Abstract
Background
SMAD4 is a potent tumor suppressor. SMAD4 loss increases genomic instability and plays a critical role in the DNA damage response that leads to skin cancer development. We aimed to investigate SMAD4 methylation effects on mRNA and protein expression of SMAD4 in cancer and healthy tissues from patients with basal cell carcinoma (BCC), cutaneous squamous cell carcinoma (cSCC), and basosquamous skin cancer (BSC).
Methods and results
The study included 17 BCC, 24 cSCC and nine BSC patients. DNA and RNA were isolated from cancerous and healthy tissues following punch biopsy. Methylation-specific polymerase chain reaction (PCR) and real-time quantitative PCR methods were used to examine SMAD4 promoter methylation and SMAD4 mRNA levels, respectively. The percentage and intensity of staining of the SMAD4 protein were determined by immunohistochemistry. The percentage of SMAD4 methylation was increased in the patients with BCC (p = 0.007), cSCC (p = 0.004), and BSC (p = 0.018) compared to the healthy tissue. SMAD4 mRNA expression was decreased in the patients with BCC (p˂0.001), cSCC (p˂0.001), and BSC (p = 0.008). The staining characteristic of SMAD4 protein was negative in the cancer tissues of the patients with cSCC (p = 0.00). Lower SMAD4 mRNA levels were observed in the poorly differentiated cSCC patients (p = 0.001). The staining characteristics of the SMAD4 protein were related to age and chronic sun exposure.
Conclusions
Hypermethylation of SMAD4 and reduced SMAD4 mRNA expression were found to play a role in the pathogenesis of BCC, cSCC, and BSC. A decrease in SMAD4 protein expression level was observed only in cSCC patients. This suggests that epigenetic alterations to the SMAD4 gene are associated with cSCC.
Trial Registration
The name of the trial register: SMAD4 Methylation and Expression Levels in Non-melanocytic Skin Cancers; SMAD4 Protein Positivity.
The registration number: NCT04759261 (https://clinicaltrials.gov/ct2/results?term=NCT04759261).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The registration number: NCT04759261 (https://clinicaltrials.gov/ct2/results?term=NCT04759261).
Introduction
Non-melanocytic skin cancers (NMSCs) are the most common malignancies in humans, and their incidence is increasing. The main NMSC types are basal cell carcinoma (BCC) and cutaneous squamous cell carcinoma (cSCC), which together account for approximately 99% of all NMSCs [1]. The incidence of NMSC increases with age, and recent global statistics indicate that that more than one million new cases are diagnosed per year [2]. BCC accounts for 80–85% of NMSC cases, with cSCC making up 15–20% of such cases [3]. Basosquamous carcinoma (BSC) is a clinically aggressive skin tumor with characteristics of both BCC and cSCC [4]. It occurs in 1.2–2.7% of all skin carcinomas [5]. Although BSC has the histological characteristics of both BCC and cSCC, the molecular and genetic mechanisms underlying this type of cancer have not been fully elucidated [6].
The pathogenesis of NMSCs is multifactorial [7]. The primary risk factor for the development of skin cancer is cumulative exposure to ultraviolet radiation (UVR) [8]. UVR-induced DNA damage and high UVR intensity are the main environmental factors involved in the etiology of skin cancer. Genetic factors (such as specific mechanisms, genes, and the tumor microenvironment) play a role in the development of NMSCs [3]. Hanahan described how cells surrounding a tumor can undergo epigenetic reprogramming, especially when recruited by molecules and growth factors that alter the function of their internal signaling pathways [9]. In recent years, epigenetic modifiers have been discovered, notably in more than 50% of cSCC mutations [10]. One of the best-known epigenetic modifications is DNA methylation, which alters the expression of key genes involved in various biological processes [11].
Transforming growth factor beta (TGF-β) signaling is a growth inhibitor for keratinocytes in the skin and a profibrotic factor in the dermis [12]. Exposure to UVR suppresses TGF-β signaling and abrogates the growth inhibitory effect of TGF-β; this is reflected in the clinical course in the early stages by the formation of actinic keratosis, a precancerous condition associated with epidermal hyperplasia [13]. With the progressive suppression of TGF-β signaling and under the influence of UVR, long-term gene mutations in TGF-β signaling components (such as TGFbRI, TGFbRII, small mothers against decapentaplegic homolog 2 (SMAD2), and SMAD4) occur in both epidermal keratinocytes and stem cells [14]. SMAD4 is the core mediator of the canonical TGF-β signaling pathway [15]. Loss of SMAD4 plays a critical role in the response to genomic instability resulting from DNA damage. This is quite evident in skin cancer, and SMAD4 is thought to play an important role in the progression of various tumor types [16].
The aim of this study was to investigate the relationship between potential promoter methylations in the SMAD4 gene, which is an important component of the TGF-β signaling pathway, SMAD4 mRNA expression, and histopathological changes in SMAD4 protein expression in healthy (adjacent tissue) and cancerous keratinocytes. It is also thought to contribute to the molecular pathogenesis of BSC.
Materials and methods
Patients and tissue preparation
Based on the clinical presentation and dermoscopic features of patients admitted to the Department of Dermatology and Venereology, Trakya University Faculty of Medicine, between March 2020 and 2021, a total of 50 patients with suspected BCC, cSCC, and BSC following a preliminary diagnosis were enrolled in the study. Patients with mucosal involvement who had previously undergone surgery for NMSC at an existing tumor site and/or had received radiotherapy for NMSC were excluded from the study. Approval for the study was granted by the Scientific Research Ethics Committee of the Trakya University Faculty of Medicine (approval number: 18/14, date: 06/11/2019). All the subjects who agreed to participate in the study gave their written informed consent.
The patients’ sociodemographic and clinical characteristics (i.e., age, gender, profession [17], Fitzpatrick skin type, tumor site [18], tumor duration, history of NMSCs, family history of skin cancer, immunosuppression, chronic sun exposure [8], tobacco and alcohol use, number of patients with ≥ 2 BCCs) were recorded. Macrophotographs were obtained from the patients, followed by polarized and unpolarized dermoscopic images (PhotoFinder®, x20). The review by Reiter et al. [19] was used to evaluate the dermoscopic characteristics of BCC, and the publication by Sgouros et al. [20] was used to evaluate the dermoscopic characteristics of cSCC. Both the clinical and dermoscopic characteristics were evaluated, and a biopsy was performed with the appropriate preliminary diagnosis. The histopathological diagnosis was confirmed by total excision in all the patients, and those patients in whom the results of the punch biopsy and total excision did not correlate were excluded from the study.
In patients who agreed to participate in the study, the most diagnostic areas of the tumoral lesions were identified and marked using a dermoscope and two incisional biopsies were obtained using a 4 mm punch biopsy instrument (Kai® Medical, Japan). The material obtained from the first biopsy was sent to the Biophysics Laboratory of Trakya University, and the material obtained from the second biopsy was sent to the university’s Pathology Laboratory. To obtain sufficient tissue material, tumor lesions smaller than 10 mm were not included in the study.
As this was designed as an epigenetic study, another biopsy was obtained from healthy tissue adjacent to the tumors, as both tissues should be similarly affected by environmental factors. Healthy tissue specimens were collected from an adjacent area of healthy skin approximately 5 mm from the tumor margin using a 5 mm punch biopsy instrument (Kai® Medical, Japan). The collected material was divided into two halves; one half of the tissue was sent to the Trakya University Biophysics Laboratory and the other half to the Trakya University Pathology Laboratory. The lesion-free skin region was selected based on the study by Gambichler et al. [21]. Since an epigenetic study was to be performed, a biopsy was taken from the adjacent tissue because tumor and healthy tissues should be similarly affected by environmental factors.
DNA preparation and bisulfite modification
The materials obtained from the tumor and healthy tissues were placed in tubes for DNA isolation and bisulfite modification and stored at − 80 °C until the study day. The DNA was isolated according to the manufacturer’s instructions using standard protocols (Invitrogen™ PureLink™ Genomic DNA Mini Kit, Thermo Fisher Scientific Inc., USA). The DNA was analyzed after quantification at 280 and 260 nm wavelengths using the spectrophotometric method. After DNA isolation, approximately 250 ng of genomic DNA was denatured through treatment with the EpiJET Bisulfite Conversion Kit for Sodium Hydroxide and modified with sodium bisulfite (Thermo Fisher Scientific Inc., USA) [22,23,24]. The specimens were stored at + 4 °C for further analysis.
Methylation analysis
After the chemical modification of the DNA, the DNA samples were used for methylation analysis of the SMAD4 gene. The program MethPrimer V1.1 beta was used to identify potential methylation sites and methylation-specific primer sequences for the SMAD4 gene sequence (www.urogene.org/methprimer/) [25]. Polymerase chain reactions (PCR) were performed on the DNA samples using methylation-specific and un-methylation-specific primers. The PCR conditions for the two sets of reactions were as follows: 95 °C for 10 min, then 35 cycles at 95 °C for 45 s, 55 °C (methylated–unmethylated) for 30 s, and 72 °C for 30 s, followed by a final extension of 5 min at 72 °C.
The media for methylated and unmethylated specific PCR were as follows: PCR buffer, 1x; MgCl2, 2 mM; DMSO, 5% (v/v); dNTP, 12.5 mM; primer forward, 10 nM; primer reverse, 10 nM; Taq polymerase, 1U (5U/µL). The template DNA (100 ng) was made up to 50 µL of dH2O. The methylated and unmethylated human DNA were used as positive and negative control DNAs (S8001 CpGenome™ Human Methylated and Non-Methylated DNA Standard Set, Sigma-Aldrich, USA). The PCR products were then evaluated on a 2% agarose gel under ultraviolet light, and the cancer tissues were compared with the healthy tissues for methylation, and the methylation percentage was determined [23,24,25]. The methylation-specific primers for the promoter of the SMAD4 gene region in the methylation-specific PCR method are listed in Supplementary Table 1.
Quantitative real-time PCR
The total RNA from 50 mg each of the healthy and cancer skin tissues was extracted using the PureLink™ RNA Mini Kit according to the manufacturer’s instructions (Thermo Fisher Scientific Inc., USA). Real-time PCR analysis was then performed to investigate the mRNA level of SMAD4 in the skin tissues. For this purpose, a StepOne™ Real-Time PCR System was used for cDNA synthesis using the Protocol for cDNA High-Capacity cDNA Reverse Transcription Kit (Thermo Fisher Scientific Inc., USA). The TaqMan assay used for SMAD4 expression analysis was Hs00929647_m1 (Thermo Fisher Scientific Inc., USA). A PCR efficiency of 100% ± 2% was guaranteed for all the assays. For cDNA, programming the thermocycler conditions with one of the thermocyclers at 25 °C for 10 min, 37 °C for 120 min, 85 °C for 5 min. Later for qPCR; 5 µL TaqMan Gene Expression Mix (Thermo Fisher Scientific Inc., USA) was used for the 0.5 µL probe assay, 2 µL cDNA and 2.5 µL H2O AMPIGENE® qPCR Probe Mix Hi-ROX (Enzo Life Sciences AG, Switzerland) were used. The real-time PCR conditions were 95 °C for 2 min, and the PCR stage consisted of 45 cycles at 95 °C for 5 s and 60 °C for 20 s. Each sample was analyzed in duplicate, and the mean the threshold cycle (Ct) value was used for further analysis. A Ct value greater than 35 was considered too low for quantification. The most relevant housekeeping gene was selected, and the relative expression was calculated using the ∆∆Ct (delta delta Ct) method and normalized to β-actin as the reference gene. We used the software programs embedded in the StepOne™ Real-Time PCR System (Thermo Fisher Scientific Inc., USA) for qPCR experiments most typically use the ∆∆Ct (2ΔΔCT) approach for relative quantification. The cycle at which the fluorescence intensity reaches a specific level is known as the Ct. This approach employs a reference gene as the normalizer to directly determine the relative gene expression in the target and control samples from the Ct data produced by a qPCR instrument. The Ct difference between the target and reference genes is the first Ct in the 2 ΔΔCT method [26, 27].
Histology and immunostaining
The tissue specimens were fixed in 10% neutral-buffered formalin, blocked in paraffin, sectioned, stained with hematoxylin and eosin, and examined by light microscopy. The diagnoses of the main types of NMSC were made by an independent pathologist based on previously determined histopathological criteria [28]. The patients diagnosed with cSCC were divided into three subgroups according to the degree of histopathological differentiation [29]. Immunohistochemical staining of the SMAD4 was performed by plating sections from the case blocks on slides with lysine (positively charged). Detection of the SMAD4 antibody was performed according to the manufacturer’s instructions (clone B-8, Santa Cruz Biotechnology Inc., Germany). The scoring system of He et al. was used to evaluate the SMAD4 immunohistochemical expression, and the percentage and intensity of SMAD4 staining in the lesional and healthy tissues were considered [30]. No points were given for the number of positively stained cells < 5%, 1 point was allocated for 6–25% positively stained cells, 2 points for 26–50%, 3 points for 51–75%, and 4 points for > 75%. When evaluating the staining intensity of the preparations, colorless preparations received 0 points, pallide-flavens preparations received 1 point, yellow-stained preparations received 2 points, and brown-stained preparations received 3 points. The positivity values were divided into four groups according to the total score obtained by multiplying the number of positively stained cells by the staining intensity score: negative (0 points), + positive (1–4 points, weak), ++ positive (5–8 points, intermediate), and +++ positive (9–12 points, strong). The healthy tissue harvested 5 mm from the lesional skin served as the control tissue and negative control antibodies for immunohistochemical staining.
Statistical analysis
The program SPSS version 22 was used to analyze the data. The data were expressed as arithmetic mean, standard deviation, number, and percentage. Pearson’s chi-square and Fisher’s exact tests were performed to compare the categorical data between the groups. Because the continuous data were not normally distributed (Kolmogorov–Smirnov and Shapiro–Wilk, p˂0.05), the Mann–Whitney U test was used for comparisons between the two groups, and the Kruskal–Wallis H test was used for comparisons between more than two groups (post hoc comparisons using the Mann–Whitney U test with Bonferroni correction). A p-value < 0.05 was considered statistically significant.
Results
Patient characteristics
The sociodemographic and clinical characteristics of the patients are summarized in Supplementary Table 2. Among the total 50 NMSC patients examined, 74.0% were over 65 years of age (n = 37), 76.0% were male (n = 38), and 88.0% had a history of chronic sun exposure (n = 44). Of the tumors, 92.0% were located in the head and neck region (n = 46).
Methylation status and mRNA expression of SMAD4 in the NMSC and healthy tissues
The mean value of the percentage of SMAD4 methylation in the patients with NMSC was 24.12 ± 25.9 in the patients with BCC, 27.67 ± 24 in the patients with cSCC, and 36.44 ± 19.35 in the patients with BSC. A statistically significant difference in the percentage of SMAD4 methylation was observed in all three groups compared with the healthy tissues (Supplementary Tables 3, Figs. 1 and 2).
PCR results of methylated (M) and unmethylated (U) primer sets on genomic DNA for the SMAD4 gene: lane 1 DNA marker; lane 2, positive control; lane 3–4 BCC; lane 5–6 cSCC; lane 7–8 BSC. The fragment length of methylated DNA was 164 bp and that of unmethylated DNA was 163 bp. PCR Polymerase chain reaction; BCC basal cell carcinoma; cSCC cutaneous squamous cell carcinoma, BSC basosquamous carcinoma, bp base pair
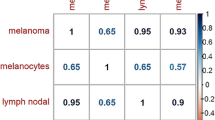
Diagram showing significant (p < 0.05) increase in methylation degree of SMAD4 in cancer tissues from patients with with BCC, cSCC and BSC compared to healthy(adjacent) tissues. In bold: statistically significant. aWilcoxon Signed Ranks analysis. BCC basal cell carcinoma, cSCC cutaneous squamous cell carcinoma, BSC basosquamous carcinoma
In the patients with NMSC, the mean value of SMAD4 mRNA expression was 0.18 ± 0.1 in the patients with BCC, 0.32 ± 0.12 in those with cSCC, and 0.22 ± 0 in those with BSC. A statistically significant difference in SMAD4 mRNA expression values was found in all three groups compared with the healthy tissues (Supplementary Tables 4, Fig. 3).
Diagram showing the significant (p < 0.05) reduction in mean mRNA expression of SMAD4 in cancer tissues from patients with BCC, cSCC and BSC compared to healthy (adjacent) tissues. In bold: statistically significant. aWilcoxon Signed Ranks analysis. BCC basal cell carcinoma, cSCC cutaneous squamous cell carcinoma, BSC basosquamous carcinoma
Histopathologically, the degree of differentiation in the patients with cSCC was compared with the mean values of the SMAD4 methylation percentages and mRNA expression, and it was concluded that the SMAD4 mRNA levels were statistically lower in the poorly differentiated group than in the well or moderately differentiated group (p = 0.001) (Table 1).
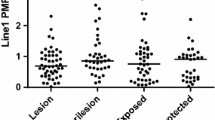
When the SMAD4 methylation status and mRNA expression levels were compared between tumoral lesions located in sun-exposed and non-sun-exposed areas, no statistically significant differences were detected (Supplementary Tables 5 and 6).
Immunohistochemical analysis of SMAD4 protein levels in the NMSC and healthy tissues
The SMAD4 protein levels were analyzed in a total of 50 NMSC specimens, of which 17 were BCC (34.0%), 24 were cSCC (48.0%), and nine were BSC (18.0%). Approximately 50 specimens of healthy tissue served as the control group. Representative photographs are shown in Fig. 4.
Histopathological analysis of cancer and healthy tissues showing BCC, SCC and BSC. a1–a3 Visualization of BCC, SCC, and BSC at 200× magnification by H and E staining, SMAD4 positivity in healthy tissues; b1 2 positive SMAD4 in healthy tissues in the patient with BCC, b2 3 positive SMAD4 in healthy tissues in the patient with SCC, b3 1 positive SMAD4 in healthy tissues in the patient with BSC (DABx200), SMAD4 positivity in cancer tissues; c1 1 positive SMAD4 in cancer tissues in the patient with BCC, c2 3 positive SMAD4 in cancer tissues in the patient with SCC, c3 3 positive SMAD4 in cancer tissues in the patient with BSC. White bars 100 μm for all panels
While positive staining of the SMAD4 protein was observed in ΔΔCt 0.0% of the healthy tissues from the patients with cSCC, staining was detected in the cancer tissues from the same patients (p = 0.00). In the patients with BCC and BSC, no statistically significant differences were found between the healthy and cancer tissues in terms of the staining characteristics of the SMAD4 protein (Table 2).
In the tissues of the patients with NMSC, the degree of immunohistochemical staining of SMAD4 and the sociodemographic and clinical characteristics of the patients were compared. Supplementary Table 7 shows that the SMAD4 staining characteristics were related to age and chronic sun exposure. While 93.8% of the cancer tissues from the patients over 65 years of age showed negative staining (n = 15), 42.3% of the cancer tissues from the patients younger than 65 years showed positive staining (n = 11; p = 0.029). Negative staining of SMAD4 was detected in 96.8% (n = 30) of the patients with a history of chronic sun exposure (p = 0.032).
Discussion
The loss of SMAD4 in squamous cell carcinomas impairs the DNA damage response and repair mechanisms, increases genomic instability, and plays an important role as an initiator of SCC. Moreover, the overexpression of TGF-β, which increases with the loss of SMAD4, leads to alterations in the tumor microenvironment, thus causing the progression of SCC [31]. Homozygous deletions (30%) and mutations associated with a loss of heterozygosity (20%) result in a lack of SMAD4 protein expression [32]. Hoot et al. showed that the downregulation of SMAD4 at a gene or protein level due to a loss of heterozygosity results in cSCC [33]. Mitra et al. found that mice with keratinocyte-specific SMAD4 deletion exhibit increased DNA damage and susceptibility to UV-induced skin cancer with chronic UVR exposure [34].
Promoter hypermethylation may precede genetic mutations and genomic instability in tumor development. It is therefore not only critical for carcinogenesis, but also represents a potential therapeutic target [35]. Hypermethylation of the promoter of the SMAD4 gene has been studied in various cancer types [36].
Squamous cell carcinoma subtypes share many phenotypic and molecular characteristics. Studies have shown that both head and neck SCC (HNSCC) and cSCC share common molecular markers [37], and overexpression of TGF-β has been observed in these two cancers [38]. Studies of HNSCCs have shown that SMAD4 expression decreases at rates ranging from 12 to 86%. The variability of these ratios is explained as follows: (1) different criteria are used in studies to define “reduced SMAD4 expression,” and (2) unrelated normal tissue instead of adjacent non-malignant tissue in tissue samples may be used as SMAD4 positive controls [31]. In the study by Bornstein et al., the SMAD4 protein was found to be lost in both the cancerous and normal mucosa of HNSCC [39]. In some cases, the loss of heterozygosity is due to mutations in the SMAD4 gene; in other cases, a decrease in SMAD4 expression may be observed due to methylation in the SMAD4 gene. Numerous studies have been conducted on DNA hypermethylation in HNSCC, and this epigenetic process has been found to be associated with the prognosis of the disease [11, 40]. A review published in 2020 highlighted that hypermethylation had been observed in many genes during the development of cSCC [41]. In our study, mRNA expression of SMAD4 was decreased in the cancer tissues compared with the healthy tissues in the patients with cSCC, while the methylation of SMAD4 was increased. Previous studies have reported hypermethylation in both HNSCCs and cSCCs. Our study is thus the first to show that the SMAD4 gene is hypermethylated in NMSCs.
Lin et al. showed that SMAD4 is involved in HNSCC cell migration and invasion via the suppression of endogenous and exogenous SMAD4 expression in patients with HNSCC [42]. Hervas-Marin et al. compared the methylation status of patients with low- and high-risk SCC and found a different methylation status between the two pathological stages, with hypomethylation in the low-risk stage of cSCC and hypermethylation in the high-risk cSCC stage [43]. In our study, a correlation was found between the differentiation degrees used in the risk assessment of cSCC and the determination of prognosis and SMAD4 mRNA expression levels. These results support the finding that low mRNA expression levels indicate a poor prognosis. In contrast to the data in the literature, a change in the percentage of SMAD4 methylation was not associated with the degree of differentiation of cSCC in our study. Because cSCC has a multifactorial etiopathogenesis, we believe that further studies on this topic are justified.
SMAD4 is known to be expressed in all layers of the epidermis, both in basal proliferative keratinocytes and suprabasal differentiated keratinocytes [38]. In one study, the immunohistochemical loss of SMAD4 expression in cSCC was reported to be 62% [34]. In our study, negative staining of the SMAD4 protein in cancer tissues was found in the patients with cSCC, and these results are consistent with those in the literature.
Studies on BCC are limited in the literature. In terms of the assessment of the methylation status of BCC, Heitzer et al. found the hypermethylated protein patched homolog (PTCH) promoter in only a small number of cases [44]. In a study by Brinkhuizen et al. on BCC subtypes, components of the Sonic Hedgehog and WNT signaling pathways, which play important roles in BCC etiopathogenesis, were shown to be hypermethylated [45]. In our study, hypermethylation of the SMAD4 gene was detected in both the BCC and BCS patients.
In their study, Gambichler et al. determined that the mRNA expression of SMAD4 did not change significantly between lesional and non-lesional skin, and immunohistochemically, no differences in the staining of SMAD4 were detected between lesional and non-lesional skin [21]. Qiao et al. conducted a study on conditional SMAD4 knockout mice and found that the mutant mice developed visible skin tumors at 3–13 months of age; histological analysis of 20 tumors revealed that most were SCC and rarely BCC [46]. In the study by Lange et al., in which they examined SMAD4 protein expression by immunohistochemistry, SMAD4 expression was decreased in the patients with BCC compared to those with benign epithelial tumors. However, the authors did not specify the BCC subtypes [47]. The study by Gambichler et al. was designed to investigate nodular and superficial subtypes of BCC [21]. In our study, no change in the staining level of the SMAD4 protein was detected by immunohistochemistry in either the BCC or BSC tissues compared with the healthy tissue. This again proved that BSC should be considered a subgroup of BCC.
A recent fascinating study by Chiang pointed out that the genomic alterations characteristic of BSC are related to the signaling pathways leading to the development of early stage BCC and those associated with late-stage SCC [4]. In our study, the SMAD4 methylation and mRNA expression levels in the patients with BSC were consistent with those in the patients with BCC and SCC.
To understand the pathogenesis of BCC, the measurements of mRNA and protein levels should be complementary [48]. From the data we obtained, unlike in cSCC, the change in mRNA expression was not commensurate with the change in protein expression in BCC and BSC. This can be explained by the different biological mechanisms that separate the mRNA and protein levels in tissues and the differences between the mRNA and protein measurement methods [49, 50].
The staining characteristics of SMAD4 protein have been found to be increased in NMSC tissues, in patients over 65 years of age, and in patients with chronic sun exposure. These results can be interpreted as follows: (1) the incidence of NMSCs increases with age [1], and (2) under the influence of UVR, gene mutations such as SMAD4 may occur in the long term [14].
Our study had some limitations: (1) It was performed in a small population. (2) A healthy population without a history of skin cancer was not used as the control group. (3) Mutations such as deletion in the SMAD4 gene and the loss of heterozygosity could not be examined.
Conclusion
The mRNA expression level of the SMAD4 gene and its changing protein expression are particularly important for the early diagnosis and prognosis of cSCC. Our study found that SMAD4 methylation increased and SMAD4 mRNA expression decreased in the major subtypes of NMSC. Furthermore, the staining characteristic of SMAD4 protein expression changed only in the patients with cSCC. To our knowledge, this study provides new insights into the pathogenesis of NMSC via the methylation of SMAD4. Despite the limitations mentioned above, we consider this a preliminary study that may guide further research to elucidate the role of SMAD4 in patients with NMSC.
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files.
References
Ciążyńska M, Kamińska-Winciorek G, Lange D et al (2021) The incidence and clinical analysis of non-melanoma skin cancer. Sci Rep 11:4337. https://doi.org/10.1038/s41598-021-83502-8
Sung H, Ferlay J, Siegel RL et al (2021) Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Zambrano-Román M, Padilla-Gutiérrez JR, Valle Y, Muñoz-Valle JF, Valdés-Alvarado E (2022) Non-melanoma skin cancer: a genetic update and future perspectives. Cancers 14:2371. https://doi.org/10.3390/cancers14102371
Chiang A, Tan CZ, Kuonen F et al (2019) Genetic mutations underlying phenotypic plasticity in basosquamous carcinoma. J Invest Dermatol 139:2263-2271e5. https://doi.org/10.1016/j.jid.2019.03.1163
Volkenstein S, Wohlschlaeger J, Liebau J et al (2010) Basosquamous carcinoma–a rare but aggressive skin malignancy. J Plast Reconstr Aesthet Surg 63:e304–306. https://doi.org/10.1016/j.bjps.2009.05.058
Tarapore E, Atwood SX (2019) Defining the genetics of basosquamous carcinoma. J Invest Dermatol 139:2258–2260. https://doi.org/10.1016/j.jid.2019.04.011
Cives M, Mannavola F, Lospalluti L et al (2020) Non-melanoma skin cancers: biological and clinical features. Int J Mol Sci 21:5394. https://doi.org/10.3390/ijms21155394
Calzavara-Pinton P, Ortel B, Venturini M (2015) Non-melanoma skin cancer, sun exposure and sun protection. G Ital Dermatol Venereol 150:369–378
Hanahan D (2022) Hallmarks of cancer: new dimensions. Cancer Discov 12:31–46. https://doi.org/10.1158/2159-8290.Cd-21-1059
Pacella G, Capell BC (2021) Epigenetic and metabolic interplay in cutaneous squamous cell carcinoma. Exp Dermatol 30:1115–1125. https://doi.org/10.1111/exd.14354
Zhou C, Ye M, Ni S et al (2018) DNA methylation biomarkers for head and neck squamous cell carcinoma. Epigenetics 13:398–409. https://doi.org/10.1080/15592294.2018.1465790
Ke Y, Wang XJ (2021) TGFβ signaling in photoaging and UV-Induced skin cancer. J Invest Dermatol 141:1104–1110. https://doi.org/10.1016/j.jid.2020.11.007
Shao Y, Zhang J, Zhang R, Wan J, Zhang W, Yu B (2012) Examination of Smad2 and Smad4 copy-number variations in skin cancers. Clin Transl Oncol 14:138–142. https://doi.org/10.1007/s12094-012-0773-7
Kim Y, He YY (2014) Ultraviolet radiation-induced non-melanoma skin cancer: regulation of DNA damage repair and inflammation. Genes Dis 1:188–198. https://doi.org/10.1016/j.gendis.2014.08.005
Zhao M, Mishra L, Deng CX (2018) The role of TGF-β/SMAD4 signaling in cancer. Int J Biol Sci 14:111–123. https://doi.org/10.7150/ijbs.23230
Yang L, Mao C, Teng Y et al (2005) Targeted disruption of Smad4 in mouse epidermis results in failure of hair follicle cycling and formation of skin tumors. Cancer Res 65:8671–8678. https://doi.org/10.1158/0008-5472.Can-05-0800
Cai H, Sobue T, Kitamura T et al (2020) Epidemiology of nonmelanoma skin cancer in Japan: occupational type, lifestyle, and family history of cancer. Cancer Sci 111:4257–4265. https://doi.org/10.1111/cas.14619
Khalesi M, Whiteman DC, Rosendahl C et al (2015) Basal cell carcinomas on sun-protected vs. sun-exposed body sites: a comparison of phenotypic and environmental risk factors. Photodermatol Photoimmunol Photomed 31:202–211. https://doi.org/10.1111/phpp.12170
Reiter O, Mimouni I, Dusza S, Halpern AC, Leshem YA, Marghoob AA (2021) Dermoscopic features of basal cell carcinoma and its subtypes: a systematic review. J Am Acad Dermatol 85:653–664. https://doi.org/10.1016/j.jaad.2019.11.008
Sgouros D, Theofili M, Damaskou V et al (2021) Dermoscopy as a tool in differentiating cutaneous squamous cell carcinoma from its variants. Dermatol Pract Concept 11:e2021050. https://doi.org/10.5826/dpc.1102a50
Gambichler T, Skrygan M, Kaczmarczyk JM et al (2007) Increased expression of TGF-beta/Smad proteins in basal cell carcinoma. Eur J Med Res 12:509–514
Ozkan U, Ozcelik F, Yildiz M, Budak M (2019) Lipoprotein(a) gene polymorphism increases a risk factor for aortic valve calcification. J Cardiovasc Dev Dis 6:31. https://doi.org/10.3390/jcdd6030031
Gao T, Nie Y, Guo J (2012) Hypermethylation of the gene LARP2 for noninvasive prenatal diagnosis of β-thalassemia based on DNA methylation profile. Mol Biol Rep 39:6591–6598. https://doi.org/10.1007/s11033-012-1489-z
Guo W, Dong Z, Guo Y, Kuang G, Yang Z, Shan B (2012) Concordant repression and aberrant methylation of transforming growth factor-beta signaling pathway genes occurs early in gastric cardia adenocarcinoma. Mol Biol Rep 39:9453–9462. https://doi.org/10.1007/s11033-012-1810-x
Li LC, Dahiya R (2002) MethPrimer: designing primers for methylation PCRs. Bioinformatics 18:1427–1431. https://doi.org/10.1093/bioinformatics/18.11.1427
Rao X, Huang X, Zhou Z, Lin X (2013) An improvement of the 2ˆ(-delta delta CT) method for quantitative real-time polymerase chain reaction data analysis. Biostat Bioinforma Biomath 3:71–85
Lin Z, Zhang L, Zhou J, Zheng J (2019) Silencing Smad4 attenuates sensitivity of colorectal cancer cells to cetuximab by promoting epithelialmesenchymal transition. Mol Med Rep 20:3735–3745. https://doi.org/10.3892/mmr.2019.10597
Xavier-Junior JCC, Ocanha-Xavier JP (2019) WHO,(2018) Classification of Skin Tumors. Am J Dermatopathol 41:699–700. https://doi.org/10.1097/dad.0000000000001446
Woo WA, Suarez MFB, Keohane SG (2021) Summary of the updated 2020 guidelines for cutaneous squamous cell carcinoma. Clin Exp Dermatol 46:1174–1177. https://doi.org/10.1111/ced.14587
He SM, Zhao ZW, Wang Y et al (2011) Reduced expression of SMAD4 in gliomas correlates with progression and survival of patients. J Exp Clin Cancer Res 30:70. https://doi.org/10.1186/1756-9966-30-70
Hernandez AL, Young CD, Wang JH, Wang XJ (2019) Lessons learned from SMAD4 loss in squamous cell carcinomas. Mol Carcinog 58:1648–1655. https://doi.org/10.1002/mc.23049
Bellizzi AM (2020) An algorithmic immunohistochemical approach to define tumor type and assign site of origin. Adv Anat Pathol 27:114–163. https://doi.org/10.1097/pap.0000000000000256
Hoot KE, Lighthall J, Han G et al (2008) Keratinocyte-specific Smad2 ablation results in increased epithelial-mesenchymal transition during skin cancer formation and progression. J Clin Invest 118:2722–2732. https://doi.org/10.1172/jci33713
Mitra D, Fernandez P, Bian L et al (2013) Smad4 loss in mouse keratinocytes leads to increased susceptibility to UV carcinogenesis with reduced Ercc1-mediated DNA repair. J Invest Dermatol 133:2609–2616. https://doi.org/10.1038/jid.2013.213
Hatziapostolou M, Iliopoulos D (2011) Epigenetic aberrations during oncogenesis. Cell Mol Life Sci 68:1681–1702. https://doi.org/10.1007/s00018-010-0624-z
Swellam M, Saad EA, Sabry S, Denewer A, Abdel Malak C, Abouzid A (2021) Alterations of PTEN and SMAD4 methylation in diagnosis of breast cancer: implications of methyl II PCR assay. J Genet Eng Biotechnol 19:54. https://doi.org/10.1186/s43141-021-00154-x
Yan W, Wistuba II, Emmert-Buck MR, Erickson HS (2011) Squamous cell carcinoma—similarities and differences among anatomical sites. Am J Cancer Res 1:275–300
Wu F, Weigel KJ, Zhou H, Wang XJ (2018) Paradoxical roles of TGF-β signaling in suppressing and promoting squamous cell carcinoma. Acta Biochim Biophys Sin (Shanghai) 50:98–105. https://doi.org/10.1093/abbs/gmx127
Bornstein S, White R, Malkoski S et al (2009) Smad4 loss in mice causes spontaneous head and neck cancer with increased genomic instability and inflammation. J Clin Invest 119:3408–3419. https://doi.org/10.1172/jci38854
Franzen A, Bootz F, Dietrich D (2020) Prognostic and predictive methylation biomarkers in HNSCC: Epigenomic diagnostics for head and neck squamous cell carcinoma (HNSCC). HNO 68:911–915. https://doi.org/10.1007/s00106-020-00902-4
Nikolouzakis TK, Falzone L, Lasithiotakis K et al (2020) Current and future trends in molecular biomarkers for diagnostic, prognostic, and predictive purposes in non-melanoma skin cancer. J Clin Med 9:2868. https://doi.org/10.3390/jcm9092868
Lin LH, Chang KW, Cheng HW, Liu CJ (2019) SMAD4 somatic mutations in head and neck carcinoma are associated with tumor progression. Front Oncol 9:1379. https://doi.org/10.3389/fonc.2019.01379
Hervás-Marín D, Higgins F, Sanmartín O et al (2019) Genome wide DNA methylation profiling identifies specific epigenetic features in high-risk cutaneous squamous cell carcinoma. PLoS ONE 14:e0223341. https://doi.org/10.1371/journal.pone.0223341
Heitzer E, Bambach I, Dandachi N, Horn M, Wolf P (2010) PTCH promoter methylation at low level in sporadic basal cell carcinoma analysed by three different approaches. Exp Dermatol 19:926–928. https://doi.org/10.1111/j.1600-0625.2010.01120.x
Brinkhuizen T, van den Hurk K, Winnepenninckx VJ et al (2012) Epigenetic changes in basal cell carcinoma affect SHH and WNT signaling components. PLoS ONE 7:e51710. https://doi.org/10.1371/journal.pone.0051710
Qiao W, Li AG, Owens P, Xu X, Wang XJ, Deng CX (2006) Hair follicle defects and squamous cell carcinoma formation in Smad4 conditional knockout mouse skin. Oncogene 25:207–217. https://doi.org/10.1038/sj.onc.1209029
Lange D, Persson U, Wollina U et al (1999) Expression of TGF-beta related smad proteins in human epithelial skin tumors. Int J Oncol 14:1049–1056. https://doi.org/10.3892/ijo.14.6.1049
Pietkiewicz P, Gornowicz-Porowska J, Bowszyc-Dmochowska M et al (2014) Discordant expression of desmoglein 2 and 3 at the mRNA and protein levels in nodular and superficial basal cell carcinoma revealed by immunohistochemistry and fluorescent in situ hybridization. Clin Exp Dermatol 39:628–635. https://doi.org/10.1111/ced.12355
Li JJ, Biggin MD (2015) Gene expression. Statistics requantitates the central dogma. Science 347:1066–1067. https://doi.org/10.1126/science.aaa8332
Fortelny N, Overall CM, Pavlidis P, Freue GVC (2017) Can we predict protein from mRNA levels? Nature 547:E19–E20. https://doi.org/10.1038/nature22293
Funding
This study was supported by the Trakya University Research Project Fund (2020/55).
Author information
Authors and Affiliations
Contributions
YGÜ and MB designed the research study. MB and EUK performed the research. YGÜ and MB carried out the data analysis and writing—review and editing. All the authors discussed the results and contributed to the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors agree that there are no conflicts of interest to declare.
Ethical approval
Approval for the study was granted by the Scientific Research Ethics Committee of the Trakya University Faculty of Medicine (approval number: 18/14, date: 06/11/2019).
Informed consent
All the subjects who agreed to participate in the study gave their written informed consent.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gürsel Ürün, Y., Budak, M. & Usturalı Keskin, E. Methylation status, mRNA and protein expression of the SMAD4 gene in patients with non-melanocytic skin cancers. Mol Biol Rep 50, 7295–7304 (2023). https://doi.org/10.1007/s11033-023-08656-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08656-2