Abstract
Cell division cycle 7 (CDC7) is a serine/threonine kinase, which plays a vital role in the replication initiation of DNA synthesis. Overexpression of the CDC7 in various tumor growths and in cell proliferation makes it a promising target for treatment of cancers. To investigate the binding between the CDC7 and furanone inhibitors, and in order to design highly potent inhibitors, a three-dimensional quantitative structure activity relationship (3D-QSAR) with molecular docking was performed. The optimum CoMSIA model showed significant statistical quality on all validation methods with a determination coefficient (R2 = 0.945), bootstrapping R2 mean (BS-R2 = 0.960), and leave-one-out cross-validation (Q2) coefficient of 0.545. The predictability of this model was evaluated by external validation using a test set of nine compounds with a predicted determination coefficient R2test of 0.96, besides the mean absolute error (MAE) of the test set was 0.258 log units. The extracted contour maps were used to identify the important regions, where the modification was necessary to design a new molecule with improved activity. Furthermore, a good consistency between the molecular docking and contour maps strongly demonstrates that the molecular modeling is reliable. Based on those obtained results, we designed several new potent CDC7 inhibitors, and their inhibitory activities were validated by the molecular models. Additionally, those newly designed inhibitors showed promising results in the preliminary in silico ADMET evaluations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
CDC7 is a serine/threonine kinase that is involved mainly in the DNA replication [1, 2] as well as a number of other important roles in various chromosomes transactions as the replication checkpoint regulation and centromeric heterochromatin formation [3, 4]. Since CDC7 is essential for DNA replication in terms of cancer, a preponderance of evidence indicates that high levels of CDC7 are reported to be linked to the proliferative capacity of tumor cells [5]. Overexpression of CDC7 was found to be in the different type of cancers as ovarian cancer [6], breast cancer [7], and pancreatic cancer [8]. Several studies exhibit that the CDC7 depletion does not trigger apoptotic cell death in normal human cells; in contract, the lack of CDC7 in cancer cell phase leads to apoptotic cell death [9]. This selective cytotoxic behavior on cancer cells [10] makes CDC7 an attractive target for designing new drugs, which could kill directly proliferating cancer cells without affecting normal.
Recently, a significant interest has arisen by several research groups to develop small-molecule inhibitors of CDC7 kinase with the low-nanomolar [11, 12], but till now, no compounds have been approved by the Food and Drug Administration (FDA) as drug as a CDC7 inhibitor. Therefore, there is always a need for designing newly potent CDC7 inhibitors.
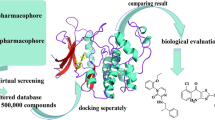
Computer-Aided-Drug-Design (CADD) could help to surmount these difficulties and shed light on new molecules for further research, while taking advantage of the spectacular improvement in the computer speed and capacity of computing molecular properties. Those can be exploited to build a quantitative structure activity relationship (QSAR) model to enable prediction of the biological activity and the efficacy of newly proposed chemical entities by means of cheminformatics methods. Recently, a series of some potent CDC7 inhibitors have been designed and reported by Irie et al. [13]; so far, this study represents the first predictive model for the CDC7 binding site based on the reported activities of this series of substituted furanone. That prompted us to aim an in silico study to highlight the structural factors controlling different active and inactive sites in the furanone scaffold to design new molecules with improved CDC7 inhibitory activity values.
To design new inhibitors, CADD including structure-based and ligand-based have proven to be valuable approaches in expediting and saving resources in drug discovery and pharmaceutical sciences [14]. Structure-based approach, which includes molecular docking, is used to predict the conceivable orientation of a ligand in the active site of a receptor as well as conformational changes of molecules. While ligand-based approaches include QSAR studies [15], which consist of mathematically derived rules that quantitatively describe activity in terms of molecular attributes, one of the popular methods in QSAR is comparative molecular similarity indexes analysis (CoMSIA) [16] that correlates changes in the 3D structural factors of chemical groups such as steric, electrostatic, and hydrophobic properties with the biological activity [17]. 3D-QSAR studies have been successfully used to explore the SARs of different chemical entities including inhibitors of PIM-1 kinase [18] and flavonoids binding site in GABAA receptors [19]. Accordingly, 3D-QSAR studies eliminate problems such as limitation in the prediction of stereochemistry of tested dataset and lack of recognition ability in search of newly active compounds [20] encountered with classical 2D-QSAR studies.
In present article, we report a 3D-QSAR study using CoMSIA methodology and molecular docking on a series of 31 CDC7 inhibitors [18] to illuminate some main patterns between CDC7 kinase and furanones and identify their key substituents and their mode of action. The results obtained from the 3D-QSAR model and molecular docking support each other. Consequently, several potent CDC7 inhibitors were designed, optimized, and their inhibitory activities were demonstrated by 3D-QSAR, molecular docking, and ADMET proprieties.
Material and methods
Data collection
A dataset of 31 furanone derivatives with their inhibitory activity against CDC7 kinase based on the studies of Irie et al. [18] were compiled to perform this study. For QSAR analysis, the in vitro half maximal inhibitory concentrations IC50 (nM) were converted into the corresponding pIC50 values (i.e., pIC50 is the negative logarithm of IC50 (pIC50 = −log(IC50)) and are listed with their corresponding structures in Table 1. The dataset was split rondamly into a training set (22 molecules) to build the quantitative model, and the remaining molecules were used to test the performance of the proposed model (test set).
Molecular modeling and molecular alignment
All modeling studies were performed using the SYBYL-X 2.0 molecular modeling package (Tripos Inc., St. Louis, USA). Three-dimensional structures of the studied compounds were built using the SKETCH option in SYBYL; then, they were minimized under the Tripos standard force field [21] with Gasteiger-Hückel atomic partial charges [22] by the Powell method with a gradient convergence criterion of 0.01 kcal/mol Å. The results obtained by CoMSIA closely relate to molecular alignment techniques. Thus, molecular alignment is regarded a sensitive step in the development of any 3D-QSAR study [23]. Figure. 1 depicts the proposed alignment; all molecules were aligned by distil-rigid alignment technique available in SYBYL, based on the most potent inhibitor 10, which was selected as a template to fit other molecules in training and test sets of furanone derivatives. Compound 10 was chosen to align the dataset in 3D-QSAR studies and to serve as a template molecule to visualize the contour maps.
CoMSIA studies
The comparative molecular similarity indexes analysis (CoMSIA) [16] method was carried out in SYBYL-X 2.0. All analyses were performed in a 3D regularly spaced grid of 2.0 Å in all Cartesians directions. A sp3 carbon with a Van Der Waals radius of 1.52 Å, net + 1.0 charge, hydrophobic interaction, hydrogen-bond donor, and acceptor properties of + 1.0 was used as a probe at every lattice point of the grid box to generate, respectively, the steric, electrostatic, hydrophobic, hydrogen bond donor, and acceptor fields from similar active molecules, to develop a CoMSIA model. In the present study, the value of attenuation factor, which controls the Gaussian function’s steepness, was set by default to 0.3 [24].
Partial least square analysis
Because of the enormous variables obtained from the fields’ calculations, the partial least square (PLS) regression method [25] was performed to evaluate a linear correlation between the 3D-QSAR descriptors and the biological activity values. In the first step, the cross (Q2) validation was performed by using a leave-one-out (LOO) [26] method where one compound is eliminated from the training set and its activity is predicted from the developed model using the residual (N-1) compounds. The same way is repeated until all compounds have been eliminated once. The model with the highest value of Q2 with the lowest standard error of estimate (Scv) and a minimal number of components was accepted. In order to reduce noise and increase the speed of the analytical process, the column filtering value (σ) was set to 2.0 kcal/mol. In the next step after determining the optimum number of components, they were used to derive the final PLS model with no validation method [27, 28] to create the maximum determination coefficient (R2).
Validation and predictive power of the model
The main objective of any QSAR study is to obtain a model with the highest predictive and generalization abilities. So, to evaluate the predictability of the developed 3D-QSAR models, nine compounds were used as a testing set [29]. These molecules were aligned using the same methods described above, and then, their inhibitory activities were predicted using the generated 3D-QSAR model from the training set. In addition, the mean absolute error (MAE) was applied to evaluate the model [30] using the MAE-based metrics (MAE ≤ 0.1 × training set range and MAE + 3 × σAE ≤ 0.2 × training set range) based on prediction errors.
Y-randomization test
The generated model was further validated by the Y-randomization method [31]. The activities of the studied molecules (pIC50) are randomly shuffled many times, and after every iteration, a new QSAR model is developed. The new QSAR models are expected to have lower Q2 and R2 values than those of the original model. This technique is performed to eliminate the possibility of chance correlation. If higher values of the Q2 and R2 are obtained, it means that an acceptable 3D-QSAR cannot be generated for this data set because of structural redundancy and chance correlation.
Model acceptability criteria
According to Alexander Tropsha and Alexander Golbraikh, a predictive model must satisfy a set of statistical criteria. A QSAR model was considered predictive if the following conditions are satisfied: (i) Q2 > 0.50; (ii) R2 > 0.60 [29, 32].
Molecular docking
Molecular docking is regarded among the most important method in discovering novel small-molecule drugs [33,32,35]. In our study, this technique was performed using Autodock vina and Surflex-dock program, which were used to verify whether both would give the same binding mode or not. If they give the same binding mode, we can say that this is the best possible position of furanone in the binding site of CDC kinase. As known, those two programs use different searching algorithms and scoring functions to look for the appropriate position of the ligand into the active site of the receptor. The Autodock vina is using the Lamarckian Genetic Algorithm (LGA) [36], while the Surflex is using incremental construction algorithm (matching algorithm). The crystal structure of CDC7 was downloaded from the protein data bank (PDB entry code: 4f9b). No one of the understudy ligands is complexed with this protein in PDB; so, its original ligand was removed. The PDB file was prepared using Discovery Studio 2016, such as all ligands, co-factors, and solvent molecules that were removed from the model. Before docking, hydrogen atoms of the receptor were added to the prepared structure. For Surflex study, the definition of active site definition was performed based on the original ligand in the crystal. While for Autodock vina [36] study, a grid box (x = 16, y = 16, z = 16 at 1 Angstrom spacing) was set to cover the binding site. We chose compound 10 as the subject to dock into the active pocket under the conditions previously cited [34], and results were analyzed using Discovery Studio 2016 software [37].
Macromolecule preparation
The crystal structure of CDC7 was downloaded from the protein data bank (PDB entry code: 4f9b). No one of the understudy ligands is complexed with this protein in PDB; so, its original ligand was removed. The PDB file was prepared using Discovery Studio 2016, such as all ligands, co-factors, and solvent molecules were removed from the model. Before docking, hydrogen atoms of the receptor were added to the prepared structure. The definition of active site definition was performed based on the original ligand in the crystal. We chose compound 10 as the subject to dock into the active pocket under the conditions previously cited.
Ligand preparation
For Surflex study, the selected compounds for docking were modeled in the same way as for the 3D-QSAR studies. Three-dimensional structures were built using the SKETCH option in SYBYL; then, they were minimized under the Tripos standard force field [21] with Gasteiger-Hückel atomic partial charges [22] by the Powell method with a gradient convergence criterion of 0.01 kcal/mol Å. For AutoDock Vina study, an extended PDB format, termed PDBQT, is used for coordinate files, which includes atomic partial charges and atom types. Torsion angles were calculated to assign the fixable and non-bonded rotation of molecules [36].
Applicability domain
The utility of any QSAR model is its accurate prediction ability for new chemical compounds. So, once the QSAR model is generated, its domain of applicability (AD) must be defined. A QSAR model is applicable only within its training domain, and only the prediction for new chemicals falling within its applicability domain can be considered trustworthy and not model extrapolations. Applicability domain was defined through the degree of similarity of the designed compound to those in the training set molecules. Assessment was done by comparing the value for each descriptor of the compound against the range of descriptor values for the compounds in the training set, using SYBYL software. The number of the out-of-range descriptors found in the test compound and the total contribution of such extrapolated points to the prediction is calculated. The sum of extrapolated terms (SUM) is the total contribution made to the prediction for the compound by the out-of-range descriptors. Additionally, standard error of prediction (SEP) was calculated during the 3D-QSAR model generation. If SUM of the test or predicted compound is larger than the SEP for the developed cross-validated model, then the extrapolation is probably too far outside the model to get a reliable prediction [38]. SUM was calculated for all newly designed compounds.
Synthetic accessibility
The best designed hits were screened for their synthetic accessibility by using SwissADME [39]. It provides a score on a scale from 1 (very easy to synthesize) to 10 (challenging and complex to synthesize). To measure this score for each compound, many criteria such as the number of stereocenters, complexity of the molecular structure, complexity of the ring system, similar to a huge number of already synthesized compounds, and the potential for using important synthetic reactions were taken into consideration.
In silico pharmacokinetics (PK)/pharmacodynamics (PD) evaluation
In silico ADMET (adsorption, distribution, metabolism, excretion, and toxicity), prediction is considered as a first step in the direction to analyze the new chemical entities to reduce wasting time on lead compounds which would be toxic or metabolized by the body into an inactive form and unable to pass through membranes. For this purpose, the best-designed hits were evaluated for their in silico pharmacokinetics parameters; in addition, their expected metabolic products and sites of metabolism for phase-i and phase-II metabolisms were also estimated through MetaPrint2D-React software, to check if those compounds are able to reach advanced stages in clinical studies, which enhance their chances to reach the stage of being candidate drugs in future.
Results and discussion
The predicted and experimental activity values and their residual values for both the training and test sets from 3D-QSAR (CoMSIA) model are given in Table 1.
CoMSIA results
Based on CoMSIA descriptors available in SYBYL, a 3D-QSAR model was proposed to explain and predict quantitatively the hydrophobic, electrostatic, steric, and acceptor fields’ effects of substituents on the CDC7 inhibitory activity of a series of 32 furanone derivatives.
Different combinations of the five fields were generated and results are listed in Table 2. They showed Q2 ranging from 0.395 to 0.581 and R2 values of 0.573 to 0.94. In general, statistical results with Q2 value ≥ 0.5 and R2 value ≥ 0.6 are considered reasonable and meaningful. These results indicate that CoMSIA were robust in generating statistically significant models. The model CoMSIA/SEA, which contains four fields (steric, electrostatic, and acceptor) and exhibits good statistical keys, was considered as the best one. Hence, all the following studies were based on the combination of S, E, and A fields. The cross-validated determination coefficient Q2 value of the training set and non-cross-validated determination coefficient R2 are 0.545 and 0.945, respectively. The optimal number of principal components used to generate the CoMSIA model is four, which is reasonable considering the number of molecules used to build the model [40]. The standard error of estimate is 0.252. The bootstrapping R2 mean (BS-R2) and bootstrapping standard deviation (BS-SD) values obtained from 1000 runs of BS are 0.96 and 0.196, respectively. Finally, the prediction ability of the proposed model was confirmed using external validation, the R2test value obtained is 0.96. The MAE and (MAE(training set) + 3 * 휎) of the test set were found to be 0.258 and 0.621, respectively, which are less than the thresholds 0.3125 (0.1 * training set range which is) and 0.2* training set range; consequently, the 3D-QSAR model in our study was reliable according to the MAE-based metrics proposed by Roy et al. [30]. These statistics results indicated the good stability and the powerful predictive ability of the proposed CoMSIA model.
Q2 cross-validated determination coefficient, N optimum number of components obtained from cross-validated PLS analysis, R2 non-cross-validated determination coefficient, Scv standard error of the estimate, F-t F-test value, R2test external validation determination coefficient
Graphical interpretation of 3D-QSAR models
3D-QSAR contour maps were generated to visualize the data contents of the derived CoMSIA model, which provide the information about the favorable and unfavorable regions for the biological activity in the studied compounds. Changes in the structure of the molecule lead to changes in its physico-chemical properties, which might increase or decrease the biological activity. Steric, electrostatic, hydrophobic, and hydrogen bond acceptor contour maps of CoMSIA are shown in Fig. 2. Compound 10 is the most active of the series; therefore, it was taken as reference structure for the generation of contour maps.
Std* coeff. contour maps of CoMSIA analysis with 2 Å grid spacing in combination with compound 10. a Steric contour map: green contours (80% contribution) indicate regions where bulky groups increase activity, while yellow contours (20% contribution) indicate regions where bulky groups decrease activity. b Electrostatic contour map. Electrostatic fields: blue contours (80% contribution) indicate regions where electron-donating groups increase activity, while red contours (20% contribution) indicate regions where electron-withdrawing groups increase activity. c Hydrogen-bond acceptor contour map. The magenta contours (80% contribution) for hydrogen-bond acceptor groups increase activity; red contours (20% contribution) indicate the disfavored region
CoMSIA contour map
In CoMSIA model shown in Fig. 2, we will highlight and interpret the distribution of the steric, electrostatic, and hydrogen-bond acceptor fields, which represent the fractions of 31.0, 38.5, and 30.5%, respectively.
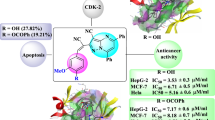
Figure 2a shows steric contour maps of the most active compound 10 and represents that areas around phenyl ring and ethyl of the ester group, in addition to areas near 4 and 5 positions of the azaindole moiety, are favorable in terms of steric field, which suggests that bulky groups introduced in these positions are helpful for increasing activity. For exemple, compounds 19 (pIC50 = 8.420) and 25 (pIC50 = 8.455) exhibit high activities.
The appearance of a small yellow contour near the R substituent of the A-substructure (Table 1) indicates that substitution by bulky groups at this location will result in a lower inhibitory activity. For example, compounds 9 (pIC50 = 8.420) and 10 (pIC50 = 6.050) exhibit low activities how they hold bulky groups at this position.
In the electrostatic contour maps depicted in Fig. 2b, a medium-sized blue areas on the 3-position of the phenyl ring are shown. This observation would tend to suggest that positive charged groups at these parts of the molecule are favorable for the biological activity. That can be interpreted by the fact that compounds 12 (pIC50 = 7.920), 15 (pIC50 = 7.5376), and 26 (pIC50 = 7.744) with negative charged groups at position 3 of phenyl show lower activities than compounds 1 (pIC50 = 8.420) and 6 (pIC50 = 8.481) which are unsubstituted at the same position, which imply that substituting a very small electron deficient group in this area can raise the activity of a molecule.
The CoMSIA contour maps of the H-bond acceptor are shown in Fig. 2c; the magenta contour maps indicate the areas where hydrogen bond accepting groups increased activity and red contour maps indicate areas where hydrogen bond accepting groups decreased inhibitory activity. As it turns out in Fig. 2c, the magenta contours of hydrogen bond accepting groups are located near the N atom of the pyridine ring in the azaindole moiety, which suggests the requirement of hydrogen bond accepting groups at this position to enhance the inhibitory activity. This can be expounded by the example, that is, compound 1 (pIC50 = 8.420) with an azaindole moiety has higher activity than that of compound 4 (pIC50 = 5.920) with OH group near the magenta area. In contrast, red contours which are related to the groups that do not have the ability of accepting hydrogen bonds are located on the 5-position of the phenyl ring. In this way, if there are hydrogen bond acceptors on one side of azaindole moiety, then the activity of compounds can be increased.
Summary of the structure-activity relationships
Some structural features between CDC7 and furanone inhibitors derived from the all above analyses are illustrated in Fig. 3. From the above discussions, we can infer the following conclusions: (1) bulky groups like phenyl ring with more positive charges groups at 3-position could improve the inhibitory activity. (2) Hydrogen acceptor groups near the nitrogen atom of azaindole moiety would be beneficial for the bioactivity. (3) Bulky, electron-withdrawing groups on the 4 and 5 positions of the azaindole moiety could raise the activity of the molecule. The conclusion above can offer a way to design highly effective CDC7 inhibitors.
External validation
Validation of the developed model is an essential part of any QSAR study. Thus, a true and trustworthy model should be able to predict a precise activity in the external test set [29]. That is why the final developed 3D-QSAR model from a training set of 22 furanone derivatives was used to predict the activity of nine remaining molecules; the parameters of the performance of the generated models are shown in Table 2.
Y-randomization
The Y-randomization method was carried out to validate the 3D-QSAR model. Several random shuffles of the dependent variable were performed; then after every shuffle, a 3D-QSAR was developed and the obtained results are shown in Table 3. The low Q2 and R2 values obtained after every shuffle indicate that the good result in our original 3D-QSAR model are not due to a chance correlation of the training set.
Docking results
Molecular docking study of the most active compound (10) was carried out using AUTODOCK and Surflex program to clarify the probable binding modes between furanone derivatives and CDC7 kinase. The crystal structure of CDC7 (PDB ID: 4f9b) with a resolution of 2.5 Å was used for our docking study. Both used programs gave the same binding mode and found that compound 10 forms four hydrogen bonds with Ser 70, Lys 90, Pro135, and Leu137 (Fig. 4 and Table 5), which provides straightforward knowledge for further structural optimization.
Identification of binding modes
In the docking analysis, four hydrogen bonds between compound 10 and CDC7 binding pocket were formed. Two hydrogen bonds 3.10 (Leu 137-NH---N) and 2.70 A (Pro 135-O---HN) were mediated by the 7-azaindole moiety, which is known as a hinge binder as it occupies the ATP binding site, while the third 3.07 A (Lys 90-NH3+ − --O=C) was formed by the O of the furanone’s carbonyl, and the fourth 3.05 A (Ser 70-OH ---O=C) was formed between the carbonyl of the ester group and the OH of the serine, which can be seen in green dotted lines. Docking results show that the N of the pyridine in the azaindole moiety and are in favor of the making hydrogen bond interaction and enhancement activity, which correlate well with the CoMSIA results.
The anilinic ring and the ester group are located in an exposed solvent area. Additionally, the CoMSIA suggests that bulky groups are supposed to enhance the activity. These observations between docking and CoMSIA model are in concordance, because substituents on these positions are exposed to the solvent and tolerated to be bulky.
From the analysis of the various properties of inhibitor 10 in Tables 5 and 7, it can be concluded that it fulfills the Lipinski’s rule [41]. Additionally, similar ligand–CDC7 interactions in which the inhibitor 10 forming interaction with CDC7 active site residue were reported in several earlier studies, namely, 0SY in (PDB: 4f9c) and ADP in (PDB: 4f99). Therefore, that can prove our docking process was reasonable. While the main cause of its mediated CDC7 inhibition by foranone derivatives is due to their shape, which contain the heteroaromatic azaindole hinge-binder and the carbonyl group on the furanone that allow them to make several hydrogen bonds within the ATP binding site.
Design for new CDC7 inhibitors containing furanone scaffold
The findings of 3D-QSAR (CoMSIA) model and molecular docking have provided the overall substitution pattern required around the furanone pharmacophore. Here, three potent substituted furanone analogs have been designed in order to improve the CDC7 inhibitory activity as well as the drug-like properties of the compounds including pharmacokinetics and toxicity. All newly designed molecules have good predicted activities. These newly designed molecules were aligned to the database using inhibitor 10 as template and their pIC50 values were predicted in addition to their SUM to check if they fall in the AD of the proposed model. For the proposed model to predict the CDC7 inhibitory activity, all newly designed compounds have SUM < SEP, so their predicted values are regarded reliable.
To ensure the viable drug designing, predicted compounds were evaluated for synthetic accessibility. Later, results of their synthetic accessibility were compared with that of the original synthesized CDC7 inhibitors. The SwissADME scores [39] of newly designed and preexisted compounds were found similar, which indicate that the synthetic ability of designed leads and results were compared with that of the most potent compound from the studied series (inhibitor 10) which is 3.69.
Binding model for designed inhibitors
The newly designed molecules were docked into the CDC7 active site. All of compounds were perfectly placed in the active site and formed hydrogen bonds with Pro 135 and Leu 137 from the hinge region and keep the same conformation at the binding pocket of CDC7 as the inhibitor 10. Binding energy of all compounds was found to be better than this of inhibitor 10 (− 7.9 kcal/mol), indicating that designed compounds have good binding affinity for CDC7 (Table 4). Among the three mentioned compounds, F2 demonstrated the lowest binding energy, so it was chosen for more studies. Owing to the introduction of the amino group at the ester sidechain, in addition to its hydrogen bond interactions with the hinge region residue and Lys 90, F2 can form one other hydrogen bond with Asp 196 as shown in Figs. 4 and 5 and Table 5.
ADMET predictions
Metabolism is the biotransformation of drugs and xenobiotic compounds to facilitate their excretion, which may produce different metabolites with different pharmacological and physicochemical properties. Therefore, the metabolism plays a critical role in the bioavailability of drugs and drug-drug interactions, and as known, metabolic processes are mainly catalyzed by the so-called phase I (oxidation, reduction, and hydrolysis) and II (sulfo-conjugation among others) enzymes, which are, for the most part, produced in the liver, and the cytochrome P450 (CYP450) enzymes are probably the most important class of enzyme to study this. Thus, exploring those parameters may offer new strategies that might bridge the gap between early stage drug discovery, preclinical and clinical trials, besides to get a deep insight on how they react in the body, which allows medicinal chemist to introduce new functional groups on the molecule to dodge the metabolic pathways susceptible to give toxic or very polar compounds that can be eliminated very easily from the body. Accordingly, that can help to synthesize metabolically stable drugs, as well as to avoid drug-drug interactions. For this purpose, the compounds were submitted to learn the potential of compounds having substrate or inhibitor of CYPs along with CYP human liver microsomes (HLM) (Table 7). All new designed compounds were found to be neither substrate nor inhibitors of CYP3A4 and 2DC.
Each time when a lead compound with high biological activity is identified, there is no guarantee that this compound with the best interactions with a target is necessarily a good medicine. Thus, it is essential to predict the ADMET parameters of the leads and optimize these parameters early on to reduce potential problems later during clinical studies. Accordingly, the ADMET parameters of the newly designed compounds and the most potent inhibitor from the studied dataset (inhibitor 21) were calculated using the pkCSM [42] and SwissADME [39] online tools. Water solubility is given in log (mol/l) (insoluble < − 10 < poorly soluble < − 6 < moderately < − 4 < soluble < − 2 < very soluble < 0 < highly soluble), intestinal absorbance value below 30% indicates poor absorbance. Low value of total clearance (logCLtot) means high drug half lifetime. For a given compound, a logBB < − 1 considered poorly distributed to the brain. Positive result in Ames test suggests that compound could be mutagenic. Prediction of ADMET parameters is listed in Table 7.
Drug molecules encounter several different membrane barriers in their journey through the body, such as gastrointestinal epithelial cells, blood-brain barrier, and the target cell. Prediction of permeability through those barriers can assist to interpret the pharmacokinetics results and in understanding the behavior of such kind of chemicals in the body.
The octanol–water partition coefficient (log P) is a vital parameter that shows the drug hydrophobicity and expresses that compounds with higher hydrophobicity present an increased metabolism and low absorption that may inadvertently be increased probability of binding to undesired hydrophobic macromolecules, hence increase the potential toxicity. Similarly, rapid renal clearance is associated with small and hydrophilic compounds.
As depicted in Tables 6 and 7, results were within the acceptable range for the designed compounds in terms of lipophilicity and PSA. Additionally, all compounds fulfill the Lipinski’s rule [41] and the aqueous solubility predictions show that they are all moderate soluble in water, which implicates a good oral bioavailability.
The drugs, which are orally administered, must be absorbed by the intestine; results show that all compounds have a good calculated intestinal absorbance; therefore, they can be easily absorbed by the intestine.
The compound-induced toxicity was predicted for Ames test, to check the safety of the designed compounds, as shown in Table 7, results specify that all newly designed compounds were non-mutagenic; additionally, the low values for brain-blood partition coefficient were found indicating that they will have a very low potential to cross the brain-blood brain barrier thereby eliminating the possibility of CNS related toxicity.
These results suggest that the newly designed compounds showed promising properties, in terms of intestinal adsorption, volume of distribution, blood-brain barrier permeability and toxicity, and present high biological activities, and are therefore potentially interesting candidates for further studies.
Conclusion
CDC7 kinase has emerged as a promising drug target for various cancer types. In this study, computer-aided drug design techniques, CoMSIA and molecular docking analyses, were conducted based on 31 furanone derivatives. CoMSIA (Q2 = 0.545, R2 = 0.945) was developed and validated using different validation techniques, the best model displayed significant statistical quality and excellent predictive ability. The graphical contour maps analysis indicated significant steric, electrostatic, and acceptor hydrogen potential contributions and suggested sufficient information for understanding the structure-activity relationship, thus aided in the further design and development of novel furanone derivatives with CDC7 improved inhibitory profile. Molecular docking results indicate that the hydrogen bond interactions with Lys 90, Pro135, and Leu 137 contribute mainly to the inhibitory activity, which highlight the importance of the heteroaromatic azaindole hinge-binder and carbonyl group of the furanone moiety for inhibitory effect. Connection of hydrophobic groups such as phenyl with electron-withdrawing groups on 2 and 4-positions to the furanone moiety is favorable. Long chain on the furanone containing a carbonyl and a terminal amine is necessary to enhance the inhibitory activity. Further, all those outcomes showed insight into the key structural features required for the CDC7 inhibitory behavior in the studied furanone derivatives. Thus, those obtained key structural factors were used to design three inhibitors by modifying the furanone scaffold, and then got a better result in terms of biological activity, in silico ADME, and toxicity. The new insights regarding the interaction mechanisms between ligand and CDC7 and the information provided from the contour maps could serve as guidelines in designing of novel potent CDC7 inhibitors.
References
Labib K (2010) Howdo Cdc7 and cyclin-dependent kinases trigger the initiation of chromosome replication in eukaryotic cells? Genes Dev 24:1208–1219. https://doi.org/10.1101/gad.1933010
Masai H, Arai K-I (2002) Cdc7 kinase complex: a key regulator in the initiation of DNA replication. J Cell Physiol 190:287–296. https://doi.org/10.1002/jcp.10070
Hayano M, Kanoh Y, Matsumoto S, Masai H (2011) Mrc1 marks early-firing origins and coordinates timing and efficiency of initiation in fission yeast. Mol Cell Biol 31:2380–2391. https://doi.org/10.1128/MCB.01239-10
Matsumoto S, Masai H (2013) Regulation of chromosome dynamics by Hsk1/Cdc7 kinase. Biochem Soc Trans 41:1712–1719. https://doi.org/10.1042/BST20130217
Montagnoli A, Moll J, Colotta F (2010) Targeting cell division cycle 7 kinase: a new approach for cancer therapy. Clin Cancer Res 16:4503–4508. https://doi.org/10.1158/1078-0432.CCR-10-0185
Kulkarni AA, Kingsbury SR, Tudzarova S et al (2009) Cdc7 kinase is a predictor of survival and a novel therapeutic target in epithelial ovarian carcinoma. Clin Cancer Res 15:2417–2425. https://doi.org/10.1158/1078-0432.CCR-08-1276
Choschzick M, Lebeau A, Marx AH et al (2010) Overexpression of cell division cycle 7 homolog is associated with gene amplification frequency in breast cancer. Hum Pathol 41:358–365. https://doi.org/10.1016/j.humpath.2009.08.008
Huggett MT, Tudzarova S, Proctor I et al (2016) Cdc7 is a potent anti-cancer target in pancreatic cancer due to abrogation of the DNA origin activation checkpoint. Oncotarget 7:18495–18507. https://doi.org/10.18632/oncotarget.7611
Kim JM, Kakusho N, Yamada M et al (2008) Cdc7 kinase mediates Claspin phosphorylation in DNA replication checkpoint. Oncogene 27:3475–3482. https://doi.org/10.1038/sj.onc.1210994
Feng D, Tu Z, Wu W, Liang C (2003) Inhibiting the expression of DNA replication-initiation proteins induces apoptosis in human cancer cells. Cancer Res 63:7356–7364
Swords R, Mahalingam D, O’Dwyer M et al (2010) Cdc7 kinase – a new target for drug development. Eur J Cancer 46:33–40. https://doi.org/10.1016/j.ejca.2009.09.020
Kurasawa O, Oguro Y, Miyazaki T et al (2017) Identification of a new class of potent Cdc7 inhibitors designed by putative pharmacophore model: synthesis and biological evaluation of 2,3-dihydrothieno[3,2- d ]pyrimidin-4(1 H )-ones. Bioorg Med Chem 25:2133–2147. https://doi.org/10.1016/j.bmc.2017.02.021
Irie T, Asami T, Sawa A et al (2017) Discovery of novel furanone derivatives as potent Cdc7 kinase inhibitors. Eur J Med Chem 130:406–418. https://doi.org/10.1016/j.ejmech.2017.02.030
Roy K, Kar S, Das RN (2015) Understanding the basics of QSAR for applications in pharmaceutical sciences and risk assessment. Underst Basics QSAR Appl Pharm Sci Risk Assess. https://doi.org/10.1016/C2014-0-00286-9
Dearden JC (2016) The history and development of quantitative structure-activity relationships (QSARs). Int J Quant Struct Relat 1:1–44. https://doi.org/10.4018/IJQSPR.2016010101
Klebe G, Abraham U, Mietzner T (1994) Molecular similarity indices in a comparative analysis (CoMSIA) of drug molecules to correlate and predict their biological activity. J Med Chem 37:4130–4146. https://doi.org/10.1021/jm00050a010
Damale MG, Harke SN, Kalam Khan FA et al (2014) Recent advances in multidimensional QSAR (4D-6D): a critical review. Mini Rev Med Chem 14:35–55
Holder S, Lilly M, Brown ML (2007) Comparative molecular field analysis of flavonoid inhibitors of the PIM-1 kinase. Bioorg Med Chem 15:6463–6473. https://doi.org/10.1016/j.bmc.2007.06.025
Huang X, Liu T, Gu J et al (2001) 3D-QSAR model of flavonoids binding at benzodiazepine site in GABA a receptors. J Med Chem 44:1883–1891. https://doi.org/10.1021/jm000557p
Veras L da S, Arakawa M, Funatsu K, Takahata Y (2010) 2D and 3D QSAR studies of the receptor binding affinity of progestins. J Braz Chem Soc 21:872–881. https://doi.org/10.1590/S0103-50532010000500015
Clark M, Cramer RD, Van Opdenbosch N (1989) Validation of the general purpose tripos 5.2 force field. J Comput Chem 10:982–1012. https://doi.org/10.1002/jcc.540100804
Purcell WP, Singer JA (1967) A brief review and table of semiempirical parameters used in the Hueckel molecular orbital method. J Chem Eng Data 12:235–246. https://doi.org/10.1021/je60033a020
AbdulHameed MDM, Hamza A, Liu J, Zhan C-G (2008) Combined 3D-QSAR modeling and molecular docking study on Indolinone derivatives as inhibitors of 3-phosphoinositide-dependent protein Kinase-1. J Chem Inf Model 48:1760–1772. https://doi.org/10.1021/ci800147v
Zheng J, Xiao G, Guo J et al (2011) Exploring QSARs for 5-lipoxygenase (5-LO) inhibitory activity of 2-substituted 5-Hydroxyindole-3-carboxylates by CoMFA and CoMSIA. Chem Biol Drug Des 78:314–321. https://doi.org/10.1111/j.1747-0285.2011.01146.x
Wold S (1991) Validation of QSAR’s. Quant Struct Relat 10:191–193. https://doi.org/10.1002/qsar.19910100302
Kubinyi H (2003) Comparative molecular field analysis (CoMFA). Handb Chemoinform 1555–1574. https://doi.org/10.1002/9783527618279.ch44d
Cruciani G, Baroni M, Clementi S et al (1992) Predictive ability of regression models. Part I: standard deviation of prediction errors (SDEP). J Chemom 6:335–346. https://doi.org/10.1002/cem.1180060604
Baroni M, Clementi S, Cruciani G, Costantino G, Riganelli D, Oberrauch E (1992) Predictive ability of regression models. Part II: Selection of the best predictive PLS model. J Chemom 6:347–356
Golbraikh A, Tropsha A (2002) Beware of q2! J Mol Graph Model 20:269–276. https://doi.org/10.1016/S1093-3263(01)00123-1
Roy K, Das RN, Ambure P, Aher RB (2016) Be aware of error measures. Further studies on validation of predictive QSAR models. Chemom Intell Lab Syst. https://doi.org/10.1016/j.chemolab.2016.01.008
Rücker C, Rücker G, Meringer M (2007) Y-randomization and its variants in QSPR/QSAR. J Chem Inf Model 47:2345–2357. https://doi.org/10.1021/ci700157b
Tropsha A, Gramatica P, Gombar VK (2003) The importance of being earnest: validation is the absolute essential for successful application and interpretation of QSPR models. QSAR Comb Sci 22:69–77. https://doi.org/10.1002/qsar.200390007
Jain AN (2007) Surflex-dock 2.1: robust performance from ligand energetic modeling, ring flexibility, and knowledge-based search. J Comput Aided Mol Des 21:281–306. https://doi.org/10.1007/s10822-007-9114-2
Jain AN (2003) Surflex: fully automatic flexible molecular docking using a molecular similarity-based search engine. J Med Chem 46:499–511. https://doi.org/10.1021/jm020406h
Wang J, Kollman PA, Kuntz ID (1999) Flexible ligand docking: a multistep strategy approach. Proteins 36:1–19
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461. https://doi.org/10.1002/jcc.21334
(2016) Dassault Systèmes BIOVIA, Discovery Studio Modeling Environment, Release 2017, San Diego: Dassault Systèmes. http://accelrys.com/products/collaborative-science/biovia-discovery-studio/. Accessed 25 Feb 2017
Gupta P, Garg P, Roy N (2012) Identification of novel HIV-1 integrase inhibitors using shape-based screening, QSAR, and docking approach. Chem Biol Drug Des 79:835–849. https://doi.org/10.1111/j.1747-0285.2012.01326.x
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7:42717. https://doi.org/10.1038/srep42717
Hickey JP, Passino-reader DR (1991) Linear solvation energy Relationships : “ rules of thumb ” for estimation of variable values. 25:1753–1760
Lipinski CA (2004) Lead- and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol 1:337–341. https://doi.org/10.1016/j.ddtec.2004.11.007
Pires DE V, Blundell TL, Ascher DB pkCSM: Predicting Small-Molecule Pharmacokinetic and Toxicity Properties Using Graph-Based Signatures. https://doi.org/10.1021/acs.jmedchem.5b00104
Acknowledgments
We are grateful to the “Association Marocaine des Chimistes Théoriciens” (AMCT) and “Moroccan Centre of Scientific and Technique research” (CNRST) for their pertinent help concerning the programs.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests
The authors declare that they have no conflict of interest.
Ethical approval
This chapter does not contain any studies with human participants or animals performed by any of the authors.”
Rights and permissions
About this article
Cite this article
Aouidate, A., Ghaleb, A., Ghamali, M. et al. Furanone derivatives as new inhibitors of CDC7 kinase: development of structure activity relationship model using 3D QSAR, molecular docking, and in silico ADMET. Struct Chem 29, 1031–1043 (2018). https://doi.org/10.1007/s11224-018-1086-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-018-1086-4