremoval of BPA. For this, BPA solutions at different concentrations (5, 10, and 20 ppm) were subjected to bioassays at various time durations, and the resulting solutions were subjected to UV absorbance measurements. The performance of both free and immobilized cells regarding BPA percentage removal was compared, as shown in Fig. 2. As can be seen, in the first 4 h, free and immobilized cells practically showed the same removal efficiency. However, after 24 h, the removal rates of the PAN@H2 NFM were generally higher than that of the free cells. The BPA removal rates of PAN@H2 NFM reached at 192 h the maximum of 88, 79, and 74% at 5, 10, and 20 ppm of BPA, respectively.
At sequence, comparative analysis of the TLC chemical profiles of the ethyl acetate extracts obtained from the biotransformation and control experiments showed that free Phanerochaete sp. H2 cells can transform BPA after eight days of incubation. No variation was detected in the pH of the culture media, which was maintained at 6.5 throughout the biotransformation. To characterize the chemical structures of the derivatives produced from the BPA biotransformation by the PAN@H2 NFM, the ethyl acetate crude extract was analyzed by UHPLC-HRMS. As shown in Fig. 3, the peak related to BPA appears as a broad peak of low intensity at around 24 min. In addition, three peaks related to more polar fungal BPA metabolites were also detected.
Critical analysis of the ESI mass spectra (Fig. 4) of fungal metabolites of BPA allowed the chemical characterization of the biotransformation process. The ESI mass spectrum (Fig. 4C) of the main derivative of BPA (3) contained a base peak at m/z 243.1012 [M-H]− (calculated for [M-H]− 243.1027) that corroborates with the molecular formula C15H16O3. The MS/MS spectrum of the compound 3 shows some main fragments displayed in Scheme 2. The ion at m/z 227 resulted from the loss of CH4. The ion at m/z 149 was due to the loss of phenol. Subsequent cleavage of the hydroxyphenyl-alkyl bond of the ion at m/z 149 generated the ion at m/z 109. Alternatively, if initial ionization occurs at the mono-hydroxylated ring, the ion at m/z 133 can be formed after the loss of catechol. Cleavage of the hydroxyphenyl-alkyl bond of the ion at m/z 133 generated the ion at m/z 93. All the fragmentations were proposed according to previously reported studies that employed labeled bisphenols (Magi et al. 2010; Zhao et al. 2016). The fragmentation mechanism for alkylphenols, such as BPA, is charge-remote fragmentation, a type of gas phase dissociation where the bond cleavage occurs far from the charge site because of the charge stability. Therefore, it does not move into the fragmenting portion of the ion (Magi et al. 2010). The proposition of the chemical structure of the diagnostic ion at m/z 149 confirmed the transformation at the phenolic ring of BPA by Phanerochaete sp. H2.
At the same rationale for mass data analysis of derivative 3, the chemical structures of derivatives 1 and 2 were proposed. The MS/MS spectra of 1 (Fig. 4A) and 2 (Fig. 4B) showed base peaks at m/z 229.0857 [M-H]− (calculated for [M-H]− 229.0870) and m/z 241.0857 [M-H]− (calculated for [M-H]− 241.0870), respectively. In the proposed fragmentation mechanism of 1 and 2, the radical ions at m/z 214 and 226, respectively, are formed by the loss of methyl radical. The fragments at m/z 135 and 133, which have been seen at spectra of 1 and 2, respectively, are due to the loss of phenol from 1 or ortho-benzoquinone from 2. Cleavages of the hydroxyphenyl-alkyl bonds of the ions at m/z 135 and 133 generated the ion at m/z 109 and 93, respectively.
Based on the above results, the biotransformation of BPA by Phanerochaete sp. H2 led to three metabolites, whose chemical structures are depicted in Scheme 3.
Discussion
Our research group has previously studied different fungi strains’ capabilities in transforming compounds (Silva Conceição et al. 2021; Pereira dos Santos et al. 2022). A wide variety of microorganisms are helpful in biocatalysis as they can transform several chemicals, providing many catalysts in small volumes and high turnover rates of enzymes and cofactors (Wu et al. 2021). Endophytic microorganisms asymptomatically occur in the internal vegetal tissues (Bacon and White 2000). Specialized studies have provided evidence that the interaction between endophytic community members can play a significant role in the onset of enzymes (Qi-he et al. 2009). Endophytes are an unexplored and least studied group of microbes (Elango et al. 2020), and their employment as enzymatic sources for bioremediation processes are very promising for discovering new enzymes (dos Santos and de Oliveira Silva 2019; Pietro-Souza et al. 2020).
Herein, we assayed endophytic fungi isolated from Handroanthus impetiginosus leaves as potential biocatalyst capable of efficiently removing BPA from aqueous media. The endophyte Phanerochaete sp. H2 has been selected and identified through sequencing of its ITS region, which has been chosen as the official barcode for molecular identification of fungi due to its ease of amplification, widespread use, and appropriately large barcode gap (Raja et al. 2017). Fungi from the Phanerochaete genus belong to the Basidiomycota phylum, and they have been used for direct BPA degradation or through the application of their extracellular enzymes (Cajthaml et al. 2009; Gassara et al. 2013; Wang et al. 2014). The Basidiomycota phylum includes ligninolytic species, also known as white-rot fungi, of great importance due to their different applications in the biodegradation of phenolic compounds (Martínková et al. 2016). A recent survey highlighted some Basidiomycota fungi used in BPA degradation (Torres-García et al. 2022).
Fungal immobilization on nanostructured platforms such as the ones proposed here holds the potential to make bioremediation processes more robust due to the prevention of foaming, biomass aging, and microbial contamination and may enhance the enzymatic activity (Spina et al. 2018). To improve the BPA bioremoval efficiency, Phanerochaete sp. H2 was immobilized onto PAN NFM. SEM images confirmed the biofilm formation. The formed biofilm was interlocked with nanofibers forming a cohesive structure typical of this type of structure (Hu et al. 2019). Moreover, it was possible to observe the presence of porosity in the fungal biofilm, which contributes to the increase of surface area and mass transfer (Hu et al. 2019; Madadi and Bester 2021).
The performance of both free and immobilized cells regarding BPA percentage removal was compared. The results showed that the PAN@H2 had superior removal properties, which can be ascribed to the adsorption-catalysis synergy. The enhanced performance of the PAN@H2 NFM could be attributed to the synergistic adsorption/biotransformation of BPA. Decreased removal efficiency with increasing BPA concentration may be related to the partial blocking of the membrane pores, thereby reducing the liquid permeability and the removal rates (Zdarta et al. 2022).
Despite the exceptional fungal ability to flourish in the presence of recalcitrant compounds, the mechanism of biodegradation by fungi remains poorly understood. Elucidating the metabolic pathway that fungi employ to transform a hazardous compound into an environmentally friendly one helps to rationalize the directions of a bioremediation procedure. Therefore, for future applications, evaluating the mechanisms of biodegradation that microorganisms use during the remediation is essential (Sosa-Martínez et al. 2020). Negative ESI spectra of three BPA derivatives (1–3) and their respective MS/MS spectra were interpreted.The fragmentation pathways were proposed, which permitted the characterization of the chemical structures. Modifications at the phenolic and alkyl chains were verified at the transformation of the BPA scaffold by fungus.
The Ecological Structure Activity Relationships (ECOSAR) program based on quantitative structure-activity relationship (QSAR) has been widely applied to evaluate the toxicity of degradation products (Khan et al. 2019). A recent study reported the removal of bisphenol E (an analog of BPA) by ferrate(VI) in water (Tian et al. 2022). In sequence, the acute and chronic toxicity of its degradation products to three aquatic species (fish, daphnia, and green algae) were evaluated by the ECOSAR program. Among the derivatives of bisphenol E, the authors reported one with the same chemical structure we identified as the BPA derivative 1, which was formed after the oxidation of bisphenol compounds to produce di-hydroxylated products. Interestingly, the toxicity of product 1 was decreased compared with the parent BPE molecule (Tian et al. 2022).
Conclusions
In the present study, an endophytic fungus (Panerochaete sp. H2) isolated from H. impetiginosus leaves was screened as a potential biocatalyst capable of efficient bioremediation of BPA from aqueous media. To improve the bioremediation efficiency, the fungus was immobilized onto PAN NFM. The results showed that the PAN@H2 had superior removal properties compared to the free cells, which can be ascribed to the adsorption-catalysis synergy. The chemical structures of the three BPA derivatives (1–3) were identified based on the proposed fragmentation pathways and accurate mass data. Modifications at the phenolic and alkyl chains were verified at the transformation of the BPA scaffold by fungus. The characterization of BPA derivatives contributed to understanding the chemistry involved in its biodegradation process, which is mandatory for rational planning of its bioremediation, prediction of the end products, and guidance of future application of the methodology developed herein.
Data Availability
The datasets generated during and/or analysed during the current study are available from the corresponding author upon reasonable request.
References
Aamir S, Sutar S, Singh SK, Baghela A (2015) A rapid and efficient method of fungal genomic DNA extraction, suitable for PCR based molecular methods. Plant Pathol Quar 5:74–81. https://doi.org/10.5943/ppq/5/2/6
Ahn H, Rehman JU, Kim T et al (2020) Fungal mycelia functionalization with halloysite nanotubes for hyphal spreading and sorption behavior regulation: a new bio-ceramic hybrid for enhanced water treatment. Water Res 186:116380. https://doi.org/10.1016/j.watres.2020.116380
Alabi OA, Ologbonjaye KI, Sorungbe AA et al (2021) Bisphenol A-induced alterations in different stages of spermatogenesis and systemic toxicity in albino mice (Mus musculus). J Heal Pollut 11. https://doi.org/10.5696/2156-9614-11.29.210307
Alam R, Mahmood RA, Islam S et al (2023) Understanding the biodegradation pathways of azo dyes by immobilized white-rot fungus, Trametes hirsuta D7, using UPLC-PDA-FTICR MS supported by in silico simulations and toxicity assessment. Chemosphere 313:137505. https://doi.org/10.1016/j.chemosphere.2022.137505
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Bacon CW, White JF (2000) Microbial endophytes. Marcel Dekker, New York, USA
Balusamy B, Sarioglu OF, Senthamizhan A, Uyar T (2019) Rational design and development of Electrospun Nanofibrous Biohybrid Composites. ACS Appl Bio Mater. https://doi.org/10.1021/acsabm.9b00308
Beltrán-Flores E, Pla-Ferriol M, Martínez-Alonso M et al (2022) Fungal bioremediation of agricultural wastewater in a long-term treatment: biomass stabilization by immobilization strategy. J Hazard Mater 439:129614. https://doi.org/10.1016/j.jhazmat.2022.129614
Cajthaml T, Křesinová Z, Svobodová K, Möder M (2009) Biodegradation of endocrine-disrupting compounds and suppression of estrogenic activity by ligninolytic fungi. Chemosphere 75:745–750. https://doi.org/10.1016/j.chemosphere.2009.01.034
Chen L, Zhang X, Zhang M et al (2022) Removal of heavy-metal pollutants by white rot fungi: mechanisms, achievements, and perspectives. J Clean Prod 354:131681. https://doi.org/10.1016/j.jclepro.2022.131681
Chen S, Zhu M, Guo X et al (2023) Coupling of Fenton reaction and white rot fungi for the degradation of organic pollutants. Ecotoxicol Environ Saf 254:114697. https://doi.org/10.1016/j.ecoenv.2023.114697
Delius J, Emmerich M, Özyurt V, Hamscher G (2022) Biotransformation of Tetracyclines by Fungi: Challenges and Future Research Perspectives. J Agric Food Chem 70:1454–1460. https://doi.org/10.1021/acs.jafc.1c05121
Dhangar K, Kumar M (2020) Tricks and tracks in removal of emerging contaminants from the wastewater through hybrid treatment systems: a review. Sci Total Environ 738:140320. https://doi.org/10.1016/j.scitotenv.2020.140320
do Nascimento JS, Silva FM, Magallanes-Noguera CA et al (2020) Natural trypanocidal product produced by endophytic fungi through co-culturing. Folia Microbiol (Praha) 65:323–328. https://doi.org/10.1007/s12223-019-00727-x
dos Santos VHP, de Oliveira Silva E (2019) Endophytic fungi from the brazilian flora and their employment in biotransformation reactions. Quim Nova 42:784–791. https://doi.org/10.21577/0100-4042.20170380
El-Gendi H, Saleh AK, Badierah R et al (2021) A comprehensive insight into fungal enzymes: structure, classification, and their role in mankind’s Challenges. J Fungi 8:23. https://doi.org/10.3390/jof8010023
Elango D, Manikandan V, Jayanthi P et al (2020) Selection and characterization of extracellular enzyme production by an endophytic fungi aspergillus sojae and its bio-efficacy analysis against cotton leaf worm, Spodoptera litura. Curr Plant Biol 23:100153. https://doi.org/10.1016/j.cpb.2020.100153
Facure MHM, Mercante LA, Correa DS (2022) Polyacrylonitrile/Reduced Graphene Oxide Free-Standing Nanofibrous membranes for detecting endocrine disruptors. ACS Appl Nano Mater 5:6376–6384. https://doi.org/10.1021/acsanm.2c00484
Felsenstein J (1985) Confidence limits on phylogenies: an Approach using the bootstrap. Evol (N Y) 39:783. https://doi.org/10.2307/2408678
Gassara F, Brar SK, Verma M, Tyagi RD (2013) Bisphenol a degradation in water by ligninolytic enzymes. Chemosphere 92:1356–1360. https://doi.org/10.1016/j.chemosphere.2013.02.071
George J, Anand SS, Senthil Kumar P et al (2022) Biocatalytic polymeric membranes to decrease biofilm fouling and remove organic contaminants in wastewater: a review. Springer International Publishing
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hu MX, Li JN, Guo Q et al (2019) Probiotics Biofilm-Integrated Electrospun Nanofiber membranes: a new starter culture for fermented milk production. J Agric Food Chem 67:3198–3208. https://doi.org/10.1021/acs.jafc.8b05024
Khan K, Benfenati E, Roy K (2019) Consensus QSAR modeling of toxicity of pharmaceuticals to different aquatic organisms: ranking and prioritization of the DrugBank database compounds. Ecotoxicol Environ Saf 168:287–297. https://doi.org/10.1016/j.ecoenv.2018.10.060
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120. https://doi.org/10.1007/BF01731581
Liao C, Kannan K (2019) Species-specific accumulation and temporal trends of bisphenols and benzophenones in mollusks from the chinese Bohai Sea during 2006–2015. Sci Total Environ 653:168–175. https://doi.org/10.1016/j.scitotenv.2018.10.271
Madadi R, Bester K (2021) Fungi and biochar applications in bioremediation of organic micropollutants from aquatic media. Mar Pollut Bull 166:112247. https://doi.org/10.1016/j.marpolbul.2021.112247
Magi E, Scapolla C, Di Carro M, Liscio C (2010) Determination of endocrine-disrupting compounds in drinking waters by fast liquid chromatography-tandem mass spectrometry. J Mass Spectrom 45:1003–1011. https://doi.org/10.1002/jms.1781
Mansilha C, Silva P, Rocha S et al (2013) Bisphenol A migration from plastic materials: direct insight of ecotoxicity in Daphnia magna. Environ Sci Pollut Res 20:6007–6018. https://doi.org/10.1007/s11356-013-1614-0
Martínková L, Kotik M, Marková E, Homolka L (2016) Biodegradation of phenolic compounds by Basidiomycota and its phenol oxidases: a review. Chemosphere 149:373–382. https://doi.org/10.1016/j.chemosphere.2016.01.022
Mercante LA, Pavinatto A, Pereira TS et al (2021) Nanofibers interfaces for biosensing: design and applications. Sens Actuators Rep 3:100048. https://doi.org/10.1016/j.snr.2021.100048
Mir-Tutusaus JA, Baccar R, Caminal G, Sarrà M (2018) Can white-rot fungi be a real wastewater treatment alternative for organic micropollutants removal? A review. Water Res 138:137–151. https://doi.org/10.1016/j.watres.2018.02.056
Mondal A, Burchat N, Sampath H (2021) Palmitate exacerbates bisphenol a toxicity via induction of ER stress and mitochondrial dysfunction. Biochim Biophys Acta - Mol Cell Biol Lipids 1866:158816. https://doi.org/10.1016/j.bbalip.2020.158816
Naveira C, Rodrigues N, Santos FS et al (2021) Acute toxicity of Bisphenol A (BPA) to tropical marine and estuarine species from different trophic groups. Environ Pollut 268:115911. https://doi.org/10.1016/j.envpol.2020.115911
Oliveira JMS, de Lima e Silva MR, Issa CG et al (2020) Intermittent aeration strategy for azo dye biodegradation: a suitable alternative to conventional biological treatments? J Hazard Mater 385:121558. https://doi.org/10.1016/j.jhazmat.2019.121558
Pereira dos Santos VH, Luiz JHH, dos Anjos JP, de Oliveira Silva E (2022) Oxidative potential of two brazilian endophytic fungi from Handroanthus impetiginosus towards progesterone. Steroids 187:109101. https://doi.org/10.1016/j.steroids.2022.109101
Pietro-Souza W, de Campos Pereira F, Mello IS et al (2020) Mercury resistance and bioremediation mediated by endophytic fungi. Chemosphere 240:124874. https://doi.org/10.1016/j.chemosphere.2019.124874
Purohit J, Chattopadhyay A, Biswas MK, Singh NK (2018) Mycoremediation of agricultural soil: Bioprospection for Sustainable Development. In: Prasad R (ed) Mycoremediation and environmental sustainability. Springer International Publishing, Cham, Switzerland, pp 91–120
Qi-he C, Jing L, Hai-feng Z et al (2009) The betulinic acid production from betulin through biotransformation by fungi. Enzyme Microb Technol 45:175–180. https://doi.org/10.1016/j.enzmictec.2009.06.005
Raja HA, Miller AN, Pearce CJ, Oberlies NH (2017) Fungal identification using molecular tools: a primer for the Natural Products Research Community. J Nat Prod 80:756–770. https://doi.org/10.1021/acs.jnatprod.6b01085
Roccuzzo S, Beckerman AP, Trögl J (2021) New perspectives on the bioremediation of endocrine disrupting compounds from wastewater using algae-, bacteria- and fungi-based technologies. Int J Environ Sci Technol 18:89–106. https://doi.org/10.1007/s13762-020-02691-3
Rodríguez Couto S (2009) Dye removal by immobilised fungi. Biotechnol Adv 27:227–235. https://doi.org/10.1016/j.biotechadv.2008.12.001
Sharma S, Tiwari S, Hasan A et al (2018) Recent advances in conventional and contemporary methods for remediation of heavy metal-contaminated soils. 3 Biotech 8:216. https://doi.org/10.1007/s13205-018-1237-8
Shi Y, Sun Y, Gao B et al (2018) Retention and Transport of Bisphenol A and Bisphenol S in Saturated Limestone Porous Media. Water Air Soil Pollut 229:260. https://doi.org/10.1007/s11270-018-3911-1
Silva Conceição JC, Vieira TM, Miller Crotti AE et al (2021) Aspergillus brasiliensis-mediated biotransformation of methyl p-coumarate via phenyloxiran moiety: a predictive model for environmental bioremediation. Int Biodeterior Biodegradation 158:105167. https://doi.org/10.1016/j.ibiod.2020.105167
Sosa-Martínez JD, Balagurusamy N, Montañez J et al (2020) Synthetic dyes biodegradation by fungal ligninolytic enzymes: process optimization, metabolites evaluation and toxicity assessment. J Hazard Mater 400:123254. https://doi.org/10.1016/j.jhazmat.2020.123254
Spina F, Tigini V, Romagnolo A, Varese G (2018) Bioremediation of Landfill Leachate with Fungi: Autochthonous vs. Allochthonous Strains Life 8:27. https://doi.org/10.3390/life8030027
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4.0. Mol Biol Evol 24:1596–1599. https://doi.org/10.1093/molbev/msm092
Thompson JD, Gibson TJ, Plewniak F et al (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882. https://doi.org/10.1093/nar/25.24.4876
Tian B, Wu N, Pan X et al (2022) Ferrate(VI) oxidation of bisphenol E–Kinetics, removal performance, and dihydroxylation mechanism. Water Res 210:118025. https://doi.org/10.1016/j.watres.2021.118025
Torres-García JL, Ahuactzin-Pérez M, Fernández FJ, Cortés-Espinosa DV (2022) Bisphenol A in the environment and recent advances in biodegradation by fungi. Chemosphere 303:134940. https://doi.org/10.1016/j.chemosphere.2022.134940
Wang J, Yamada Y, Notake A et al (2014) Metabolism of bisphenol A by hyper lignin-degrading fungus Phanerochaete sordida YK-624 under non-ligninolytic condition. Chemosphere 109:128–133. https://doi.org/10.1016/j.chemosphere.2014.01.029
Wang J, Xie Y, Hou J et al (2022) Biodegradation of bisphenol A by alginate immobilized Phanerochaete chrysosporium beads: continuous cyclic treatment and degradation pathway analysis. Biochem Eng J 177:108212. https://doi.org/10.1016/j.bej.2021.108212
White TJ, Bruns T, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocol: a guide to methods and applications. Academic Press, San Diego, CA, pp 315–322
Wu S, Snajdrova R, Moore JC et al (2021) Biocatalysis: enzymatic synthesis for Industrial Applications. Angew Chemie Int Ed 60:88–119. https://doi.org/10.1002/anie.202006648
Yang W, Li Q, Guo S et al (2022) Rational design of aspergillus flavus A5p1-immobilized cell system to enhance the decolorization of reactive blue 4 (RB4). Chin J Chem Eng 52:37–44. https://doi.org/10.1016/j.cjche.2021.11.028
Zdarta J, Jesionowski T, Meyer AS, Pinelo M (2022) Removal of tetracycline in enzymatic membrane reactor: enzymatic conversion as the predominant mechanism over adsorption and membrane rejection. J Environ Chem Eng 10:106973. https://doi.org/10.1016/j.jece.2021.106973
Zhao H, Xiang L, Li J et al (2016) Investigation on fragmentation pathways of bisphenols by using electrospray ionization orbitrap mass spectrometry. Rapid Commun Mass Spectrom 30:1901–1913. https://doi.org/10.1002/rcm.7666
Zhuo R, Fan F (2021) A comprehensive insight into the application of white rot fungi and their lignocellulolytic enzymes in the removal of organic pollutants. Sci Total Environ 778:146132. https://doi.org/10.1016/j.scitotenv.2021.146132
Zielinska M, Wojnowska-Baryła I, Cydzik-Kwiatkowska A (2019) Bisphenol a removal from Water and Wastewater, 2th edn. Springe, Switzerland
Funding
The authors thank the financial support from Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq, grant number: 445982/2020-9), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES - Brazil) – código de financiamento 001, Rede Agronano (EMBRAPA) and Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP, grant number: 2018/22214-6). JCSC thanks the Fundação de Amparo à Pesquisa do Estado da Bahia (FAPESB) for his scholarship.
Author information
Authors and Affiliations
Contributions
João C. S. Conceição and Augusto D. Alvarenga: Methodology, Investigation, Data curation; Luiza A. Mercante: Conceptualization, Formal analysis, Writing (review and editing); Daniel S. Correa: Project administration, Writing (review and editing); Eliane O. Silva: Design of manuscript, Formal analysis, Writing (original draft). All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Conceição, J.C.S., Alvarega, A.D., Mercante, L.A. et al. Endophytic fungus from Handroanthus impetiginosus immobilized on electrospun nanofibrous membrane for bioremoval of bisphenol A. World J Microbiol Biotechnol 39, 261 (2023). https://doi.org/10.1007/s11274-023-03715-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03715-z