Abstract
Shiga toxin-producing Escherichia coli (STEC) are zoonotic food pathogens associated with foodborne diarrheal illness, hemorrhagic colitis, and complications such as hemolytic uremic syndrome (HUS). The ability to adhere to epithelial cells is an important virulence trait, and pathogenicity islands (PAIs) play an important role on it. Some STEC carrying a PAI named locus of enterocyte effacement (LEE-positive) have been frequently associated to HUS; however, STEC that do not carry LEE (LEE-negative) have also been associated with this outcome. The burden of disease caused by LEE-negative STEC has increased recently in several countries like Argentina, Chile, and Paraguay. A new PAI –the Locus of Adhesion and Autoagregation (LAA)—has been associated to severe disease in humans. In this study, we aimed to analyze the distribution of LAA and its possible predictor, the gene hes, in LEE-negative STEC strains isolated from Chile and Paraguay from different sources. The presence of the different LAA modules and hes were detected by PCR. LAA was found in 41.6% and 41.0% of strains isolated from Chile and Paraguay, respectively. Strains were isolated from diverse origins and belonged to several serogroups including O91, O103, and O113. The hes gene was detected in 50% of the isolates from Paraguay and Chile. Therefore, the detection of LAA and hes in STEC could complement current genetic evaluation schemes, allowing to classify LEE negative STEC strains as LAA-positive or LAA-negative STEC strains.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Foodborne pathogens are an important public health issue in many countries and are responsible for a substantial burden of disease in the developed world. In developing countries, such as Argentina, Paraguay, and Chile, this problem is of great concern [1]. One of the microorganisms involved in foodborne diseases is Shiga toxin-producing Escherichia coli (STEC), a diverse group of bacteria capable of causing severe human diseases such as hemorrhagic colitis (HC) and the hemolytic-uremic syndrome (HUS) [2]. Cattle are the main reservoir of STEC; however, pigs, dairy products, nuts, seeds, water -which has increasingly become a concern as a source of contamination for fruits and vegetables-, and person to person contact have been shown to commonly participate in transmission [3,4,5]. STEC surveillance systems are different in each Latin American country. In general, disease incidence data relies primarily on either the foodborne disease surveillance system (Argentina and Paraguay) and/or through the acute diarrheal surveillance system (Argentina, Chile, and Paraguay, among others). Argentina has the highest number of cases caused by STEC worldwide: HUS incidence is 8.4 cases per 100,000 children less than 5 years old [6]. On the other hand, the incidence is 3.4 cases per 100,000 children in Chile (Metropolitan Region) [7]. In both countries HUS represents one of the main causes of acute renal failure in children with lethality ranges from 2 to 6% [8]. In Paraguay, routine surveillance is performed with all the stool cultures that are sent to the Central Public Health Laboratory. The incidence of HUS is low. Between 2013 and 2015, eight STEC diarrhea cases and ten HUS cases without STEC isolation were reported [8, 9].
STEC refers to the pathotype capable of producing Shiga toxin type 1 (Stx1), type 2 (Stx2), or both, which are encoded by stx1 and stx2 genes, respectively [2]. Epidemiological studies suggest that STEC strains encoding stx2 are more virulent than those harboring stx1 only [10]. Although Stx1 is less cytotoxic than Stx2, it may potentially cause disease in humans, but the information about its clinical implications is limited [11]. In addition, some STEC strains carry the Locus of Enterocyte Effacement (LEE), a pathogenicity island (PAI) which encodes genes necessary to produce attaching and effacing (A/E) lesions on the intestinal epithelium. LEE is also carried by enteropathogenic E. coli (EPEC) [12] and Escherichia albertii strains which might have been misidentified as LEE-positive STEC or EPEC because they carry the eae gene [13]. However, STEC strains lacking LEE (LEE-negative), such as those belonging to serogroups O91, O113 and O174 [14], have also been isolated from cases of severe illness, including HUS [15]. A great number of adhesins, including the pO113-encoded autoagglutinating adhesin (Saa), have been linked to the pathogenesis of LEE-negative STEC [16]. Montero et al. [17] described and characterized a member of the Heat resistant agglutinin family (Hra Family), named Hemagglutinin from Shiga toxin-producing E. coli (Hes) that participates in several colonization-associated phenotypes, including hemagglutination, adhesion and autoaggregation. Hes is encoded by a gene located in a PAI of 86-kb chromosomal mosaic element named the Locus of Adhesion and Autoaggregation (LAA), which contains 80 genes organized into four modules: module I (hes and other genes), module II (iha, lesP and others genes), module III (pagC, tpsA, tpsB and other genes) and module IV (agn43 and other genes). Moreover, LAA may be present as a “complete” (4 modules) or “incomplete” (with less than 4 modules) structure [17].
LAA presence is associated with severe human disease and although the mechanisms used by LEE-negative STEC strains to colonize the human intestine are not clear yet [17]. Recently, the role of LAA in intestinal colonization was demonstrated in a murine model of STEC infection, suggesting that LAA may be also involved in the adherence of STEC to the human intestine [18, 19]. Moreover, the association of LAA with stx1a, stx2a, stx2d and cdtB toxin genes that cause severe disease has been demonstrated [18]. In a previous study in Argentina, our group has demonstrated the presence of LAA in 46% of LEE-negative STEC strains isolated from different sources [20]. Because the incidence of HUS caused by LEE-negative STEC strains has been increased in several countries [8], in this study, we aimed to analyze the distribution of LAA and hes in LEE-negative STEC strains isolated in Chile and Paraguay from different sources.
Materials and Methods
Strains and Serotyping
A total of 128 LEE-negative STEC strains were analyzed: 72 from Chile and 56 from Paraguay. STEC strains were isolated from beef cattle (n = 94), meat (n = 31), cheese (n = 2) and wild bird (n = 1). These isolates were previously analyzed for the presence of stx1, stx2, and saa genes by PCR (Table 1) [21]. The serogroup was determined by micro-agglutination test described by Guinée et al. [22] and modified by Blanco et al. [23] (Table 1).
PCR Amplification
STEC strains were characterized by multiplex PCR to detect LAA modules I, II, and III [17]. Additionally, the presence of agn43, as a marker of module IV, and hes were characterized by monoplex PCR [20, 24]. One LEE positive STEC strain was used as a negative control (O157:H7 EDL933). PCR reactions (multiplex and monoplex) were performed and standardized in a total volume of 50 μl by using a T-17 thermal cycler (Ivema). The reaction mixture contained 500 mM KCl, 100 mM Tris–HCl pH 9, Triton X-100, 25 mM MgCl2, 200 μM 4 deoxynucleotides (dATP, dGTP, dCTP, dTTP), 1U of TaqDNA Polymerase Highway® (Inbio), and 5 μl of DNA. The DNA was obtained by boiling bacteria suspended in sterile water for 10 min as previously described [25]. The LAA and hes primers were reported by Montero et al. [17]. Amplification products were separated by electrophoresis on 2% agarose gels containing 0.8 μg/ml of ethidium bromide in running buffer and were visualized in a UV transilluminator.
Results
STEC strains were considered as LAA-complete when they harbored modules I, II, III, and IV and were considered as LAA-incomplete when less than four modules were detected.
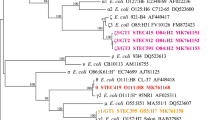
Complete LAA structure was identified in strains from Paraguay and Chile in similar frequencies: 41.0% (23/56) and 41.6% (30/72), respectively (Figs. 1 and 2). Similarly, the gene hes was detected in 50% of the isolates from Paraguay (28/56) and Chile (36/72) (Table 1). STEC strains carrying LAA (complete or incomplete) were distributed in 23 serogroups. The most frequent were O91 (4/4, 100%), O113 (5/6, 83%), O103 (6/9, 67%) and O174 (2/5, 40%) (Table 1).
Regarding toxins, the stx2 gene was detected in 30% (7/23) and 73% (22/30) of STEC strains harboring LAA from Paraguay and Chile, respectively (Table 1).
Discussion
Although E. coli O157:H7 is the most prevalent serotype associated with HUS, there is growing concern on the global emergence of LEE-negative STEC that have been associated with outbreaks and/or severe human illness [26]. LEE-negative STEC strains have been detected in humans, animals, food, and the environment. Still, the pathogenic mechanisms used by this group of strains to colonize humans are yet to be elucidated [27]. LAA is a PAI of an emerging group of STEC strains that cause severe diseases in humans. In fact, the complete LAA island is present in LEE-negative STEC strains which have been isolated from cases of hemorrhagic colitis and HUS [17].
In this study, the complete LAA was widely distributed in STEC strains from different origins and belonging to several serogroups. LEE-negative STEC strains isolated from Chile (41.6%) and Paraguay (41.0%) harbored the complete LAA island with a frequency of detection that is similar to that previously reported Argentina (46%) [20].
Stx2-producing strains are more often associated with HUS than strains that produce Stx1 [28]. In this study, we detected stx2 in 30% and 73% of the isolated LAA positive obtained from Paraguay and Chile, respectively. Likewise, Colello et al. [20] found a significant association between the presence of a complete LAA island and stx2 in strains isolated in Argentina. In addition, LEE-negative STEC strains of serogroups O91, O113 and O174 have been isolated from cases of severe illness [29], and in our study most of the isolates belonging to these serogroups harbored LAA. PAIs like LAA has had a remarkable role in the emergence of LEE-negative STEC strains and may contribute to the evolution and virulence of pathogenic E. coli [18].
Montero et al. [17] suggested that hes is a potential genetic marker for LAA, and this raises a new possible epidemiological scenario [17] for STEC since it is widely distributed in LEE-negative STEC strains. Our results revealed that hes was present in all LAA positive strains, carrying the complete island. In concordance with Colello et al. [20], we also observed that hes was one of the most prevalent genes in LEE-negative STEC strains [18, 20].
Therefore, our work provides new data about the presence of LAA and the hes gene in STEC strains from different sources and isolated in two Latin-American countries. These results support previous findings that suggest that the current STEC classification based on the presence of LEE might be insufficient to detect human pathogenic STEC strains, and that hes detection could complement current genetic evaluation schemes to detect strains representing risk [17, 30].
Concluding, to define a STEC strain as pathogenic is complex because there are not combinations of markers that can predict the potential of a STEC strain to cause human disease. In LEE positive strains, stx2 and eae are predictive markers for severe disease. In LEE-negative STEC strains, the scenario is even more complex because there are not additional virulence factors to stx2 associated with severe disease. In this context, hes could be used as a marker allowing classify the LEE negative STEC strains as LAA positive or negative STEC strains, and therefore, inferring the potential risk of LEE-negative strains.
References
Schlundt J, Toyofuku H, Jansen J, Herbst S (2004) Emerging food-borne zoonoses. Rev Sci Tech 23(2):513–534
Paton JC, Paton AW (1998) Pathogenesis and diagnosis of Shiga toxin-producing Escherichia coli infections. Clin Microbiol Rev 11(3):450–479
Organization WH (2018) Shiga toxin-producing Escherichia coli (STEC) and food: Attribution, Characterization, and Monitoring
Haymaker J, Sharma M, Parveen S, Hashem F, May EB, Handy ET, White C, East C, Bradshaw R, Micallef SA (2019) Prevalence of Shiga-toxigenic and atypical enteropathogenic Escherichia coli in untreated surface water and reclaimed water in the Mid-Atlantic US. Environ Res 172:630–636
Smith-Palmer A, Hawkins G, Couper S, Maxwell H, Reynolds B, Harkins V, Allison L, Hanson M (2018) I1 Global spread of STEC and managing the consequences. BMJ Publishing Group Ltd
Rivas Marta CIaBECG (2016) Enterohemorrahagic (Shiga Toxin-Producing) Escherichia coli, vol 2. springer International Publishing Switzerland edn. https://doi.org/10.1007/978-3-319-45092-6
Prado V, Cavagnaro F (2008) Hemolytic uremic syndrome associated to Shiga-toxin producing Escherichia coli in Chilean children: Clinical and Epidemiological Aspects. Rev Chilena Infectol 25(6):435–444
Torres AG (2017) Escherichia coli diseases in Latin America-a ‘One Health’multidisciplinary approach’. Pathog Dis. https://doi.org/10.1093/femspd/ftx012
Weiler N, Orrego M, Alvarez M, Huber C (2017) Detección molecular de Escherichia coli diarreogénica en pacientes pediátricos con síndrome diarreico agudo en Paraguay. Mem Inst Investig Cienc Salud. https://doi.org/10.1016/j.ijfoodmicro.2011.10.032
Lee M-S, Koo S, Jeong D, Tesh V (2016) Shiga toxins as multi-functional proteins: induction of host cellular stress responses, role in pathogenesis and therapeutic applications. Toxins 8(3):77
Amézquita-López BA, Soto-Beltrán M, Lee BG, Yambao JC, Quiñones B (2018) Isolation, genotyping and antimicrobial resistance of Shiga toxin-producing Escherichia coli. J Microbiol Immunol Infect 51(4):425–434
Spears KJ, Roe AJ, Gally DL (2006) A comparison of enteropathogenic and enterohaemorrhagic Escherichia coli pathogenesis. FEMS Microbiol Lett 255(2):187–202
Ooka T, Seto K, Kawano K, Kobayashi H, Etoh Y, Ichihara S, Kaneko A, Isobe J, Yamaguchi K, Horikawa K (2012) Clinical significance of Escherichia albertii. Emerg Infect Dis 18(3):488
Galli L, Miliwebsky E, Irino K, Leotta G, Rivas M (2010) Virulence profile comparison between LEE-negative Shiga Toxin-producing Escherichia coli (STEC) strains isolated from cattle and humans. Vet Microbiol 143(2–4):307–313
Herold S, Paton JC, Paton AW (2009) Sab, a novel autotransporter of locus of enterocyte effacement-negative Shiga-toxigenic Escherichia coli O113: H21, contributes to adherence and biofilm formation. Infect Immun 77(8):3234–3243
Paton AW, Srimanote P, Woodrow MC, Paton JC (2001) Characterization of Saa, a Novel autoagglutinating adhesin produced by locus of enterocyte effacement-negative Shiga-ToxigenicEscherichia coli strains that are virulent for humans. Infect Immun 69(11):6999–7009
Montero DA, Velasco J, Del Canto F, Puente JL, Padola NL, Rasko DA, Farfán M, Salazar JC, Vidal R (2017) Locus of adhesion and autoaggregation (LAA), a pathogenicity island present in emerging Shiga Toxin–producing Escherichia coli strains. Sci Rep 7(1):7011
Montero DA, Canto FD, Velasco J, Colello R, Padola NL, Salazar JC, Martin CS, Oñate A, Blanco J, Rasko DA (2019) Cumulative acquisition of pathogenicity islands has shaped virulence potential and contributed to the emergence of LEE-negative Shiga toxin-producing Escherichia coli strains. Emerg Microbes Infect 8(1):486–502. https://doi.org/10.1080/22221751.2019.1595985
Montero DA, Del Canto F, Salazar JC, Cespedes S, Cádiz L, Arenas-Salinas M, Reyes J, Oñate Á, Vidal RM (2020) Immunization of mice with chimeric antigens displaying selected epitopes confers protection against intestinal colonization and renal damage caused by Shiga toxin-producing Escherichia coli. npj Vaccines 5(1):1–13
Colello R, Vélez MV, González J, Montero DA, Bustamante AV, Del Canto F, Etcheverría AI, Vidal R, Padola NL (2018) First report of the distribution of locus of adhesion and autoaggregation (LAA) pathogenicity island in LEE-negative Shiga toxin-producing Escherichia coli isolates from Argentina. Microb Pathog 123:259–263
Toro M, Rivera D, Jiménez MF, Díaz L, Navarrete P, Reyes-Jara A (2018) Isolation and characterization of non-O157 Shiga toxin-producing Escherichia coli (STEC) isolated from retail ground beef in Santiago, Chile. Food Microbiol 75:55–60
Guinée P, Jansen W, Wadström T, Sellwood R (1981) Escherichia coli associated with neonatal diarrhoea in piglets and calves. In: DeLeeuw PW, Guinée PAM (eds) Laboratory diagnosis in neonatal calf and pig diarrhea. Springer, Berlin
Blanco J, Blanco M, Alonso MP, Blanco JE, Garabal J, González EA (1992) Serogroups of Escherichia coli strains producing cytotoxic necrotizing factors CNF1 and CNF2. FEMS Microbiol Lett 96(2–3):155–159
Montero D, Orellana P, Gutiérrez D, Araya D, Salazar JC, Prado V, Oñate Á, del Canto F, Vidal R (2014) Immunoproteomic analysis to identify Shiga toxin-producing Escherichia coli outer membrane proteins expressed during human infection. Infect Immun 82(11):4767–4777
Parma A, Viñas M, Sanz M (1996) Improvement of the polymerase chain reaction to detect Escherichia coli Shiga-like toxin II gene from clinical isolates. J Microbiol Methods 26(1–2):81–85
Farfan MJ, Torres AG (2012) Molecular mechanisms that mediate colonization of Shiga toxin-producing Escherichia coli strains. Infect Immun 80(3):903–913
Doughty S, Sloan J, Bennett-Wood V, Robertson M, Robins-Browne RM, Hartland EL (2002) Identification of a novel fimbrial gene cluster related to long polar fimbriae in locus of enterocyte effacement-negative strains of enterohemorrhagic Escherichia coli. Infect Immun 70(12):6761–6769
Ethelberg S, Olsen KE, Scheutz F, Jensen C, Schiellerup P, Engberg J, Petersen AM, Olesen B, Gerner-Smidt P, Mølbak K (2004) Virulence factors for hemolytic uremic syndrome. Denmark Emerg Infect Dis 10(5):842
Brooks JT, Sowers EG, Wells JG, Greene KD, Griffin PM, Hoekstra RM, Strockbine NA (2005) Non-O157 Shiga toxin–producing Escherichia coli infections in the United States, 1983–2002. J Infect Dis 192(8):1422–1429
Joint F, Additives WECoF, Organization WH (2002) Evaluation of certain mycotoxins in food: fifty-sixth report of the Joint FAO/WHO Expert Committee on Food Additives. World Health Organization
Author information
Authors and Affiliations
Contributions
MVV, conducting experiments, data analysis and interpretation, writing of the manuscript. RC. Data analysis and interpretation, writing and revising of the manuscript. AIE, Study design, data interpretation, revising of the manuscript. RV. data analysis, editing of the manuscript. DAM, data interpretation, revising of the manuscript. PA contributing with strains isolated from Paraguay, revising of the manuscript. RMGF, data analysis, revising of the manuscript. MT, data analysis revising of the manuscript. NLP, study design, data interpretation, data analysis, revising of the manuscript. All authors approved the final manuscript version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Vélez, M.V., Colello, R., Etcheverría, A.I. et al. Distribution of Locus of Adhesion and Autoaggregation and hes Gene in STEC Strains from Countries of Latin America. Curr Microbiol 77, 2111–2117 (2020). https://doi.org/10.1007/s00284-020-02062-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-02062-8