Abstract
Shiga toxin-producing Escherichia coli (STEC) are recognized as being responsible for many cases of foodborne diseases worldwide. Cattle are the main reservoir of STEC, shedding the microorganisms in their feces. The serogroup STEC O91 has been associated with hemorrhagic colitis and hemolytic uremic syndrome. Locus of Adhesion and Autoaggregation (LAA) and its hes gene are related to the pathogenicity of STEC and the ability to form biofilms. Considering the frequent isolation of STEC O91, the biofilm-forming ability, and the possible role of hes in the pathogenicity of STEC, we propose to evaluate the ability of STEC to form biofilms and to evaluate the expression of hes before and after of biofilm formation. All strains were classified as strong biofilm-forming. The hes expression showed variability between strains before and after biofilm formation, and this may be due to other genes carried by each strain. This study is the first to report the relationship between biofilm formation, and hes expression and proposes that the analysis and diagnosis of LAA, especially hes as STEC O91 virulence factors, could elucidate these unknown mechanisms. Considering that there is no specific treatment for HUS, only supportive care, it is necessary to know the survival and virulence mechanisms of STEC O91.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Shiga toxin-producing Escherichia coli (STEC) are zoonotic pathogens associated with human diseases such as hemorrhagic colitis (HC) and hemolytic-uremic syndrome (HUS) (Toma et al. 2004). Cattle are the main reservoir of STEC, shedding the microorganisms in their faces (Fernández and Padola 2012). Moreover, chickens and pigs have also been reservoirs from STEC (Alonso et al. 2016; Colello et al. 2016; Haymaker et al. 2019; Smith-Palmer et al. 2018). The consumption of contaminated water, which is increasingly concerning as a source of contamination for fruits and vegetables, along with the consumption of undercooked meat, and direct contact with animal reservoirs and the environment, are some of the potential routes of human exposure to STEC (Angel Villegas et al. 2013; EFSA 2020). STEC has been associated with more than two million acute illnesses worldwide (FAO & WHO 2018). Shiga toxins (Stx) are recognized as the main virulence factor in the pathogenicity of STEC; however, to produce damage, the bacteria have to adhere to the gastrointestinal tract of the host (Tarr and Chandler 2005). The Locus of Enterocyte and Effacement (LEE) encodes genes that participate in the adherence and lesion of the intestinal epithelium (Spears et al. 2006). STEC strains lacking LEE (LEE-negative) have also been isolated from cases of illness. Furthermore, the mechanisms by which LEE-negative STEC strains adhere to the host intestinal epithelium remain (Herold et al. 2009). Hemagglutinin from STEC (Hes) is a protein encoded by the hes gene, that could participate as virulence factors in colonization, adhesion, and autoaggregation. The hes gen is located in the first of four modules of the Locus of Adhesion and Autoaggregation (LAA). LAA contains 80 genes organized into four modules, being able to be present as a complete (4 modules) or incomplete (with less than 4 modules) structure. In addition, it has been demonstrated that LAA plays an essential role in the emergence of LEE-negative STEC strains, contributing to the evolution of the virulence of this pathogen. These authors propose the investigation of hes as a potential marker of LAA (Montero et al. 2017). In previous studies, Colello et al. (2018) and Vélez et al. (2020) demonstrated that hes was present in many LAA-positive STEC strains. LAA has been related to different parameters of pathogenicity as biofilm formation (Montero et al. 2017; Vélez et al. 2021). Biofilms are a community of bacterial cells enclosed in a self-producing matrix and adhered to biotic or abiotic surfaces (Donlan and Costerton 2002) and it has been recognized that biofilms can produce contamination and spoilage in food and pose a risk to consumer health (Angel Villegas et al. 2013; Vélez et al. 2022). Moreover, biofilm formation causes significant monetary loss in the food industry (Lindsay and Holy 2006). Nüesch-Inderbinen et al. (2018) found that LAA, as a complete structure, was identified in STEC O91 as highly pathogenic and isolated from clinical cases. The serogroup O91 ranked in the top five of the serogroups non-O157 related to human infection (Bielaszewska et al. 2009; EFSA 2012). Considering the frequent isolation of STEC O91, the biofilm formation capacity, and the possible role of hes in the pathogenicity of LAA-positive STEC, this study aimed to analyze the ability of STEC O91 to form biofilms and to evaluate the expression of hes in these mechanisms.
Materials and methods
Bacterial strains
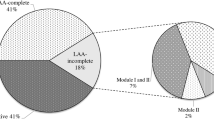
A total of 21 LEE-negative STEC strains belonging to serogroup O91 were isolated from cattle and food of animal origin and, in previous studies, the virulence profiles were analyzed by PCR (Table 1) (Colello et al. 2016; Etcheverría et al. 2010; Fernández et al. 2012; Padola et al. 2004; and Parma et al. 2000). Seventeen strains carried the four LAA modules (named LAA-positive), three strains carried some modules (named LAA-incomplete), and one strain does not carried any module (LAA-negative) (Table 1). Moreover, two mutants of O91 strains were used. One of them with LAA deleted (named O91∆LAA), and another with hes deleted (named O91∆hes). Two transformed strains were used, one strain without LAA with the insertion of hes plasmid (named O91∆LAApVB1_hes), and an E. coli non-adherent HB101 with the insertion of hes plasmid (named E. coli HB101pVB1_hes) (Montero et al. 2017).
Biofilm assay
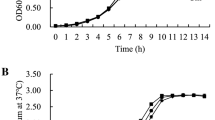
This assay was performed with all bacterial strains (wild-type and mutants) according to Cáceres et al. (2019). It was carried out in 96-well polystyrene flat-bottom culture plate (Greiner Bio-One, CELLSTAR®). The strains were grown in Luria Bertani (LB, Britania) broth at 37 °C for 18 h. The cultures were standardized, the optical density (OD) of each culture was measured and, dilutions were made adjusting the OD to 0.5 (equivalent to 2.5 × 108 CFU/mL). An aliquot of 10 µL was inoculated in each well by triplicate containing 190 µL of LB. It was incubated at 37 °C without shaking for 48 h with media renewal after 24 h. The microplates were washed once with double distilled water, fixed with methanol 100% (Biopack), 200 µL for 15 min, and stained with 200 µL of crystal violet (Biopack) 0.1% (p/v) in water. Then, the plates were washed with water three times, and the remaining dye was solubilized with 200 µL of 96% ethanol (Biopack). The biofilm formation was estimated by measuring the optical density at 570 nm (OD570) using the microplate reader (Labsystem Multi- scan EX – ICTSI Instrumentacion Cientifica Tecnica S. L). The OD of each strain was corrected by a cutting OD (ODc) (sum of the OD average of the control wells − 3 wells with non-inoculated sterile medium- and three times their standard deviation). Based on the results obtained, the strains were classified into four categories: non-biofilm forming (NBF): the OD of the strain is below the ODc (OD ≤ ODc); weak biofilm formers (WBF): the OD of the strain is between the ODc and the OD value corresponding to double the same (ODc < OD ≤ 2ODc); moderate biofilm formers (MBF): the OD of the strain is between twice the ODc value and the OD value corresponding to quadruple of the same (2ODc < OD ≤ 4ODc); and, strong biofilm formers (SBF): the OD of the strain is above four times the ODc (4ODc < OD) (Gómez et al. 2013). Biofilm formation was assessed with three replicates for each strain in three independent experiments.
Statistical analyzes
To assess the difference between OD for the O91 strains, a general mixed model with a fixed effect that was “strain” and a random effect “independent experiments” was fitted. When significant differences were found, a Tukey test was used to determine differences between levels of the fixed variable. The analysis was carried out with ‘R’ version 3.6.2 (R Development Core Team). We used MuMIn (Barton 2022) and lme4 (Bates et al. 2015).
Scanning electron microscopy analysis
Biofilm formation by E. coli HB101 and its transform E. coli HB101pBV1_hes, the wild-type strain VO 7-4-4, and its mutant O91∆LAA were observed by scanning electron microscopy (SEM). The study was carried out on 12-well polystyrene plates on which sterile metallic coupons were placed. The biofilm formation was realized with the methodology described in Sect. 2.2. The coupons were removed at three different times of the biofilm formation (6, 24, and 48 h). Each sample were washed with PBS to remove non-adherence cells, and fixed with a methanol solution 100% (Biopack) by 15 min. Subsequently, the samples were dehydrated. The dehydration was carried out by exposing them to 4 ethanol/water solutions from lowest to highest concentration (50%, 60%, 80% and 100%). Each solution was left to act for 15 min. Once dehydrated, coupons were critical point dried using liquid carbon dioxide within the pressurized chamber at 10 °C. The samples were metalized with Au 24 and observed under a Zeiss Evo10 SEM microscope.
Expression of hes gene before and after biofilm formation
Bacterial growth and RNA extraction
Five O91 strains were analyzed in two conditions, before and after biofilm formation: four strains carrying LAA (strains 3, 4, 5, 14) and one carrying only hes (strain 20) (Table 1). The strains were selected based on the isolation origin variability (cattle, dairy cows, calves, and food). A non-adherent strain was transformed by a plasmid with the hes gene (E. coli HB101pVB1_hes). This strain does not carry another set of genes; therefore, this strain will allow us to compare the exclusive participation of hes. To study the expression of the hes gene before biofilm formation, cultures were grown overnight (ON) at 37 ºC in LB broth with shaking and proceeding with the RNA extraction. The RNA was extracted to study the hes expression after the biofilm formation from a plate of 96-wells following the protocol described above. After 48 h of incubation, the biofilm formed was mechanically removed with a sterile spatula and collected in an Eppendorf ARNase-free for subsequent RNA extraction treatment.
Following both growths (ON and biofilm), total RNA was extracted using the TRIzol™ protocol. Additional treatment with DNase I (Roche Diagnostics GmbH) was performed before reverse transcription to eliminate genomic DNA contamination. One µg of RNA was incubated with 20 U of DNase I for 1 h at 37 ºC and 12 min at 72 ºC for inactivation. Afterward, cDNA was synthesized using the high-capacity cDNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA) according to the manufacturer’s instructions. Controls without reverse transcriptase were carried out for each sample.
Quantitative real-time PCR analysis
The transcription levels of the hes were evaluated by quantitative real-time PCR assays (qPCR). Two assays in three independent events were performed. The primers used to detect transcripts of the housekeeping gene, tufA, were taken from de Sablet et al. (2008) and hes was detected with primer design by Colello et al. (2023). Relative quantification reactions were performed on OneStep Plus Real-Time PCR System (Applied Biosystems). Each 20 µL qPCR mix contained 4 µL of 1/5 diluted cDNA template, 10 µL of 2X SYBR Green master mix (FastStart Universal SYBR Green Master, Roche), and 300 nM of each primer. A no-template control was included in each run to assess reagent contamination. The relative quantification was performed compared to the basal hes transcription of the E. coli HB101pVB1_hes strain. In this strain, the expression was induced by adding 2mM toluic acid (10 µL) and were grown in LB. Finally, the transcription levels relative to the control strain were calculated by the ΔΔCT method Pfaffl (2001) using the efficiency corresponding to each gene. A standard curve method was used to determine the amplification efficiency for the genes. Five dilutions of the control cDNA were made in a series of 5, that were amplified in duplicate. The determination of the amplification efficiency was carried out by a linear regression model according to the equation E = 10 [− 1 /slope]. Relative standard curves and was determined using the Relative Expression Software Tool (Pfaffl et al. 2002). Datasets were logarithmically transformed.
Results
Quantification of biofilm formation by O91
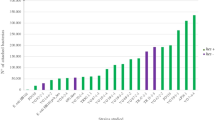
All O91 STEC strains could form biofilms and were classified as SBF, including mutant strains, based on the means of OD570 (Fig. 1). Although all strains were classified as SBF, high variability was found between wild-type strains (OD570 = 1.34 for the lowest and OD570 = 3.09 for the highest) with significant differences between them (p < 0.05; Table 1). Comparing the O91 STEC wild-type with their mutant strains, the average OD570 of M3 (O91H21∆LAApVB1_hes) was higher than the averages recorded for the O91 strains, while the M2 (O91∆hes) presented the lowest average. M3 and M2 differed significantly (p < 0.05; Fig. 2). This result confirms the participate of hes in biofilm formation in this assay.
Biofilm formation of STEC O91
OD: OD of biofilm formation / STEC strains. Green = mutants / Orange = wild-type LAA-incomplete / Red = wild-type LAA-negative / Grey = wild-type LAA-positive. M1: O91∆LAA, M2: O91∆hes, M3: O91H21∆LAApVB1_hes, 1–21: wild-types O91 strains. The line represents ODc > 0,620 (SBF).
OD average of the groups studied
Strains: M1: O91∆LAA; M2: O91∆hes; M3: O91H21∆LAApVB1_hes; N-/I: LAA-negative and LAA-incomplete wild-type strains; N+: LAA-positive wild-type strains. OD: Optic Density related to biofilm formation. Different letters indicate significant differences between means (p < 0.05). Errors bars indicate standard errors
Analyzed of biofilm formation with SEM
The images revealed that in the biofilm formed after 24 and 48 h of incubation, E. coli HB101pBV1_hes formed more biofilm than E. coli HB101 (Fig. 3). Moreover, the transformed strain increased the biofilm formation as the incubation time increased, unlike the E. coli HB101. Regarding VO 7-4-4 and its mutant O91∆LAA, LAA deletion modified the biofilm morphology. In the O91∆LAA strain, the biofilm formation was observed as flat and unstructured through time. In contrast, the wild-type strain formed biofilm as a dense network of mono or multi-layer cells embedded within a matrix of extracellular polymer material (Fig. 4).
Hes expression related to biofilm formation
Heterogeneous transcription levels of hes before and after biofilm formation by O91 strains were observed. The transcription levels of each isolate were expressed as fold change values relative to the control strain (E. coli HB101pVB1_hes) (Fig. 5). Regarding the normalization of the relative quantification reaction, for the construction of the standard curves for the target and the housekeeping genes, the amplification efficiencies of each gene were 98% (hes) and 99% (tufA). All the strains analyzed in both conditions showed detectable transcription levels of hes. Strains 3, 5, and 20 expressed higher levels of hes before biofilm formation than E. coli HB101pVB1_hes. In only one case, the expression of hes was higher after biofilm formation (strain 14) (Fig. 5). On the other hand, in strain 4, hes showed considerably low transcription levels in both conditions. The strain LAA-incomplete (strain 20), which carries only hes, showed markedly higher transcription levels (49-fold higher transcription than the control E. coli HB101pVB1_hes) (Table 1).
Expression of hes before and after the biofilm formation
Numbers 3, 4, 5, 14, and, 20 are O91 STEC strains analyzed. Strain E. coli HB101pVB1_hes was used as a control strain. Those strains with fold change values > 0 have higher transcription levels than the control, and those with fold change values < 0, have lower transcription levels than the control (Table 1)
Discussion
The biofilm formation by pathogens such as STEC is a mechanism that could participate in the infection of animals and humans (García and Percival 2011). Previous studies have analyzed non-O157 strains, detecting the potential expression of genes that participate in the infection/colonization of the host, but did not express them under basal conditions (Cadona et al. 2020). The genetic variation can be challenging in detecting hypervirulent STEC strains (Carter et al. 2022).
Previous studies have recognized that the biofilm formation capacity could be one of the factors for the persistence of STEC because this matrix offers the ability to survive in adverse conditions promoting infection to the host (Angel Villegas et al. 2013; Vélez et al. 2021; Wang et al. 2012). Another characteristic of biofilm formation is to allow the horizontal transference of genes. One example is the diverse distribution of LAA modules in O91 STEC strains, which could suggest that the acquisition of modules contributes to the high diversity of those strains and could occur on the biofilm (Nüesch-Inderbinen et al. 2021).
Hes is a member protein of the Heat-resistant agglutinin family produced by LEE-negative STEC strains (Montero et al. 2017). A former study showed that 53% of LEE-negative strains carried hes (Vélez et al. 2021) being the O91 strains one of the most prevalent for LAA and hes (Colello et al. 2018; Montero et al. 2017). An increase in clinical STEC O91 has been documented, but molecular data by which this serogroup causes the human disease is lacking (Nüesch-Inderbinen et al. 2018). This study proposes that the analysis and diagnosis of LAA, especially hes as a virulence factor of STEC O91 could clarify unknown mechanisms. STEC O91 strains were classified as SBF, in agreement with previous studies that have confirmed that LEE-negative strains are biofilm former (Vélez et al. 2021; Wang et al. 2012). In this study, the strain O91∆hes was form less biofilms than O91:H21∆LAA (OD570: 1.39 and OD570: 1.58, respectively) and the mutant with the insertion of a plasmid with hes O91:H21∆LAApVB1_hes form more biofilm than the others mutant (OD570: 2.76). This result corroborates that biofilm formation is affected by the presence of LAA and hes, since significant differences were detected between the previously described mutant strains (p < 0.001). Moreover, we could observe by SEM that the insertion of hes promoted the biofilms formation, and the deletion of LAA modified the structure of the biofilm in comparison with the wild-type strain. The biofilms formed by the wild-type strain (VO 7-4-4) present three-dimensional structures, which provide an opportunity for the microorganisms to accommodate each other and also to thrive under hostile conditions of pH, oxygen availability, and redox potential (Sauer et al. 2022).
The expression of hes was variable between strains before and after biofilm formation, confirming the participation of hes in concordance with Montero et al. (2017). The study of the expression of hes in biofilms within 48 h of maturation is highly relevant since one of the characteristics of biofilms is to allow gene transfer being able to acquire the modules of LAA under certain conditions (Montero et al. 2017; Nüesch-Inderbinen et al. 2018). Until now, our results demonstrate the expression of genes belonging to LAA in biofilm formation processes for the first time.
The participation of hes and LAA in the biofilm formation was expected, taking into account previously collected information (Montero et al. 2017; Vélez et al. 2021). With this gene expression study, we confirm the function of hes and LAA in one of the mechanisms STEC reaches the host.
Further studies are required to define how virulence gene expression contributes to biofilm formation. Although several factors involved in biofilm formation are related to colonization, elements in the survival of LEE-negative STEC within biofilms remain unknown. Hes participates in the growth of STEC, on some occasions, from the first hours of culture (ON). In addition to the intrinsic factors of the bacterium, gene expression may be influenced by extrinsic factors as different micro-environmental situations within a biofilm that may be responsible for the appearance of different phenotypes in a population (Roberfroid et al. 2016).
The biofilm formation processes are mediated by different genes and their production is affected by environmental conditions (Vogeleer et al. 2016). Considering that we analyzed wild-type strains that carried a wide variety of genes, the variability of the expression could be due to other genes carried by each strain.
There are discrepancies between the different studies that relate the ability to form biofilms with specific parameters. Naves et al. (2008) suggest that this could be because the strains behave in different ways in vitro, probably due to their genetic composition and the conditions of each laboratory. Therefore, it is necessary that when investigating this type of structure, the bacterial strain, the test conditions, and the surface involved are considered independently. Despite these potential limitations, the study in laboratories with different conditions found in the food industry, whether environment, surface, or temperature, is a practical and valuable tool to understand the behavior of bacteria that can be transmitted by food and be harmful to human health.
In concordance with Vogeleer et al. (2016) who distinguished that despite the development of new strategies to eradicate biofilms formed by pathogens, no practical solutions to remove STEC biofilms, this research focuses on the identification of hes as a factor that promotes STEC survival, especially in STEC O91. Prevention measures and control strategies to avoid biofilm formation are essential factors in reducing the transmission of STEC. Identifying STEC in different ecological niches is necessary to reduce food contamination and its transmission to humans after ingestion (Vélez et al. 2021).
In conclusion, this study confirms the strong ability to form biofilms by STEC O91 and that LAA and hes is a gene that participates in biofilm formation. It is relevant considering that the incidence of STEC O91 infections requires more attention than others because they have been little studied, and the HUS cases by this serogroup are increasing. Therefore, further research is needed to understand better the mechanism of biofilm formation and simultaneous expression of genes in the pathogenicity of LAA-positive STEC strains.
Data availability
Data is available.
References
Alonso MZ, Krüger A, Sanz ME, Padola NL, Lucchesi PMA (2016) Serotipos, perfiles de virulencia y subtipos de stx en Escherichia coli productor de toxina shiga aislados de productos de pollo. Rev Argent Microbiol 48(4):325–328. https://doi.org/10.1016/j.ram.2016.04.009
Angel Villegas N, Baronetti J, Albesa I, Polifroni R, Parma AE, Etcheverría A, Becerra M, Padola N, Paraje M (2013) Relevance of biofilms in the pathogenesis of Shiga-toxin-producing Escherichia coli infection. The Scientific World Journal, 2013. https://doi.org/10.1155/2013/607258
Barton K (2022) Package ‘MuMIn’ Version 1.46.0. R Package, Version 1.46.0
Bates D, Mächler M, Bolker BM, Walker SC (2015) Fitting linear mixed-effects models using lme4. J Stat Softw 67(1). https://doi.org/10.18637/jss.v067.i01
Bielaszewska M, Stoewe F, Fruth A, Zhang W, Prager R, Brockmeyer J, Mellmann A, Karch H, Friedrich AW (2009) Shiga toxin, cytolethal distending toxin, and hemolysin repertoires in clinical Escherichia coli O91 isolates. J Clin Microbiol 47(7):2061–2066. https://doi.org/10.1128/JCM.00201-09
Cáceres ME, Etcheverría AI, Padola NL (2019) Effects of the culture medium and the methodology applied on the biofilm formation of diarrheagenic Escherichia coli strains. Rev Argent Microbiol 51(3):208–213. https://doi.org/10.1016/j.ram.2018.04.007
Cadona JS, Burgán J, González J, Bustamante AV, Sanso AM (2020) Differential expression of the virulence gene nleB among Shiga toxin-producing Escherichia coli strains. Heliyon. https://doi.org/10.1016/j.heliyon.2020.e04277
Carter MQ, Laniohan N, Lo CC, Chain PSG (2022) Comparative Genomics Applied to Systematically Assess Pathogenicity Potential in Shiga Toxin-Producing Escherichia coli O145:H28. Microorganisms 10(5). https://doi.org/10.3390/microorganisms10050866
Colello R, Cáceres ME, Ruiz MJ, Sanz M, Etcheverría AI, Padola NL (2016) From farm to table: follow-up of Shiga toxin-producing Escherichia coli throughout the pork production chain in Argentina. Front Microbiol 7(FEB):1–7. https://doi.org/10.3389/fmicb.2016.00093
Colello R, Vélez MV, González J, Montero DA, Bustamante AV, Del Canto F, Etcheverría AI, Vidal R, Padola NL (2018) First report of the distribution of locus of adhesion and autoaggregation (LAA) pathogenicity island in LEE-negative Shiga toxin-producing Escherichia coli isolates from Argentina. Microb Pathog 123(July):259–263. https://doi.org/10.1016/j.micpath.2018.07.011
Colello R, Vélez MV, Nieto Farias MV, Rodríguez M, Montero D, Vidal R, Etcheverría AI, Padola NL (2023) Expression of hes, iha, and tpsA codified in Locus of Adhesion and Autoaggregation and their involvement in the capability of shiga toxin-producing Escherichia coli strains to adhere to epithelial cells. BMC Res Not 16(1):163. https://doi.org/10.1186/s13104-023-06433-9
de Sablet T, Bertin Y, Vareille M, Girardeau JP, Garrivier A, Gobert AP, Martin C (2008) Differential expression of stx2 variants in Shiga toxin-producing Escherichia coli belonging to seropathotypes A and C. Microbiology 154(1):176–186. https://doi.org/10.1099/mic.0.2007/009704-0
Donlan RM, Costerton JW (2002) Biofilms: survival mechanisms of clinically relevant microorganisms. Clin Microb Rev 15(2):167–193. https://doi.org/10.1128/CMR.15.2.167-193.2002
EFSA. (2012) The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2010. Euro Surveillance: Bulletin Européen Sur Les maladies Transmissibles. Eur Commun Disease Bull, 17(10), 1–442. https://doi.org/10.2903/j.efsa.2012.2597
EFSA (2020) Pathogenicity assessment of Shiga toxin-producing Escherichia coli (STEC) and the public health risk posed by contamination of food with STEC. EFSA J 18(1):1–105. https://doi.org/10.2903/j.efsa.2020.5967
Etcheverría AI, Padola NL, Sanz ME, Polifroni R, Krüger A, Passucci J, Rodríguez EM, Taraborelli AL, Ballerio M, Parma AE (2010) Occurrence of Shiga toxin-producing E. Coli (STEC) on carcasses and retail beef cuts in the marketing chain of beef in Argentina. Meat Sci 86(2):418–421. https://doi.org/10.1016/j.meatsci.2010.05.027
FAO & WHO, (2018). Shiga toxin-producing Escherichia coli (STEC) and food: attribution, characterization, and monitoring in Microbiological risks. Assesment series. https://apps.who.int/iris/bitstream/handle/10665/272871/9789241514279-eng.pdf
Fernández D & Padola NL (2012). Escherichia coli verocitotoxigénico: Varias cuestiones...y los tambos también. Rev Argent Microbiol, 44(4), 312–323. ISSN 0325–7541.
Fernández D, Sanz ME, Parma AE, Padola NL (2012) Short communication: characterization of Shiga toxin-producing Escherichia coli isolated from newborn, milkfed, and growing calves in Argentina. J Dairy Sci 95(9):5340–5343. https://doi.org/10.3168/jds.2011-5140
García AB, Percival SL (2011) Zoonotic infections: the role of Biofilms. Biofilm and Veterinary Medicine. https://doi.org/10.1007/978-3-642-21289-5_3
Gómez J, Gómez-Lus ML, Bas P, Ramos C, Cafini F, Maestre JR, Prieto J (2013) ¿Es la cuantificación del biofilm un elemento diferenciador en la patogénia de bacilos gramnegativos? Rev Esp De Quimiot 26(2):97–102
Haymaker J, Sharma M, Parveen S, Hashem F, May EB, Handy ET, White C, East C, Bradshaw R, Micallef SA, Callahan MT, Allard S, Anderson B, Craighead S, Gartley S, Vanore A, Kniel KE, Solaiman S, Bui A, Sapkota AR (2019) Prevalence of Shiga-toxigenic and atypical enteropathogenic Escherichia coli in untreated surface water and reclaimed water in the Mid Atlantic U.S. Environ Res 172:630–636. https://doi.org/10.1016/j.envres.2019.02.019
Herold S, Paton JC, Paton AW (2009) Sab, a novel autotransporter of Locus of Enterocyte Effacement-negative shiga-toxigenic Escherichia coli O113:H21, contributes to adherence and biofilm formation. Infect Inmun 77(8):3234–3243. https://doi.org/10.1128/IAI.00031-09
Lindsay D, Holy A, Von (2006) What food safety professionals should know about bacterial biofilms. Br Food J 108(1):27–37. https://doi.org/10.1108/00070700610637616
Montero D, Velasco J, Del Canto F, Puente JL, Padola NL, Rasko DA, Farfán M, Salazar JC, Vidal R (2017) Locus of adhesion and autoaggregation (LAA), a pathogenicity island present in emerging Shiga Toxin-producing Escherichia coli strains. Sci Rep 7(1):1–13. https://doi.org/10.1038/s41598-017-06999-y
Naves P, del Prado G, Huelves L, Gracia M, Ruiz V, Blanco J, Dahbi G, Blanco M, del Carmen Ponte M, Soriano F (2008) Correlation between virulence factors and in vitro biofilm formation by Escherichia coli strains. Microb Pathog. 2008;45:86–91
Nüesch-Inderbinen M, Cernela N, Wüthrich D, Egli A, Stephan R (2018) Genetic characterization of Shiga toxin producing Escherichia coli belonging to the emerging hybrid pathotype O80:H2 isolated from humans 2010–2017 in Switzerland. Int J Med Microb 308(5):534–538. https://doi.org/10.1016/j.ijmm.2018.05.007
Nüesch-Inderbinen M, Stevens MJ, Cernela N, Müller A, Biggel M, Stephan R (2021). Distribution of virulence factors, antimicrobial resistance genes and phylogenetic relatedness among shiga toxin-producing Escherichia coli serogroup O91 from human infections. Int J Med Microbiol 311(8):151541. https://doi.org/10.1016/j.ijmm.2021.151541
Padola NL, Sanz ME, Blanco JE, Blanco M, Blanco J, Etcheverria AI, Arroyo GH, Usera MA, Parma AE (2004) Serotypes and virulence genes of bovine shigatoxigenic Escherichia coli (STEC) isolated from a feedlot in Argentina. Vet Microb 100(1–2):3–9. https://doi.org/10.1016/S0378-1135(03)00127-5
Parma AE, Sanz ME, Blanco JE, Blanco J, Viñas MR, Blanco M, Padola NL, Etcheverría AI (2000) Virulence genotypes and serotypes of verotoxigenic Escherichia coli isolated from cattle and foods in Argentina: Importance in public health. Eu J Epidemiol 16(8):757–762. https://doi.org/10.1023/A:1026746016896
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acid Reserarch 29:2007. https://doi.org/10.1111/j.1365-2966.2012.21196.x
Pfaffl MW, Graham WH, Dempfle L (2002) Erratum: Corrigendum to Modeling of the planetary ball-milling process: The case study of ceramic powders J. Eur. Ceram. Soc. 36 (9) (2016) 2205–2212] (https://doi.org/10.1016/j.jeurceramsoc.2015.09.032) (S0955221915301515) https://doi.org/10.1016/j.jeurceramsoc.2016.09.026
Roberfroid S, Vanderleyden J, Steenackers H (2016) Gene expression variability in clonal populations: causes and consequences. Crit Rev Microb 42(6):969–984. https://doi.org/10.3109/1040841X.2015.1122571
Sauer K, Stoodley P, Goeres DM, Hall-Stoodley L, Burmølle M, Stewart PS, Bjarnsholt T (2022) The biofilm life cycle: expanding the conceptual model of biofilm formation. Nat Rev Microbiol 20(10):608–620. https://doi.org/10.1038/s41579-022-00767-0
Smith-Palmer A, Hawkins G, Couper S, Maxwell H, Reynolds B, Harkins V, Allison L, Hanson M (2018) Global spread of stec and managing the consequences. Arch Dis Childhood 103(1). https://doi.org/10.1136/archdischild-2018-rcpch.469. A196 LP-A197
Spears KJ, Roe AJ, Gally DL (2006) A comparison of enteropathogenic and enterohaemorrhagic Escherichia coli pathogenesis. FEMS Microb Let 255(2):187–202. https://doi.org/10.1111/j.1574-6968.2006.00119.x
Tarr PI, Gordon CA, Chandler WL (2005) Seminar shiga-toxin-producing. Epidemiology, 1073–1086
Toma C, Espinosa EM, Song T, Miliwebsky E, Chinen I, Iyoda S, Iwanaga M, Rivas M (2004) Distribution of putative adhesins in different seropathotypes of Shiga toxin-producing Escherichia coli. J Clin Microbiol 42(11):4937–4946. https://doi.org/10.1128/JCM.42.11.4937-4946.2004
Vélez MV, Colello R, Etcheverría AI, Vidal RM, Montero DA, Acuña P, Guillén Fretes RM, Toro M, Padola NL (2020) Distribution of Locus of Adhesion and Autoaggregation and hes gene in STEC Strains from countries of Latin America. Curr Microbiol 77(9):2111–2117. https://doi.org/10.1007/s00284-020-02062-8
Vélez MV, Colello R, Etcheverría S, Etcheverría AI, Padola NL (2021) Biofilm formation by LEE-negative Shiga toxin–producing Escherichia coli strains. Microb Patho 157. https://doi.org/10.1016/j.micpath.2021.105006
Vélez MV, Colello R, Etcheverría AI, Padola NL (2022) Escherichia coli productora de toxina shiga: El desafío De adherirse para sobrevivir. Rev Argent Microbiol. https://doi.org/10.1016/j.ram.2022.04.001
Vogeleer P, Tremblay YDN, Jubelin G, Jacques M, Harel J (2016) Biofilm-forming abilities of Shiga toxin-producing Escherichia coli isolates associated with human infections. App Environ Microb 82(5):1448–1458. https://doi.org/10.1128/AEM.02983-15
Wang R, Bono JL, Kalchayanand N, Shackelford S, Harhay DM (2012) Biofilm formation by shiga toxin-producing Escherichia coli O157:H7 and non-O157 strains and their tolerance to sanitizers commonly used in the food processing environment. J Food Prot 75(8):1418–1428. https://doi.org/10.4315/0362-028X.JFP-11-427
Acknowledgements
The authors thank María Rosa Ortíz for her technical assistance.
Funding
This work was supported by PICT 2015–2666, CIC, and SECAT from Argentina.
Author information
Authors and Affiliations
Contributions
Conceptualization, R. Colello., M.V. Vélez, and N.L. Padola.; Funding acquisition, N.L. Padola.; Investigation, R. Colello. and M.V. Vélez.; Methodology, M.V. Vélez and L. E. Paz.; Supervision, N.L. Padola. and A.I. Etcheverría.; Validation, M.V.Vélez and M.V. Nieto Farias.; Visualization, R. Colello., and N.L. Padola.; Writing-original draft, M.V. Vélez.; Writing-review and editing, R. Colello., M.V. Vélez., M.V. Nieto Farias., R. Vidal., A.I. Etcheverria. and N.L. Padola.
Corresponding author
Ethics declarations
Ethical approval
This declaration is not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Vélez, M.V., Colello, R., Nieto, M.V. et al. Transcription levels of hes and their involvement in the biofilm formation of Shiga toxin-producing Escherichia coli O91. Vet Res Commun 48, 1821–1830 (2024). https://doi.org/10.1007/s11259-024-10308-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11259-024-10308-0