Abstract
Stripe rust, caused by Puccinia striiformis f. sp. tritici (Pst), is a destructive foliar disease of wheat worldwide. Sustainable control of the disease is preferably achieved by deploying stripe rust resistance genes in wheat cultivars. Our previous studies have shown that Baidatou, a Chinese wheat landrace, displayed good adult-plant resistance (APR) to wheat stripe rust in Gansu Province, an epidemic region for stripe rust in China. To elucidate the genetic basis of APR to stripe rust in Baidatou, a cross between Baidatou and Mingxian 169, which is susceptible to all the known Chinese (Pst) races, was performed. Adult plants of F1, F2 and F2:3 generations derived from the cross Mingxian 169/Baidatou were inoculated in the field with the most prevalent Chinese Pst race, CYR33, in Yangling, Shaanxi Province, during 2009–2010 and 2010–2011 crop seasons, respectively. The results showed that the resistance of Baidatou to stripe rust was conferred by a single dominant gene. Six hundred and sixty simple sequence repeat (SSR) markers and 128 sequence-related amplified polymorphism (SRAP) markers were screened for association with the resistance gene to stripe rust using bulked segregant analysis. Four polymorphic SSR markers and two SRAP markers were identified to be linked to the resistance gene. A linkage map was constructed with six molecular markers and the resistance gene. The genetic distance of two flanking SSR markers to the resistance gene, temporarily designated YrBai, was 3.6 and 5.4 cM, respectively. Based on the position of the SSR markers on the wheat chromosome, YrBai was located on chromosome 6DS. According to the rust reaction patterns, SSR marker allele analysis and the pedigree of the Yr genes on chromosome 6D, YrBai is likely to be a novel APR gene against stripe rust. The specificity of the two flanking markers of YrBai was validated in 99 wheat germplasms. The gene and its flanking markers should be useful for developing wheat cultivars with durable resistance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Common wheat (Triticum aestivum L.) is one of the most important crops worldwide, providing staple foods for 35 % of the world population (Huang and Röder 2004). It is constantly challenged by many biotic stresses such as rusts, powdery mildew and fusarium head blight. Stripe rust, caused by Puccinia striiformis Westend. f. sp. tritici Eriks. (Pst), is one of the most devastating diseases of wheat in cool and temperate regions of the world (Wan et al. 2007; Stubbs 1985; Wellings and McIntosh 1990; Wellings 2011; Chen 2005). In China, it occurs most frequently in the northwestern and southwestern wheat-growing regions (Zeng and Luo 2006, 2008). Historical records show that the most destructive and widespread stripe rust epidemics occurred in 1950, 1964, 1990 and 2002, resulting in severe production losses (Wan et al. 2004, 2007). Breeding and use of resistant wheat cultivars are the most efficient, economically viable and environmental friendly means of controlling the disease (Line 2002; Ma et al. 2013c).
Resistance to rust diseases can usually be classified into seedling resistance and adult-plant resistance (APR) on the basis of the growth stage of the plant (Chen 2005, 2012). Seedling resistance can be detected at early stages of growth, and the resistance is usually effective throughout the life of the host. In contrast, plants with APR show susceptibility at the seedling stage, but resistance at adult-plant stage when inoculated with the same pathotype. Owing to its race-specific nature, seedling resistance is frequently overcome by new races of the pathogen (Line 2002; Chen 2005; Chen et al. 2010). In contrast, APR genes provide resistance to a broader range of pathogens and tend to be more durable.
More than 60 genes conferring resistance to stripe rust have been identified, characterized and officially named (McIntosh et al. 2012; Xu et al. 2013; Basnet et al. 2014; Lu et al. 2014; Ren et al. 2012), and other dozens of genes have been temporarily named (Ma et al. 2013a, b, c; Li et al. 2012; Cao et al. 2008; Li et al. 2010; Sui et al. 2009). About 12 of them have been characterized as APR genes, including Yr16, Yr18, Yr29, Yr30, Yr36, Yr39, Yr48 and Yr52 (Chen 2012).
A series of commercial cultivars with resistance genes have been bred and grown in China since the 1950s. However, most of the commercial resistance wheat cultivars were compromised with the appearance and development of new virulent races of the stripe rust pathogen (Wan et al. 2007). To date, 33 races and dozens of pathotypes have been identified in stripe rust populations in China. Most commercial cultivars are susceptible to the currently predominant races. Therefore, it is urgent to identify new stripe rust resistance genes and molecular markers for their efficient incorporation in cultivars in order to design pyramiding strategies of resistance. Wheat is a major crop in China. As a consequence of breeding programs during its long history of cultivation, numerous resistance types have been deposited in wheat germplasm collections. Examination of disease-resistant germplasm resources is invaluable for the characterization of new stripe rust resistance genes in order to increase their diversification in commercial wheat cultivars (Wan et al. 2004; Zhang et al. 2011; Ma et al. 2013c; Cao et al. 2008).
Baidatou, a Chinese winter wheat landrace, has long been cultivated in the Longnan region of Gansu Province, in which stripe rust epidemics are frequent and severe. Although many new virulent Pst races have occurred in the region over the past five decades, the cultivar is known to have maintained resistance to all Pst races in the field. In addition to its high adaptability and yield, Baidatou seems to have adult-plant resistance to stripe rust. The objective of this study was to map the gene(s) for stripe rust resistance in Baidatou and to identify the molecular markers associated with the resistance.
Materials and methods
Plant and pathogen materials
Baidatou, a Chinese wheat landrace, was obtained from Tianshui Institute of Agricultural Sciences of Gansu Province. Our preliminary study showed that it was susceptible to Chinese Pst races CYR25, CYR27, CYR29, CYR30, CYR31, CYR32, CYR33 and Sun11-4 in seedling stage under controlled greenhouse conditions, but resistant to natural populations of the pathogen at adult-plant stage in field conditions in Tianshui, Gansu Province.
Baidatou was used as the male parent to cross with Mingxian 169, which is highly susceptible to all Chinese Pst races. The F1 plants were grown in greenhouse to obtain F2 seeds, and the F2 plants were grown in 2010 in the field to obtain F2:3 families. Finally, F1, F2 and F2:3 generations derived from the cross Mingxian 169/Baidatou were used for genetic analyses and molecular mapping.
The four US wheat cultivars Fielder, Tres, Tyee and Lee were obtained from the Wheat Genetics, Quality and Physiology Laboratory of the US Department of Agriculture in Pullman, WA, USA. In total, ninety-nine wheat cultivars and breeding lines from the Yellow–Huai growing area were used to evaluate the polymorphisms of the molecular markers flanking the resistance gene in Baidatou.
The predominant Chinese Pst races CYR25, CYR27, CYR29, CYR30, CYR31, CYR32, CYR33 and Sun11-4 were provided by the Laboratory of Plant Resistance and Genetics, Northwest A & F University. The inoculum of each race was prepared by single-spore isolates and validated against a set of Chinese differential hosts.
Resistance evaluation in the greenhouse
The seedling resistance tests were conducted under controlled greenhouse conditions. About 10 seeds of each parent were used. Seedling tests were performed in two-leaf stage plants about 10 days after planting. Seedlings were inoculated separately with Pst races CYR25, CYR27, CYR29, CYR30, CYR31, CYR32, CYR33 and Sun11-4. The inoculum of a mixture of urediniospore and talc at a 1:50 ratio was dusted onto wheat leaves, and plants were then kept in a dark dew chamber at 10 °C for 24 h. The inoculated plants were transferred to an environmentally controlled greenhouse with 16-h light/8-h dark at 12–17 °C. Infection-type (IT) data were recorded 17–18 days after inoculation when rust had fully developed on the susceptible check Mingxian 169. Infection type was based on a 0–4 scale as described by Ma et al. (2013c). Plants with IT 0–2+ were considered to be resistant, and those with IT 3−–4 susceptible. The adult-plant tests were conducted in plots with three replicates under controlled greenhouse conditions. Inoculations were performed with race CYR33 at the booting stage in spring. Stripe rust infection types (IT) were scored when disease severity was 10 % on the susceptible control, and data were then collected at weekly intervals.
Resistance evaluation in the field
In the 2009–2010 crop season, the parents, seven F1 and the 160 F2 plants of Mingxian 169/Baidatou were sown at a nursery in the experimental station in Yangling, Shaanxi Province, where the weather conditions are favorable for spreading or stripe rust infection. In the 2010–2011 sowing season, the F2:3 lines derived from the F2 plants were sown at the same location. Thirty seeds were evenly planted in a 1.0-m row in a randomized complete block design, with each line repeated three times. The susceptible cultivar Mingxian 169 was planted every 10 rows and around the test plots to increase the uniformity of inoculum throughout the field. Standard fertilizer and cropping practices were used for the field management. Inoculation was also performed with race CYR33, the prevalent race in the region, at the beginning of stem extension stage. Stripe rust IT was recorded when disease severity was 30 and 80 % on the susceptible control. If all the plants in a row had uniform IT, the line was considered to be homozygous susceptible when IT was 3–4 or resistant when IT was 0–2. When the infection type was different between individual plants in a row, the line was considered to be non-homozygous.
SSR analysis
Genomic DNA was extracted from healthy leaves using the CTAB method as modified by Yan et al. (2003). One hundred and fifty F2:3 lines from Mingxian 169/Baidatou were used as a mapping population. Based on the F3 disease phenotypic data, DNA from 15 homozygous resistant lines and 15 homozygous susceptible lines were used for constructing resistant and susceptible bulks, respectively. Each bulk was composed of equal amounts of F2:3 plant DNA (Michelmore et al. 1991).
The simple sequence repeat (SSR) procedure was used to identify markers linked to the resistance loci. SSR primer sequences were obtained from the GrainGenes 2.0 Web site (http://wheat.pw.usda.gov). A total of 660 wheat SSR markers on different chromosomes were screened on the two parental lines and the resistant and susceptible bulks to detect polymorphisms. A 15-μL reaction mixture consisted of 150 ng of template DNA, 1.5 μL 10× PCR buffer, 5 mM of MgCl2, 1.5 unit of Taq DNA polymerase, 0.25 mM each of dCTP, dGTP, dTTP and dATP, and 7.5 µM of each primer. PCR amplification of SSRs was performed in an MJ Research PTC-200 thermal cycler. The PCR program consisted of initial denaturation at 94 °C for 5 min, followed by 35 cycles of 30-s denaturation at 94 °C, 1-min annealing at 50–68 °C depending on primers and 45-s extension at 72 °C, with a final extension at 72 °C for 10 min. After amplification, 6 μL formamide loading buffer [98 % formamide, 10 mM EDTA (pH 8.0), 0.5 % (W/V) xylene cyanol and 0.5 % (W/V) bromophenol blue] was added to the PCR product. About 5–7 μL mixture of the PCR product and loading buffer for each sample was loaded for electrophoresis on 8 % polyacrylamide gels. The gel was stained and visualized as previously described (Bassam et al. 1991).
Data analysis and linkage map construction
Segregation of markers and resistance gene loci was tested for goodness-of-fit to the expected ratio using the chi-squared tests with the Excel data analysis of Microsoft Office 2007. A genetic linkage map of sequence-related amplified polymorphism (SRAP) markers, SSR markers and the resistance loci was constructed using MAPMAKER version 3.0b (Lincoln et al. 1992). The recombination frequency was converted to map distance (in centimorgans) according to the Kosambi mapping function (Kosambi 1943). The markers and resistance gene loci were organized into a linkage group with the ‘group’ command at a minimum LOD = 3 and maximum distance between two loci of 37.5 cM. The linkage map was drawn using the software Mapdraw version 2.1 (Liu and Meng 2003).
Determination of polymorphism of flanking markers in wheat genotypes
The usefulness of a molecular marker in resistance breeding largely depends upon the polymorphism between the resistance gene donor and genotypes without the gene. To determine the usefulness of flanking markers Xcfd19 and Xbarc54 in marker-assisted selection, 99 wheat genotypes (Supplementary Table 3), including currently grown spring and winter cultivars, advanced breeding lines and stripe rust differentials, were evaluated. The closest SSR markers to the resistance gene in Baidatou were used to test four wheat cultivars, Fielder, Lee, Tres and Tyee, which contained stripe rust resistance genes on wheat chromosome 6D.
Results
Characterization of stripe rust resistance in Baidatou
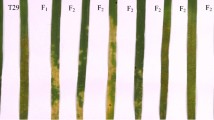
In the tests under controlled greenhouse conditions, Baidatou was susceptible (IT 3–4) to Pst races CYR25, CYR27, CY29, CYR30, CYR31, CYR32, CYR33 and Sun11-4 at the seedling stage. However, adult plants of Baidatou were highly resistant (IT 0;). The control Mingxian 169 was consistently susceptible (IT 4) with abundant uredinia in all tests. In the field tests, Baidatou was resistant (IT 0; to 2) at adult plant stage in Yangling. In contrast, Mingxian 169 was susceptible (IT 4) in all tests (Supplementary Table 1). The data were consistent with the previous evaluations of more than 10 races from 2005 to 2010 at Yangling and Tianshui (unpublished). These results confirmed that Baidatou has the typical adult-plant resistance effective against a wide range of races.
When adult plants of the parents, F1 and F2 were tested with race CYR33 under field conditions, Mingxian 169 was susceptible (IT 4) with abundant uredinia and higher severity, whereas Baidatou was resistant (IT 0;) (Table 1). All seven F1 plants had IT 0; (necrotic flecks, without sporulation). The 160 inoculated F2 plants were divided into 123 resistant (IT 0–2+) and 37 susceptible (IT 3−–4), a 3:1 ratio (χ 2 = 0.21, P = 0.58) (Table 1). We harvested F2:3 seeds from 155 of the 160 F2 plants. Of them, 35 were homozygous resistant, 83 segregating and 27 homozygous susceptible lines, showing a 1:2:1 (resistant/segregating/susceptible) ratio (χ 2 = 0.83, P = 0.66). All these results together indicated that a single dominant gene conferred the stripe rust resistance to CYR33 in Baidatou. We tentatively designated this gene as YrBai.
Mapping of the adult resistance gene YrBai
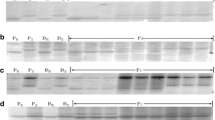
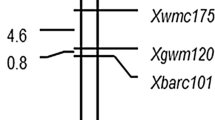
Of 660 SSR markers screened, four pairs of SSR primers produced bands appearing to be associated with stripe rust response in the bulked segregant analysis (BSA). The associations were confirmed by genotyping all 155 F2:3 plants. A total of 128 SRAP markers were also screened in the BSA to enrich marker density; two of them were polymorphic and associated with the resistance locus. When genotyped on all the F2:3 population, the four SSR markers and two SRAP markers fitted a 3:1 (dominant) or 1:2:1 (co-dominant) ratio, indicating that each of the six markers inherited as the single locus (Supplementary Table 2). The polymorphism of SSR markers Xbarc54 and Xcfd19 is shown in Supplementary Fig. 1 as an example. Using these markers, a linkage group spanning 37.8 cM was constructed around the resistance gene locus (Fig. 1). Two SSR markers (Xbarc54 and Xcfd19) were the closest markers flanking YrBai, with map distances of 3.6 and 2.1 cM, respectively. Based on the wheat SSR consensus map (http://wheat.pw.usda.gov), the four linked SSR markers were located on the short arm of chromosome 6D, assigning YrBai on chromosome 6DS.
Polymorphism of markers flanking the YrBai locus in various wheat genotypes
To determine the usefulness of the two flanking markers, Xbarc54 and Xcfd19, in marker-assisted selection, a total of 99 winter wheat genotypes were tested for polymorphism (Supplementary Table 3). Of the 99 genotypes, 80 (80.8 %) and 82 (82.8 %) did not have the alleles that were present in Baidatou at Xbarc54 and Xcfa19, respectively. Thirteen wheat genotypes had only the Xbarc54 allele of Baidatou. For these 13 cultivars, Xcfd19 should be used for selection. In addition, 11 wheat genotypes had only the Xcfd19 target allele. For these 11 cultivars, Xbarc54 should be used. A total of six genotypes (Womai 0608, Danshi 802, Zhongmai 895, Luo 3429, Huaimai 05155 and Wenhang 6) susceptible to Pst race CYR33 showed the same alleles as Baidatou at both loci. Thus, the two flanking markers could not be directly used in marker-assisted selection for YrBai in crosses of these genotypes with Baidatou as donor due to lack of polymorphism of the markers in these cultivars. Because the six winter wheat genotypes are separately susceptible to CYR33 at seedling and adult stages, they cannot have YrBai. Single nucleotide polymorphisms (SNPs) should be developed to incorporate the gene YrBai into these six cultivars in marker-assisted selection.
Because stripe rust resistance genes in wheat cultivars Fielder (Yr20), Lee (Yr23), Tres (YrTr1) and Tyee (YrTye) were previously mapped on chromosome 6D, they were compared with Baidatou using the two closest flanking SSR markers (Xbarc54 and Xcfd19). The SSR analyses showed that the target allele of Xbarc54 and Xcfd19 was present only in Baidatou (Fig. 2). According to the greenhouse test, Fielder carrying Yr20, Lee carrying Yr23, Tres carrying YrTr1 and Tyee carrying YrTye were separately susceptible to CYR33 at seedling and adult stages (Supplementary Table 1). The results indicated that the resistance gene in Baidatou may be different from these four Yr genes.
Discussion
Baidatou is an old Chinese winter wheat landrace, and since the 1950s it has been cultivated over a large area in Tianshui of Gansu Province in China. It was highly resistant at the adult-plant stage in different locations in the field, although it was susceptible to most predominant Chinese Pst races at the seedling stage. In the present study, we have shown that Baidatou has a typical adult-plant resistance to stripe rust. We have identified a single dominant gene conferring adult-plant resistance in Baidatou and mapped the gene on wheat chromosome 6DS, tentatively designating it YrBai. Understanding of the resistance gene in Baidatou is very helpful for enriching the resources of disease-resistance genes for wheat breeding programs, providing a valuable resistance source for the sustainable control of wheat stripe rust.
Yr20, Yr23, YrTr1 and YrTye are located on chromosome 6D (Chen et al. 1995a, b). In this study, we identified a new resistance gene YrBai in wheat landrace Baidatou. Comparing the SSR genotypes, YrBai is different from the four other identified Yr genes. Moreover, the four other Yr genes had been located on chromosome 6D only by the allelic analysis and monosomic analysis methods. However, there are no published articles on the fine chromosome location for these genes. From previous studies (Chen et al. 1995a, b), Yr20, Yr23, YrTr1 and YrTye are different based on diallel cross (Chen and Line 1992). From pedigree, Fielder and Lee originated from North America, while Baidatou is a winter wheat landrace in Gansu Province. All these results together strongly suggest that YrBai is different from those other genes on wheat chromosome 6D.
To obtain durable and high-level rust resistance, the ideal approach is to pyramid effective all-stage resistance into a single wheat cultivar (Chen 2012). Stripe rust prefers cool temperatures, and it usually infects wheat from seedling stage to milky stage. Introgressing all-stage resistance genes into adult-plant resistance cultivars can prevent rust infection in wheat at the early growth stage and reduce rust inoculums in the epidemic region. This will also slow down the occurrence of new virulence combinations by genetic recombination (Smith et al. 2002). YrBai is an adult-plant resistance gene conferring a high level of resistance to stripe rust in a hot epidemic region of southern Gansu Province. Introgression of YrBai from Baidatou into wheat has become crucial to the continuing needs for sources of resistance to stripe rust in wheat. Because resistance to stripe rust in many Chinese cultivars has a relatively narrow genetic base, the availability of YrBai in wheat and identification of flanking markers should be beneficial for increasing the overall diversity available for wheat breeding.
References
Basnet B, Singh R, Ibrahim A, Herrera-Foessel S, Huerta-Espino J, Lan C, Rudd J (2014) Characterization of Yr54 and other genes associated with adult plant resistance to yellow rust and leaf rust in common wheat Quaiu 3. Mol Breed 33:385–399
Bassam BJ, Caetano-Anolles G, Gresshoff PM (1991) Fast and sensitive silver staining of DNA in polyacrylamide gels. Anal Biochem 196(1):80–83
Cao Z, Deng Z, Wang M, Wang X, Jing J, Zhang X, Shang H, Li Z (2008) Inheritance and molecular mapping of an alien stripe-rust resistance gene from a wheat-Psathyrostachys huashanica translocation line. Plant Sci 174(5):544–549
Chen X (2005) Epidemiology and control of stripe rust [Puccinia striiformis f. sp. tritici] on wheat. Can J Plant Pathol 27(3):314–337
Chen X (2012) High-temperature adult-plant resistance, key for sustainable control of stripe rust. Am J Plant Sci 4:608–627
Chen X, Line RF (1992) Genes for resistance to stripe rust in ‘Tres’ wheat. Crop Sci 32(3):692–696
Chen X, Jones S, Line R (1995a) Chromosomal location of genes for stripe rust resistance in spring wheat cultivars Compair, Fielder, Lee, and Lemhi and interactions of aneuploid wheats with races of Puccinia striiformis. Phytopathology 85(3):375–381
Chen X, Line RF, Jones SS (1995b) Chromosomal location of genes for resistance to Puccinia striiformis in winter wheat cultivars Heines VII, Clement, Moro, Tyee, Tres, and Daws. Phytopathology 85(11):1362–1367
Chen X, Penman L, Wan A, Cheng P (2010) Virulence races of Puccinia striiformis f. sp. tritici in 2006 and 2007 and development of wheat stripe rust and distributions, dynamics, and evolutionary relationships of races from 2000 to 2007 in the United States. Can J Plant Pathol 32(3):315–333
Huang X-Q, Röder MS (2004) Molecular mapping of powdery mildew resistance genes in wheat: a review. Euphytica 137(2):203–223
Kosambi D (1943) The estimation of map distances from recombination values. Ann Hum Genet 12(1):172–175
Li Q, Hu M, Chen J, Jing J, Wang B, Zhou X (2010) Inheritance and molecular mapping of genes for all-stage resistance to stripe rust in wheat cultivar N. Strampelli. Can J Plant Sci 90(4):529–536
Li Q, Huang J, Hou L, Liu P, Jing J, Wang B, Kang Z (2012) Genetic and molecular mapping of stripe rust resistance gene in wheat—Psathyrostachys huashanica translocation line H9020-1-6-8-3. Plant Dis 96(10):1482–1487
Lincoln SE, Daly MJ, Lander ES (eds) (1992) Constructing genetic linkage maps with Mapmarker/EXP3.0. In: A Whitehead Institute for Biomedical Research Technical Report, 3rd edn. Whitehead Institute, Cambridge
Line RF (2002) Stripe rust of wheat and barley in North America: a retrospective historical review 1. Annu Rev Phytopathol 40(1):75–118
Liu R, Meng J (2003) MapDraw: a microsoft excel macro for drawing genetic linkage maps based on given genetic linkage data. Hereditas 25(3):317
Lu Y, Wang M, Chen X, See D, Chao S, Jing J (2014) Mapping of Yr62 and a small-effect QTL for high-temperature adult-plant resistance to stripe rust in spring wheat PI 192252. Theor Appl Genet 127(6):1449–1459. doi:10.1007/s00122-014-2312-0
Ma D, Hou L, Tang M, Wang H, Li Q, Jing J (2013a) Genetic analysis and molecular mapping of a stripe rust resistance gene YrH9014 in wheat line H9014-14-4-6-1. J Integr Agric 12(4):638–645
Ma D, Jing J, Hou D, Li Q, Zhou X, Du J, Lu Q (2013b) Genetics and molecular mapping of a high-temperature resistance gene to stripe rust in seeding-stage in winter wheat cultivar Lantian 1. J Integr Agric 12(6):1018–1025
Ma D, Zhou X, Hou L, Bai Y, Li Q, Wang H, Tang M, Jing J (2013c) Genetic analysis and molecular mapping of a stripe rust resistance gene derived from Psathynrostachys huashanica Keng in wheat line H9014-121-5-5-9. Mol Breed 32:365–372
McIntosh R, Yamazaki Y, Dubcovsky J, Rogers W, Morris C, Somers D, Appels R, Devos K (2012) MacGene 2012: catalog of gene symbols for wheat. http://www.shigen.nig.ac.jp/wheat/komugi/genes/download.jsp. Accessed 4 Aug 2013
Michelmore RW, Paran I, Kesseli R (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci USA 88(21):9828–9832
Ren R, Wang M, Chen X, Zhang Z (2012) Characterization and molecular mapping of Yr52 for high-temperature adult-plant resistance to stripe rust in spring wheat germplasm PI 183527. Theor Appl Genet 125(5):847–857
Smith P, Koebner R, Boyd L (2002) The development of a STS marker linked to a yellow rust resistance derived from the wheat cultivar Moro. Theor Appl Genet 104(8):1278–1282
Stubbs R (1985) Stripe rust. In: Roelfs AP, Bushnell WR (eds) The cereal rusts, vol II. Academic Press, New York, pp 61–101
Sui X, Wang M, Chen X (2009) Molecular mapping of a stripe rust resistance gene in spring wheat cultivar Zak. Phytopathology 99(10):1209–1215
Wan A, Zhao Z, Chen X, He Z, Jin S, Jia Q, Yao G, Yang J, Wang B, Li G (2004) Wheat stripe rust epidemic and virulence of Puccinia striiformis f. sp. tritici in China in 2002. Plant Dis 88(8):896–904
Wan A, Chen X, He Z (2007) Wheat stripe rust in China. Crop Pasture Sci 58(6):605–619
Wellings CR (2011) Global status of stripe rust: a review of historical and current threats. Euphytica 179(1):129–141
Wellings CR, McIntosh R (1990) Puccinia striiformis f. sp. tritici in Australasia: pathogenic changes during the first 10 years. Plant Pathol 39(2):316–325
Xu L, Wang M, Cheng P, Kang Z, Hulbert S, Chen X (2013) Molecular mapping of Yr53, a new gene for stripe rust resistance in durum wheat accession PI 480148 and its transfer to common wheat. Theor Appl Genet 126(2):523–533
Yan G, Chen X, Line R, Wellings C (2003) Resistance gene analog polymorphism markers co-segregating with the Yr5 gene for resistance to wheat stripe rust. Theor Appl Genet 106:636–643
Zeng S, Luo Y (2006) Long-distance spread and interregional epidemics of wheat stripe rust in China. Plant Dis 90(8):980–988
Zeng S, Luo Y (2008) Systems analysis of wheat stripe rust epidemics in China. Eur J Plant Pathol 121(4):425–438
Zhang S, Xu Z, Wang R, Li Q, Yao Q, Jing J (2011) Genetics and molecular mapping of stripe rust resistance gene YrShan515 in Chinese wheat cultivar Shan 515. Agric Sci China 10(4):553–559
Acknowledgments
The authors are grateful to Dr. Meinan Wang for critical reviews of this manuscript, and to Dr. Simón-Mateo Carmen and Dr. Zeguang Liu for English language editing of the manuscript. The research was supported by the National Basic Research Program of China (2013CB127700), the National High Technology Research and Development Program (863 Program) funding (2012AA101503), the 111 Project from the Education Ministry of China (No. B07049) and the National Science and Technology Support Program (2012BAD19B04).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Dongfang Ma and Qiang Li made equal contributions to this study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ma, D., Li, Q., Tang, M. et al. Mapping of gene conferring adult-plant resistance to stripe rust in Chinese wheat landrace Baidatou. Mol Breeding 35, 157 (2015). https://doi.org/10.1007/s11032-015-0244-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-015-0244-2