Abstract
Squalene is a triterpene hydrocarbon, a biochemical precursor for all steroids in plants and animals. It is a principal component of human surface lipids, in particular of sebum. Squalene has several applications in the food, pharmaceutical, and medical sectors. It is essentially used as a dietary supplement, vaccine adjuvant, moisturizer, cardio-protective agent, anti-tumor agent and natural antioxidant. With the increased demand for squalene along with regulations on shark-derived squalene, there is a need to find alternatives for squalene production which are low-cost as well as sustainable. Microbial platforms are being considered as a potential option to meet such challenges. Considerable progress has been made using both wild-type and engineered microbial strains for improved productivity and yields of squalene. Native strains for squalene production are usually limited by low growth rates and lesser titers. Metabolic engineering, which is a rational strain engineering tool, has enabled the development of microbial strains such as Saccharomyces cerevisiae and Yarrowia lipolytica, to overproduce the squalene in high titers. This review focuses on key strain engineering strategies involving both in-silico and in-vitro techniques. Emphasis is made on gene manipulations for improved precursor pool, enzyme modifications, cofactor regeneration, up-regulation of limiting reactions, and downregulation of competing reactions during squalene production. Process strategies and challenges related to both upstream and downstream during mass cultivation are detailed.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
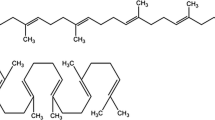
Squalene is a naturally occurring linear triterpene hydrocarbon (C30H50) comprising six double bonds (Fig. 1). Squalene was named after the shark family "Squalidae" (Tsujimoto 1916). The molecular weight of squalene is 410.7 g/mol, it has a melting point of − 75 °C, a boiling point of 285 °C, and a refractive index of 1.49 (Popa et al. 2015). Squalene has several applications in the food, chemical, and healthcare sectors. It is used as a dietary supplement (Naziri et al. 2011a), moisturizer, cardio-protective agent, anti-tumor agent, a natural antioxidant in addition to its antifungal (Smith 2000), antibacterial, and antivirulence activities (Bavisetty et al. 2015). It has been used as a vaccine adjuvant in seasonal influenza and COVID-19 vaccines (Tateno et al. 2020). Squalene intake in the Mediterranean region and the United States is in the 200–400 mg/day and 30 mg/day ranges, respectively (Newmark 1999). Squalane, a stable hydrogenated derivative of natural squalene is favored in the cosmetic industries as it penetrates human skin and imparts smoothness without greasy feels (Kim and Karadeniz 2012). With the increased demand for squalene along with regulations on shark-derived squalene, there is a need to find alternatives for squalene production which are low-cost as well as sustainable (Paramasivan 2019). In contrast to plant sterols such as sitosterol and stigmasterol, and animal sterols such as cholesterol, ergosterol acts as a predominant sterol in the cell wall of the fungus. Squalene is a ergosterol pathway intermediate in S. cerevisiae. It is produced in yeast through the mevalonate (MVA) biosynthesis process and in bacteria through the methylerythritol 4-phosphate (MEP) biosynthesis process (Fig. 1). However, squalene, as an intermediary chemical, is produced at lower quantities in wild-type strains than other macromolecules and is usually below the detection level.
Squalene biosynthesis pathway in yeast (the MVA pathway) and bacteria (the MEP pathway). CDP-ME, 4-diphosphocytidyl-2-C-methylerythritol; CDP-MEP, CDP-2-C-methyl-D-erythritol 2-phosphate; DMAPP, dimethylallyl pyrophosphate; DXP, 1-Deoxy-D-xylulose 5-phosphate; dxr-1-deoxy-D-xylulose-5-phosphate reductoisomerase; dxs, 1-deoxyxylulose-5-phosphate synthase; ERG8, Phosphomevalonate kinase; ERG10, Acetyl-CoA Cacetyltransferase; ERG13, Hydroxy methyl glutaryl-CoA synthase; ERG12, Mevalonate kinase; ERG19, Mevalonate pyrophosphate decarboxylase; ERG9, Squalene synthase; ERG20, farnesyl pyrophosphate synthase; FPP, Farnesyl Pyrophosphate; GPP, Geranyl Pyrophosphate; HMB-PP, 4-Hydroxy-3-methyl-but-2-enyl pyrophosphate; IDI1, isopentenyl pyrophosphate: dimethylallyl pyrophosphate isomerase; IPP, Isopentenyl Pyrophosphate; ispD-2-C-methyl-D-erythritol-4-phosphate cytidylyl transferase; ispE, 4-diphophocy-tidyl-2-C-methyl-D-erythritol kinase; ispF, 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase; ispG, HMB-PP synthase; ispH, HMB-PP reductase; HMGR, Hydroxy methyl glutaryl CoA reductase; MEP, 2-C-methylerythritol 4-phosphate; MEcPP, 2-C-methyl-D-erythritol-2,4-cyclodiphosphate
Squalene is a biomolecule of economic importance, and efforts are undertaken in recent years to synthesize it utilizing S. cerevisiae as a cell factory (Table 1) (Mantzouridou and Tsimidou 2010; Naziri et al. 2011b; Rasool et al. 2016a; Han et al. 2018; Wei et al. 2018; Paramasivan et al. 2019). In the last decade, only a few patents on the generation of squalene in modified yeast have been granted (Walker et al. 2009; Choi et al. 2012). Amyris, Inc., a biotech firm, has commercially synthesized squalene using a modified yeast strain (Fisher et al. 2013). Biotechnological synthesis of squalene has also been pursued in recent years in various other species such as Escherichia coli, Synechocystis sp, Yarrowia lipolytica, and Kluyveromyces lactis other than S. cerevisiae (Table 2) (Bergquist et al. 2014; Drozdíková et al. 2015; Huang et al. 2018). Squalene synthesis in microbes, plants, and mammals and their applications has been recently reviewed (Gohil et al. 2019). In the current review, metabolic engineering and process strategies for squalene enhancement in different microbes have been comprehensively discussed with emphasis on squalene synthesis in yeast. The discussed engineering strategies could also be effectively applied for the synthesis of different terpenes in yeast as well as in other microbes.
Genetic modifications to improve squalene synthesis
Precursor pool rebalancing
In S. cerevisiae, 3-hydroxy-3-methyl-glutaryl-coenzyme A (HMG-CoA) reductase is translated as two different isomers, Hmg1p and Hmg2p. Hmg1p accounts for nearly 83 percent of overall enzyme activity in the wild-type strain (Basson et al. 1986). Further analysis revealed that these two variants have a membrane-binding region (1-552 amino acids) that covers the ER (endoplasmic reticulum) as well as the catalytic domain (553-1046 amino acids) (Basson et al. 1988). HMG-CoA reductase is a major regulatory enzyme in the ergosterol biosynthesis pathway and its overexpression has proven essential in most metabolic engineering efforts of S. cerevisiae towards terpene synthesis (Rodwell et al. 1976). Later, it was revealed that upregulation of the catalytic domain of HMG1, known as tHMG1, enhanced squalene levels in S. cerevisiae (Donald et al. 1997; Polakowski et al. 1998). Overexpression of membrane-bound Hmg1p results in the development of ER membrane stacks, also known as karmellae (Wright et al. 1988). To make a soluble and non-membrane bound, a truncated form of the HMG-CoA reductase enzyme (tHMG1) has been overexpressed (Polakowski et al. 1998). Moreover to avoid feed-back inihibition of ergosterol, it was expressed under a constitutive promoter in their studies. This theory established the basis for the upregulation of tHMG1 in an array of studies targeting the synthesis of terpenes in S. cerevisiae (Donald et al. 1997; Polakowski et al. 1998; Thompson et al. 2014; Zhuang and Chappell 2015; Kwak et al. 2017; Han et al. 2018; Paramasivan et al. 2019; Liu et al. 2020a, b; Zhu et al. 2021) (Table 1). Overexpression of tHMG1 encoding solely the catalytic domain of protein to circumvent regulation did not raise ergosterol content as predicted, but it did boost squalene production by nearly 30 times in S. cerevisiae (Thompson et al. 2014). Full-length HMG1 overexpression has also been attempted and has been shown to increase squalene levels despite the feedback regulation (Tokuhiro et al. 2009). It might be attributable to the multi-copy integration of this gene in their studies; however, the exact theory is unclear. Hmg2p, a major isoform of this reductase in low oxygen conditions, has therefore turned into a target for upregulation during anaerobic growth to synthesize squalene (Mantzouridou and Tsimidou 2010).
Upregulation of the MVA pathway
Upregulation of structural genes, particularly those in the sterol pathway ahead of squalene, have been identified as key targets for squalene production in S. cerevisiae (Fig. 1) (Rasool et al. 2016a; Li et al. 2020; Liu et al. 2020a, b; Zhou et al. 2021; Zhu et al. 2021; Meng et al. 2020; Sun et al. 2020). The genes include ERG10 coding for acetyl-CoA C-acetyltransferase, ERG13 coding for HMG-CoA synthase, HMGR, ERG12 coding for mevalonate kinase, ERG8 coding for phosphomevalonate kinase, IDI1 coding for isopentenyl pyrophosphate: dimethylallyl pyrophosphate isomerase, ERG20 coding for farnesyl pyrophosphate synthase, and ERG9 coding for squalene synthase. In a study by Rasool et al. (2016a), overexpression of the structural genes in the MVA pathway such as HMG1, IDI1, ERG20, and ERG9 has been attempted during the synthesis of squalene and has enhanced the squalene up to 10-folds (Table 1; Figs. 1, 2a). Further, the presence of squalene monooxygenase inhibitor, terbinafine, together with overexpression of the MVA pathway genes has increased squalene titer to 304.49 mg/l (Rasool et al. 2016a). Overexpression of the MVA pathway genes combined with the carbon source controlled fed-batch fermentation has resulted in a strain with squalene synthesis ability of 9.5 g/l (Table 1) (Li et al. 2020).
Strain improvement strategies and tools applied towards squalene synthesis in microbes. (a) Improvement of precursor pool by episomal expression of genes and chromosomal integration of genes towards product flux improvement; (b) Deletion of competing reactions by chromosomal gene replacement; (c) Site-directed mutagenesis for expression of mutant genes; (d)Cofactor regeneration and heterologous gene expression; (e) In-silico genome scale modeling towards gene knockout predictions; (f) Compartmentalization of pathways by expression of genes with specific signaling sequences; (g) Bioprocess strategies by reactor cultivation and medium standardization; (h) Adaptive laboratory evolution. acs – Acetyl-CoA synthetase; Ac-CoA – Acetyl-CoA; FPP – Farnesyl pyrophosphate; HMG-CoA – Hydroxy Methyl Glutaryl-CoA; INO2 – Inositol requiring transcription factor; MTS – Mitochondrial targeting sequence; PTS – Peroxisomal targeting sequence; tHMG1-truncated HMG-CoA reductase
Knockout of competing pathways
Deletion of the competing reactions is an effective method for constructing an efficient microbial chassis (Pickens et al. 2014) (Fig. 2b). This approach is referred to as either knockout which is typically applied by deleting a gene engaged in the competing reactions or knockdown which is reducing the levels of gene expression by introducing a weak promoter, and occasionally by allowing a system to regulate expression through the use of a programmable or customizable promoter (referred as repression). Through a random mutation in the ERG1 gene, Kamimura et al. (1994) achieved a squalene production of 5 mg/g DCW (dry cell weight) by disrupting the ergosterol biosynthesis pathway in S. cerevisiae. Garaiová et al. (2014) investigated squalene increase in the mutant strains of ERG1 grown in the medium supplemented with terbinafine, ergosterol biosynthesis inhibitor and obtained a squalene production of 1000 µg per 109 cells. Knockout of the ERG6 gene coding for delta (24)-sterol C-methyltransferase along with a site-specific mutation in HMG2 (K6R) has enhanced squalene to 20-fold (Mantzouridou and Tsimidou 2010). Likewise, knockout of ERG6 gene along with knockdown of ERG11 gene involved in ergosterol biosynthesis has enhanced squalene up to 43 mg/g DCW in S. cerevisiae (Paramasivan et al. 2018). In another study, knockdown of downstream genes involved in ergosterol synthesis and the alcohol dehydrogenase genes involved in ethanol biosynthesis combined with several other strategies has enhanced squalene to 304.49 mg/l (Rasool et al. 2016a). Also, knockout of the SHC gene coding for squalene hopene cyclase in the Rhodopseudomonas palustris and the cyanobacterial Synechocystis sp has enhanced squalene up to 70-fold and 27-fold, respectively (Englund et al. 2014; Xu et al. 2016) (Table 2). In Yarrowia lipolytica, knockout of PEX10 encoding a peroxisomal membrane E3 ubiquitin ligase, and URE2 encoding a transcriptional regulator involved in the repression of nitrogen catabolite has enhanced squalene yield (Wei et al. 2021). Rox1p is a repressor transcriptional factor protein involved in the regulation of ergosterol biosynthesis (Henry et al. 2002). ROX1 gene deletion has enhanced the mevalonate and terpenoid synthesis in Saccharomyces cerevisiae (Özaydın et al. 2013). So far, this approach has not been demonstrated in yeast squalene synthesis.
Enzyme engineering towards squalene enhancement
Site-specific mutagenesis is a critical enzyme engineering technique to improve target compounds in the cells (Fig. 2c). Directed evolution improves enzyme characteristics such as stability, solubility, and specificity, resulting in the generation of customized enzymes (Kaur and Sharma 2006). Site-directed mutagenesis aims to introduce deliberate mutations at a specific codon in the genes, resulting in a changed codon. This method has shown to be an invaluable tool in synthetic biology for improving protein characteristics, and it has been used to promote squalene synthesis in yeast and other microbes. As squalene epoxide synthesis reaction is catalyzed by the ERG1 gene, attempts were made to modify the ERG1 gene to reduce the flux towards the downstream ergosterol biosynthesis pathway. In a study, glutamine Q at position 443 was mutagenized to UAG in Erg1p that in turn increased squalene up to 30–40-fold in the engineered S. cerevisiae (Garaiová et al. 2014) (Table 1). Several other point mutational studies have been made in the ERG1 gene. In a study by Zhou et al. (2021), glycine G at position 30 was mutagenized to serine S in Erg1p to reduce the specific activity. The study resulted in a strain with squalene production ability up to 3.5 g/l (Table 1). Site-specific mutations have also been made in another critical gene, HMG2. Primarily, lysine K at positions 6, and 357 on Hmg2p were identified to be crucial for protein degradation, and so these lysines were substituted to evaluate the Hmgp2 stability (Gardner and Hampton 1999). In a study by Mantzouridou and Tsimidou (2010), lysine K at position 6 has been mutagenized to asparagine R in Hmg2p which resulted in a strain that produced 5.2 mg/g DCW of squalene (Table 1). Hmg2P (K6R) has also been successful in improving several other terpenes (Paramasivan and Mutturi 2017a, 2017b). During the synthesis of squalene, Garaiová et al. (2014) downregulated squalene epoxidase coded by ERG1 and expressed multiple enzyme mutant variants of the ERG1 gene. Based on reverse engineering approaches, a causal mutation for high levels of squalene accumulation in yeast cells has been identified in Paramasivan et al. (2021) when S. cerevisiae cells were adapted in the presence of terbinafine. In this study it was demonstrated that phenylalanine at position 420 of Erg1p was replaced with isoleucine (F420I), which resulted in improved squalene and terbinafine resistance. Under aerobic growth, Upc2p (uptake control protein) mutants are involved in sterol uptake (Vik Å and Rine 2001). Mutations in Upc2p such as G888A has been demonstrated to enhance external sterol (ergosterol) uptake, downregulation of internal ergosterol formation, and thereby realigning the allocation of resources for improvement in specific terpene synthesis. So far, this strategy has not been applied towards squalene synthesis studies in yeast.
Heterologous gene expression towards squalene synthesis
This section describes the heterologous expression-based metabolic engineering approaches that have been developed to augment squalene synthesis in S. cerevisiae and other microbes (Fig. 2d). Heterologous enzymes are sometimes more efficient in catalysis when compared to the native enzyme. In this regard, several attempts have been made to enhance the squalene synthesis in microbes by heterologous expression of non-native enzymes. In a study, as an alternative to ERG20, heterologous expression of IspA coding for farnesyl diphosphate synthase from E. coli was carried to obtain squalene levels of 1.24 g/l in S. cerevisiae (Choi et al. 2012) (Table 1). In another study by Zhuang and Chappell (2015), squalene synthase from Botryococcus braunii was expressed in S. cerevisiae which led to squalene synthesis of nearly 270 mg/l. Expression of squalene synthase from humans has also become another predominant strategy towards squalene improvement in host cells such as E. coli. (Furubayashi et al. 2014; Katabami et al. 2015; Xu et al. 2019). Human SQS has a high catalytic efficiency towards FPP (kcat/Km = 0.51 μM−1 s−1). Similarly, squalene synthase from T. elongatus BP-1 has an even higher catalytic efficiency (kcat/Km = 1.8 μM−1 s−1) and has been expressed in E. coli for increased squalene synthesis (Katabami et al. 2015). Also, the squalene synthase from S. cerevisiae (ScSQS) along with the entire MVA pathway genes has been expressed in E. coli towards squalene enhancement (Sun et al. 2020). In another study by Ghimire et al. (2009), hopD (farnesyl diphosphate synthase) and hopAB (squalene/phytoene synthases) genes from Streptomyces peucetius have been heterologously expressed to augment squalene synthesis in yeast cells leading to an increased titer of 11.8 mg/l in E. coli. A novel heterologous pathway of squalene biosynthesis from Zymomonas mobilis and Rhodopseudomonas palustris has been expressed and characterized in E. coli. The pathway consists of three genes, hpnD, hpnC, and hpnE expressing enzymes that convert FPP to PSPP (presqualene diphosphate), PSPP to HSQ (hydroxysqualene) and HSQ to squalene, respectively (Pan et al. 2015).
Manipulations in the central carbon metabolism
Since the biosynthesis of a metabolite includes multiple reactions within the cell, improvements in essential upstream molecules generated may affect enhancing downstream pathway components. Acetyl-CoA is an example of an early intermediate metabolite that contributes to the TCA cycle, sterol, fatty acids, and polyketides biosynthesis. In S. cerevisiae, cytoplasmic acetyl-CoA is formed via a PDH-bypass, which predominantly promotes sterol and fatty acid pathway (Shiba et al. 2007; Chen et al. 2013; Krivoruchko et al. 2015). Pulling the flux towards squalene biosynthesis exhausts acetyl-CoA supply in S. cerevisiae cells impairing its functions. To overcome this, the ACS1 gene coding for acetyl-CoA synthetase has been overexpressed in a study targeting squalene improvement where ACS1 combined with the ERG10 gene expression has significantly enhanced squalene biosynthesis in yeast (Zhu et al. 2021). Cytoplasmic acetyl-CoA augmentation is one of the primary strategies applied to improve squalene synthesis. Erg10p is the foremost enzyme in the MVA pathway that pulls the flux towards squalene synthesis by converting acetyl-CoA to acetoacetyl-CoA. ERG10 gene overexpression has been attempted in several studies that aim to improve squalene synthesis (Rasool et al. 2016b; Kwak et al. 2017; Li et al. 2020; Liu et al. 2020a, b; Zhou et al. 2021; Zhu et al. 2021). Overexpression of tHMG1 along with the ERG10 gene in a xylose-fermenting strain has improved the squalene levels to 150 mg/l using xylose as a carbon source (Kwak et al. 2017). Likewise, endogenous gene atoB coding for acetyl-CoA acetyltransferase has been overexpressed in an attempt to enhance squalene synthesis in E. coli (Katabami et al. 2015). Yet another promising strategy to improve acetyl-CoA is by conversion of ethanol to acetyl-CoA. In an interesting study by Li et al. (2020), the ADH2 gene that converts ethanol to acetyl-CoA has been overexpressed together with the heterologous expression of ADA from Dickeya zeae which converts acetaldehyde to acetyl-CoA in S. cerevisiae. As the ADA gene coding enzyme does not require ATP consumption, the metabolic burden on the cells during squalene synthesis is reduced. In addition, ACL1 and ACL2 genes coding for ATP citrate lyase from Yarrowia lipolytica that converts citrate to acetyl-CoA have been overexpressed in S. cerevisiae towards squalene synthesis. This strategy has proven to enhance the squalene content in the cells significantly (Liu et al. 2020a, b).
Co-factor regeneration
In yeast cells, NADPH is one of the ubiquitous cofactors (Agledal et al. 2010). Several enzymes in the sterol pathway of S. cerevisiae require NADPH as a cofactor. Overexpression of the MVA pathway genes such as tHMG1, HMG2, and ERG9 would deplete cytosolic NADPH. Hence, the expression of NADPH regeneration-related genes has become a popular approach to stimulate terpenes and other metabolite syntheses (Brown et al. 2015) (Fig. 2d). In yeast cells, NADH kinases and NADP+-dependent dehydrogenases are responsible for NADPH synthesis, and regeneration respectively (Shi et al 2005, 2011). Precisely, NADH kinase is involved in the phosphorylation of NADH to NADPH while the dehydrogenases are involved in the reduction of NADP+ to NADPH. POS5 gene codes for the enzyme that harbors both NAD kinase as well as NADH kinase activity (Strand et al. 2003) while another enzyme glucose-6-phosphate dehydrogenase coded by ZWF1 has NADP+-dependent dehydrogenase activity. In S. cerevisiae, ZWF1 is a prime target for promoting cytosolic NADPH regeneration. ZWF1 gene overexpression enhanced squalene synthesis to 2.3-fold, whereas POS5 overexpression with mitochondrial targeting sequence (MTS) has enhanced squalene production to sixfold upon overexpression in yeast cells (Paramasivan & Mutturi, 2017a, b). In the same study, ZWF1 and POS5 (with MTS) gene expression combined with tHMG1 expression has synergistically enhanced squalene to 22.5-fold and 27.5-fold respectively. In another study, POS5 gene expression was combined with tHMG1 expression and knockout of ADK1 and LYS1 genes identified by in-silico based genome-scale models towards squalene increase (Fig. 2e). The genetic modifications combined with fed-batch cultivation have enhanced squalene levels to 1.9 g/l. Similarly, zwf expression in E. coli combined with several other strategies has enhanced squalene synthesis to 52 mg/l with net increment up to 21-folds (Xu et al. 2019) (Table 1). In addition, to reduce the metabolic burden caused by the NADPH cofactor, an alternative NADH-based enzyme has been used as a target towards squalene synthesis (Li et al. 2020; Liu et al. 2020a, b). Since NADH is more readily available compared to NADPH, NADH-based HMGR from Silicibacter pomeroyi has been heterologously expressed in S. cerevisiae for squalene improvement. NADH-HMGR overexpression combined with several other strategies has resulted in a strain with squalene levels of 9.5 g/l (Li et al. 2020) and 11 g/l (Liu et al. 2020a, b) (Table 1).
Promoter based engineering strategies towards squalene synthesis
Promoters are critical for the control of gene expression thereby regulating the metabolic pathways in S. cerevisiae. Promoter-based engineering strategies are gaining an advantage in recent times as they could significantly affect gene expression levels. ERG1 gene codes for squalene monooxygenase in the MVA pathway downstream of squalene in S. cerevisiae. Being an essential gene, disruption of ERG1 leads to a lethal phenotype and therefore repression of ERG1 gene by promoter replacement has become a prominent strategy in the development of cell factories towards squalene biosynthesis (Hull et al. 2014; Manzoor et al. 2020; Liu et al. 2020a, b; Zhu et al. 2021). Also, the usage of repressible promoters provides better economic feasibility when compared to the enzyme inhibitors at an industrial level. Initially, doxycycline repressible tet07-CYC1 promoter has been utilized to control the ERG1 expression by substituting the native ERG1 promoter at the chromosome. In a study by Hull et al. (2014), ERG1 gene under repressible tet07-CYC1 promoter system along with grass juice as a carbon source has enhanced squalene up to 18 mg/l. Later, two iron and copper repressible promoters PAFT1 and PCTR1, respectively are characterized and used to repress the ERG1 gene in S. cerevisiae towards squalene bioproduction. ERG1 repression with metal ion repressible PCTR1 promoter has enhanced squalene up to twofold when compared to PAFT1 promoter (Manzoor et al. 2020). In the same study, TFBSTEF1 and TFBSHHF2 were engineered into different constitutive promoters thereby enhancing the strength of the yeast constitutive promoters. The engineered promoters such as PHHF2, PIRA1, PRHO1, and PPET9 with TFBSTEF1 were utilized to drive the transcription of HMG1, IDI1, ERG20, and ERG9 genes respectively. This study resulted in a strain with squalene improvement to nearly 74-fold (Manzoor et al. 2020). Further, PHXT1 is a glucose inducible promoter and causes repression when the glucose levels in the medium become low or zero. The native promoter of ERG1 when replaced with the HXT1 promoter combined with multiple other strategies has significantly enhanced squalene titer to 11 g/l (Liu et al. 2020a, b). Similarly, the PHXT1 has been used to repress the ERG1 which in combination with the compartmentalization of genes in mitochondria leads to the highest production of squalene reported to date which is 21.1 g/l (Table 1; Zhu et al. 2021). Another noteworthy and most commonly used repressible promoter is the methionine repressible promoter (PMET3) which has been extensively used to repress the ERG9 gene towards other terpene production in yeast (Westfall et al. 2012). However tunable PMET3 has not been used towards squalene synthesis so far. GAL80 negatively regulates gal regulon by binding to PGAL4 in the absence of galactose as well as in the presence of carbon sources such as glucose. Since galactose is an expensive carbon source compared to glucose, GAL80 gene knockout is critical to express the genes under the strong galactose inducible promoters PGAL1 and PGAL10 (Westfall et al. 2012). Thus, Li et al. (2020) have overexpressed the mevalonate pathway gene tHMG1 under the galactose inducible promoter in the GAL80 deletion strain where the gene activation happens in response to glucose as the negative regulator is not present. This leads to a 33-fold improvement in squalene synthesis. In an interesting investigation by Zhou et al. (2021), the SRE (sterol response element) motif in ERG1 and ERG11 genes was replaced with the heterologous marO motif to downregulate the expression of both the genes thereby improving squalene and lanosterol accumulation, respectively. Replacement of the SRE motif reduces the feedback regulation by SREBP (sterol response element-binding proteins) at high sterol levels in the cells while marO insertion helps in controlling the metabolic flux. marO is a cis-element of mar operator bound by the transcriptional repressor protein, marR in E. coli. marO engineered near the TATA box of ERG1 and ERG11 promoter could efficiently repress the respective gene expression proving to be a promising alternative for gene repression studies in yeast S. cerevisiae.
Compartmentalization for squalene synthesis
Owing to the abundant availability of precursors and cofactors such as acetyl-CoA and redox equivalents, respectively, harnessing organelles such as mitochondria, ER, and peroxisomes are considered as a viable strategy towards squalene production (Fig. 2f). In a study by Zhu et al. (2021), to surpass the cell toxicity and the metabolic strain imposed by the phosphorylated intermediates of the MVA pathway, a partial pathway from acetyl-CoA to MVA has been incorporated into the mitochondria by merging the mitochondrial targeting signal sequence with overexpression genes. Upon combining the cytoplasmic and mitochondrial engineering with a two-stage fed-batch fermentation, the squalene titers have synergistically improved to 21.1 g/l with 437.1 mg/g DCW (Table 1). Similarly, introducing the MVA pathway in the peroxisomes by fusing the genes with peroxisomal targeting sequence has enhanced the squalene titers up to 11 g/l upon combining with PHXT1-ERG1 and a two-stage fermentation (Table 1) (Liu et al. 2020a, b). Another noteworthy ER-based metabolic engineering has enhanced squalene levels up to 71-fold where the overexpression of the INO2 gene has expanded the ER functions. The key function of ER is protein folding and overexpression of genes causes metabolic burden in the ER. Therefore, the size regulatory factor gene INO2 has been overexpressed in the yeast to restore the normal ER functions (Kim et al. 2019). Hence, organelle engineering could be considered as a novel and effective alternative to improve squalene production.
Storage of squalene in lipid droplets and cell membrane
Squalene is stored in the cell membrane, peroxisomes, and lipid droplets in yeast cells. A lipid droplet found in all eukaryotic cells is a lipid storage organelle that is made of sterol esters, triacylglycerols, and squalene. It consists of neutral lipids and serves as a depot for non-polar storage lipids such as squalene and fatty acids. Neutral lipid production and storage is a complex and dynamic process. Lipid droplets are ER-derived storage organelles whose quantity, size, and content vary depending on nutritional requirements and growth circumstances (Farese and Walther 2009; Fei et al. 2009). Triacylglycerides and steryl esters are inert and are found in the lipid bodies unless they are hydrolyzed while squalene is either sequestered in lipid particles or involved in membrane biogenesis (Spanova et al. 2010; Grillitsch et al. 2011). In a study by Wei et al. (2018), the DGA1 gene coding for the diacylglycerol acyltransferase gene involved in fatty acid metabolism has been overexpressed in S. cerevisiae which enhanced squalene up to 250-fold upon the combination with the tHMG1 gene. DGA1 gene expression has proven to enhance lipid body formation in yeast cells thereby increasing the storage ability of the cells. Cell membrane engineering has been carried out to enhance the storage of squalene in E. coli. In this regard, tsr gene coding for a serine chemoreceptor protein has been overexpressed which in turn enhanced the fatty acid composition in membrane lipid composition leading to the formation of a multi-layered structure. The engineered strain has synthesized squalene up to 612 mg/l which is the maximum known squalene synthesis to date in E. coli (Meng et al. 2020) (Table 2).
Process strategies for improving squalene synthesis
Bioprocess conditions play a key role in the high-level production of compounds in microbial fermentation (Fig. 2g). Under anaerobic fermentative conditions, Bhattacharjee et al. (2001) have generated squalene in a commercial yeast strain with a yield of up to 41 µg/g DCW. Besides, under optimized semi-anaerobic cultivation, inoculation levels, and growth conditions, Mantzouridou et al. (2009) reached a maximum squalene yield of 3.1 mg/l. Subsequently, overexpression of tHMG1 and DGA1 and growth under nitrogen-limited conditions have led to a squalene titer of 445.6 mg/l in galactose-based fed-batch cultivation (Wei et al. 2018). In another study by Han et al. (2018), tHMG1 and bacterial ispA gene in tandem with terbinafine-based inhibition of squalene epoxidase have enhanced the squalene synthesis up to 2 g/l in glucose-based fed-batch cultivation. In some cases, the carbon sources to supplement the growth have been varied. For instance, grass juice and cane molasses have been used as a feedstock to increase squalene synthesis to 18 mg/l and 3.5 g/l respectively (Hull et al. 2014; Zhou et al. 2021). In an interesting study by Kwak et al (2017), xylose utilizing engineered strain was subjected to growth under xylose-based fed-batch cultivation. This study resulted in a strain with 532 mg/l of squalene titer. On the other hand, open loop-based exponential feeding of glucose in fed-batch cultivation has resulted in 1.9 g/l of squalene production using an engineered S. cerevisiae (Paramasivan et al. 2019). An optimized two-stage fed-batch cultivation using glucose and ethanol as a carbon source has been used to achieve a squalene titer of 11 g/l in an engineered strain (Liu et al. 2020a, b). C/N ratio, pH, and temperature optimization have led to a significantly high squalene production of 2.4 g/l in the yeast-like fungus Pseudozyma sp. SD301, a native producer of squalene (Song et al. 2015).
Adaptive laboratory evolution and downstream inhibition by terbinafine
Adaptive evolution is the use of selection pressure to leverage the innate capacity of an organism to undergo mutations toward the desired phenotype (Conrad et al. 2014) (Fig. 2h). It entails a process of continuous evolution with a selective gain for the variant exhibiting the desired trait (Sonderegger and Sauer 2003; Çakar et al. 2005). Adaptive laboratory evolution in combination with sequencing is a critical strategy in metabolic engineering for biomolecule synthesis (Conrad et al. 2014). Lowering the activity of the downstream enzyme, squalene epoxidase is a key technique for enhancing squalene levels and may be accomplished using a specialized non-competitive enzymatic inhibitor, terbinafine (Ta et al. 2012; Garaiová et al. 2014; Rasool et al. 2016a). Terbinafine has been extensively used in various studies and in different microbes to increase the squalene accumulation (Fan et al. 2010; Naziri et al. 2011a, b; Drozdıkova et al. 2015; Han et al. 2018). Naziri et al. (2011a, b) pioneered the cultivation of S. cerevisiae in terbinafine which led to at least ten times improvement in squalene. Following that, Garaiová et al. (2014) have produced 1000 µg of squalene per 109 cells in the presence of terbinafine with an engineered strain with Erg1p (Q443UAG). An engineered yeast strain has been subjected to grow in the presence of terbinafine as a selection pressure to enhance the squalene compound. This study resulted in a strain that produced 198 mg/l which was a 16.5-fold enhancement in comparison to the parent strain (Paramasivan et al. 2021). Further, terbinafine addition to the medium for the native squalene producer Aurantiochytrium mangrovei has increased squalene up to 40% (Fan et al. 2010). Besides, terbinafine has also been used to enhance squalene content in Kluyveromyces lactis (Drozdıkova et al. 2015). Han et al. (2018) have cultivated an engineered yeast strain in a fed-batch fermentation medium in the presence of terbinafine which has improved squalene synthesis up to twofold. However, large-scale cultivation of S. cerevisiae in the presence of terbinafine towards squalene synthesis could be an expensive strategy. Hence some studies reported the site-specific mutagenesis of squalene epoxidase and replacement of the promoter for ERG1 gene to downregulate the activity of this enzyme (Garaiová et al. 2014; Hull et al. 2014). Such strategies resulted in improved squalene accumulation as discussed earlier.
Engineering towards squalene synthesis in other microbes
The genes involved in the MEP pathway are upregulated in combination with the heterologous expression of squalene synthase from Bacillus megaterium has enhanced the squalene synthesis up to 29-fold in Bacillus subtilis (Song et al. 2020) (Table 2). Overexpression of dxs and idi genes, repression of idsA gene, and heterologous expression of ERG9 from S. cerevisiae has led to improved squalene levels (105 mg/l) in Corynebacterium glutamicum (Park et al. 2018) (Table 2). Further, when protein engineering by expression of human squalene synthase was coupled with pathway engineering, fed-batch cultivation, and dodecane extraction, the C. glutamicum strain generated 1.5 g/l (Park et al. 2021) (Table 2). Squalene synthase from S. cerevisiae along with the overexpression of the MVA pathway genes and tsr gene has increased squalene in the engineered E. coli (Meng et al. 2020) (Table 2). Cyanobacterial species such as Synechococcus elongatus has been utilized for squalene synthesis by heterologous expression of dxs, idi, ispA from E. coli, and overexpression of the fusion protein CpcB1-ScSQS along with light optimization in photobioreactor which has increased the squalene content up to 79.2 mg/g DCW (Choi et al. 2017) (Table 2). Metabolic engineering of a novel species Rhodopseudomonas palustris has enhanced squalene up to 23.3 mg/g DCW with 178-fold net increment by the knockout of shc and crtB genes as well as the overexpression of dxs, crtE, and hpnD (Xu et al. 2021) (Table 2). Overexpression of ylHMG1 and ylDGA1 along with fermentation optimization enhanced squalene to 731 mg/l in Yarrowia lipolytica. This is the highest reported so far in that species (Tang et al. 2021) (Table 2). In a study by Kang et al (2021), overexpression of squalene synthase, inhibition of squalene-hopene cyclase by ferulenol, knockout of pds-crtN-ald gene cluster, and fed-batch cultivation with potassium nitrate supplementation has improved squalene levels to 31.3 mg/l in Methylomonas sp. DH-1 (Kang et al. 2021) (Table 2).
Conclusion and future prospects
Concerns about squalene production from shark liver oil, and lower yields from plant sources, microbial platforms for squalene production have been sought. However, there exists challenges with microbial production systems as well. Some of the significant limitations of existing microbial squalene manufacturing platforms is lower squalene TRY (titer, productivity and yields) meant for industrial production. Moreover, obtaining toxin-free squalene from the microbes that naturally produce squalene such as Schizochytrium sp. and Aurantiochytrium sp. is challenging. Besides, physiology, genetics, and process constraints needs to be well studied for such organisms. Some of the prominent microbial platforms for squalene production includes S. cerevisiae, E. coli, C. glutamicum and Y. lipolytica. Metabolic engineering strategies in S. cerevisiae such as co-factor engineering, improved precursor supply, alterations outside sterol pathway, and process strategies have been thoroughly investigated and highest squalene titer of 21.1 g/l has been reported so far. However, majority of studies in S. cerevisiae focused on episomal expression, which presents a process issue when grown in a bioreactor. Amyris Inc. commercial synthesis of squalene is implemented in two stages, (i) farnesene is synthesized in the first stage using modified S. cerevisiae, and (ii) the obtained farnesene is transformed to squalene in the second stage utilizing chemical catalysis. Such hybrid production platforms can be exploited for commercial viability. Several microbial strains with high squalene productivity are developed in recent times which could serve as potential chassis for commercial production of squalene and also to compete with other low cost non-sustainable alternatives such as shark liver oil and vegetable oils. The present review on strain engineering and bioprocess strategies might be highly useful in improving squalene synthesis, leading to the development of a feasible and robust microbial platform for industrial production squalene.
References
Agledal L, Niere M, Ziegler M (2010) The phosphate makes a difference: cellular functions of NADP. Redox Rep 15:2–10. https://doi.org/10.1179/174329210X12650506623122
Basson ME, Thorsness M, Rine J (1986) Saccharomyces cerevisiae contains two functional genes encoding 3-hydroxy-3-methylglutaryl-coenzyme A reductase. Proc Natl Acad Sci USA 83:5563–5567. https://doi.org/10.1073/pnas.83.15.5563
Basson ME, Thorsness M, Finer-Moore J et al (1988) Structural and functional conservation between yeast and human 3-hydroxy-3-methylglutaryl coenzyme A reductases, the rate-limiting enzyme of sterol biosynthesis. Mol Cell Biol 8:3797–3808. https://doi.org/10.1128/MCB.8.9.3797
Bavisetty S, Mishra DP, Narayan B (2015) Inhibition of virulence of Staphylococcus aureus - a food borne pathogen - by squalene, a functional lipid. J Funct Foods 18:224–234. https://doi.org/10.1016/j.jff.2015.07.008
Bergquist J, Englund E, Pattanaik B et al (2014) Production of squalene in Synechocystis sp. PCC 6803. PLoS ONE 9:e90270. https://doi.org/10.1371/journal.pone.0090270
Bhattacharjee P, Shukla VB, Singhal RS, Kulkarni PR (2001) Studies on fermentative production of squalene. World J Microbiol Biotechnol 17:811–816. https://doi.org/10.1023/A:1013573912952
Brown S, Clastre M, Courdavault V, O’Connor SE (2015) De novo production of the plant-derived alkaloid strictosidine in yeast. Proc Natl Acad Sci 112:3205–3210. https://doi.org/10.1073/pnas.1423555112
Çakar ZP, Seker UOS, Tamerler C et al (2005) Evolutionary engineering of multiple-stress resistant Saccharomyces cerevisiae. FEMS Yeast Res 5:569–578. https://doi.org/10.1016/j.femsyr.2004.10.010
Chang M, Kim H, Jahng K (2008) The isolation and characterization of Pseudozyma sp. JCC 207, a novel producer of squalene. Appl Microbiol Biotechnol 78:963–972. https://doi.org/10.1007/s00253-008-1395-4
Chen G, Fan KW, Lu FP et al (2010) Optimization of nitrogen source for enhanced production of squalene from thraustochytrid Aurantiochytrium sp. New Biotechnol 27:382–389. https://doi.org/10.1016/j.nbt.2010.04.005
Chen Y, Daviet L, Schalk M et al (2013) Establishing a platform cell factory through engineering of yeast acetyl-CoA metabolism. Metab Eng 15:48–54. https://doi.org/10.1016/j.ymben.2012.11.002
Choi SY, Wang J, Kwak HS et al (2017) Improvement of squalene production from CO in Synechococcus elongatus PCC 7942 by metabolic engineering and scalable production in a photobioreactor. ACS Synth Biol 6:1289–1295. https://doi.org/10.1021/acssynbio.7b00083
Choi ES, Whang JI, Ryu K, Kim S won (2012) Modified yeast strain and a method for producing squalene using the same. US0322129
Choi SY, Sim SJ, Ko SC et al (2020) Scalable cultivation of engineered cyanobacteria for squalene production from industrial flue gas in a closed photobioreactor. J Agric Food Chem 68:10050–10055. https://doi.org/10.1021/acs.jafc.0c03133
Conrad TM, Lewis NE, Palsson BO (2014) Microbial laboratory evolution in the era of genome-scale science. Mol Syst Biol 7:509–509. https://doi.org/10.1038/msb.2011.42
Donald KAG, Hampton RY, Fritz IB (1997) Effects of overproduction of the catalytic domain of 3-hydroxy-3- methylglutaryl coenzyme A reductase on squalene synthesis in Saccharomyces cerevisiae. Appl Environ Microbiol 63:3341–3344. https://doi.org/10.1128/aem.63.9.3341-3344.1997
Drozdíková E, Garaiová M, Csáky Z, Obernauerová M, Hapala I (2015) Production of squalene by lactose-fermenting yeast Kluyveromyces lactis with reduced squalene epoxidase activity. Lett Appl Microbiol 61:77–84. https://doi.org/10.1111/lam.12425
Ebert BE, Czarnotta E, Blank LM (2018) Physiologic and metabolic characterization of Saccharomyces cerevisiae reveals limitations in the synthesis of the triterpene squalene. FEMS Yeast Res 18:foy077. https://doi.org/10.1093/femsyr/foy077
Englund E, Pattanaik B, Ubhayasekera SJK et al (2014) Production of squalene in Synechocystis sp. PCC 6803. PLoS ONE 9:e90270. https://doi.org/10.1371/journal.pone.0090270
Fagundes MB, Vendruscolo RG, Maroneze MM et al (2019) Towards a sustainable route for the production of squalene using cyanobacteria. Waste Biomass Valoriz 10:1295–1302. https://doi.org/10.1007/s12649-017-0191-8
Fan KW, Tsunehiro F, Feng A (2010) Enhanced production of squalene in the thraustochytrid Aurantiochytrium mangrovei by medium optimization and treatment with terbinafine. World J Microbiol Biotechnol 26:1303–1309. https://doi.org/10.1007/s11274-009-0301-2
Farese RV, Walther TC (2009) Lipid droplets finally get a little R-E-S-P-E-C-T. Cell 139:855–860. https://doi.org/10.1016/j.cell.2009.11.005
Fei W, Wang H, Fu X et al (2009) Conditions of endoplasmic reticulum stress stimulate lipid droplet formation in Saccharomyces cerevisiae. Biochem J 424:61–67. https://doi.org/10.1042/BJ20090785
Fisher K, Schofer SJ, Kanne DB (2013) Squalane and isosqualane compositions and methods for preparing the same. United States U.S. Patent 8,586,814 B2. (19 Nov 2013)
Furubayashi M, Li L, Katabami A et al (2014) Construction of carotenoid biosynthetic pathways using squalene synthase. FEBS Lett 588:436–442. https://doi.org/10.1016/j.febslet.2013.12.003
Garaiová M, Veronika Z, Zuzana S et al (2014) Squalene epoxidase as a target for manipulation of squalene levels in the yeast Saccharomyces cerevisiae. FEMS Yeast Res 14:310–323. https://doi.org/10.1111/1567-1364.12107
Gardner RG, Hampton RY (1999) A highly conserved signal controls degradation of 3-hydroxy-3- methylglutaryl-coenzyme a (HMG-CoA) reductase in eukaryotes. J Biol Chem 274:31671–31678. https://doi.org/10.1074/jbc.274.44.31671
Ghimire GP, Hei CL, Jae KS (2009) Improved squalene production via modulation of the methylerythritol 4-phosphate pathway and heterologous expression of genes from Streptomyces peucetius ATCC 27952 in Escherichia coli. Appl Environ Microbiol 75:7291–7293. https://doi.org/10.1128/AEM.01402-09
Gohil N, Bhattacharjee G, Khambhati K et al (2019) Engineering strategies in microorganisms for the enhanced production of squalene: advances, challenges and opportunities. Front Bioeng Biotechnol 7:50. https://doi.org/10.3389/fbioe.2019.00050
Grillitsch K, Connerth M, Köfeler H et al (2011) Lipid particles/droplets of the yeast Saccharomyces cerevisiae revisited: lipidome meets proteome. Biochim Biophys Acta 1811:1165–1175. https://doi.org/10.1016/j.bbalip.2011.07.015
Han JY, Hwa S, Jae S et al (2018) High level recombinant production of squalene using selected Saccharomyces cerevisiae strains. J Ind Microbiol Biotechnol 45:239–251. https://doi.org/10.1007/s10295-018-2018-4
Henry KW, Nickels JT, Edlind TD (2002) ROX1 and ERG regulation in Saccharomyces cerevisiae: implications for antifungal susceptibility. Eukaryot Cell 6:1041–1044. https://doi.org/10.1128/EC.1.6.1041-1044.2002
Hoang MH, Ha NC, Thom LT et al (2014) Extraction of squalene as value-added product from the residual biomass of Schizochytrium mangrovei PQ6 during biodiesel producing process. J Biosci Bioeng 118:632–639. https://doi.org/10.1016/j.jbiosc.2014.05.015
Huang Y, Jian X, Lv Y et al (2018) Enhanced squalene biosynthesis in Yarrowia lipolytica based on metabolically engineered acetyl-CoA metabolism. J Biotechnol 281:106–114. https://doi.org/10.1016/j.jbiotec.2018.07.001
Hull CM, Loveridge EJ, Rolley NJ et al (2014) Co-production of ethanol and squalene using a Saccharomyces cerevisiae ERG1 (squalene epoxidase) mutant and agro-industrial feedstock. Biotechnol Biofuels 7:1–9. https://doi.org/10.1186/s13068-014-0133-7
Kajikawa M, Kinohira S, Ando A et al (2015) Accumulation of squalene in a microalga Chlamydomonas reinhardtii by genetic modification of squalene synthase and squalene epoxidase genes. PLoS ONE 10:e0120446. https://doi.org/10.1371/journal.pone.0120446
Kamimura N, Hidaka M, Masaki H, Uozumi T (1994) Construction of squalene-accumulating Saccharomyces cerevisiae mutants by gene disruption through homologous recombination. Appl Microbiol Biotechnol 42:353–357. https://doi.org/10.1007/BF00902741
Kang CK, Jeong SW, Jo JH et al (2021) High-level squalene production from methane using a metabolically engineered Methylomonas sp. DH-1 strain. ACS Sustain Chem Eng 9:16485–16493. https://doi.org/10.1021/acssuschemeng.1c06776
Katabami A, Li L, Iwasaki M et al (2015) Production of squalene by squalene synthases and their truncated mutants in Escherichia coli. J Biosci Bioeng 119:165–171. https://doi.org/10.1016/j.jbiosc.2014.07.013
Kaur J, Sharma R (2006) Directed evolution: an approach to engineer enzymes. Crit Rev Biotechnol 26:165–199. https://doi.org/10.1080/07388550600851423
Kaya K, Nakazawa A, Matsuura H et al (2011) Thraustochytrid Aurantiochytrium sp. 18W–13a accummulates high amounts of squalene. Biosci Biotechnol Biochem 8451:10–13. https://doi.org/10.1271/bbb.110430
Kim SK, Karadeniz F (2012) Biological importance and applications of squalene and squalane. Adv Food Nutr Res 65:223–233. https://doi.org/10.1016/B978-0-12-416003-3.00014-7
Kim JE, Jang IS, Son SH et al (2019) Tailoring the Saccharomyces cerevisiae endoplasmic reticulum for functional assembly of terpene synthesis pathway. Metab Eng 56:50–59. https://doi.org/10.1016/j.ymben.2019.08.013
Krivoruchko A, Zhang Y, Siewers V et al (2015) Microbial acetyl-CoA metabolism and metabolic engineering. Metab Eng 28:28–42. https://doi.org/10.1016/j.ymben.2014.11.009
Kwak S, Kim SR, Xu H et al (2017) Enhanced isoprenoid production from xylose by engineered Saccharomyces cerevisiae. Biotechnol Bioeng 114:2581–2591. https://doi.org/10.1002/bit.26369
Li T, Liu GS, Zhou W et al (2020) Metabolic engineering of Saccharomyces cerevisiae to overproduce squalene. J Agric Food Chem 68:2132–2138. https://doi.org/10.1021/acs.jafc.9b07419
Liu G-S, Li T, Zhou W et al (2020a) The yeast peroxisome: a dynamic storage depot and subcellular factory for squalene overproduction. Metab Eng 57:151–161. https://doi.org/10.1016/j.ymben.2019.11.001
Liu H, Wang F, Deng L, Xu P (2020b) Genetic and bioprocess engineering to improve squalene production in Yarrowia lipolytica. Biores Technol 317:123991. https://doi.org/10.1016/j.biortech.2020.123991
Mantzouridou F, Tsimidou MZ (2010) Observations on squalene accumulation in Saccharomyces cerevisiae due to the manipulation of HMG2 and ERG6. FEMS Yeast Res 10:699–707. https://doi.org/10.1111/j.1567-1364.2010.00645.x
Mantzouridou F, Naziri E, Tsimidou MZ (2009) Squalene versus ergosterol formation using Saccharomyces cerevisiae: combined effect of oxygen supply, inoculum size, and fermentation time on yield and selectivity of the bioprocess. J Agric Food Chem 57:6189–6198. https://doi.org/10.1021/jf900673n
Manzoor R, Ahmed M, Riaz N et al (2020) Self-redirection of metabolic flux toward squalene and ethanol pathways by engineered yeast. Metabolites 10:56. https://doi.org/10.3390/metabo10020056
Meng Y, Shao X, Wang Y et al (2020) Extension of cell membrane boosting squalene production in the engineered Escherichia coli. Biotechnol Bioeng 117:3499–3507. https://doi.org/10.1002/bit.27511
Nakazawa A, Matsuura H, Kose R et al (2012) Optimization of culture conditions of the thraustochytrid Aurantiochytrium sp. strain 18W–13a for squalene production. Biores Technol 109:287–291. https://doi.org/10.1016/j.biortech.2011.09.127
Nakazawa A, Kokubun Y, Matsuura H et al (2014) TLC screening of thraustochytrid strains for squalene production. J Appl Phycol 26:29–41. https://doi.org/10.1007/s10811-013-0080-x
Naziri E, Mantzouridou F, Tsimidou MZ (2011a) Squalene resources and uses point to the potential of biotechnology. Lipid Technol 23:270–273. https://doi.org/10.1002/lite.201100157
Naziri E, Mantzouridou F, Tsimidou MZ (2011b) Enhanced squalene production by wild-type Saccharomyces cerevisiae strains using safe chemical means. J Agric Food Chem 59:9980–9989. https://doi.org/10.1021/jf201328a
Newmark HL (1999) Squalene, olive oil, and cancer risk review and hypothesis. Ann N Y Acad Sci 889:193–203. https://doi.org/10.1111/j.1749-6632.1999.tb08735.x
Özaydın B, Burd H, Lee TS et al (2013) Carotenoid-based phenotypic screen of the yeast deletion collection reveals new genes with roles in isoprenoid production. Metab Eng 15:174–183. https://doi.org/10.1016/j.ymben.2012.07.010
Pan JJ, Solbiati JO, Ramamoorthy G et al (2015) Biosynthesis of squalene from farnesyl diphosphate in Bacteria: three steps catalyzed by three enzymes. ACS Cent Sci 1:77–82. https://doi.org/10.1021/acscentsci.5b00115
Paramsivan K (2019) Metabolic engineering of Saccharomyces cerevisiae for overproduction of squalene. Academy of Scientific and Innovative Research, Ghaziabad
Paramasivan K, Mutturi S (2017a) Progress in terpene synthesis strategies through engineering of Saccharomyces cerevisiae. Crit Rev Biotechnol 37:974–989. https://doi.org/10.1080/07388551.2017.1299679
Paramasivan K, Mutturi S (2017b) Regeneration of NADPH coupled with HMG-CoA reductase activity increases squalene synthesis in Saccharomyces cerevisiae. J Agric Food Chem 65:8162–8170. https://doi.org/10.1021/acs.jafc.7b02945
Paramasivan K, Rajagopal K, Mutturi S (2018) Studies on squalene biosynthesis and the standardization of its extraction methodology from Saccharomyces cerevisiae. Appl Biochem Biotechnol 187:691–707. https://doi.org/10.1007/s12010-018-2845-9
Paramasivan K, Kumar HNP, Mutturri S (2019) Systems-based Saccharomyces cerevisiae strain design for improved squalene synthesis. Biochem Eng J 148:37–45. https://doi.org/10.1016/j.bej.2019.04.025
Paramasivan K, Aneesha A, Gupta N, Mutturi S (2021) Adaptive evolution of engineered yeast for squalene production improvement and its genome-wide analysis. Yeast 38:424–437. https://doi.org/10.1002/yea.3559
Park J, Yu BJ, Choi J, Woo HM (2018) Heterologous production of squalene from glucose in engineered Corynebacterium glutamicum using multiplex CRISPR interference and high-throughput fermentation. J Agric Food Chem 67:308–319. https://doi.org/10.1021/acs.jafc.8b05818
Park J, Kang DH, Woo HM (2021) Microbial bioprocess for extracellular squalene production and formulation of nanoemulsions. ACS Sustain Chem Eng 9:14263–14276. https://doi.org/10.1021/acssuschemeng.1c05453
Patel A, Rova U, Christakopoulos P et al (2019) Simultaneous production of DHA and squalene from Aurantiochytrium sp. grown on forest biomass hydrolysates. Biotechnol Biofuels 12:1–2. https://doi.org/10.1186/s13068-019-1593-6
Pickens LB, Tang Y, Chooi Y-H (2014) Metabolic engineering for the production of natural products. Annu Rev Chem Biomol Eng 2:211–236. https://doi.org/10.1146/annurev-chembioeng-061010-114209
Polakowski T, Stahl U, Lang C (1998) Overexpression of a cytosolic hydroxymethylglutaryl-CoA reductase leads to squalene accumulation in yeast. Appl Microbiol Biotechnol 49:66–71. https://doi.org/10.1007/s002530051138
Popa O, Băbeanu NE, Popa I et al (2015) Methods for obtaining and determination of squalene from natural sources. Biomed Res Int. https://doi.org/10.1155/2015/367202
Potijun S, Jaingam S, Sanevas N, Vajrodaya S, Sirikhachornkit A (2021) Green microalgae strain improvement for the production of sterols and squalene. Plants 10(8):1673. https://doi.org/10.3390/plants1008167
Rasool A, Saad Ahmed M, Li C (2016a) Overproduction of squalene synergistically downregulates ethanol production in Saccharomyces cerevisiae. Chem Eng Sci. https://doi.org/10.1016/j.ces.2016.06.014
Rasool A, Zhang G, Li Z, Li C (2016b) Engineering of the terpenoid pathway in Saccharomyces cerevisiae co-overproduces squalene and the non-terpenoid compound oleic acid. Chem Eng Sci 152:457–467. https://doi.org/10.1016/j.ces.2016.06.004
Rodwell VW, Nordstrom JL, Mitschelen JJ (1976) Regulation of HMG-CoA reductase. Adv Lipid Res 14:1–74. https://doi.org/10.1016/B978-0-12-024914-5.50008-5
Saengwong A, Yongmanitchai W, Chonudomkul D (2018) Screening and optimization of squalene production from microalgae Aurantiochytrium sp. Chiang Mai J Sci 45:680–691
Shakeri S, Khoshbasirat F, Maleki M (2021) Rhodosporidium sp. DR37: a novel strain for production of squalene in optimized cultivation conditions. Biotechnol Biofuels 14:1–4. https://doi.org/10.1186/s13068-021-01947-5
Shi F, Kawai S, Mori S et al (2005) Identification of ATP-NADH kinase isozymes and their contribution to supply of NADP(H) in Saccharomyces cerevisiae. FEBS J 272:3337–3349. https://doi.org/10.1111/j.1742-4658.2005.04749.x
Shi F, Li Z, Sun M, Li Y (2011) Role of mitochondrial NADH kinase and NADPH supply in the respiratory chain activity of Saccharomyces cerevisiae. Acta Biochim Biophys Sin 43:989–995. https://doi.org/10.1093/abbs/gmr092
Shiba Y, Paradise EM, Kirby J et al (2007) Engineering of the pyruvate dehydrogenase bypass in Saccharomyces cerevisiae for high-level production of isoprenoids. Metab Eng 9:160–168. https://doi.org/10.1016/j.ymben.2006.10.005
Smith TJ (2000) Squalene: potential chemo preventive agent. Expert Opin Investig Drugs 9:1841–1848. https://doi.org/10.1517/13543784.9.8.1841
Sonderegger M, Sauer U (2003) Evolutionary engineering of Saccharomyces cerevisiae for anaerobic growth on xylose. Appl Environ Microbiol 69:1990–1998. https://doi.org/10.1128/AEM.69.4.1990
Song X, Wang X, Tan Y et al (2015) High production of squalene using a newly isolated yeast-like strain. J Agric Food Chem 63:8445–8451. https://doi.org/10.1021/acs.jafc.5b03539
Song Y, Guan Z, van Merkerk R et al (2020) Production of squalene in Bacillus subtilis by squalene synthase screening and metabolic engineering. J Agric Food Chem 68:4447–4455. https://doi.org/10.1021/acs.jafc.0c00375
Spanova M, Czabany T, Zellnig GN et al (2010) Effect of lipid particle biogenesis on the subcellular distribution of squalene in the yeast Saccharomyces cerevisiae. J Biol Chem 285:6127–6133. https://doi.org/10.1074/jbc.M109.074229
Strand MK, Stuart GR, Longley MJ et al (2003) POS5 gene of Saccharomyces cerevisiae encodes a mitochondrial NADH kinase required for stability of mitochondrial DNA. Eukaryot Cell 2:809–820. https://doi.org/10.1128/EC.2.4.809-820.2003
Sun C, Ding Y, Cheng B, Zong Y (2020) Using engineered Escherichia coli to synthesize squalene with optimized manipulation of squalene synthase and mevalonate pathway. bioRxiv. https://doi.org/10.1101/2020.10.28.360032
Ta MT, Kapterian TS, Fei W et al (2012) Accumulation of squalene is associated with the clustering of lipid droplets. FEBS J 279:4231–4244. https://doi.org/10.1111/febs.12015
Tang WY, Wang DP, Tian Y et al (2021) Metabolic engineering of Yarrowia lipolytica for improving squalene production. Biores Technol 323:124652. https://doi.org/10.1016/j.biortech.2020.124652
Tateno M, Stone BJ, Srodulski SJ et al (2020) Synthetic biology-derived triterpenes as efficacious immunomodulating adjuvants. Sci Rep 10:17090. https://doi.org/10.1038/s41598-020-73868-6
Thompson A, Kwak S, Jin Y (2014) Squalene production using Saccharomyces cerevisiae. i-ACES 1:57–63
Tokuhiro K, Muramatsu M, Ohto C et al (2009) Overproduction of geranylgeraniol by metabolically engineered Saccharomyces cerevisiae. Appl Environ Microbiol 75:5536–5543. https://doi.org/10.1128/AEM.00277-09
Tsujimoto M (1916) A highly unsaturated hydrocarbon in shark liver oil. J Ind Eng Chem 8:889–896
Vik Å, Rine J (2001) Upc2p and Ecm22p, dual regulators of sterol biosynthesis in Saccharomyces cerevisiae. Mol Cell Biol 21:6395–6405. https://doi.org/10.1128/MCB.21.19.6395-6405.2001
Walker KA, Knuth ME, Fong NM, Beetham PR (2009) Production of squalene using yeast U.S. Patent Application No. 12/471,273 (3 Dec 2009)
Wei L, Kwak S, Lane JLS et al (2018) Improved squalene production through increasing lipid contents in Saccharomyces cerevisiae. Biotechnol Bioeng 115:1793–1800. https://doi.org/10.1002/bit.26595
Wei LJ, Cao X, Liu JJ et al (2021) Increased accumulation of squalene in engineered Yarrowia lipolytica through deletion of PEX10 and URE2. Appl Environ Microbiol 87:e00481-e521. https://doi.org/10.1128/AEM.00481-21
Westfall PJ, Pitera DJ, Lenihan JR et al (2012) From the Cover: PNAS Plus: production of amorphadiene in yeast, and its conversion to dihydroartemisinic acid, precursor to the antimalarial agent artemisinin. Proc Natl Acad Sci 109:E111–E118. https://doi.org/10.1073/pnas.1110740109
Wright R, Basson M, D’Ari L, Rine J (1988) Increased amounts of HMG-CoA reductase induce “karmellae”: a proliferation of stacked membrane pairs surrounding the yeast nucleus. J Cell Biol 107:101–114. https://doi.org/10.1083/jcb.107.1.101
Xu W, Chai C, Shao L et al (2016) Metabolic engineering of Rhodopseudomonas palustris for squalene production. J Ind Microbiol Biotechnol 43:719–725. https://doi.org/10.1007/s10295-016-1745-7
Xu W, Yao J, Liu L et al (2019) Improving squalene production by enhancing the NADPH/NADP+ ratio, modifying the isoprenoid-feeding module and blocking the menaquinone pathway in Escherichia coli. Biotechnol Biofuels 12:1–9. https://doi.org/10.1186/s13068-019-1415-x
Xu W, Wang D, Fan J et al (2021) Improving squalene production by blocking the competitive branched pathways and expressing rate-limiting enzymes in Rhodopseudomonas palustris. Biotechnol Appl Biochem. https://doi.org/10.1002/bab.2222
Yue CJ, Jiang Y (2009) Impact of methyl jasmonate on squalene biosynthesis in microalga Schizochytrium mangrovei. Process Biochem 44:923–927. https://doi.org/10.1016/j.procbio.2009.03.016
Zhang A, Xie Y, He Y et al (2019) Bio-based squalene production by Aurantiochytrium sp. through optimization of culture conditions, and elucidation of the putative biosynthetic pathway genes. Bioresour Technol 287:121415. https://doi.org/10.1016/j.biortech.2019.121415
Zhou C, Li M, Lu S et al (2021) Engineering of cis-element in Saccharomyces cerevisiae for efficient accumulation of value-added compound squalene via downregulation of the downstream metabolic flux. J Agric Food Chem 69:12474–12484. https://doi.org/10.1021/acs.jafc.1c04978
Zhu Z-T, Du M-M, Gao B et al (2021) Metabolic compartmentalization in yeast mitochondria: burden and solution for squalene overproduction. Metab Eng 68:232–245. https://doi.org/10.1016/j.ymben.2021.10.011
Zhuang X, Chappell J (2015) Building terpene production platforms in yeast. Biotechnol Bioeng 112:1854–1864. https://doi.org/10.1002/bit.25588
Acknowledgements
SM acknowledges the Science and Engineering Research Board (SERB), India for funding the project [CRG/2019/003323]. KP acknowledges the Department of Biotechnology India, and the Department of Science and Technology (DST), India for awarding the fellowship. The Director, CSIR—Central Food Technological Research Institute (CFTRI), Mysuru, India is also acknowledged for supporting this work.
Funding
This work was supported by the Science and Engineering Research Board (SERB) [CRG/2019/003323].
Author information
Authors and Affiliations
Contributions
KP and SM wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
Not applicable.
Consent for publication
All authors agreed to manuscript publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Paramasivan, K., Mutturi, S. Recent advances in the microbial production of squalene. World J Microbiol Biotechnol 38, 91 (2022). https://doi.org/10.1007/s11274-022-03273-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-022-03273-w