Abstract
Stripe rust, which is caused by Puccinia striiformis f. sp. Tritici (Pst), is one of the most destructive wheat (Triticum aestivum L.) diseases worldwide. To control stripe rust, the best strategy is breeding and growing novel resistant cultivars. Qubaichun (QBC), a Tibetan wheat landrace, displays near-immune resistance to wheat stripe rust in western China. Previously, our studies have shown that the stripe rust resistance of QBC is controlled by a dominant gene at the seedling stage and two independent genes at the adult-plant stage. These two genes comprise an all-stage resistance (ASR) gene and a durable adult resistance gene, which was identified as Yr18. The unknown ASR gene is temporarily named Yrqbc. To map this gene, a segregating population of QBC × Chinese Spring (CS) was generated. SSR analysis, BSR-seq and Infinium 660 K iSelect SNP genotyping were successively performed. The results show that Yrqbc finely mapped to a 5.1 cM genetic interval between molecular markers A009200 and A009192, and the genetic distance to the marker A009200 was 0.1 cM. Furthermore, Yrqbc was confirmed to be Yr5 by sequencing. Diagnostic markers are used for detection Yr5 and Yr18 in 323 new cultivars (lines) worldwide. The results show that 27 cultivars (lines) carry the durable adult resistance gene Yr18, and only one material carrying the ASR resistance gene Yr5. In order to analyse the effectiveness and practicality of transferring the pyramided of Yr18 and Yr5, these two stripe rust resistance genes were introduced into elite cultivars which lost resistance. Field test results verified that the combination of Yr5 and Yr18 could provide effective resistance to stripe rust for the whole life of wheat, which provided a genetic foundation for the near immunity of elite cultivars. As a result, this Tibetan landrace could be used for developing high-level, durable resistant wheat cultivars.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Common wheat (Triticum aestivum L.), one of the most important crops worldwide, provides staple food for 35–40% of the global population (https://www.fao.org/faostat/zh/?#data). However, its yield has been strongly restricted by Pst, which is one of the most severe wheat diseases worldwide (Chen 2005). This disease can cause up to 100% of crop yield losses, although the losses are typically in the range of 10–70% (Chen 2005). China has the largest stripe rust epidemic region in the world in terms of wheat acreage affected by the disease (Wan et al. 2007). Sichuan Province is the inoculum base, the centre of diversity, and a major over-summering area of the pathogen (Zeng and Luo 2006; Wan et al. 2007). For example, CYR34, first monitored in this area, has now become the first major epidemic race in China (Liu et al. 2010). CYR34 presents a high frequency of 17–65% and has led to severe yield losses. It has expanded to nine provinces in China and exhibits a further expansion trend, which poses a great threat to wheat safety, production, and disease resistance breeding (Jin et al. 2018). Therefore, the prevention and control of stripe rust races are imminently needed. The most effective, economical and environmentally friendly control strategy is to plant resistant varieties (Chen 2005). However, breeding a new high-yield, high-resistance cultivated variety takes a long time, and most resistance genes have lost their resistance to the new races of stripe rust because of the monoculture deployment of resistant cultivars in widespread areas (Ellis et al. 2014). Therefore, it is imperative to discover novel germplasms for their efficient incorporation into cultivars to design a “pyramiding strategy” of resistance (Elisabeth et al. 2017; Zhang et al. 2001).
To develop high-level, durable resistant cultivars, two types of resistant genes are combined by breeders when using a “pyramiding strategy” (Johnson 1981). Generally, one is the all-stage resistance (ASR) gene, and the other is a durable resistance gene (Chen 2013). The ASR gene protects wheat from infection at the seedling stage and can also play an important role in the HTAP gene at the adult stage via different molecular mechanisms (Chen et al. 2013). No studies have reported the effectiveness of the natural pyramiding of ASR and HTAP or slowing rust resistance gene using known genes except for those parts of QTLs (Guo et al. 2008). Previously, the winter wheat cultivar Druchamp was reported to contain both HTAP QTLs and ASR genes and exhibits durable resistance since it was introduced from France to the United States in the late 1940s (Hou et al. 2015).
“Pyramiding strategy” has been widely implemented for disease resistance in other crops to achieve high-level or durable resistance (Elisabeth et al. 2017; Werner et al. 2005). In soybean, Rsv1, Rsv3, and Rsv4 were pyramided for mosaic virus (SMV) resistance (Shi et al. 2009). In pea, sym9 and sym10 were pyramided to promote nodulation processes during rhizobial or endomycorrhizal endosymbiosis (Schneider et al. 2002). In rice, pi21, pi34 and pi35 were aggregated to enhance rice blast resistance (Nobuko et al. 2015). This “pyramiding strategy” has also been reported in wheat for other diseases (Bariana et al. 2007). For example, Lr41, Lr42 and Lr43 were transferred from Triticum tauschii to common wheat to improve wheat leaf rust resistance (Cox et al. 1994). For stripe rust disease, Yr64 and Yr15, this two genes were estimated to be linked in repulsion and separated by 7.8 cM, both on the chromosome 1BS, were pyramided to achieve high-level and durable resistance, however, it is very difficult to pyramid genes linked in repulsion from different genotypes, but this approach may accelerate the pyramiding and selection processes in breeding programs (Yanmin et al. 2019). Moreover, two HTAP genes—even four to five HTAP genes—were aggregated to improve resistance (Yan et al. 2014).
Previously, QBC, a wheat landrace from Tibet, was collected and analysed. QBC is a landrace formed by natural for a long time due to its geographical location and growing history. Phenotypic identification has indicated that QBC displayed near-immune resistance to stripe rust in field test in Sichuan Province, where Pst is severe and naturally occurring. To analyse the resistance genes in QBC, a segregating population [QBC × Chuanmai 28 (CM 28), with CM28 being highly susceptible to stripe rust] was generated. The results suggested the presence of one dominant gene for stripe rust resistance in the test population at the seedling stage (Supplementary Table 1). However, at the adult stage, the result showed that the stripe rust resistance of QBC was controlled by two dominant genes (Zhou et al. 2015). The segregating ratio of these two populations (QBC × CM 28 direct and reciprocal crossing) was 12:3:1 (resistant:intermediate resistant:susceptible) (Supplementary Table 2). Moreover, QBC exhibited a phenotype of apical necrosis in field investigations, which indicated that it possibly harboured a durable adult resistance gene (Singh et al. 2005; Rosewarne et al. 2006). The durable adult resistance gene was identified as Yr18 by using functional marker analysis (Lagudah et al. 2009).
Previous studies have confirmed that QBC resistance is controlled by two independent dominant genes. One is the durable adult resistance gene (Yr18), and the other is an unknown ASR gene, temporarily named Yrqbc. The objectives of the present study were (1) to finely map Yrqbc and develop co-segregating molecular markers for stripe rust resistance-assisted breeding; (2) to confirme that Yr5 and Yrqbc are the same gene by using Yr5-specific maker and sequencing; (3) to analyse the effectiveness and practicality of the combination of two resistance genes in QBC for durable and high-level resistance breeding. This study is expected to generate information on the mechanism of “pyramiding strategy” and incorporate resistance genes from QBC into elite cultivars.
Materials and methods
Plant material and stripe rust pathogens
QBC, CS, Huixianhong (HXH, which is highly susceptible to stripe rust and was an induced cultivar) and 323 elite cultivars germplasms which included 271 Chinese wheat cultivars (lines) (32 Northern winter wheat area cultivars, 99 Huanghuai winter wheat area cultivars, 103 Southwest winter wheat area cultivars, 15 middle and lower reaches of the Yangtze River cultivars, and 5 Qinghai-Tibet spring/winter wheat area cultivars), 42 CIMMTY cultivars (lines), 10 European cultivars and Australian cultivars. All these germplasms were provided by the Chengdu Institute of Biology, Chinese Academy of Sciences. The Chinese wheat Pst races CYR32, CYR33, CYR34, Su11-4 and Su11-7 were provided by the College of Plant Protection, Northwest A&F University, for the identification of stripe rust resistance. QBC was used as a parent for the generation of two segregating populations, with each reciprocally crossed with CS (which is susceptible to stripe rust at the seedling stage but moderately resistant at the adult stage). The F1, F2 and F2:3 generations derived from the reciprocal crosses of QBC × CS were planted in the field in Chengdu.
Identification of stripe rust resistance
In the growing season, QBC, CS, CM28, 12 elite cultivars (Chuanmai 42, Chuanmai 104, Chuanmai 107, Chuanyu 12, Chuanyu 16, Chuanyu 20, Mianyang 26, Mianyang 28, Mianmai 367, Zhongkemai 47, Zhongkemai 138 and Yangmai 158), and 323 elite cultivars (lines) as well as the F1 and F2 generations of the reciprocal crosses of QBC × CS were sown in the field in Chengdu. Thirteen seeds were planted in a 1.5 m row, with 10 cm spacing between rows. HXH, susceptible control or induced lines, was planted around the nursery lines every 10 rows. Standard fertilizer and cropping practices were applied for field management. Inoculation was performed with mixed races of CYR32, CYR33, CYR34, Su11-4 and Su11-7 by the field spraying method at the beginning of the stem elongation stage. Stripe rust ITs were recorded when the disease was in a sufficient infection situation. Infection types 0–3, 4–6 and 7–9 were considered resistant, intermediate and susceptible, respectively (Line and Qayoum 1992).
DNA extraction and polymerase chain reaction (PCR)
At the seedling stage, the healthy leaves of QBC, CM28, CS, the F1 and F2 generations of their crossing combinations, and 323 wheat cultivars (lines) were collected for extraction of genomic DNA using the improved CTAB method (Rogers and Bendich 1985).
PCR was performed on a Mastercycler Nexus SX1 PCR instrument. The reaction volume was 10 μL, which consisted of 5 μL of 2 × master mix, 0.1 μL of forward primer (10 μM), 0.1 μL of reverse primer(10 μM), 0.8 μL of DNA template and 4 μL of ddH2O. The PCR programme consisted of initial denaturation at 94 °C for 4 min, followed by 35 cycles of 30 s of denaturation at 94 °C, 30 s to 1 min of annealing at 52–68 °C (depending on the primers) and 45 s of extension at 72 °C, with a final extension at 72 °C for 10 min. Two microlitres of PCR product was then loaded for electrophoresis on 8% polyacrylamide gels for SSR analysis (https://maswheat.ucdavis.edu).
SSR analysis
Two hundred eight F2:3 lines from the direct cross of CS × QBC were used for mapping the resistance gene Yrqbc. Based on the method of bulked segregant analysis (BSA) and the F3 disease phenotypic data, DNA from 15 homozygous resistant lines and 15 homozygous susceptible lines was used for constructing resistant and susceptible bulks, respectively. Each bulk was composed of equal amounts of DNA. A total of 371 pairs of SSR markers were screened within the two parental lines and the two bulks to detect polymorphisms (Gupta et al. 2002; Qi et al. 2004; Röder et al. 1998; Somers et al. 2004; Sourdille et al. 2004). The SSR primers were synthesized according to the published sequence of wheat SSR primers available at https://wheat.pw.usda.gov/GG3/, https://www.wheatgenome.org/, https://www.ncbi.nlm.nih.gov/ and other databases. Fragment analysis of the PCR products was carried out on 8% non-denaturing polyacrylamide gels (39 acrylamide: 1 bisacrylamide). After electrophoresis, the gels were silver stained and imaged. Mapmaker/EXP 3.0b software was used to calculate the linkage between markers and stripe rust resistance genes, and a linkage map was constructed by MapDraw 2.0 software.
BSR-seq and Infinium 660 K iSelect SNP genotyping
Bulked segregant analysis (BSA) combined with transcriptome sequencing (RNA-seq) was applied for enriching the makers of this gene (Ramirez-Gonzalez et al. 2015; Cheng et al. 2016). Two mixture pools were constructed by selecting individuals with extreme traits in each population. According to the phenotypic identification results, 20 homozygous resistant/susceptible individuals were screened from the F2 population of QBC × CS. Total RNA from the mixture pools was extracted for transcriptome sequencing. The transcriptome data were ultimately analysed by the classic Bayesian algorithm, and SNP markers were developed to predict the genome segment of the target gene. RNA-seq of the bulked pools was implemented to identify SNPs associated with Yrqbc and then were classified based on the results from the sequenced bulks. The bulk frequency ratio (BFR) of the SNPs between the resistant and susceptible bulks was calculated, and those showing six-fold enrichment/depletion in the corresponding bulks were selected.
Furthermore, SNP genotyping of the parents of the CS × QBC population was carried out using a wheat 660 K array (Cui et al. 2017). Non-polymorphic and low-quality SNPs were excluded, which included those with more than 20% deletion value, nonsignificant clustering and less than 5% allele frequency. In addition, in reference to the version 1.0 genome sequence (IWGSC) of Chinese Spring, the different SNPs within the flanking sequences and genomes were queried via local BLAST searches. This information was then used to develop KASP markers.
Detection of Yr5 and Yr18 in cultivars (lines)
The Yr5 gene-specific marker was used to detect Yr5 in the 323 cultivars (lines) and QBC. These cultivars (lines) were the major new cultivars released in China for commercial production and planting in wheat areas. PCR amplification was conducted at 63 °C for 35 cycles. The primer combinations L34SPF/L34DINT13R2 and L34DINT9F/L34MINUSR were used for Yr18 detection. This reaction amplified two bands of contrasting size that could easily be resolved in a 1% agarose gel: a 751 bp fragment specific for the + Yr18 allele and a 523 bp fragment specific for the − Yr18 allele (Lagudah et al. 2009).
Transferring of Yr5 and Yr18 into elite cultivars
In order to verify the validity of pyramiding Yr5 and Yr18, and the feasibility of QBC breeding, we screened 12 elite cultivars (Chuanmai 42, Chuanmai 104, Chuanmai 107, Chuanyu 12, Chuanyu 16, Chuanyu 20, Mianyang 26, Mianyang 28, Mianmai 367, Zhongkemai 47, Zhongkemai 138 and Yangmai 158) were selected as receptor parents, and the IT value survey results of these cultivars (lines) were all highly susceptible. Then, using the backcross breeding method combined with the marker-aided selection (MAS) method, the resistance genes, Yr5 and Yr18, in QBC were introduced into the elite cultivars (lines) selected above. Field test were record the phenotype of their offspring.
Results
Localization of Yrqbc
A genetic map was constructed from a set of 281 SSR markers by using bulked segregant analysis (BSA). Only one marker (Xwmc175) was detected to be polymorphic between the parental lines and exhibited linkage with the resistant and susceptible bulks. Xwmc175 is located on the long arm of chromosome 2B. According to the chromosome specificity of SSR, Xwmc175 was identified as a co-dominant marker. To enrich the location of Yrqbc on chromosome 2BL, 19 SSR makers (Xcfd73, Xgwm501, Xgwm191, Xgwm120, Xgwm16, Xwmc332, Xwmc627, Xwmc149, XbarcM139, Xbarc1064, Xbarc167, Xbarc91, XbarcM147, Xbarc128, Xbarc101, Xgwm388, Xwmc435, Xwmc499, Xbarc1027) close to Xwmc175 were used for mapping. The results indicated that five primers (Xwmc332, Xbarc167, Xbarc101, Xbarc91 and Xgwm120) were polymorphic.
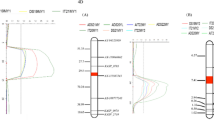
To further confirm the genetic linkage between the six markers and Yrqbc, the F2 segregating population of QBC × CS was investigated. By the chi-square test, the ratio of amplification of these markers was in accordance with the theoretical separation ratio of 1:2:1. This result suggests that these markers are reliable for genetic linkage map construction. As a result, the linkage map was constructed around Yrqbc by using these markers (Fig. 1; Supplementary Table 3). The results indicated that two SSR markers, Xwmc332 and Xwmc175, were the closest flanking markers to Yrqbc, with distances of 6.7 and 10.1 cM, respectively.
Fine mapping of Yrqbc
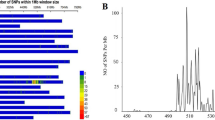
The target gene (Yrqbc) was mapped to the end of the 2BL chromosome by analysing the BSR-Seq data. The associated SNPs are mainly concentrated in the 633–653 Mb, 673–693 Mb and 723–763 Mb regions of 2BL (the total length of 2B is 801 Mb) (Supplementary Fig. 3). A total of 92 candidate SNPs in this region were polymorphic between the bulks. Six SNPs could be used to construct genetic linkage maps after genotyping.
A wheat 660 K array was used to map the resistance gene. Based on the above results, the genetic distance of the two flanking SSR markers (Xwmc175 and Xwmc332) to Yrqbc refer to 670–730 Mb on 2BL. In addition, 1714 SNPs in this interval were detected. With respect to the wheat genome version 1.0 (IWGSC) of Chinese Spring, the 1716 SNP flanking sequences (a total of 200–100 bp in the front and 100 bp in the back) were queried via local BLAST searches. A total of 457 specific SNPs and the flanking sequences were unique in the Chinese Spring reference genome. As a result, 14 SNPs were found to be linked to Yrqbc by genotyping.
Compared to Chinese Spring reference genome, the resistance gene Yrqbc was mapped to a 5.1 cM genetic interval between molecular markers A009200 and A009192 by screening the segregating population of the polymorphic markers, and the genetic distance was 0.1 cM and 5.0 cM, respectively (Fig. 2; Supplementary Table 4).
Yr5 identification
The physical location of Yrqbc indicates that it may be allelic to the previously reported Yr5 in the same region. However, the associated markers of Yr5 suggested that Yrqbc and Yr5 may not be the same gene according to a previous report. For example, Xbarc349, Yr5STS7/8, S19M93-140, S23M41-310, Xwgp19, Xwgp26 and other markers were not polymorphic in the F2 segregating population of QBC × CS (Murphy et al. 2009; Yan et al. 2003). Compared to markers in previous linkage maps, the only molecular marker with polymorphism was Xwmc175 (Supplementary Fig. 1) (Xu et al. 2013). Yr5 was cloned and reported when we sought to further enrich the linkage map. According to the specific Yr5 marker (Clemence et al. 2018), the results indicated that Yrqbc may be identical to Yr5 (Supplementary Fig. 2). Furthermore, no difference was detected between the two sequences of Yrqbc and Yr5 by sequencing using the improved primers.
Yr5 and Yr18 utilization in cultivars
Afterward, 323 wheat cultivars (lines) were collected, including 42 CIMMYT cultivars (lines), 6 European cultivars (lines), 4 Australian cultivars (lines), and 271 cultivars (lines) of major wheat regions in China. The Yr18 and Yr5 diagnostic markers were used to detect whether these cultivars (lines) contained Yr18 or Yr5. The results showed that 27 cultivars (lines) carry the Yr18 gene and that only one cultivar (line), Zhongkemai 90, has the gene Yr5. According to the design principle of the marker Yr5-insertion, we found that 85 cultivars (lines) contain the Yr5 allele, which has a 774 bp deletion compared to Yr5 (Supplementary Fig. 3). Due to this 774 bp deletion, these cultivars did not have stripe rust resistance. None of the 323 cultivars (lines) had the naturally pyramided of the two genes.
Utilization analysis of QBC
12 Chinese high-yield elite cultivars were screened, which contains neither the Yr5 nor Yr18 gene (Supplementary Table 6). Backcross breeding and marker-aided selection (MAS) methods were used for introducing resistance gene into elite cultivars (lines). The results showed that, due to the complexity of the spontaneous structure of the stripe rust fungus and the genetic background of the parents, the resistance of the single stripe rust resistance gene (Yr5 or Yr18) does not reach the immune effect (IT = 0; or 1). Only when the derived lines have two genes (Yr5 and Yr18) to achieve the immune effect (IT = 0) (Fig. 3). The other agronomic traits of QBC were very close to these 12 elite cultivars, except for the plant height (Supplementary Table 7). This will greatly shorten the breeding cycle as only disease resistance and dwarf individuals are needed to screen in breeding program. Therefore, after the fourth generation, most of the derived lines have stabilized their disease resistance traits and other agronomic traits and no separation will occur (Fig. 4).
The field investigation results of of QBC × elite cultivars. There were no urediospore of stripe rust on the leaves of Yr5 + Yr18 + plants, and its IT was recorded “0”. The urediospore occurs in individual plants with only one gene; their ITs were recorded “1” or “2”. The IT values of neither Yr5 nor Yr18 plants were recorded as 8 or 9
Discussion
QBC, a Tibetan wheat landrace, has been cultivated in Tibet and Sichuan for many years and displays near-immune resistance to wheat stripe rust. According to the epidemic of stripe rust in China, Sichuan is the inoculum base, the centre of diversity and a major over-summering area of this pathogen (Zeng and Luo 2006). Moreover, CYR34, the first major epidemic race, has been detected in this region, and few wheat cultivars show resistance to this species (Liu et al. 2010). As a result, it is urgent to transfer the genes in QBC to elite cultivars to improve wheat resistance to rust in this region.
To utilize the resistance genes, mapping populations were constructed. It was previously shown that there are two independent and dominant stripe rust resistance genes in QBC. In addition, the leaves showed a phenotype of apical necrosis in the adult stage, which suggested it possibly carried a durable adult resistance gene. This gene was ultimately identified as Yr18 by functional marker analysis. As a result, the other ASR gene needed to be identified. A combination of several biological strategies was applied to map this gene. BSA-SSR, BSR-seq, a wheat 660 K SNP array and KASP technology were used for fine mapping. The results indicated that this gene is located in the 5.1 cM genetic interval between molecular markers A009200 and A009192. In this region, several resistance genes have been reported, such as Yr5, Yr7 and YrSP. Specifically, Yr7 and YrSP provide susceptibility to epidemic races in Sichuan, especially CYR34. Furthermore, functional marker analysis and sequencing were applied, and the results indicated that Yrqbc was identical to Yr5. As a result, the two independent and dominant genes in QBC were Yr5 and Yr18.
In recent years, pyramiding of high-level and durable resistance to stripe rust has been considered an effective method for controlling this disease. Cultivation of durable, resistant varieties has become the mainstream of international disease resistance breeding (Chen 2013; Hou et al. 2015). Plants with durable resistance genes are usually susceptible to disease at the seedling stage and are moderately susceptible to stripe rust at the adult stage. Yield will also decrease when the disease is severe. Furthermore, spring wheat is very susceptible to stripe rust from the seedling stage to the milky stage, as the temperature and humidity are suitable for stripe rust growth. As a result, the ASR gene needs to be implemented. ASR genes can prevent rust infection at the early growth stage and can reduce rust inocula in epidemic regions. The presence of this gene will also slow the occurrence of new virulence combinations by genetic recombination. In general, the ideal condition of stripe rust resistance is the simultaneous presence of ASR such as Yr5, Yr15, Yr26 and HTAP/slowing rust resistance genes such as Yr18, Yr29, Yr36, Yr39 (Chen 2013).
Crop breeding generally starts from landraces which should be the basis for the improvement of varieties in various regions. They are native, long lasting and best adapted to the local natural environment and production conditions as well as the corresponding production potential. The phenotypic identification results showed that QBC provides immune resistance to all epidemic Chinese Pst races identified in the field and greenhouse, which indicated that QBC is an excellent landrace for wheat breeding in the winter wheat region of southwest China. This phenomenon may be explained by the combination of the ASR and durable adult resistance genes. In QBC, the ASR gene is Yr5, and the durable adult resistance gene is Yr18. Yr5 showed a high level of resistance, and Yr18 provides partial and durable resistance against the devastating fungal pathogens leaf rust, stripe rust, and powdery mildew. As a result, this excellent resistance material can be used as a parent to improve resistance to stripe rust. In China, resistance to stripe rust has improved in recent years. However, the disease-resistant types are relatively individual. Moreover, few cultivars show resistance to CYR34 (Liu et al. 2010). Therefore, it is necessary to further broaden the source of resistant materials and improve the diversity of disease resistance genes. Moreover, the aggregation breeding of disease resistance genes throughout the whole growth period and in the adult stage is important.
Breeders and researchers have been searching for durable and high-level resistance for many years (Johnson 1981). It is believed that a “pyramiding strategy” is an effective and easy way to achieve this goal (Mallick et al. 2015; Yanmin et al. 2019). However, this strategy is not easily practicable. Many studies have reported that pyramiding genes are easily lost during hybridization (Liu et al. 2018). As a result, utilizing cultivars that already contain pyramided genes is practical. QBC, a landrace from Tibet, has been planted for decades in Sichuan. It shows durable and high-level resistance to stripe rust not only in the adult stage but also in the seedling stage. The two resistance genes, Yr5 and Yr18, are naturally pyramided. These pyramided two types of resistant genes (ASR and durable adult resistance) show enhanced resistance to stripe rust according to the results of the resistance of a single gene. In turn, the pyramiding of resistance genes was able to achieve durable and high-level resistance.
Elite cultivars quickly lost their resistance in China due to the emergence of new stripe rust races. Yr9, Yr10, Yr17, Yr24 and other previously widely used resistance genes in production nearly lost their resistance. Even Yr5 has been identified as a moderate resistance gene in some rust disease epidemic years. As a result, the application of additional resistance gene resources in China is urgently needed. However, few practical applications of pyramided resistance genes, such as Yr5 and Yr18, have been successful in breeding so far (Singh et al. 2008; Zeng et al. 2014; Maccaferri et al. 2015; Kankwatsa et al. 2017). In 2014, 134 Sichuan wheat varieties (lines) were evaluated, and only a few genes, such as Yr5, Yr10, Yr15 and Yr50, maintained resistance to rust in field identifications (Ren et al. 2014). Moreover, only 17.9% of the varieties (lines) contained adult-plant resistance genes. The occurrence of Yr5 and Yr18 was very low among 672 wheat accessions, and no pyramided Yr5 and Yr18 genes were detected (Zheng et al. 2017). Furthermore, the combination of Yr5 and Yr18 is seldom used in the main cultivars in major wheat-growing provinces in China (Jin et al. 2018; Miaomiao et al. 2018; Dong et al. 2012; Zhou et al. 2017; Wang et al. 2018). Our results of 323 cultivars (lines) marker analysis also demonstrate that the Yr5 and Yr18 resistance genes are used relatively infrequently in wheat cultivars (lines) worldwide, and no cultivars (lines) have both genes at same time. If the outbreak of stripe rust occurs at this time, it will cause immeasurable losses to production. Therefore, implementation of a “pyramiding strategy” should be accelerated. The application of QBC may accelerate the deployment of Yr5 and Yr18 and could improve the disadvantage of monoculture deployment of resistance genes. Many research teams have initiated research on the pyramiding of resistance genes on chromosomes 2B and 7B.
In recent years, the large-scale use of wheat backbone parents has increased the homogenization of varieties and significantly reduced gene polymorphism, which has become one of the important bottlenecks restricting wheat breeding all over the world. QBC, a landrace from Tibet, contains the naturally pyramided Yr5 and Yr18 genes. Yr5 serves as an ASR gene and Yr18 serves as a adult-plant gene, which together provide high-level and durable resistance to all races. It is effective and practical to improve the resistance of local elite varieties by introducing pyramided genes. Therefore, this Tibetan landrace could be used as an excellent germplasm to develop high-level and durable resistant varieties in breeding program.
References
Bariana HS, Brown GN, Bansal UK, Miah H, Standen GE, Lu M (2007) Breeding triple rust resistant wheat cultivars for Australia using conventional and marker-assisted selection technologies. Aust J Agric Res 58:576–587. https://doi.org/10.1071/AR07124
Chen XM (2005) Epidemiology and control of stripe rust [Puccinia striiformis f. sp. tritici] on wheat. Can J Plant Path 27:314–337. https://doi.org/10.1080/07060660509507230
Chen X (2013) Review article: high-temperature adult-plant resistance, key for sustainable control of stripe rust. Am J Plant Sci 04:608–627
Chen XM, Tristan C, Xueling H, Meinan W, Andrea D (2013) Understanding molecular mechanisms of durable and non-durable resistance to stripe rust in wheat using a transcriptomics approach. Curr Genom 14:111–126. https://doi.org/10.2174/1389202911314020004
Cheng Z, Wang P, Xu Y (2016) Bulked sample analysis in genetics, genomics and crop improvement. Plant Biotechnol J 14:1941–1955. https://doi.org/10.1111/pbi.12559
Clemence M et al (2018) BED-domain-containing immune receptors confer diverse resistance spectra to yellow rust. Nat Plants 4:662–668
Cox TS, Raupp WJ, Gill BS (1994) Leaf rust-resistance genes Lr41, Lr42, and Lr43 transferred from Triticum tauschii to common wheat. Crop Sci 34:339–343. https://doi.org/10.2135/cropsci1994.0011183X003400020005x
Cui F et al (2017) Utilization of a Wheat660K SNP array-derived high-density genetic map for high-resolution mapping of a major QTL for kernel number. Sci Rep 7:3788
Dong SJ, Xu WG, Hu L et al (2012) Molecular detection of stripe rust resistant genes in 43 wheat varieties planted in Henan Province. Acta Agric Boreal Sin 27:157–162. https://doi.org/10.3969/j.issn.1000-7091.2012.05.030
Elisabeth LM, DePope V-PC, Van der Wanf W (2017) Achieving durable resistance against plant diseases: scenario analyses with a national-ccale spatially explicit model for a wind-dispersed. Plant Pathog Phytopathol 107:580–589. https://doi.org/10.1094/PHYTO-05-16-0207-R
Ellis JG, Lagudah ES, Spielmeyer W, Dodds PN (2014) The past, present and future of breeding rust resistant wheat. Front Plant Sci 5:641. https://doi.org/10.3389/fpls.2014.00641
Guo Q, Zhang ZJ, Xu YB, Li GH, Feng J, Zhou Y (2008) Quantitative trait loci for high-temperature adult-plant and slow-rusting resistance to Puccinia striiformis f. sp. tritici in wheat cultivars. Phytopathology 98:803–809. https://doi.org/10.1094/PHYTO-98-7-0803
Gupta P et al (2002) Genetic mapping of 66 new microsatellite (SSR) loci in bread wheat. Theor Appl Genet 105:413–422. https://doi.org/10.1007/s00122-002-0865-9
Hou L, Chen X, Wang M, See DR, Chao S, Bulli P, Jing J (2015) Mapping a large number of QTL for durable resistance to stripe rust in winter wheat druchamp using SSR and SNP markers. PLoS ONE 10:e0126794. https://doi.org/10.1371/journal.pone.0126794
Jin H, Jia Q, Bo Z, Sun Z, Huang M, Jin S (2018) Epidemic forecasting of the new strains G22-9(CYR34) and G22-14 of Puccinia striiformis f. sp. tritici in wheat in Gansu Province. J Plant Prot
Johnson R (1981) Durable resistance: definition of, genetic control, and attainment in plant breeding. Phytopathology 71:567–568
Kankwatsa P, Singh D, Thomson PC, Babiker EM, Bonman JM, Newcomb M, Park RF (2017) Characterization and genome-wide association mapping of resistance to leaf rust, stem rust and stripe rust in a geographically diverse collection of spring wheat landraces. Mol Breed 37:113. https://doi.org/10.1007/s11032-017-0707-8
Lagudah ES et al (2009) Gene-specific markers for the wheat gene Lr34/Yr18/Pm38 which confers resistance to multiple fungal pathogens. Theor Appl Genet 119:889–898
Line RF, Qayoum A (1992) Virulence, aggressiveness, evolution and distribution of races of Puccinia striiformis(the cause of stripe rust of wheat) in North America, 1968–8. US Dept Agric Tech Bull 1788:44
Liu TG, Peng YL, Chen WQ, Zhang ZY (2010) First detection of virulence in Puccinia striiformis f. sp. tritici in China to resistance genes Yr24(=Yr26) present in wheat cultivar Chuanmai 42. Plant Dis 94:1163–1163. https://doi.org/10.1094/PDIS-94-9-1163C
Liu L, Wang MN, Feng JY, See DR, Chao SM, Chen XM (2018) Combination of all-stage and high-temperature adult-plant resistance QTL confers high-level, durable resistance to stripe rust in winter wheat cultivar Madsen. Theor Appl Genet 131:1835–1849. https://doi.org/10.1007/s00122-018-3116-4
Maccaferri M et al (2015) A genome-wide association study of resistance to stripe rust (Puccinia striiformis f. sp. tritici) in a worldwide collection of hexaploid spring wheat (Triticum aestivum L.). G3 5:449–465. https://doi.org/10.1534/g3.114.014563
Mallick N, Vinod SJB, Tomar RS, Sivasamy M, Prabhu KV (2015) Marker-assisted backcross breeding to combine multiple rust resistance in wheat. Plant Breed 134:172–177. https://doi.org/10.1111/pbr.12242
Miaomiao H, Zhenyu S, Shiqin C, Qiuzhen J, Taiguo L, Wanquan C (2018) Evaluation of the resistance of 223 wheat landraces in Gansu Province to stripe rust and molecular detection. J Plant Prot
Murphy LR, Santra D, Kidwell K, Yan GP, Chen XM, Campbell KG (2009) Linkage maps of wheat stripe rust resistance genes Yr5 and Yr15 for use in marker-assisted selection. Crop Sci 49:1786–1790. https://doi.org/10.2135/cropsci2008.10.0621
Nobuko Y, Takayuki M, Keiko H, Shinzo K, Yoshikatsu F (2015) Effects of pyramiding quantitative resistance genes Pi21, Pi34, and Pi35 on rice leaf blast disease: an international journal of applied plant pathology. Plant Dis. https://doi.org/10.1094/PDIS-02-14-0214-RE
Qi LL et al (2004) A chromosome bin map of 16,000 expressed sequence tag loci and distribution of genes among the three genomes of polyploid wheat. Genetics 168:701–712. https://doi.org/10.1534/genetics.104.034868
Röder MS, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149:2007
Ramirez-Gonzalez RH et al (2015) RNA-Seq bulked segregant analysis enables the identification of high-resolution genetic markers for breeding in hexaploid wheat. Plant Biotechnol J 13:613–624. https://doi.org/10.1111/pbi.12281
Ren Y, Li SR, Zhou Q, Du XY et al (2014) Evaluation of resistance to stripe rust of 134 wheat cultivars and lines from Sichuan Province. J Triticeae Crops 34:847–853
Rogers SO, Bendich AJ (1985) Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol Biol 5:69–76
Rosewarne GM et al (2006) Leaf tip necrosis, molecular markers and β1-proteasome subunits associated with the slow rusting resistance genes Lr46/Yr29. Theor Appl Genet 112:500–508. https://doi.org/10.1007/s00122-005-0153-6
Schneider A, Walker S, Sagan M, Duc G, Ellis T, Downie J (2002) Mapping of the nodulation loci sym9 and sym10 of pea (Pisum sativum L.). Theor Appl Genet 104:1312–1316
Shi A, Chen P, Li D, Zheng C, Bo Z, Hou A (2009) Pyramiding multiple genes for resistance to soybean mosaic virus in soybean using molecular markers. Mol Breed 23:113
Singh D, Park RF, Mcintosh RA, Bariana HS (2008) Characterisation of stem rust and stripe rust seedling resistance genes in selected wheat cultivars from the United Kingdom. J Plant Pathol 90:553–562
Singh RP, Huerta-Espino J, William HM (2005) Genetics and breeding for durable resistance to leaf and stripe rusts in wheat Turkish. J Agric For 29:121–127
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum asetivum L.). Theor Appl Genet 109:1105–1114. https://doi.org/10.1007/s00122-004-1740-7
Sourdille P et al (2004) Microsatellite-based deletion bin system for the establishment of genetic-physical map relationships in wheat (Triticum aestivum L.). Funct Integr Genom 4:12–25
Wan AM, Chen XM, He ZH (2007) Wheat stripe rust in China. Aust J Agric Res 58:605–619. https://doi.org/10.1071/AR06142
Wang SH, Gong KY, Chu BY, Sun QY, Luo Y, Zhan-Hong MA (2018) Molecular detection of stripe rust resistance gene(s) in 100 wheat cultivars(lines) from Sichuan Province in China. Acta Phytopathol Sin 164:946–958
Werner K, Friedt W, Ordon F (2005) Strategies for pyramiding resistance genes against the barley yellow mosaic virus complex (BaMMV, BaYMV, BaYMV-2). Mol Breed 16:45–55
Xu LS, Wang MN, Cheng P, Kang ZS, Hulbert SH, Chen XM (2013) Molecular mapping of Yr53, a new gene for stripe rust resistance in durum wheat accession PI 480148 and its transfer to common wheat. Theor Appl Genet 126:523–533
Yan GP, Chen XM, Line RF, Wellings CR (2003) Resistance gene-analog polymorphism markers co-segregating with the Yr5 gene for resistance to wheat stripe rust. Theor Appl Genet 106:636–643
Yan L, Meinan W, Xianming C, Deven S, Shiaoman C, Jinxue J (2014) Mapping of Yr62 and a small-effect QTL for high-temperature adult-plant resistance to stripe rust in spring wheat PI 192252. Theor Appl Genet 127:1449–1459. https://doi.org/10.1007/s00122-014-2312-0
Yanmin Q, Yan L, Meinan W, Xing L, See DR, Diaoguo A, Xianming C (2019) Development, validation, and re-selection of wheat lines with pyramided genes Yr64 and Yr15 linked on the short arm of chromosome 1B for resistance to stripe rust. Plant Dis 103:51–58. https://doi.org/10.1094/PDIS-03-18-0470-RE
Zeng SM, Luo Y (2006) Long-distance spread and interregional epidemics of wheat stripe rust in China. Plant Dis 90:980–988. https://doi.org/10.1094/PD-90-0980
Zeng QD et al (2014) Stripe rust resistance and genes in Chinese wheat cultivars and breeding lines. Euphytica 196:271–284. https://doi.org/10.1007/s10681-013-1030-z
Zhang ZJ, Yang GH, Li GH, Jin SL, Yang XB (2001) Transgressive segregation, heritability, and number of genes controlling durable resistance to stripe rust in one Chinese and two Italian wheat cultivars. Phytopathology 91:680–686. https://doi.org/10.1094/PHYTO.2001.91.7.680
Zheng S et al (2017) Evaluating the contribution of Yr genes to stripe rust resistance breeding through marker-assisted detection in wheat. Euphytica 213:50. https://doi.org/10.1007/s10681-016-1828-6
Zhou C, Zhibin XU, Feng B, Xiang C, Wang T (2015) Genetic analysis of stripe rust resistance in Tibetan wheat landrace Qubaichun. J Triticeae Crops. https://doi.org/10.1371/journal.pone.0203283
Zhou X et al (2017) Molecular detection of stripe rust resistant genes Yr5 and Yr10 in 38 Wheat Lines Gansu. Agric Sci Technol. https://doi.org/10.1111/jph.12515
Acknowledgements
We thank Tibet Academy of Agriculture and Animal Husbandry Sciences for providing Qubaichun seeds. The KASP assay was conducted by China Golden Marker (Beijing) Biotech Co., Ltd. Thanks to Professor Zhiyong Liu from Institute of genetics, CAS for BSR-seq data analysis.
Funding
This work was supported by grants from the National Natural Science Foundation of China (Grant No. 31671679) and Youth Innovation Promotion Association, CAS (Grant No. 2018403).
Author information
Authors and Affiliations
Contributions
T-W coordinated the project, conceived and designed experiments. B-F conducted the bioinformatics work, generated and analyzed data, and edits the manuscript. Z-B X and F-X L collected the samples; F-W, G-S J and Q-Z performed the laboratory work. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10722_2020_938_MOESM1_ESM.pptx
Supplementary file1 (PPTX 506 kb) Supplementary Figure 1 The integrated genetic linkage map of stripe rust resistance gene and its co-segregated markers on 2BL. Supplementary Figure 2 The ASR gene in QBC was identified by Yr5 functional marker. The amplified fragment size of PCR was 1281 bp. Material 1-7 was a randomly selected F2 generation of QBC × CM28 with IT of "0", Material 8 was a cultivar containing Yr5 gene, Material 9 was QBC, Material 10 was CS. The results showed that the ASR gene in QBC was Yr5. Supplementary Figure 3 The analysis results of BSR-seq, SNPs are mainly concentrated in the 633~653 Mb, 673~693 Mb and 723~763 Mb regions of 2BL
Rights and permissions
About this article
Cite this article
Cao, J., Xu, Z., Fan, X. et al. Genetic mapping and utilization analysis of stripe rust resistance genes in a Tibetan wheat (Triticum aestivum L.) landrace Qubaichun. Genet Resour Crop Evol 67, 1765–1775 (2020). https://doi.org/10.1007/s10722-020-00938-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-020-00938-z