Abstract
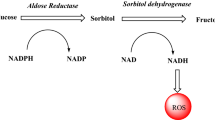
In the polyol pathway, aldose reductase (AR) catalyzes the formation of sorbitol from glucose. In order to detoxify some dangerous aldehydes, AR is essential. However, due to the effects of the active polyol pathway, AR overexpression in the hyperglycemic state leads to microvascular and macrovascular diabetic problems. As a result, AR inhibition has been recognized as a potential treatment for issues linked to diabetes and has been studied by numerous researchers worldwide. In the present study, a series of acyl hydrazones were obtained from the reaction of vanillin derivatized with acyl groups and phenolic Mannich bases with hydrazides containing pharmacological groups such as morpholine, piperazine, and tetrahydroisoquinoline. The resulting 21 novel acyl hydrazone compounds were investigated as an inhibitor of the AR enzyme. All the novel acyl hydrazones derived from vanillin demonstrated activity in nanomolar levels as AR inhibitors with IC50 and KI values in the range of 94.21 ± 2.33 to 430.00 ± 2.33 nM and 49.22 ± 3.64 to 897.20 ± 43.63 nM, respectively. Compounds 11c and 10b against AR enzyme activity were identified as highly potent inhibitors and showed 17.38 and 10.78-fold more effectiveness than standard drug epalrestat. The synthesized molecules’ absorption, distribution, metabolism, and excretion (ADME) effects were also assessed. The probable-binding mechanisms of these inhibitors against AR were investigated using molecular-docking simulations.
Graphical abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The acyl hydrazone skeleton is an important intermediate used in the synthesis of interesting biologically active heterocyclic compounds and is also used as a ligand in the formation of metal complexes [1, 2]. Hydrazone-type compounds containing azomethine protons constitute a significant class of compounds for novel drug research [3], such as anti-antimicrobial [4], inflammatory [5], antioxidants [6], antitubercular [7], analgesic [8], anti-candida [9], α-glucosidase [10], anticancer [11], and antiproliferative [12] activities in medicinal fields [13,14,15]. Acyl hydrazone fragments bound to heterocyclic systems were displayed to provide enhanced activity. This bioactivity of acyl hydrazones is explained by their tendency to form a hydrogen bond with the molecular target [16,17,18]. In addition, it has been reported by studies in the literature that acyl hydrazones have lower toxicity than hydrazides due to the blockage effects of NH2 groups. These research findings further increased the importance of synthesizing acyl hydrazone-derived compounds [19].
A three-component Mannich condensation builds a class of compounds known as Mannich bases on one pot reaction of primary or secondary amine, an aldehyde reagent, and structurally diverse substrates containing at least one active hydrogen atom [20]. Mannich base also acts as essential pharmacophores for synthesizing novel compounds in medicinal chemistry [21]. Examples of clinically useful Mannich bases containing aminoalkyl scaffold in their structure are drugs such as trihexyphenidyl, procyclidine, ranitidine, and biperiden [22, 23]. Phenolic compounds are one of the most critical carbons Mannich bases because they contain active hydrogen. In the last decades, there have been many studies on the biological activities of phenolic Mannich bases, such as cytotoxic [24, 25], anticancer [26, 27], anticonvulsant [28], anti-inflammatory [29], antifungal [30], and carbonic anhydrase inhibitory activities [31, 32].
Morpholine and piperazines are privileged backbones that are widely used as a core structural element or substituent in effective drugs such as Noroxin (antibiotic), Clozaril (an atypical antipsychotic medication), Iressa (breast, lung, and other cancers), and Moclobemide (depression and social anxiety) [33, 34]. They can also improve the pharmacokinetic features of molecules, such as metabolic stability and solubility in water [35]. Another core structure, tetrahydroisoquinoline (THQ), forms the main backbone of many natural products (saframycin, naphthyridinomycin/bioxalomycin, and quinocarcin/tetrazomine) and bioactive compounds [36]. From this perspective, with the potential of the drugs, it would be helpful to design and synthesize some novel acyl hydrazone derivatives incorporating piperazine, morpholine, and tetrahydroquinoline and screen them for potential biological activities.
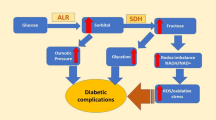
Aldose reductase (AKR1B1, AR with EC number 1.1.1.2)1 is the first rate-limiting enzyme in the polyol pathway, belongs to the Aldo–keto reductase superfamily, and is a monomer comprising 315 amino acid residues [37,38,39,40,41]. This overproduction of the AR and sorbitol dehydrogenase on the polyol pathway and depletion in reduced NADP+ and the oxidized NAD+, which are cofactors of this process, causes various metabolic processes disturbances such as the nephropathy, retinopathy, cataracts, and neuropathy [42,43,44,45,46]. The aforementioned metabolic abnormalities are the primary targets of diabetic complications in those tissues involved in insulin-independent glucose uptake and are responsible for early tissue damage in the organs [47,48,49,50,51,52,53].
We investigated the synthesis, characterization, and biological activity of a series of novel acyl hydrazones to uncover novel AR inhibitors in the current work. In addition, we conducted in silico experiments, including absorption, distribution, metabolism, and excretion (ADME), density functional theory (DFT), and molecular docking, to evaluate the inhibitory mechanisms of those compounds against the target mentioned above, AR.
Experimental
Chemistry
The chemicals used in this study were supplied from Sigma Aldrich (Germany). Melting points were determined on WRS-2A Microprocessor Melting-point Apparatus and are uncorrected. IR spectra of compounds were recorded using ALPHA-P BRUKER FT-IR Spectrophotometer.1H NMR spectra were recorded on Bruker (400 MHz) spectrometer. 13C NMR spectra were recorded on Bruker (100 MHz) spectrometer. Chemical shifts are reported as δ in ppm relative to tetramethylsilane (TMS) (δ 0.00 singlets) in deuterated chloroform (CDCl3). High-resolution mass spectrometry measurements were recorded on Agilent 6530 Accurate-Mass Q-TOF LC/MS.
General procedure for synthesis of compounds 2a–b
Synthesis of 2a–b was performed according to the previously reported method [54].
4-Formyl-2-methoxyphenyl furan-2-carboxylate (2a)
White solid, yield 83%, mp: 101–103 °C (lit. 101–103 °C) [54].
4-Formyl-2-methoxyphenyl thiophene-2-carboxylate (2b)
White solid, yield: 81%, mp: 89–91 °C. IR (ATR, cm−1): νmax 3108, 3070, 2843, 1744, 1682, 1456, 1262, 1054, 854, 724. 1H NMR (400 MHz, CDCl3, δ/ppm): δ 10.0 (s, 1H), 8.0 (d, J = 3.8 Hz, 1H), 7.7 (d, J = 5.0 Hz, 1H), 7.5 (dt, J = 7.9, 1.6 Hz, 2H), 7.4 (d, J = 7.9 Hz, 1H), 7.2 (d, J = 4.9 Hz, 1H), 3.9 (s, 3H). 13C NMR (100 MHz, CDCl3, δ/ppm): δ 191.0 159.5, 152.2, 144.7, 135.7, 135.2, 133.9, 132.0, 128.2, 124.7, 123.6, 111.0, 56.2. HRMS (Q-TOF) m/z calcd for C15H14O5S [M + H]+: 262.0323, Found: 262.0341.
General procedure for synthesis of compounds 3a–e
Synthesis of 3a–e was carried out according to the previously reported method [55].
4-Hydroxy-3-methoxy-5-(morpholinomethyl)benzaldehyde (3a)
White solid; yield: 85%, mp: 99–100 °C; IR (ATR, cm−1) νmax 2945, 2866, 2829, 2733, 1647, 1592, 1270, 1120, 868, 705; 1H NMR (400 MHz, CDCl3) δ 9.79 (s, 1H), 7.36 (m, 1H), 7.19 (m, 1H) 3.95 (s, 3H), 3.82 (s, 2H), 3.78 (m, 4H), 2.63 (m, 4H); 13C NMR (100 MHz, CDCl3)δ 190.6, 153.5, 148.6, 128.5, 125.5, 120.3, 109.8, 66.6, 61.1, 56.0, 52.7.
4-Hydroxy-3-methoxy-5-((4-phenylpiperazin-1-yl)methyl)benzaldehyde (3b)
White solid; yield: 90%, mp: 156 °C (lit: 156–157 °C) [56]; IR (ATR, cm−1) νmax 2959, 2938, 2827, 2737, 1677, 1586, 1315, 1235, 1141, 760, 691; 1H NMR (400 MHz, CDCl3) δ10.90 (brs, 1H), 9.81 (s, 1H), 7.39–7.38 (m, 1H), 7.32–7.27 (m, 2H), 7.22–7.17 (m, 1H) 6.95–6.86 (m, 3H), 3.96 (s, 3H), 3.89 (s, 2H), 3.28 (m, 4H), 2.80 (m,4H); 13C NMR (100 MHz, CDCl3)δ 190.6, 153.7, 150.7, 148.7, 129.2, 128.4, 125.5, 120.6, 120.4, 116.5, 109.9, 60.8, 56.0, 52.5, 49.2.
4-Hydroxy-3-methoxy-5-((3-methylpiperidin-1-yl)methyl)benzaldehyde (3c)
Light brown solid; yield: 88%, mp: 142–144 °C; IR (ATR, cm−1) νmax 2946, 2922, 2853, 2748, 1651, 1592, 1271, 1147, 864, 707; 1H NMR (400 MHz, CDCl3) δ11.55 (s, 1H), 9.76 (s, 1H), 7.33 (m, 1H), 7.15 (m, 1H) 3.94 (s, 3H), 3.78 (m, 2H), 2.96–2.90 (m, 2H), 2.11 (t, J = 10.5 Hz, 1H), 1.83–1.58 (m, 5H), 0.98–0.95 (m, 1H), 0.89 (d, J = 6.3 Hz, 3H),13C NMR (100 MHz, CDCl3)δ 190.6, 154.8, 148.6, 127.8, 125.4, 120.8, 109.5, 61.2, 60.7, 55.9, 53.2, 32.2, 31.0, 25.0, 19.2.
3-((3,4-Dihydroisoquinolin-2(1H)-yl)methyl)-4-hydroxy-5-methoxybenzaldehyde (3d)
Light yellow solid; yield: 88%, mp: 181–183 °C; IR (ATR, cm−1) νmax 3053, 2956, 2817, 2750, 1649, 1590, 1274, 749; 1H NMR (400 MHz, DMSO) δ 9.79 (s, 1H), 7.45 (s, 1H), 7.37 (s, 1H), 7.24–6.89 (m, 4H), 3.89 (s, 2H), 3.86 (s, 3H), 3.69 (s, 2H), 2.86 (t, J = 5.4 Hz, 2H), 2.80 (t, J = 5.4 Hz, 2H); 13C NMR (101 MHz, DMSO) δ 191.0, 152.7, 148.0, 133.8, 133.5, 128.5, 127.8, 126.5, 126.3, 125.7 (2C), 122.9, 109.8, 57.5, 55.7, 54.7, 49.7, 28.2.
N-ethyl-4-(5-formyl-2-hydroxy-3-methoxybenzyl)piperazine-1-carboxamide (3e)
White solid, yield 80%, mp: 86–87 °C; IR (ATR, cm−1) νmax 2978, 2948, 2823, 2735, 1735, 1681, 1589, 1237, 1138, 863, 694; 1H NMR (400 MHz, CDCl3) δ 9.79 (s, 1H), 7.36 (m, 1H), 7.18 (m, 1H), 4.15 (q, J = 7.1 Hz, 2H), 3.95 (s, 3H), 3.83 (m, 1H), 3.56 (m, 4H), 2.58 (m, 4H), 1.27 (t, J = 7.1 Hz, 3H); 13C NMR (100 MHz, CDCl3) δ 190.6, 155.2, 153.4, 148.6, 128.5, 125.4, 120.4, 109.9, 61.6, 60.8, 56.0, 52.2, 43.4, 43.3, 14.6.
General procedure for synthesis of compounds 6a–c
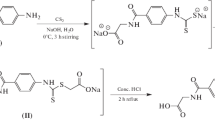
In a 50 mL round-bottom flask, the related secondary amine (10 mmol) and triethylamine (11 mmol, 1.11 g, 1.53 mL) were dissolved in 20 mL of THF and the solution was then put into an ice bath. Ethylchloroacetate (10 mmol, 1.23 g) in 20 mL of THF was added to this solution dropwise and stirred for three hours at room temperature. After completion, the solvent was removed under reduced pressure. It was washed with cold water to remove the triethylammonium chloride salt from the oily mixture obtained. Then the crude product was dissolved in 20 mL of ethanol and hydrazinium hydroxide (80%, 25 mmol) was added to the solution. The reaction mixture was refluxed for two hours. The solvent was removed under reduced pressure and the product was washed with cold water. The crude product was used for next step without any purification.
2-(4-Phenylpiperazin-1-yl)acetohydrazide (6a)
White solid, yield 88%, mp: 76–78 °C (lit. 75 °C) [57].
2-(3,4-Dihydroisoquinolin-2(1H)-yl)acetohydrazide (6b)
White solid, yield 92%, mp: 85–87 °C [58]. 1H NMR (400 MHz, DMSO) δ 8.95 (s, 1H), 7.22–6.87 (m, 4H), 4.25 (brs, 2H), 3.61 (s, 2H), 3.10 (s, 2H), 2.82 (t, J = 5.8 Hz, 2H), 2.71 (t, J = 5.9 Hz, 2H). 13C NMR (101 MHz, DMSO) δ 168.3, 134.6, 133.9, 128.4, 126.2, 125.9, 125.4, 59.7, 55.2, 50.6, 28.6.
2-Morpholinoacetohydrazide (6c)
White solid, yield 75%, mp: 95–98 °C (lit. 99–101 °C) [59].
General procedure for synthesis of compounds 7a–c, 11a–c, 12a–c, and 13a–c
The corresponding aldehyde 2a–b, 3a–e (10 mmol) and acetohydrazide derivative 6a–c (10 mmol) were dissolved in absolute ethanol (20 mL), and 4–5 drops of acetic acid was added. Reaction mixture was refluxed for 1–2 h. Reaction was monitored by TLC. After completion, half of the solvent volume was removed under reduced pressure. The mixture was left in the freezer overnight, and the formed solid was filtered off. The crude product was recrystallized from ethanol.
2-Methoxy-4-((2-(2-(4-phenylpiperazin-1-yl)acetyl)hydrazono)methyl)phenylfuran-2-carboxylate (7a)

Beige solid, yield 77%, mp: 178–180 °C, IR (ATR, cm−1) νmax 3205, 3065, 2936, 1729, 1657, 1597, 1232, 1070, 745; 1H NMR (400 MHz, CDCl3) δ 10.21 (s, 1H), 8.25 (s, 1H), 7.69 (s, 1H), 7.57 (s, 1H), 7.42 (d, J = 3.4 Hz, 1H), 7.33–7.28 (m, 3H), 7.25–7.14 (m, 2H), 7.01–6.84 (m, 2H), 6.61 (dd, J = 3.4, 1.6 Hz, 1H), 3.89 (s, 3H), 3.35–3.33 (m, J = 10.8 Hz, 6H), 2.82 (brs, 4H). 13C NMR (101 MHz, CDCl3) δ 166.0, 156.2, 151.8, 150.9, 148.0, 147.3, 143.6, 141.2, 132.7, 129.2, 123.1, 121.8, 120.2, 119.8, 116.3, 112.3, 109.9, 61.0, 56.2, 53.6, 49.3. HRMS (Q-TOF) m/z: [M + H]+ calcd for C25H26N4O5, 463.1981; found 463.1977.
4-((2-(2-(3,4-Dihydroisoquinolin-2(1H)-yl)acetyl)hydrazono)methyl)-2-methoxyphenyl furan-2-carboxylate (7b)

Beige solid, yield 81%, mp: 105–107 °C, IR (ATR, cm−1) νmax 3145, 3028, 2922, 1742, 1668, 1546, 1270, 1077, 746; 1H NMR (400 MHz, CDCl3) δ 10.30 (s, 1H), 8.13 (s, 1H), 7.68 (s, 1H), 7.56 (s, 1H), 7.40 (d, J = 3.4 Hz, 1H), 7.25–7.10 (m, 5H), 7.05 (d, J = 6.0 Hz, 1H), 6.60 (dd, J = 3.5, 1.7 Hz, 1H), 3.86 (s, 3H), 3.79 (s, 2H), 3.38 (s, 2H), 3.00 (t, J = 5.6 Hz, 2H), 2.91 (t, J = 5.7 Hz, 2H). 13C NMR (100 MHz, CDCl3) δ 166.5, 156.2, 151.7, 147.8, 147.3, 143.6, 141.1, 133.5, 128.8, 126.7, 126.6, 126.0, 123.0, 121.8, 119.8, 112.3, 109.9, 61.0, 56.2, 51.6, 29.2. HRMS (Q-TOF) m/z: [M + H]+ calcd for C24H23N3O5, 434.1716; found 434.1710.
2-Methoxy-4-((2-(2-morpholinoacetyl)hydrazono)methyl)phenyl furan-2-carboxylate (7c)

Beige solid, yield 75%, mp: 114–116 °C, IR (ATR, cm−1) νmax 3153, 3108, 2955, 1743, 1658, 1572, 1267, 1067, 745; 1H NMR (400 MHz, CDCl3) δ 10.15 (s, 1H), 8.22 (s, 1H), 7.68 (s, 1H), 7.53 (s, 1H), 7.40 (d, J = 3.3 Hz, 1H), 7.18 (m, 2H), 6.60 (d, J = 4.9 Hz, 1H), 3.86 (s, 3H), 3.77 (t, J = 4.4 Hz, 4H), 3.21 (s, 2H), 2.62 (t, J = 4.4 Hz, 4H). 13C NMR (100 MHz, CDCl3) δ 166.0, 156.2, 151.7, 148.0, 147.3, 143.5, 141.1, 132.7, 123.0, 121.7, 119.9, 112.3, 109.9, 66.8, 61.4, 56.2, 53.9. HRMS (Q-TOF) m/z: [M + H]+ calcd for C19H21N3O6, 388.1509; found 388.1503.
2-Methoxy-4-((2-(2-(4-phenylpiperazin-1-yl)acetyl)hydrazono)methyl)phenyl thiophene-2-carboxylate (8a)

White solid, yield 79%, mp: 175–177 °C, IR (ATR, cm−1) νmax 3193, 3050, 2929, 1734, 1654, 1587, 1253, 1071, 755; 1H NMR (400 MHz, CDCl3) δ 10.21 (s, 1H), 8.24 (s, 1H), 8.00 (d, J = 3.8 Hz, 1H), 7.69 (d, J = 5.0 Hz, 1H), 7.57 (s, 1H), 7.38–7.25 (m, 3H), 7.26–7.13 (m, 3H), 7.03–6.90 (m, 3H), 3.89 (s, 3H), 3.30–3.28 (m, 6H), 2.81 (t, J = 4.0 Hz, 4H). 13C NMR (100 MHz, CDCl3) δ 166.0, 159.9, 151.9, 150.9, 148.0, 141.6, 134.9, 133.7, 132.2, 129.2, 128.1, 123.1, 121.8, 120.2, 116.3, 110.0, 61.0, 56.3, 53.6, 49.3. HRMS (Q-TOF) m/z: [M + H]+ calcd for C25H26N4O4S, 479.1753; found 479.1747.
4-((2-(2-(3,4-Dihydroisoquinolin-2(1H)-yl)acetyl)hydrazono)methyl)-2-methoxyphenyl thiophene-2-carboxylate (8b)

White solid, yield 82%, mp: 110–112 °C, IR (ATR, cm−1) νmax 3172, 3073, 2915, 1734, 1661, 1561, 1251, 1089, 734; 1H NMR (400 MHz, CDCl3) δ 10.34 (s, 1H), 8.13 (s, 1H), 7.99 (d, J = 4.7 Hz, 1H), 7.67 (d, J = 5.0 Hz, 1H), 7.55 (s, 1H), 7.25–7.12 (m, 6H), 7.05 (d, J = 6.2 Hz, 1H), 3.86 (s, 3H), 3.81 (s, 2H), 3.39 (s, 2H), 3.00 (t, J = 5.5 Hz, 2H), 2.92 (t, J = 5.7 Hz, 2H). 13C NMR (100 MHz, CDCl3) δ 166.4, 159.9, 151.8, 147.9, 141.6, 134.9, 133.7, 132.3, 128.8, 128.1, 126.7, 126.6, 126.1, 123.1, 121.8, 109.9, 61.0, 56.2, 51.6, 29.1. HRMS (Q-TOF) m/z: [M + H]+ calcd for C24H23N3O4S, 450.1488; found 450.1483.
2-Methoxy-4-((2-(2-morpholinoacetyl)hydrazono)methyl)phenyl thiophene-2-carboxylate (8c)

White solid, yield 77%, mp: 140–142 °C, IR (ATR, cm−1) νmax 3177, 3053, 2956, 1735, 1655, 1572, 1247, 1082, 740; 1H NMR (400 MHz, CDCl3) δ 10.18 (s, 1H), 8.20 (s, 1H), 7.99 (d, J = 4.0 Hz, 1H), 7.68 (d, J = 4.0 Hz, 1H), 7.21–7.16 (m, 3H), 3.85 (s, 3H), 3.77 (t, J = 4.4 Hz, 4H), 3.22 (s, 2H), 2.62 (t, J = 4.4 Hz, 4H). 13C NMR (10o MHz, CDCl3) δ 166.0, 160.0, 151.8, 148.1, 141.6, 135.0, 133.7, 132.6, 132.5, 128.1, 123.1, 121.7, 110.0, 66.8, 61.4, 56.2, 53.9. HRMS (Q-TOF) m/z: [M + H]+ calcd for C19H21N3O5S, 404.1280; found 404.1274.
N′-(4-hydroxy-3-methoxy-5-(morpholinomethyl)benzylidene)-2-(4-phenylpiperazin-1-yl) acetohydrazide (9a)

White solid, yield 88%, mp: 187–189 °C, IR (ATR, cm−1) νmax 3187, 3060, 2942, 1660, 1594, 1258, 1077, 761; 1H NMR (400 MHz, CDCl3) δ 10.06 (s, 1H), 8.08 (s, 1H), 7.30–7.26 (m, 3H), 7.00 (s, 1H), 6.94–6.87 (m, 3H), 3.90 (s, 3H), 3.75–3.72 (m, 6H) 3.25 (brs, 4H), 2.76 (t, J = 4.0 Hz, 4H), 2.58 (brs, 4H). 13C NMR (100 MHz, CDCl3) δ 165.8, 150.9, 149.7, 148.8, 148.3, 129.2, 124.6, 121.8, 120.6, 116,2, 109.4, 66.7, 61.0, 56.1, 53.6, 52.8, 49.3. HRMS (Q-TOF) m/z: [M + H]+ calcd for C25H33N5O4, 468.2611; found 468.2605.
2-(3,4-Dihydroisoquinolin-2(1H)-yl)-N'-(4-hydroxy-3-methoxy-5-(morpholinomethyl)benzylidene) acetohydrazide (9b)

White solid, yield 91%, mp: 209–211 °C, IR (ATR, cm−1) νmax 3186, 3056, 2960, 1656, 1592, 1268, 1080, 742; 1H NMR (400 MHz, CDCl3) δ 10.13 (s, 1H), 7.99 (s, 1H), 7.26 (s, 1H), 7.21–7.16 (m, 3H), 7.05 (s, 1H), 6.97 (s, 1H), 3.90 (s, 3H), 3.78 (s, 2H), 3.75 (t, J = 4.0 Hz, 4H) 3.72 (s, 2H), 3.36 (s, 2H), 2.99 (t, J = 5.5 Hz, 2H), 2.90 (t, J = 5.5 Hz, 2H), 2.58 (brs, 4H). 13C NMR (100 MHz, CDCl3) δ 166.0, 149.7, 148.7, 148.3, 133.8, 133.4, 128.8, 126.7, 126.6, 126.0, 124.7, 121.8, 120.6, 109.4, 66.7, 61.0, 56.2, 52.8, 51.6, 29.2. HRMS (Q-TOF) m/z: [M + H]+ calcd for C24H30N4O4, 439.2345; found 439.2339.
N′-(4-hydroxy-3-methoxy-5-(morpholinomethyl)benzylidene)-2-morpholinoacetohydrazide (9c)

White solid, yield 85%, mp: 183–185 °C, IR (ATR, cm−1) νmax 3206, 3045, 2956, 1664, 1591, 1267, 1078; 1H NMR (400 MHz, CDCl3) δ 9.98 (s, 1H), 8.05 (s, 1H), 7.22 (s, 1H), 6.95 (s, 1H), 3.86 (s, 3H), 3.72 (t, J = 4.4 Hz, 8H), 3.15 (s, 2H), 2.56 (t, J = 4.4 Hz, 8H). 13C NMR (100 MHz, CDCl3) δ 165.7, 149.7, 148.8, 148.2, 124.6, 121.8, 120.6, 109.3, 66.9, 66.6, 61.4, 61.1, 56.0, 53.8, 52.8. HRMS (Q-TOF) m/z: [M + H]+ calcd for C19H28N4O5, 393.2138; found 393.2132.
N′-(4-hydroxy-3-methoxy-5-((4-phenylpiperazin-1-yl)methyl)benzylidene)-2-(4-phenylpiperazin-1-yl)acetohydrazide (10a)

White solid, yield 91%, mp: 208–210 °C, IR (ATR, cm−1) νmax 3192, 2997, 2935, 1659, 1594, 1225, 1081, 762; 1H NMR (400 MHz, CDCl3) δ 10.06 (s, 1H), 8.11 (s, 1H), 7.33–7.27 (m, 5H), 7.05 (s, 1H), 6.97–6.90 (m, 6H), 3.94 (s, 3H), 3.81 (s, 2H), 3.28–3.26 (m, 10H), 2.79 (t, J = 4.4 Hz, 8H). 13C NMR (100 MHz, CDCl3) δ 165.8, 150.9, 150.8, 149.9, 148.9, 148.3, 129.2, 124,6, 121.8, 120.9, 120.4, 120.2, 116.5, 116.2, 109.4, 61.1, 60.8, 56.1, 53.6, 52.5, 49.3, 49.2. HRMS (Q-TOF) m/z: [M + H]+ calcd for C31H38N6O3, 543.3084; found 543.3080.
2-(3,4-Dihydroisoquinolin-2(1H)-yl)-N′-(4-hydroxy-3-methoxy-5-((4-phenylpiperazin-1-yl)methyl)benzylidene)acetohydrazide (10b)

White solid, yield 93%, mp: 214–216 °C, IR (ATR, cm−1) νmax 3203, 3067, 2916, 1666, 1595, 1226, 1078, 739; 1H NMR (400 MHz, CDCl3) δ 10.14 (s, 1H), 8.01 (s, 1H), 7.31–7.26 (m, 3H), 7.22–7.17 (m, 3H), 7.05 (s, 1H), 7.01 (s, 1H), 6.94–6.90 (m, 3H), 3.93 (s, 3H), 3.80 (s, 4H), 3.38 (s, 2H), 3.26 (brs, 4H), 3.00 (t, J = 5.5 Hz, 2H), 2.91 (t, J = 5.5 Hz, 2H), 2.76 (brs, 4H). 13C NMR (100 MHz, CDCl3) δ 166.0, 150.8, 149.8, 148.7, 148.3, 133.8, 133.5, 129.2, 128.8, 126.7, 126.6, 126.0, 124.6, 120.9, 120.3, 116.4, 109.4, 61.0, 60.8, 56.2, 56.1, 52.5, 51.6, 49.2, 29.3. HRMS (Q-TOF) m/z: [M + H]+ calcd for C30H35N5O3, 514.2818; found 514.2810.
N′-(4-hydroxy-3-methoxy-5-((4-phenylpiperazin-1-yl)methyl)benzylidene)-2-morpholinoaceto hydrazide (10c)

White solid, yield 93%, mp: 188–190 °C, IR (ATR, cm−1) νmax 3210, 3068, 2946, 1662, 1596, 1226, 1076, 758; 1H NMR (400 MHz, CDCl3) δ 10.00 (s, 1H), 8.10 (s, 1H), 7.28–7.24 (m, 3H), 7.01 (s, 1H), 6.92–6.86 (m, 3H), 3.91 (s, 3H), 3.81–3.75 (m, 6H), 3.24 (brs, 4H), 3.19 (s, 2H), 2.73 (brs, 4H), 2.60 (t, J = 4.4 Hz, 4H). 13C NMR (100 MHz, CDCl3) δ 165.7, 150.8, 149.9, 148.9, 148.3, 129.2, 124.5, 121.8, 120.9, 120.3, 116.4, 109.3, 66.9, 61.5, 60.8, 56.1, 53.9, 52.4, 49.2. HRMS (Q-TOF) m/z: [M + H]+ calcd for C25H33N5O4, 468.2611; found 468.2603.
N′-(4-hydroxy-3-methoxy-5-((3-methylpiperidin-1-yl)methyl)benzylidene)-2-(4-phenylpiperazin-1-yl)acetohydrazide (11a)

Beige solid, yield 82%, mp: 134–136 °C, IR (ATR, cm−1) νmax 3208, 3064, 2926, 1657, 1593, 1232, 1076, 760; 1H NMR (400 MHz, CDCl3) δ 10.04 (s, 1H), 8.06 (s, 1H), 7.30–7.24 (m, 3H), 6.99 (s, 1H), 6.95–6.87 (m, 3H), 3.90 (s, 3H), 3.69 (d, J = 7.2 Hz, 2H), 3.25–3.23 (m, 7H), 2.89–2.85 (m, 2H), 2.76 (t, J = 4.4 Hz, 4H), 2.05 (t, J = 10.0 Hz, 1H), 1.75–1.69 (m, 4H), 0.95–0.86 (m, 4H). 13C NMR (100 MHz, CDCl3) δ 165.8, 150.9, 150.6, 149.1, 148.2, 129.2, 124.0, 121.6, 121.2, 120.1, 116.2, 109.1, 61.2, 61.0, 60.8, 56.1, 53.6, 53.2, 49.3, 32.3, 31.0, 19.4. HRMS (Q-TOF) m/z: [M + H]+ calcd for C27H37N5O3, 480.2975; found 480.2967.
2-(3,4-Dihydroisoquinolin-2(1H)-yl)-N′-(4-hydroxy-3-methoxy-5-((3-methylpiperidin-1-yl)methyl) benzylidene)acetohydrazide (11b)

Beige solid, yield 85%, mp: 170–172 °C, IR (ATR, cm−1) νmax 3201, 3070, 2923, 1661, 1591, 1227, 1081, 738; 1H NMR (400 MHz, CDCl3) δ 10.10 (s, 1H), 7.97 (s, 1H), 7.22 (s, 1H), 7.19–7.14 (m, 3H), 7.03 (d, J = 4.4 Hz, 1H), 6.95 (s, 1H), 3.88 (s, 3H), 3.76 (s, 2H), 3.67 (d, J = 7.2 Hz, 2H), 3.33 (s, 2H), 2.98–2.85 (m, 7H), 2.03 (t, J = 10.0 Hz, 1H), 1.74–1.72 (m, 4H), 0.97–0.86 (m, 4H). 13C NMR (100 MHz, CDCl3) δ 166.1, 150.5, 148.9, 148.2, 133.9, 133.5, 128.8, 126.6, 126.0, 124.0, 121.5, 121.2, 109.1, 61.3, 61.0, 60.8, 56.2, 56.0, 53.2, 51.6, 32.3, 31.0, 29.3, 25.1, 24.2, 19.4. HRMS (Q-TOF) m/z: [M + H]+ calcd for C26H34N4O3, 451.2709; found 451.2704.
N'-(4-hydroxy-3-methoxy-5-((3-methylpiperidin-1-yl)methyl)benzylidene)-2-morpholinoaceto hydrazide (11c)

Beige solid, yield 81%, mp: 130–132 °C, IR (ATR, cm−1) νmax 3208, 3074, 2928, 1663, 1592, 1246, 1078; 1H NMR (400 MHz, CDCl3) δ 9.97 (s, 1H), 8.03 (s, 1H), 7.18 (s, 1H), 6.91 (s, 1H), 3.84 (s, 3H), 3.74 (t, J = 4.4 Hz, 4H), 3.63 (d, J = 7.2 Hz, 2H), 3.13 (s, 2H), 2.94–2.54 (m, 7H), 2.00 (t, J = 10.0 Hz, 1H), 1.70–1.65 (m, 4H), 0.91–0.82 (m, 4H). 13C NMR (100 MHz, CDCl3) δ 165.6, 150.6, 149.1, 148.2, 123.9, 121.5, 121.3, 66.8, 61.5, 61.4, 61.3, 60.8, 56.0, 53.8, 53.2, 32.3, 31.0, 25.1, 19.3. HRMS (Q-TOF) m/z: [M + H]+ calcd for C21H32N4O4, 405.2502; found 405.2492.
N′-(3-((3,4-dihydroisoquinolin-2(1H)-yl)methyl)-4-hydroxy-5-methoxybenzylidene)-2-(4-phenyl piperazin-1-yl)acetohydrazide (12a)

White solid, yield 94%, mp: 232–234 °C, IR (ATR, cm−1) νmax 3209, 3063, 2944, 1668, 1592, 1228, 1079, 750; 1H NMR (400 MHz, CDCl3) δ 10.05 (s, 1H), 8.12 (s, 1H), 7.33–7.28 (m, 2H), 7.18–7.12 (m, 4H), 7.09 (s, 1H), 7.02–6.90 (m, 5H), 3.93–3.92 (m, 5H), 3.80 (s, 2H), 3.29–3.27 (m, 5H), 2.99 (t, J = 5.5 Hz, 2H), 2.91 (t, J = 5.5 Hz, 2H), 2.80 (t, J = 4.4 Hz, 4H). 13C NMR (100 MHz, CDCl3) δ 165.8, 150.9, 150.2, 149.0, 148.4, 133.3, 132.8, 129.3, 128.7, 126.7, 126.6, 126.0, 121.8, 121.1, 120.2, 116.2, 109.5, 61.1, 60.4, 56.1, 55.1, 53.6, 49.9, 49.3, 28.4. HRMS (Q-TOF) m/z: [M + H]+ calcd for C30H35N5O3, 514.2818; found 514.2811.
2-(3,4-Dihydroisoquinolin-2(1H)-yl)-N′-(3-((3,4-dihydroisoquinolin-2(1H)-yl)methyl)-4-hydroxy-5-methoxybenzylidene)acetohydrazide (12b)

White solid, yield 92%, mp: 215–217 °C, IR (ATR, cm−1) νmax 3188, 3063, 2905, 1658, 1589, 1226, 1077, 737; 1H NMR (400 MHz, CDCl3) δ 10.13 (s, 1H), 8.02 (s, 1H), 7.28 (s, 1H), 7.20–7.13 (m, 6H), 7.07–7.00 (m, 3H), 3.91 (s, 5H), 3.80 (s, 2H), 3.78 (s, 2H), 3.38 (s, 2H), 3.02–2.96 (m, 4H), 2.92–2.87 (m, 4H). 13C NMR (100 MHz, CDCl3) δ 166.0, 150.2, 148.8, 148.4, 133.9, 133.5, 133.3, 128.8, 128.7, 126.7, 126.6, 126.6, 126.0, 124.5, 121.7, 121.1, 109.5, 61.0, 60.5, 56.2, 56.1, 55.2, 51.6, 49.9, 29.3, 28.5. HRMS (Q-TOF) m/z: [M + H]+ calcd for C29H32N4O3, 485.2553; found 485.2548.
N′-(3-((3,4-dihydroisoquinolin-2(1H)-yl)methyl)-4-hydroxy-5-methoxybenzylidene)-2-morpholin oacetohydrazide (12c)

White solid, yield 90%, mp: 220–222 °C, IR (ATR, cm−1) νmax 3199, 3069, 2936, 1660, 1565, 1229, 1081, 745; 1H NMR (400 MHz, CDCl3) δ 9.97 (s, 1H), 8.13 (s, 1H), 7.30 (s, 1H), 7.20–7.12 (m, 3H), 7.08 (s, 1H), 7.02–7.00 (m, 1H), 3.93 (s, 5H), 3.80–3.78 (m, 6H), 3.21 (s, 2H), 2.98–2.91 (m, 4H), 2.63 (bs, 4H). 13C NMR (100 MHz, CDCl3) δ 165.6, 150.2, 149.0, 148.4, 133.2, 132.7, 128.7, 126.7, 126.6, 126.1, 124.5, 121.8, 121.0, 114.8, 109.5, 66.9, 61.5, 60.3, 56.1, 55.1, 53.9, 49.9, 28.4. HRMS (Q-TOF) m/z: [M + H]+ calcd for C24H30N4O4, 439.2345; found 439.2340.
Ethyl 4-(2-hydroxy-3-methoxy-5-((2-(2-(4-phenylpiperazin-1-yl)acetyl)hydrazono)methyl)benzyl) piperazine-1-carboxylate (13a)

White solid, yield 91%, mp: 181–183 °C, IR (ATR, cm−1) νmax 3186, 3056, 2922, 1698, 1657, 1594, 1239, 1089, 760; 1H NMR (400 MHz, CDCl3) δ 10.09 (s, 1H), 8.06 (s, 1H), 7.27–7.23 (m, 3H), 6.97 (s, 1H), 6.91–6.84 (m, 3H), 4.12 (q, J = 7.1 Hz, 2H), 3.89 (s, 3H), 3.51 (brs, 4H), 3.22 (m, 8H), 2.73 (t, J = 4.4, 4H), 2.50 (brs, 4H), 1.23 (t, J = 7.1, 3H). 13C NMR (100 MHz, CDCl3) δ 165.8, 155.2, 150.9, 149.6, 148.7, 148.3, 129.2, 124.7, 121.6, 120.7, 120.1, 116.1, 109.4, 61.6, 61.0, 60.8, 56.1, 53.6, 52.1, 49.2, 43.4, 14.6. HRMS (Q-TOF) m/z: [M + H]+ calcd for C28H38N6O5, 539.2982; found 539.2975.
Ethyl 4-(5-((2-(2-(3,4-dihydroisoquinolin-2(1H)-yl)acetyl)hydrazono)methyl)-2-hydroxy-3-methoxy benzyl)piperazine-1-carboxylate (13b)

White solid, yield 93%, mp: 188–191 °C, IR (ATR, cm−1) νmax 3183, 3034, 2936, 1703, 1655, 1596, 1267, 1082, 750; 1H NMR (400 MHz, CDCl3) δ 10.15 (s, 1H), 7.99 (s, 1H), 7.25 (s, 1H), 7.20–7.14 (m, 3H), 7.04 (d, J = 5.6 Hz, 1H), 6.97 (s, 1H), 4.14 (q, J = 7.1 Hz, 2H), 3.90 (s, 3H), 3.78 (s, 2H), 3.72 (s, 2H), 3.54 (brs, 4H), 3.36 (s, 2H), 2.98 (t, J = 5.5 Hz,, 2H), 2.91 (t, J = 5.5, 2H), 2.54 (brs, 4H), 1.26 (t, J = 7.1, 3H). 13C NMR (100 MHz, CDCl3) δ 166.0, 155.2, 149.5, 148.6, 148.3, 133.7, 133.4, 128.8, 126.7, 126.6, 126.0, 124.7, 121.6, 120.7, 109.5, 61.6, 61.0, 60.8, 56.2, 56.1, 52.2, 51.6, 43.4, 29.2, 14.6. HRMS (Q-TOF) m/z: [M + H]+ calcd for C27H35N5O5, 510.2716; found 510.2712.
Ethyl 4-(2-hydroxy-3-methoxy-5-((2-(2-morpholinoacetyl)hydrazono)methyl)benzyl)piperazine-1-carboxylate (13c)

White solid, yield 88%, mp: 163–165 °C, IR (ATR, cm−1) νmax 3187, 3055, 2924, 1699, 1656, 1590, 1236, 1087, 764; 1H NMR (400 MHz, CDCl3) δ 9.99 (s, 1H), 8.06 (s, 1H), 7.23 (s, 1H), 6.96 (s, 1H), 4.11 (q, J = 7.1 Hz, 2H), 3.89 (s, 3H), 3.73–3.71 (m, 5H), 3.51 (brs, 4H), 3.16 (s, 2H), 2.57 (t, J = 4.4 Hz, 4H), 2.52 (brs, 4H), 1.23 (t, J = 7.1, 3H). 13C NMR (100 MHz, CDCl3) δ 165.6, 155.2, 149.6, 148.8, 148.3, 124.7, 121.7, 120.7, 109.4, 66.9, 61.6, 61.4, 60.8, 56.1, 53.8, 52.15, 43.5, 14.6. HRMS (Q-TOF) m/z: [M + H]+ calcd for C22H33N5O6, 464.2509; found 464.2503.
Biological studies
According to prior research, the AR purification procedure was carried out utilizing the (NH4)2SO4 precipitation DE-52 cellulose ion-exchange column, Sephadex G-100 gel filtration column, and 2′5′-ADP Sepharose-4B affinity column [60,61,62,63,64,65]. The Bradford technique at 595 nm was used to determine the protein content of the samples [66,67,68]. SDS-PAGE technique was employed to ensure enzyme purity [69,70,71]. AR activity was assessed spectrophotometrically using dl-glyceraldehyde and NADPH reduction at 340 nm [72,73,74]. Activity (%) novel acyl hydrazones generated from vanillin compounds and standard inhibitor epalrestat plots were used to calculate the IC50 values, inhibitory concentrations that reduce enzyme activity by 50%. Three different inhibitory doses were applied to determine KI values and inhibition types [75, 76].
In silico studies
The Maestro ver. 13.1 [77], Protein Preparation Wizard [78], SiteMap [79], Receptor Grid Generation [80], LigPrep [81], QikProp [82], Prime MM-GBSA [83], and Jaguar [84] tools are implemented in Small-Molecule Drug Discovery Suite 2022-1 for Mac (Schrödinger, LLC, NY, USA) and were used to perform molecular docking, ADME, and DFT calculations. All compounds, including novel acyl hydrazones and the reference ligand EPR, were sketched in the 2D-sdf format using ChemDraw ver. 19.1 for Mac [85] (PerkinElmer, Inc., Waltham, MA, USA), and ligand production was performed using the LigPrep tool [86,87,88]. The QikProp module was used to estimate ADME-related parameters for these substances as described in previous studies [89,90,91]. The Protein Data Bank provided the X-ray structure of the template 4JIR [92] (Resolution: 2.00 Å; R-Values free and work: 0.210 and 0.160, respectively; Species: Homo sapiens) and wass prepared using the Protein Preparation Wizard [93,94,95]. The Receptor Grid Generation tool [96,97,98] was used to create the docking grid box. The extra-precision (XP) approach [99,100,101] was used to perform molecular-docking simulations. Also, the VSGB energy model [102,103,104] and OPLS4 force field [105, 106] were used to calculate MM-GBSA binding energies [107, 108], which predict relative binding affinities for these novel acyl hydrazones. The novel acyl hydrazones were also analyzed via Becke’s three-parameter exchange potential and Lee–Yang–Parr correlation functional (B3LYP) using a 6-31G∗∗ basic set level. With single-point calculations, the implicit solvation model of Poisson Boltzmann Finite was used. The electrostatic potentials were computed using the molecule’s van der Waals contact surface area [109, 110].
Statistical studies
Analysis of the data and drawing of graphs were realized using GraphPad Prism ver. 8 for Mac (GraphPad Software, La Jolla California USA). The inhibition constants were calculated by SigmaPlot ver. 12 for Windows (Systat Software, San Jose California USA). The fit of enzyme inhibition models was compared using the extra sum-of-squares F test and the AICc approach. The results were exhibited as mean ± standard error of the mean (95% confidence intervals). Differences between datasets were considered statistically significant when the p value was less than 0.05.
Results and discussion
Chemistry
The synthesis pathway of designed molecules was carried out using reagents and conditions as presented in Schemes 1 and 2. Briefly, compounds 2a and 2b were synthesized according to the Schotten Baumann reaction, and compounds 3a-e were synthesized according to the Mannich Reaction, with good yields.
In Scheme 2, hydrazides (6a–c) were synthesized from esters formed by the reaction of the related cyclic secondary amine with ethyl chloroacetate. In the final step, the synthesized aldehydes (2a–b, 3a–e) were treated with the synthesized hydrazides (6a–c), producing target compounds (7a–c to 13a–c) with yields ranging from 75 to 90%. The melting points of the known intermediates were compared with the values in the literature. The structures of the newly synthesized compounds were characterized by IR, 1H NMR, and 13C NMR spectroscopic methods.
In the IR spectra of the compounds 7a–c to 13a–c, NH stretching bands are observed at 3210–3145 cm−1. Aromatic C–H stretching bands are seen at 3108–3034 cm−1 and the aliphatic C–H stretching bands are observed at 2960–2905 cm−1. C=O of hydrazone moiety and CH=N stretching bands are observed at 1668–1654 cm−1 and 1597–1556 cm−1, respectively. C–O and N–H bending bands are seen at 1270–1225 cm−1 and 1089–1067 cm−1, respectively. For compounds 7a–c, 8a–c, and 13a–c, stretching bands of ester carbonyl are observed at 1743–1698 cm−1.
In the 1H NMR spectra of the compounds 7a–c to 13a–c, peaks of NH protons are seen as singlet at δ 10.34–9.97 ppm. Peaks of N=CH protons are observed as singlet at δ 8.25–7.97 ppm. The resonance signals of aromatic protons are observed at δ 7.99–6.60 ppm as singlet, doublet, triplet, and multiplet relative to their chemical environment. Peaks of OCH3 and Ph–CH2–N protons are seen as singlet at δ 3.94–3.84 and δ 3.81–3.71 ppm, respectively. Peaks of N–CH2–C=O protons are seen as singlet at δ 3.39–3.15 ppm. Aliphatic protons of morpholine, piperazine, tetrahydroisoquinoline, and piperidine moieties are observed at δ 3.75–0.82 ppm as singlet, doublet, triplet, and multiplet relative to their chemical environment. For compounds 13a–c, peaks of OCH2CH3 protons are seen as quartet at δ 4.14–4.11 ppm and peaks of OCH2CH3 protons are seen as triplet at δ 1.26–1.23 ppm. Chemical shifts, integrations, and splits are fully compatible with the structures.
In the 13C NMR spectra of target molecules, peaks of HN–C=O and CH=N carbons are seen at δ 166.5–165.6 ppm and δ 147.8–148.4 ppm, respectively. For compounds 7a–c and 8a–c, peaks of Ar–C=O carbons are observed at δ 160.0–156.2 ppm. Peaks of aromatic carbons are observed at δ 151.9–109.1 ppm. Peaks of OCH3 carbons are seen at δ 56.3–56.0 ppm, and peaks of N–CH2–C=O carbons are seen at δ 61.5–61.0 ppm. For compounds 13a–c, peaks of CH3CH2–O–C=O carbons are observed at δ 155.2 ppm. Also, for compounds 13a–c, peaks of CH3CH2–O–C=O and CH3CH2–O–C=O carbons are observed at δ 43.5–43.4 ppm and δ 14.6 ppm, respectively. Finally, aliphatic carbons of morpholine, piperazine, tetrahydroisoquinoline, and piperidine moieties are observed at δ 66.9–19.3 ppm. Chemical shifts and the number of the peaks are fully compatible with the structures.
Biological studies and structure–activity relationship
As shown in Table 1, 21 compounds exhibited relative more potent inhibitory activity against AR with KI values ranged from 49.22 ± 3.64 to 1114.00 ± 49.64 nM, and among them, compound 11c, named N'-(4-hydroxy-3-methoxy-5-((3-methylpiperidin-1-yl)methyl)benzylidene)-2-morpholinoaceto hydrazide, displayed the strongest inhibitory effect with an KI value of 49.22 ± 3.64 nM (Fig. 1). The inhibitor effects of novel acyl hydrazones derived from vanillin compounds against AR were decreased in the following order: 11c > (KI: 49.22 ± 3.64 nM) 10b > (KI: 79.36 ± 5.77 nM) 7a > (KI: 101.00 ± 8.21 nM) 10a > (KI: 145.80 ± 22.30 nM) 8a > (KI: 182.40 ± 14.35 nM) 7c > (KI: 304.00 ± 13.36 nM) 9b > (KI: 312.80 ± 53.48 nM) 12b > (KI: 338.40 ± 17.18 nM) 9c > (KI: 372.10 ± 62.45 nM) 11b > (KI: 394.00 ± 22.65 nM) 11a > (KI: 398.20 ± 15.20 nM) 12a > (KI: 437.40 ± 14.08 nM) 13b > (KI: 444.50 ± 24.74 nM) 7b > (KI: 523.00 ± 22.54 nM) 12c > (Ki: 533.50 ± 26.67 nM) 10c > (KI: 598.70 ± 19.27 nM) 8c > (KI: 746.20 ± 34.41 nM) 13a > (KI: 787.10 ± 39.32 nM) 13c > (KI: 854.00 ± 33.96 nM) 8b > (KI: 897.20 ± 43.63 nM) 9a > (KI: 1114.00 ± 49.64 nM).
There are different types of inhibition, including mixed, non-competitive, competitive, and un-competitive. It would be appropriate to state that the inhibitory potential of the molecules is due to the structural, 3D chemical structure, and conformation features that vary according to the different groups on which the backbone structure depends. When compounds 7a and 7c are compared, substitution of 4-phenylpiperazin-1-yl structure with 2-morpholino caused a threefold change in the inhibition value. The 4-phenylpiperazin-1-yl group showed a better inhibition effect in the replacement of the phenylfuran-2-carboxylate structure in the structure of compounds 7a and 7c with thiophene-2-carboxylate (8a and 8c).
When acetohydrazide compounds were compared (9c, 10c, and 11c), the inhibition effect was observed as follows, respectively: 3-methylpiperidin-1-yl (11c, KI: 49.22 ± 3.64 nM) > 5-morpholinomethyl (9c, KI: 372.10 ± 62.45 nM) > 4-phenylpiperazin-1-yl (10c, KI: 598.70 ± 19.27 nM). On the contrary, when we look at the inhibition order of 2-(4-phenylpiperazin-1-yl)acetohydrazide compounds, the inhibition effect was observed as follows, respectively: 4-phenylpiperazin-1-yl (10a) > 3-methylpiperidin-1-yl (11a) > 5-morpholinomethyl (9a). Considering the inhibition order of piperazine-1-carboxylate compounds (13a, 13b and 13c), compound 13b showed better inhibition effect (KI: 444.50 ± 24.74 nM).
Potential inhibitory effect of synthesized compounds against AR has been reported in the literature. Yapar et al. [111] synthesized the novel bis-hydrazone compounds bearing isovanillin moiety and studied inhibition effect of these compounds on AR enzyme activity. They found that the novel bis-hydrazones demonstrated in nanomolar levels as AR inhibitors with KI values in the range of 13.38–88.21 nM. Maccari et al. [112] performed inhibition effect of 5-arylidene-2,4-thiazolidinediones on AR enzyme. The authors found that a hydroxyl group on the 5-arylidene moiety led to significant inhibitory effect. Alexiou et al. [113] synthesized a series of N-(3,5-difluoro-4-hydrozyphenyl)benzenesulfonamide derivatives and studied the inhibition effect of novel compounds on AR. They enhanced these compounds compared to N-benzenesulfonylglycine lead derivative. The most potent inhibitor was found to be compound 66 with the IC50 value of 14.1 μM.
In silico studies
Table 2 summarizes the results of the determination of ADME-related parameters for novel acyl hydrazones. New acyl hydrazones were identified as hit-agents with drug-like effects based on ADME properties calculations. According to this information, the molecular weights (MWs, 392.45–542.68) and dipole moments (Dipole, in the 2.65 to 9.64) of the novel acyl hydrazones derived from vanillin compounds (7a–c and 13a–c) have reported being in the permissible values. Volume (in range 1227.48 to 1752.91), which is the total solvent-accessible volume descriptor, was determined to be in the permissible ranges for these hydrazones (7a–c and 13a–c), compared with reference values. The logP values, such as QPlogPoct, QPlogPw, QPlogPo/w, QPlogS, QPlogBB, QPlogKp, and QPlogKhs, are in ranging from 20.27 to 29.35, 12.63 to 16.76, 0.69 to 4.61, − 6.46 to − 1.26, − 1.14 to − 0.24, − 7.17 to − 3.28, and − 0.55 to 0.93, respectively, and indicates of target derivatives (7a–c and 13a–c) have the high capacity. The values of human oral absorption (HOAs) were higher than 30%, and van der Waals surface area of polar nitrogen and oxygen atoms (PSA, in the range 86.82 to 134.05) indicate that all analogs (7a–c and 13a–c) had at the acceptable values. All the acyl hydrazones have displayed normal Caco-2 cell permeability rates (except for compounds 13b and 13c; QPPCaco, in the 23.47 to 245.90), and MDCK cell permeability values (except for compound 13a, 13b, and 13c; QPPMDCK, in range 10.49 to 214.30). Indeed, all newly synthesized acyl hydrazones derived from vanillin compounds (7a–c and 13a–c) displayed good drug-like properties with zero violation of Lipinski’s rule (except for compounds 10a–b, 12a, and 13a–c) and zero or one violation of the Jorgensen’s rule (except for compounds 12a) (Table 2). Moreover, the ADME-Tox values calculated for N′-(4-hydroxy-3-methoxy-5-((3-methylpiperidin-1-yl)methyl)benzylidene)-2-morpholinoaceto hydrazide 11c might explain why, being a potent AR inhibitor, this ligand has the most AR inhibitory activity in biological experiments.
Molecular docking experiments were used to obtain substantial insight into the origins of the structure–activity connections examined for the new acyl hydrazones. Initially, the native ligand EPR ((5-[(2E)-2-methyl-3-phenylprop-2-en-1-ylidene]-4-oxo-2-thioxo-1,3-thiazolidin-3-yl)acetic acid) in the AR receptor's binding site (PDB code 4JIR) [92] was employed in the redocking computation. At a root-mean-square deviation (RMSD) of 0.10 (docking score of − 7.04 kcal/mol and MM-GBSA value of − 41.07 kcal/mol), the docked pose of EPR overlapped with the pose in the X-ray crystal structure of the AR (Fig. 2). This redocking experiment was crucial in determining which model structure would best accommodate all of the newly synthesized AR ligands. Then, using the Glide Ligand Docking tool in this series, the generated binding model was used to perform docking calculations of the most potent AR inhibitor 11c. A docking score of − 8.07 kcal/mol and MM-GBSA value of − 69.69 kcal/mol indicated compound 11c are a tight binder for AR compared to EPR. The carboxy moiety formed an H-bond with residue Trp111 (distance 2.16 Å), while the –NH group displayed π-cation interaction with Phe122. Furthermore, compound 11c monitored hydrophobic interactions with residues Trp20, Val47, Tyr48, Trp79, Phe121, Tyr209, Trp219, Ile260, Cys298, Leu300, Leu301, and Cys303 played significant roles in the binding of the ligand with 4JIR (Fig. 3).
Molecular docking of aldode reductase (AR; PDB code: 4JIR) with native ligand EPR ((5-((2E)-2-methyl-3-phenylprop-2-en-1-ylidene)-4-oxo-2-thioxo-1,3-thiazolidin-3-yl)acetic acid). A 3D ligand interaction diagram of 4JIR with native ligand EPR. B 2D docking pose of native ligand EPR with the key amino acids within the binding pocket of 4JIR
Molecular docking of aldode reductase (AR; PDB code: 4JIR) with compound 11c (N'-(4-hydroxy-3-methoxy-5-((3-methylpiperidin-1-yl)methyl)benzylidene)-2-morpholinoaceto hydrazide). A 3D ligand interaction diagram of 4JIR with compound 11c. B 2D docking pose of compound 11c with the key amino acids within the binding pocket of 4JIR
To explain the structural parameters, the DFT calculation was performed for compound 11c, which has the most potent AR inhibitory activity and was optimized at the level of B3LYP/6-31G∗∗. In chemical reactivity, derivative 11c is sparkling, and the HOMO (highest occupied molecular orbitals)-LUMO (lowest unoccupied molecular orbitals) gap increases the charge transfer of the compounds. The electron density is indicated by the intensity of the color that reflects the distinctive feature of the molecule. Because electrons can move quickly between energy levels in the HOMO and LUMO, energy gap levels reveal the delicate nature of reactivity. The energy gap of the compound 11c in the HOMO–LUMO analysis is 0,160,511 eV, and the HOMO–LUMO plot of 11c is shown in Fig. 4. From this plot, it is seen that the value of ΔE decreases in the case of complex, which further supports the binding framework and that compound 11c has significant chemical reactivity and polarizability.
Conclusion
A series of acyl hydrazones derived from vanillin were synthesized and their effects on the AR were investigated. KI values in the range of 49.22 ± 3.64 to 897.20 ± 43.63 nM. Compounds 11c and 10b against AR enzyme activity were identified as the highly potent inhibitors than epalrestat. AR is novel molecular target involved in different pathways related to the development of type II diabetes mellitus and related comorbidities. The design of effective bioavailable inhibitors for AR enzyme is still an urgent need. We expect that our findings will lead to the development of novel AR inhibitors based on inhibition and molecular docking investigations. We also hope that our compounds will be good therapeutic candidates with further investigation.
References
Areas ES, Bronsato BJdS, Pereira TM, Guedes GP, Miranda FdS, Kümmerle AE et al (2017) Novel Co(III) complexes containing fluorescent coumarin-N-acylhydrazone hybrid ligands: synthesis, crystal structures, solution studies and DFT calculations. Spectrochim Acta Part A. https://doi.org/10.1016/j.saa.2017.06.031
Aarjane M, Slassi S, Amine A (2021) Novel series of N-acylhydrazone based on acridone: synthesis, conformational and theoretical studies. J Mol Struct. https://doi.org/10.1016/j.molstruc.2020.129079
Rollas S, Küçükgüzel SG (2007) Biological activities of hydrazone derivatives. Molecules 12(8):1910–1939. https://doi.org/10.3390/12081910
Abdelrahman MA, Salama I, Gomaa MS, Elaasser MM, Abdel-Aziz MM, Soliman DH (2017) Design, synthesis and 2D QSAR study of novel pyridine and quinolone hydrazone derivatives as potential antimicrobial and antitubercular agents. Eur J Med Chem. https://doi.org/10.1016/j.ejmech.2017.07.004
Moldovan CM, Oniga O, Parvu A, Tiperciuc B, Verite P, Pîrnău A et al (2011) Synthesis and anti-inflammatory evaluation of some new acyl-hydrazones bearing 2-aryl-thiazole. Eur J Med Chem 46(2):526–534. https://doi.org/10.1016/j.ejmech.2010.11.032
Kareem HS, Ariffin A, Nordin N, Heidelberg T, Abdul-Aziz A, Kong KW et al (2015) Correlation of antioxidant activities with theoretical studies for new hydrazone compounds bearing a 3, 4, 5-trimethoxy benzyl moiety. Eur J Med Chem. https://doi.org/10.1016/j.ejmech.2015.09.016
Vavříková E, Polanc S, Kočevar M, Horváti K, Bősze S, Stolaříková J et al (2011) New fluorine-containing hydrazones active against MDR-tuberculosis. Eur J Med Chem 46(10):4937–4945. https://doi.org/10.1016/j.ejmech.2011.07.052
Hernández P, Cabrera M, Lavaggi ML, Celano L, Tiscornia I, da Costa TR et al (2012) Discovery of new orally effective analgesic and anti-inflammatory hybrid furoxanyl N-acylhydrazone derivatives. Bioorg Med Chem 20(6):2158–2171. https://doi.org/10.1016/j.bmc.2012.01.034
Carradori S, Secci D, Bolasco A, Rivanera D, Mari E, Zicari A et al (2013) Synthesis and cytotoxicity of novel (thiazol-2-yl) hydrazine derivatives as promising anti-Candida agents. Eur J Med Chem. https://doi.org/10.1016/j.ejmech.2013.04.042
Wang G, Chen M, Wang J, Peng Y, Li L, Xie Z et al (2017) Synthesis, biological evaluation and molecular docking studies of chromone hydrazone derivatives as α-glucosidase inhibitors. Bioorg Med Chem Lett 27(13):2957–2961. https://doi.org/10.1016/j.bmcl.2017.05.007
Nagender P, Kumar RN, Reddy GM, Swaroop DK, Poornachandra Y, Kumar CG et al (2016) Synthesis of novel hydrazone and azole functionalized pyrazolo [3, 4-b] pyridine derivatives as promising anticancer agents. Bioorg Med Chem Lett 26(18):4427–4432. https://doi.org/10.1016/j.bmcl.2016.08.006
Li Z-H, Yang D-X, Geng P-F, Zhang J, Wei H-M, Hu B et al (2016) Design, synthesis and biological evaluation of [1, 2, 3] triazolo [4, 5-d] pyrimidine derivatives possessing a hydrazone moiety as antiproliferative agents. Eur J Med Chem. https://doi.org/10.1016/j.ejmech.2016.10.022
Celebioglu HU, Erden Y, Hamurcu F, Taslimi P, Şentürk OS, Özmen ÜÖ et al (2021) Cytotoxic effects, carbonic anhydrase isoenzymes, α-glycosidase and acetylcholinesterase inhibitory properties, and molecular docking studies of heteroatom-containing sulfonyl hydrazone derivatives. J Biomol Struct Dyn 39(15):5539–5550. https://doi.org/10.1080/07391102.2020.1792345
Kaya Y, Erçağ A, Zorlu Y, Demir Y, Gülçin İ (2022) New Pd(II) complexes of the bisthiocarbohydrazones derived from isatin and disubstituted salicylaldehydes: synthesis, characterization, crystal structures and inhibitory properties against some metabolic enzymes. J Biol Inorg Chem 27(2):271–281. https://doi.org/10.1007/s00775-022-01932-9
Kucukoglu K, Gul HI, Taslimi P, Gulcin I, Supuran CT (2019) Investigation of inhibitory properties of some hydrazone compounds on hCA I, hCA II and AChE enzymes. Bioorg Chem. https://doi.org/10.1016/j.bioorg.2019.02.008
Todeschini AR, de Miranda ALP, da Silva KCM, Parrini SC, Barreiro EJ (1998) Synthesis and evaluation of analgesic, antiinflammatory and antiplatelet properties of new 2-pyridylarylhydrazone derivatives. Eur J Med Chem 33(3):189–199. https://doi.org/10.1016/S0223-5234(98)80008-1
Dimmock JR, Vashishtha SC, Stables JP (2000) Anticonvulsant properties of various acetylhydrazones, oxamoylhydrazones and semicarbazones derived from aromatic and unsaturated carbonyl compounds. Eur J Med Chem 35(2):241–248. https://doi.org/10.1016/S0223-5234(00)00123-9
Zhang B, Zhao YF, Zhai X, Fan WJ, Ren JL, Wu CF et al (2012) Design, synthesis and antiproliferative activities of diaryl urea derivatives bearing N-acylhydrazone moiety. Chin Chem Lett 23(8):915–918. https://doi.org/10.1016/j.cclet.2012.06.009
Buu-Hoï NP, Xuong ND, Nam NH, Binon F, Royer R (1953) Tuberculostatic hydrazides and their derivatives. J Chem Soc 278:1358–1364. https://doi.org/10.1039/JR9530001358
Roman G (2015) Mannich bases in medicinal chemistry and drug design. Eur J Med Chem. https://doi.org/10.1016/j.ejmech.2014.10.076
Martin-Escolano R, Moreno-Viguri E, Santivanez-Veliz M, Martin-Montes A, Medina-Carmona E, Paucar R et al (2018) Second generation of Mannich base-type derivatives with in vivo activity against Trypanosoma cruzi. J Med Chem 61(13):5643–5663. https://doi.org/10.1021/acs.jmedchem.8b00468
Racane L, Tralic-Kulenovic V, Fiser-Jakic L (2001) Synthesis of bis-substituted amidinobenzothiazoles as potential anti-HIV agents. Heterocycles 55(11):2085–2098. https://doi.org/10.3987/COM-01-9305
Kashiyama E, Hutchinson I, Chua M-S, Stinson SF, Phillips LR, Kaur G et al (1999) Antitumor benzothiazoles. 8. Synthesis, metabolic formation, and biological properties of the C- and N-oxidation products of antitumor 2-(4-aminophenyl) benzothiazoles. J Med Chem 42(20):4172–4184. https://doi.org/10.1021/jm990104o
Gul HI, Yerdelen KO, Gul M, Das U, Pandit B, Li PK et al (2007) Synthesis of 4′-hydroxy-3′-piperidinomethylchalcone derivatives and their cytotoxicity against PC-3 cell lines. Arch Pharm (Weinheim, Ger) 340(4):195–201. https://doi.org/10.1002/ardp.200600072
Reddy MVB, Su C-R, Chiou W-F, Liu Y-N, Chen RY-H, Bastow KF et al (2008) Design, synthesis, and biological evaluation of Mannich bases of heterocyclic chalcone analogs as cytotoxic agents. Bioorg Med Chem 16(15):7358–7370. https://doi.org/10.1016/j.bmc.2008.06.018
Dimmock J, Kumar P (1997) Anticancer and cytotoxic properties of Mannich bases. Curr Med Chem 4(1):1–22
Tugrak M, Gul HI, Sakagami H, Mete E (2017) Synthesis and anticancer properties of mono Mannich bases containing vanillin moiety. Med Chem Res 26(7):1528–1534. https://doi.org/10.1007/s00044-017-1833-x
Gul HI, Calis U, Vepsalainen J (2004) Synthesis of some mono-Mannich bases and corresponding azine derivatives and evaluation of their anticonvulsant activity. Arzneim-Forsch 54(07):359–364. https://doi.org/10.1055/s-0031-1296984
Chen G, Shan W, Wu Y, Ren L, Dong J, Ji Z (2005) Synthesis and anti-inflammatory activity of resveratrol analogs. Chem Pharm Bull 53(12):1587–1590. https://doi.org/10.1248/cpb.53.1587
Malhotra M, Sharma R, Sanduja M, Kumar R, Jain J, Deep A (2012) Synthesis, characterization and evaluation of Mannich bases as potent antifungal and hydrogen peroxide scavenging agents. Acta Pol Pharm Drug Res 69:355–361
Yamali C, Tugrak M, Gul HI, Tanc M, Supuran CT (2016) The inhibitory effects of phenolic Mannich bases on carbonic anhydrase I and II isoenzymes. J Enzyme Inhib Med Chem 31(6):1678–1681. https://doi.org/10.3109/14756366.2015.1126715
Sieger GM, Barringer WC, Krueger JE (1971) Mannich derivatives of medicinals. 2. Derivatives of some carbonic anhydrase inhibitors. J Med Chem 14(5):458–460. https://doi.org/10.1021/jm00287a027
Marvadi SK, Krishna VS, Sriram D, Kantevari S (2019) Synthesis of novel morpholine, thiomorpholine and N-substituted piperazine coupled 2-(thiophen-2-yl) dihydroquinolines as potent inhibitors of Mycobacterium tuberculosis. Eur J Med Chem. https://doi.org/10.1016/j.ejmech.2018.12.043
Özil M, Parlak C, Baltaş N (2018) A simple and efficient synthesis of benzimidazoles containing piperazine or morpholine skeleton at C-6 position as glucosidase inhibitors with antioxidant activity. Bioorg Chem. https://doi.org/10.1016/j.bioorg.2017.12.019
Patil P, Madhavachary R, Kurpiewska K, Kalinowska-Tłuścik J, Dömling A (2017) De novo assembly of highly substituted morpholines and piperazines. Org Lett 19(3):642–645. https://doi.org/10.1021/acs.orglett.6b03807
Scott JD, Williams RM (2002) Chemistry and biology of the tetrahydroisoquinoline antitumor antibiotics. Chem Rev 102(5):1669–1730. https://doi.org/10.1021/cr010212u
Hyndman D, Bauman DR, Heredia VV, Penning TM (2003) The aldo-keto reductase superfamily homepage. Chem-Biol Interact. https://doi.org/10.1016/S0009-2797(02)00193-X
Tammali R, Reddy A, Srivastava SK, Ramana KV (2011) Inhibition of aldose reductase prevents angiogenesis in vitro and in vivo. Angiogenesis 14(2):209–221. https://doi.org/10.1007/s10456-011-9206-4
Jannapureddy S, Sharma M, Yepuri G, Schmidt AM, Ramasamy R (2021) Aldose reductase: an emerging target for development of interventions for diabetic cardiovascular complications. Front Endocrin. https://doi.org/10.3389/fendo.2021.636267
Ramos RJ, Albersen M, Vringer E, Bosma M, Zwakenberg S, Zwartkruis F et al (2019) Discovery of pyridoxal reductase activity as part of human vitamin B6 metabolism. Biochim Biophys Acta, General Sub 1863:1088–1097. https://doi.org/10.1016/j.bbagen.2019.03.019
Quattrini L, La Motta C (2019) Aldose reductase inhibitors: 2013-present. Expert Opin Ther Pat 29(3):199–213. https://doi.org/10.1080/13543776.2019.1582646
Lj Y (2018) Redox imbalance stress in diabetes mellitus: Role of the polyol pathway. Anim Models Exp Med 1(1):7–13. https://doi.org/10.1002/ame2.12001
Langer HT, Afzal S, Kempa S, Spuler S (2020) Nerve damage induced skeletal muscle atrophy is associated with increased accumulation of intramuscular glucose and polyol pathway intermediates. Sci Rep 10(1):1–10. https://doi.org/10.1038/s41598-020-58213-1
Lu Q, Hao M, Wu W, Zhang N, Isaac AT, Yin J et al (2018) Antidiabetic cataract effects of GbE, rutin and quercetin are mediated by the inhibition of oxidative stress and polyol pathway. Acta Biochim Pol 65(1):35–41. https://doi.org/10.18388/abp.2016_1387
Chung SS, Ho EC, Lam KS, Chung SK (2003) Contribution of polyol pathway to diabetes-induced oxidative stress. J Am Soc Nephro 14(suppl 3):S233–S236
Oates PJ (2002) Polyol pathway and diabetic peripheral neuropathy. Int Rev Neuro. https://doi.org/10.1016/S0074-7742(02)50082-9
Yabe-Nishimura C (1998) Aldose reductase in glucose toxicity: a potential target for the prevention of diabetic complications. Pharmacol Rev 50(1):21–34
Wojnar W, Zych M, Borymski S, Kaczmarczyk-Sedlak I (2020) Chrysin reduces oxidative stress but does not affect polyol pathway in the lenses of type 1 diabetic rats. Antioxidants 9(2):160. https://doi.org/10.3390/antiox9020160
Oyama T, Miyasita Y, Watanabe H, Shirai K (2006) The role of polyol pathway in high glucose-induced endothelial cell damages. Diabet Res Clin Practic 73(3):227–234. https://doi.org/10.1016/j.diabres.2006.02.010
Li Q, Hwang YC, Ananthakrishnan R, Oates PJ, Guberski D, Ramasamy R (2008) Polyol pathway and modulation of ischemia-reperfusion injury in Type 2 diabetic BBZ rat hearts. Cardiovasc Diabet 7(1):1–11. https://doi.org/10.1186/1475-2840-7-33
Giacco F, Brownlee M (2010) Oxidative stress and diabetic complications. Circ Res 107(9):1058–1070. https://doi.org/10.1161/CIRCRESAHA.110.223545
Demir Y, Işık M, Gülçin İ, Beydemir Ş (2017) Phenolic compounds inhibit the aldose reductase enzyme from the sheep kidney. J Biochem Mol Toxicol 31(9):e21936. https://doi.org/10.1002/jbt.21935
Taslimi P, Aslan HE, Demir Y, Oztaskin N, Maraş A, Gulçin İ et al (2018) Diarylmethanon, bromophenol and diarylmethane compounds: discovery of potent aldose reductase, α-amylase and α-glycosidase inhibitors as new therapeutic approach in diabetes and functional hyperglycemia. Int J Biol Macromol. https://doi.org/10.1016/j.ijbiomac.2018.08.004
Listvan V, Listvan V, Shekel A (2002) Synthesis of cholesteryl esters of heterocyclic analogs of cinnamic acid and hetaroyloxycinnamic acids by the Wittig reaction. Chem Heterocycl Compd 38(12):1480–1483. https://doi.org/10.1023/A:1022693427914
Sharghi H, Razavi SF, Aberi M, Tavakoli F, Shekouhy M (2020) The Co2+ complex of [7-hydroxy-4-methyl-8-coumarinyl] glycine as a nanocatalyst for the synthesis and biological evaluation of new mannich bases of benzimidazoles and benzothiazoles. ChemistrySelect 5(9):2662–2671. https://doi.org/10.1002/slct.201904700
Khadilkar B, Jaisinghani H, Saraf M, Desai S (2001) Synthesis and pharmacological studies of new derivatives of dimethyl 1,4-dihydro-2,6-dimethyl-3, 5-pyridinedicarboxylate. Indian J Chem 40B:82–86
Sengupta A (1977) Synthesis of substituted piperazinyl semicarbazides and thiosemicarbazides as possible acetylcholinesterase (AChE) inhibitors. J Indian Chem Soc 54(10):961–964
Wu C, Anderson CE, Bui H, Gao D, Holland GW, Kassir J, et al (2004) Pyridine, pyrimidine, quinoline, quinazoline, and naphthalene urotensin-ii receptor antagonists. Google Patents
Di Braccio M, Grossi G, Alfei S, Ballabeni V, Tognolini M, Flammini L et al (2014) 1, 8-Naphthyridines IX. Potent anti-inflammatory and/or analgesic activity of a new group of substituted 5-amino [1, 2, 4] triazolo [4, 3-a][1, 8] naphthyridine-6-carboxamides, of some their Mannich base derivatives and of one novel substituted 5-amino-10-oxo-10H-pyrimido [1, 2-a][1, 8] naphthyridine-6-carboxamide derivative. Eur J Med Chem. https://doi.org/10.1016/j.ejmech.2014.08.069
Türkeş C, Demir Y, Beydemir Ş (2019) Anti-diabetic properties of calcium channel blockers: inhibition effects on aldose reductase enzyme activity. Appl Biochem Biotechnol 189(1):318–329. https://doi.org/10.1007/s12010-019-03009-x
Tokalı FS, Demir Y, Demircioğlu İH, Türkeş C, Kalay E, Şendil K et al (2021) Synthesis, biological evaluation, and in silico study of novel library sulfonates containing quinazolin-4 (3H)-one derivatives as potential aldose reductase inhibitors. Drug Dev Res. https://doi.org/10.1002/ddr.21887
Sever B, Altıntop MD, Demir Y, Türkeş C, Özbaş K, Çiftçi GA et al (2021) A new series of 2,4-thiazolidinediones endowed with potent aldose reductase inhibitory activity. Open Chem. https://doi.org/10.1515/chem-2021-0032
Akdağ M, Özçelik AB, Demir Y, Beydemir Ş (2022) Design, synthesis, and aldose reductase inhibitory effect of some novel carboxylic acid derivatives bearing 2-substituted-6-aryloxo-pyridazinone moiety. J Mol Struct. https://doi.org/10.1016/j.molstruc.2022.132675
Sever B, Altıntop MD, Demir Y, Pekdoğan M, Çiftçi GA, Beydemir Ş et al (2021) An extensive research on aldose reductase inhibitory effects of new 4H–1, 2, 4-triazole derivatives. J Mol Struct. https://doi.org/10.1016/j.molstruc.2020.129446
Sever B, Altıntop MD, Demir Y, Çiftçi GA, Beydemir Ş, Özdemir A (2020) Design, synthesis, in vitro and in silico investigation of aldose reductase inhibitory effects of new thiazole-based compounds. Bioorg Chem. https://doi.org/10.1016/j.bioorg.2020.104110
Bradford MM (1976) A rapid and sensitive method for the quantitation microgram quantities of a protein isolated from red cell membranes. Anal Biochem 72(1–2):248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Demir Y, Köksal Z (2020) Some sulfonamides as aldose reductase inhibitors: Therapeutic approach in diabetes. Arch Physiol Biochem. https://doi.org/10.1080/13813455.2020.1742166
Demir Y, Özaslan MS, Duran HE, Küfrevioğlu Öİ, Beydemir Ş (2019) Inhibition effects of quinones on aldose reductase: antidiabetic properties. Environ Toxicol Pharmacol. https://doi.org/10.1016/j.etap.2019.103195
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227(5259):680–685. https://doi.org/10.1038/227680a0
Demir Y, Duran HE, Durmaz L, Taslimi P, Beydemir Ş, Gulçin İ (2020) The influence of some nonsteroidal anti-inflammatory drugs on metabolic enzymes of aldose reductase, sorbitol dehydrogenase, and α-glycosidase: a perspective for metabolic disorders. Appl Biochem Biotechnol 190(2):437–447. https://doi.org/10.1007/s12010-019-03099-7
Demir Y, Taslimi P, Koçyiğit ÜM, Akkuş M, Özaslan MS, Duran HE et al (2020) Determination of the inhibition profiles of pyrazolyl–thiazole derivatives against aldose reductase and α-glycosidase and molecular docking studies. Arch Pharm (Weinheim, Ger) 353(12):2000118. https://doi.org/10.1002/ardp.202000118
Demir Y, Durmaz L, Taslimi P, Gulçin İ (2019) Antidiabetic properties of dietary phenolic compounds: Inhibition effects on α-amylase, aldose reductase, and α-glycosidase. Biotechnol Appl Biochem 66(5):781–786. https://doi.org/10.1002/bab.1781
Erdemir F, Celepci DB, Aktaş A, Gök Y, Kaya R, Taslimi P et al (2019) Novel 2-aminopyridine liganded Pd(II) N-heterocyclic carbene complexes: synthesis, characterization, crystal structure and bioactivity properties. Bioorg Chem. https://doi.org/10.1016/j.bioorg.2019.103134
Sever B, Altıntop MD, Demir Y, Yılmaz N, Çiftçi GA, Beydemir Ş et al (2021) Identification of a new class of potent aldose reductase inhibitors: Design, microwave-assisted synthesis, in vitro and in silico evaluation of 2-pyrazolines. Chem-Biol Interact. https://doi.org/10.1016/j.cbi.2021.109576
Demir Y (2020) Naphthoquinones, benzoquinones, and anthraquinones: molecular docking, ADME and inhibition studies on human serum paraoxonase-1 associated with cardiovascular diseases. Drug Dev Res 81(5):628–636. https://doi.org/10.1002/ddr.21667
Demir Y (2019) The behaviour of some antihypertension drugs on human serum paraoxonase-1: an important protector enzyme against atherosclerosis. J Pharm Pharmacol 71(10):1576–1583. https://doi.org/10.1111/jphp.13144
Askin S, Tahtaci H, Türkeş C, Demir Y, Ece A, Çiftçi GA et al (2021) Design, synthesis, characterization, in vitro and in silico evaluation of novel imidazo [2, 1-b][1, 3, 4] thiadiazoles as highly potent acetylcholinesterase and non-classical carbonic anhydrase inhibitors. Bioorg Chem. https://doi.org/10.1016/j.bioorg.2021.105009
Türkeş C (2019) Investigation of potential paraoxonase-I inhibitors by kinetic and molecular docking studies: chemotherapeutic drugs. Protein Pept Lett 26(6):392–402. https://doi.org/10.2174/0929866526666190226162225
Beydemir Ş, Türkeş C, Yalçın A (2021) Gadolinium-based contrast agents: in vitro paraoxonase 1 inhibition, in silico studies. Drug Chem Toxicol 44(5):508–517. https://doi.org/10.1080/01480545.2019.1620266
Türkeş C (2019) A potential risk factor for paraoxonase 1: in silico and in-vitro analysis of the biological activity of proton-pump inhibitors. J Pharm Pharmacol 71(10):1553–1564. https://doi.org/10.1111/jphp.13141
Işık M, Demir Y, Durgun M, Türkeş C, Necip A, Beydemir Ş (2020) Molecular docking and investigation of 4-(benzylideneamino)-and 4-(benzylamino)-benzenesulfonamide derivatives as potent AChE inhibitors. Chem Pap. https://doi.org/10.1007/s11696-019-00988-3
Akocak S, Taslimi P, Lolak N, Işık M, Durgun M, Budak Y et al (2021) Synthesis, characterization, and inhibition study of novel substituted phenylureido sulfaguanidine derivatives as α-glycosidase and cholinesterase inhibitors. Chem Biodivers 18(4):e2000958. https://doi.org/10.1002/cbdv.202000958
Istrefi Q, Türkeş C, Arslan M, Demir Y, Nixha AR, Beydemir Ş et al (2020) Sulfonamides incorporating ketene N, S-acetal bioisosteres as potent carbonic anhydrase and acetylcholinesterase inhibitors. Arch Pharm (Weinheim, Ger) 353(6):e1900383. https://doi.org/10.1002/ardp.201900383
Bochevarov AD, Harder E, Hughes TF, Greenwood JR, Braden DA, Philipp DM et al (2013) Jaguar: a high-performance quantum chemistry software program with strengths in life and materials sciences. Int J Quantum Chem 113(18):2110–2142. https://doi.org/10.1002/qua.24481
Sever B, Türkeş C, Altıntop MD, Demir Y, Beydemir Ş (2020) Thiazolyl-pyrazoline derivatives: in vitro and in silico evaluation as potential acetylcholinesterase and carbonic anhydrase inhibitors. Int J Biol Macromol. https://doi.org/10.1016/j.ijbiomac.2020.09.043
Türkeş C, Beydemir Ş (2020) Inhibition of human serum paraoxonase-I with antimycotic drugs: in vitro and in silico studies. Appl Biochem Biotechnol 190(1):252–269. https://doi.org/10.1007/s12010-019-03073-3
Taslimi P, Işık M, Türkan F, Durgun M, Türkeş C, Gülçin İ et al (2021) Benzenesulfonamide derivatives as potent acetylcholinesterase, α-glycosidase, and glutathione S-transferase inhibitors: biological evaluation and molecular docking studies. J Biomol Struct Dyn 39(15):5449–5460. https://doi.org/10.1080/07391102.2020.1790422
Kilic A, Beyazsakal L, Işık M, Türkeş C, Necip A, Takım K et al (2020) Mannich reaction derived novel boron complexes with amine-bis(phenolate) ligands: synthesis, spectroscopy and in vitro/in silico biological studies. J Organomet Chem. https://doi.org/10.1016/j.jorganchem.2020.121542
Türkeş C, Arslan M, Demir Y, Cocaj L, Nixha AR, Beydemir Ş (2019) Synthesis, biological evaluation and in silico studies of novel N-substituted phthalazine sulfonamide compounds as potent carbonic anhydrase and acetylcholinesterase inhibitors. Bioorg Chem. https://doi.org/10.1016/j.bioorg.2019.103004
Demir Y, Ceylan H, Türkeş C, Beydemir Ş (2021) Molecular docking and inhibition studies of vulpinic, carnosic and usnic acids on polyol pathway enzymes. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2021.1967195
Türkeş C, Akocak S, Işık M, Lolak N, Taslimi P, Durgun M et al (2021) Novel inhibitors with sulfamethazine backbone: synthesis and biological study of multi-target cholinesterases and α-glucosidase inhibitors. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2021.1916599
Zhang L, Zhang H, Zhao Y, Li Z, Chen S, Zhai J et al (2013) Inhibitor selectivity between aldo–keto reductase superfamily members AKR1B10 and AKR1B1: role of Trp112 (Trp111). FEBS Lett 587(22):3681–3686. https://doi.org/10.1016/j.febslet.2013.09.031
Madhavi Sastry G, Adzhigirey M, Day T, Annabhimoju R, Sherman W (2013) Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Comput-Aided Mol Des 27(3):221–234. https://doi.org/10.1007/s10822-013-9644-8
Gündoğdu S, Türkeş C, Arslan M, Demir Y, Beydemir Ş (2019) New isoindole-1, 3-dione substituted sulfonamides as potent inhibitors of carbonic anhydrase and acetylcholinesterase: design, synthesis, and biological evaluation. ChemistrySelect 4(45):13347–13355. https://doi.org/10.1002/slct.201903458
Yaşar Ü, Gönül İ, Türkeş C, Demir Y, Beydemir Ş (2021) Transition–metal complexes of bidentate Schiff-base ligands: in vitro and in silico evaluation as non-classical carbonic anhydrase and potential acetylcholinesterase inhibitors. ChemistrySelect 29(6):7278–7284. https://doi.org/10.1002/slct.202102082
Türkeş C, Demir Y, Beydemir Ş (2021) Calcium channel blockers: molecular docking and inhibition studies on carbonic anhydrase I and II isoenzymes. J Biomol Struct Dyn 39(5):1672–1680. https://doi.org/10.1080/07391102.2020.1736631
Sever B, Türkeş C, Altıntop MD, Demir Y, Çiftçi GA, Beydemir Ş (2021) Novel metabolic enzyme inhibitors designed through the molecular hybridization of thiazole and pyrazoline scaffolds. Arch Pharm (Weinheim, Ger) 354(12):e2100294. https://doi.org/10.1002/ardp.202100294
Işık M, Akocak S, Lolak N, Taslimi P, Türkeş C, Gülçin İ et al (2020) Synthesis, characterization, biological evaluation, and in silico studies of novel 1,3-diaryltriazene-substituted sulfathiazole derivatives. Arch Pharm (Weinheim, Ger) 353(9):e2000102. https://doi.org/10.1002/ardp.202000102
Friesner RA, Murphy RB, Repasky MP, Frye LL, Greenwood JR, Halgren TA et al (2006) Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein−ligand complexes. J Med Chem 49(21):6177–6196. https://doi.org/10.1021/jm051256o
Demir Y, Türkeş C, Beydemir Ş (2020) Molecular docking studies and inhibition properties of some antineoplastic agents against paraoxonase-I. Anti-Cancer Agents Med Chem 20(7):887–896. https://doi.org/10.2174/1871520620666200218110645
Türkeş C, Demir Y, Beydemir Ş (2022) Some calcium-channel blockers: kinetic and in silico studies on paraoxonase-I. J Biomol Struct Dyn 40(1):77–85. https://doi.org/10.1080/07391102.2020.1806927
Türkeş C, Kesebir Öztürk A, Demir Y, Küfrevioğlu Öİ, Beydemir Ş (2021) Calcium channel blockers: the effect of glutathione S-transferase enzyme activity and molecular docking studies. ChemistrySelect 6(40):11137–11143. https://doi.org/10.1002/slct.202103100
Kalaycı M, Türkeş C, Arslan M, Demir Y, Beydemir Ş (2021) Novel benzoic acid derivatives: synthesis and biological evaluation as multitarget acetylcholinesterase and carbonic anhydrase inhibitors. Arch Pharm (Weinheim, Ger) 354(3):2000282. https://doi.org/10.1002/ardp.202000282
Osmaniye D, Türkeş C, Demir Y, Özkay Y, Beydemir Ş, Kaplancıklı ZA (2022) Design, synthesis, and biological activity of novel dithiocarbamate-methylsulfonyl hybrids as carbonic anhydrase inhibitors. Arch Pharm (Weinheim Ger). https://doi.org/10.1002/ardp.202200132
Çalışkan B, Demir Y, Türkeş C (2021) Ophthalmic Drugs: In vitro paraoxonase 1 inhibition and molecular docking studies. Biotechnol Appl Biochem. https://doi.org/10.1002/bab.2284
Güleç Ö, Türkeş C, Arslan M, Demir Y, Yeni Y, Hacımüftüoğlu A et al (2022) Cytotoxic effect, enzyme inhibition, and in silico studies of some novel N-substituted sulfonyl amides incorporating 1,3,4-oxadiazol structural motif. Mol Divers. https://doi.org/10.1007/s11030-022-10422-8
Türkeş C, Demir Y, Beydemir Ş (2021) Infection medications: assessment in-vitro glutathione S-transferase inhibition and molecular docking study. ChemistrySelect 6(43):11915–11924. https://doi.org/10.1002/slct.202103197
Barreiro G, Guimarães CR, Tubert-Brohman I, Lyons TM, Tirado-Rives J, Jorgensen WL (2007) Search for non-nucleoside inhibitors of HIV-1 reverse transcriptase using chemical similarity, molecular docking, and MM-GB/SA scoring. J Chem Inf Model 47(6):2416–2428. https://doi.org/10.1021/ci700271z
Sasikumar G, Arulmozhi S, Ashma A, Sudha A (2019) Mixed ligand ternary complexes of Co(II), Ni(II), Cu(II) and Zn(II) and their structural characterization, electrochemical, theoretical and biological studies. J Mol Struct. https://doi.org/10.1016/j.molstruc.2019.03.031
Deswal Y, Asija S, Dubey A, Deswal L, Kumar D, Jindal DK et al (2022) Cobalt(II), nickel(II), copper(II) and zinc(II) complexes of thiadiazole based Schiff base ligands: synthesis, structural characterization, DFT, antidiabetic and molecular docking studies. J Mol Struct. https://doi.org/10.1016/j.molstruc.2021.132266
Yapar G, Duran HE, Lolak N, Akocak S, Türkeş C, Durgun M et al (2021) Biological effects of bis-hydrazone compounds bearing isovanillin moiety on the aldose reductase. Bioorg Chem. https://doi.org/10.1016/j.bioorg.2021.105473
Maccari R, Ottanà R, Curinga C, Vigorita MG, Rakowitz D, Steindl T et al (2005) Structure–activity relationships and molecular modelling of 5-arylidene-2, 4-thiazolidinediones active as aldose reductase inhibitors. Bioorg Med Chem 13(8):2809–2823. https://doi.org/10.1016/j.bmc.2005.02.026
Alexiou P, Nicolaou I, Stefek M, Kristl A, Demopoulos VJ (2008) Design and synthesis of N-(3, 5-difluoro-4-hydroxyphenyl) benzenesulfonamides as aldose reductase inhibitors. Bioorg Med Chem 16(7):3926–3932. https://doi.org/10.1016/j.bmc.2008.01.042
Acknowledgements
This work was supported by the Research Fund of Ardahan University (Grant Number 2019-008), the Research Fund of Erzincan Binali Yıldırım University (Grant Number FBA-2017-501), and the Research Fund of Anadolu University (Grant Number 2102S003).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Demir, Y., Tokalı, F.S., Kalay, E. et al. Synthesis and characterization of novel acyl hydrazones derived from vanillin as potential aldose reductase inhibitors. Mol Divers 27, 1713–1733 (2023). https://doi.org/10.1007/s11030-022-10526-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-022-10526-1