Abstract
The human P-glycoprotein (P-gp) efflux pump is of great interest for medicinal chemists because of its important role in multidrug resistance (MDR). Because of the high polyspecificity as well as the unavailability of high-resolution X-ray crystal structures of this transmembrane protein, ligand-based, and structure-based approaches which were machine learning, homology modeling, and molecular docking were combined for this study. In ligand-based approach, individual two-dimensional quantitative structure–activity relationship models were developed using different machine learning algorithms and subsequently combined into the Ensemble model which showed good performance on both the diverse training set and the validation sets. The applicability domain and the prediction quality of the developed models were also judged using the state-of-the-art methods and tools. In our structure-based approach, the P-gp structure and its binding region were predicted for a docking study to determine possible interactions between the ligands and the receptor. Based on these in silico tools, hit compounds for reversing MDR were discovered from the in-house and DrugBank databases through virtual screening using prediction models and molecular docking in an attempt to restore cancer cell sensitivity to cytotoxic drugs.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Human P-glycoprotein (P-gp), encoded by the ABCB1/MDR1 gene, is one of the most important and widely studied members of the ATP-Binding Cassette (ABC) transporter superfamily [1, 2]. The energy-dependent efflux activity with broad substrate specificity (natural products, anticancer agents, peptides, steroids, lipids, cytokines, dyes, and ionophores) as well as the widespread distribution in normal tissues (intestine, brain, testis, placenta, liver, and kidney) and tumors of this protein support its roles in human physiology and drug therapy [3]. Taking part in the physiological defense mechanism against xenobiotics such as toxins and drugs, P-gp should be considered as an antitarget in company with hERG (human ether-a-go-go related gene), Cytochrome P450s, and PXR (pregnane X-receptor) in the drug discovery and development process [4, 5]. Blocking this nontarget with inhibitors (e.g., ketoconazole) could change the concentration of a co-administered drug or its metabolite in blood (e.g., terfenadine) and consequently lead to drug–drug interactions and unwanted side effects (e.g., QT prolongation/Torsades de Pointes) [6]. Conferring a multidrug resistance (MDR) phenotype of cancer cells through active outward transport of cytotoxic drugs, P-gp has been also counted as a clinical target in chemotherapy [7]. P-gp inhibition is subjected to overcome MDR in cancer patients by increasing the intracellular accumulation of substrate drugs and thus their cytotoxicity [8].
With regard to structure, P-gp is a 170 kDa transmembrane protein [9] formed by two symmetrical halves: N-terminal and C-terminal. Each half of the molecule contains six transmembrane (TM) domains, followed by a nucleotide-binding domain (NBD). The inward facing conformation (which is stabilized by TMs 4, 5, 10, and 11) makes up a large internal cavity (\(\sim \)6,000 Å\(^{3})\) opened to both the cytoplasm and the inner leaflet for multispecific drug binding [10]. Until now, ABCB10 is the only human ABC transporter resolved for structure-based approaches, apart from X-ray structures of several other ABC transporters from prokaryotes (e.g., bacteria) and eukaryotic organisms (e.g., mice) [11, 12]. To deal with the unavailability of its crystallographic 3D structures at high-resolution, homology models of P-gp were generated using the resolved relevant structures as templates. For example, the recent work of Ambudkar et al. [13] revealed multiple active-binding sites for substrates and modulators including one primary site in a large flexible pocket in the TM domains and secondary sites from the combination of homology modeling, molecular docking, site-directed mutagenesis, and cell- and membrane-based assays.
The concept of using small molecule inhibitors (SMIs) of P-gp for resensitization of tumor cells to chemotherapeutic drugs has been universally acknowledged [14]. Over the last three decades, three generations of P-gp inhibitors have been developed and examined in preclinical and clinical studies [15]. However, no clinically used drugs for P-gp blockade are yet available and the failure of candidates may be attributed to their unfavorable properties (e.g., poor solubility, poor specificity, and toxicity) and pharmacokinetic interactions [15–19]. In this context, flavonoids (polyphenolic compounds) that are ubiquitous in nature have emerged as reversal agents of the efflux pump-mediated MDR because of their advantages, including dual effects (P-gp modulation and antitumor activity) and safety and formed the third generation, nonpharmaceutical category of P-gp inhibitors [15, 20–24]. Chemical modifications on flavonoids of herbal origin have been implemented for structure–activity relationship (SAR) studies. For example, Ecker et al. [25] have recently reported a series of new synthetic chalcones with good biological evaluation results and indicated the importance of specific groups for P-gp inhibitory activity based on 2D- and 3D-QSAR analyses.
Computer-aided drug design (CADD) methods including QSAR, molecular modeling, and docking are helpful to predict and elucidate ligand–protein interactions in the early stage of the drug discovery process [12, 26]. For this reason, the present study aimed to develop high-performance 2D-QSAR models (individual and ensemble) for predicting P-gp inhibitory activity. In addition, we endeavored to identify binding sites as well as plausible binding modes into P-gp homology model of the compounds having the best predicted pIC\(_{50}\) values from two libraries of in-house chalcones and drug-bank compounds by means of docking.
Materials and methods
The research process including machine learning, homology modeling, and molecular docking is summarized as shown in Fig. 1 and described in detail as follows:
Data sources
Databases for predicting P-gp inhibitory activity
As proven in the work of Ecker et al. [27], it was impossible to create a large chemical diverse dataset of P-gp inhibitors whose bioactivity values derived from different assays. For this reason, three SAR datasets based on the same daunorubicin efflux assay in MDR CCRF vcr1000 cells were compiled and combined: (i) 198 chemical compounds from the publication of Ecker et al. [27]; (ii) 159 tested compounds from the PubChem BioAssay with Assay Identification (AID) number 281137, comprising 141 active compounds and 18 unspecified compounds; and (iii) 303 tested compounds from the PubChem BioAssay with AID number 781331, comprising 274 active compounds and 29 unspecified compounds. In fact, they were all presented in the ChEMBL open source database [28]. In addition, 22 chalcone derivatives whose IC\(_{50}\) values were measured in a daunomycin efflux inhibition assay using the mentioned cell line were also collected from another publication of Ecker et al. [25] to broaden the chemical space of QSAR models. These four datasets were merged using MOE 2008.10 [29]. After inspection, two duplicates (CID: 13504844 and CID: 9976755) in the AID 281137 database having their structures identical to those of two other compounds (CID: 73357260 and CID: 73346637) in the AID 781331 database were removed. As a result of this process, a final dataset of 499 compounds with chemical structures and bioactivity values (IC\(_{50})\) was obtained for the generation of prediction models (Supplementary Material 2, SM2-DB1). The negative logarithm of IC\(_{50}\) (pIC\(_{50})\) was used as the dependent variable. Among them, we randomly took 99 compounds (20 %) as an external test set to assess the extrapolation capacity of generated models.
Databases for virtual screening
In this work, we utilized an in-house library of 87 chalcones for in silico screening to seek novel efflux pump inhibitors which might help to overcome MDR in tumors (Supplementary Material 2, SM2-DB2). Furthermore, another library of 6874 compounds belonging to six groups, namely approved, experimental, investigational, nutraceutical, withdrawn, and illicit, was downloaded from DrugBank [30–33] for the same purpose to exploit existing drugs (Supplementary Material 2, SM2-DB3).
Attribute calculation and selection
The 2D structures were built in ChemBioDrawUltra 12.0 [34] if not available and subsequently energy minimized in MOE [29] before attribute calculation. Chemical descriptors converting chemical compounds into descriptor vectors to be understood by computers are of great importance in predicting protein–ligand interactions [35]. On one hand, 184 2D molecular descriptors which are physicochemical properties, subdivided surface areas, atom and bond counts, Kier&Hall connectivity and Kappa shape indices, adjacency and distance matrices, pharmacophoric features, and partial charges were computed for 499 database compounds using MOE [29]. In addition, their 1444 1D-2D descriptors representing 63 different molecular properties were also calculated using PaDEL-Descriptor 2.21 [36].
Feature selection aims to remove redundant or irrelevant features for improving model quality and reducing computational cost [37]. First of all, the compounds with no missing attributes were filtered using RapidMiner 5.3.008 [38]. Subsequently, removing useless and correlated attributes and optimizing selection by a genetic algorithm (GA) were also implemented in RapidMiner [38]. Finally, the search method BestFirst in Weka 3.7.9 [39] that searches the space of attribute subsets by greedy hill-climbing augmented with a backtracking facility was used to select attributes with 10-fold cross validation. All parameters in the variable selection process were set to default.
Data partition into training and test sets
For internal validation, the database of 400 remaining compounds was divided into the training set (75 %) and the internal test set (25 %) using two tools in MOE [29], Rand and Diverse Subset. The Rand function assigning a random number between 0 and 1 to each compound was used to split the database randomly. In contrast, the Diverse Subset application assigning a rank to each compound by a ranking process whose methodology can be found in detail in the software was used to determine the farthest ones in the study population for the diverse training set.
Applicability domain determination
To predict a new compound truly using a QSAR model, determining the applicability domain (AD) is necessary. This term was interpreted in different ways [40–42] but it could be simply understood as “the response and chemical structure space in which the QSAR model makes predictions with a given reliability” [43]. There are available methods for defining the domain of applicability such as Ranges in the descriptor space; Geometrical methods; Distance-based methods; Probability density distribution; and Range of the response variable and Miscellaneous [40, 42, 44]. Recently, a new simple method has been suggested by Roy et al. [43] to identify the X-outliers in case of training set and the compounds outside the AD in case of the test set. Its principle is based on the theory of standardization approach, considering mean ±3 standard deviation (SD) as the zone of most training set compounds (99.7 %) and the remainder as the zone of dissimilar compounds. Following this method, an X-outlier (if in the training set) or outside AD (if in the test set) could be specified by computing the standardized descriptor \({S}_{\mathrm{i(k)}}\), the maximum \({S}_{\mathrm{i(k)}}\) value \(([{S}_{\mathrm{i}}]_{\mathrm{max(k)}})\), the minimum \({S}_{\mathrm{i(k)}}\) value \(([{S}_\mathrm{i}]_{\mathrm{min(k)}})\) (if necessary), the \({S}_{\mathrm{new(k)}}\) (if necessary) and comparing them with 3 (threshold value). In the present study, we employed the software “Applicability domain using standardization approach” also developed by Roy et al. [43] to execute this process.
Machine learning methods
Machine learning provides the technical basis of data mining for a variety of purposes. In drug discovery and development, machine learning tools and techniques are being increasingly applied to predict pharmacodynamic (inhibitor, substrate, antagonist, agonist, blocker, and activator) and pharmacokinetic (ADMET) properties of the compounds [45]. In the present study, we used two nodes, Numeric Predictor and Ensemble in Clementine 12.0 [46], for predicting P-gp inhibitory activity. The Numeric Predictor node can estimate the models for a maximum of six machine learning algorithms, viz Neural Network, C&R Tree, CHAID, Regression, Generalized Linear, and Support Vector Machine (SVM). And Ensemble is the node used for combining predictions from the most accurate models to avoid limitations of individual models and achieve a higher overall accuracy. The principles and the application examples of these methods were described in detail elsewhere [37, 46, 47].
Model evaluation
In this study, statistical parameters and validation criteria used to measure the QSAR model performance for predictivity were presented as follows:
Internal validation
The internal quality of models is judged by the squared correlation coefficient \({R}^{2}\) and the cross-validated squared correlation coefficient \({Q}^{2}\). Leave-one-out (LOO) is a particular case of k-fold cross validation where k equals the number of compounds in the data [48]. In the case of LOO cross validation, the \({Q}^{2}\) metric [49] is calculated according to the following formula:
In Eq. (1), \({y}_{i} \), and \({\hat{{y}}}_{i} \) are, respectively, the observed and predicted activity values and \({\bar{{y}}} \) is the mean value of \({y}_{i} \) in the training set. For an acceptable model, \({R}^{2} \ge 0.7, {Q}_{\mathrm{LOO}}^2 \ge 0.6\) and \(\left| {{R}^{2}-{Q}_{\mathrm{LOO}}^2 } \right| \le 0.1\) are required [50–53].
For further internal validation, model randomization or y-randomization was implemented on the training set. In this process, the values of the dependent variable (y) were randomly scrambled 10 times and new models were developed from each y-randomized training set. The \({R}_\mathrm{p}^2 \) metric proposed by Roy et al. [49] to ensure the models not to be developed by chance is calculated by the following formula:
In Eq. (2), \({R}^{2}\) and \({R}_\mathrm{r}^2 \) are the squared correlation coefficient of the nonrandomized model and the squared mean correlation coefficient of randomized models, respectively. For a predictive QSAR model, the value of \({R}_\mathrm{p}^2 \) should be more than 0.5 [49].
External validation
The more widely used parameters \({Q}_{\mathrm{F}1}^2 \)[54]; \({Q}_{\mathrm{F}2}^2 \) [55]; \({Q}_{\mathrm{F}3}^2 \) [56, 57]; \({r}_\mathrm{m}^2 ;{\overline{{r}_\mathrm{m}^2}} ; \Delta {r}_\mathrm{m}^2 \) [58–60]; and CCC (concordance correlation coefficient) [61] were applied to externally validate the models for their predictions on the compounds not involving the model development. Apart from them, mean absolute error (MAE) was also employed to assess the predictability on the external test set [57]. These metrics are calculated based on the following formulas:
In Eqs. (3), (4), (5), (10) and (11), \({y}_{i} \) and \({\hat{{y}}} _{i} \) are, respectively, the observed and predicted activity values, while \({\bar{{y}}} \) and \({\bar{\hat{{y}}} }\) are, respectively, the mean values of \({y}_{i} \) and \({\hat{{y}}}_{i} \). In Eq. (6) and (7), \({r}^{2}\) and \({r}_0^2 \) are, respectively, the determination coefficients in the regression function with and without intercept in case of using experimental data on the y-axis and predicted data on the x-axis, while \({r}^{\prime 2}\) and \({r}^{\prime 2}_0 \) are, respectively, the same coefficients in the opposite case. The most stringent validation criteria thresholds including \({Q}_{\mathrm{Fn}}^2 \ge 0.7; {\overline{{{r}_\mathrm{m}^2}}} \ge 0.65\); CCC \(\ge 0.85\) [62, 63]; \({r}_\mathrm{m}^2 \ge 0.5\) [49]; and \(\Delta {r}_\mathrm{m}^2 \le 0.2\) [64] were applied to verify the external predictivity of good models. Using the MAE-based criteria lately proposed by Roy et al. [65], the predictions could be considered good when MAE \(\le \)0.1 \(\times \) training set range and MAE + 3 \(\sigma \le \) 0.2 \(\times \) training set range and bad when MAE > 0.15 \(\times \) training set range or MAE + 3 \(\sigma>\) 0.25 \(\times \) training set range. In this study, we employed the software “XternalValidationPlus” also developed by Roy et al. [65] to execute this process.
Homology modeling
Due to the lack of high-resolution crystal structures of transporter proteins such as P-gp, homology modeling is considered as a feasible solution to obtain their structural information [66]. In the present study, the I-TASSER (Iterative Threading ASSEmbly Refinement) server which is one freely available online system for automated protein 3D structure prediction [67, 68] was employed to generate homology or comparative models of P-gp for docking studies. The I-TASSER methodology based on the state-of-the-art algorithms was described in detail elsewhere [69, 70] and summarized in three stages: (i) multiple threading, (ii) structural assembly, and (iii) model selection and refinement [68]. Before running, the P-gp sequence of 1280 amino acids was submitted in FASTA format [71] without assigning additional restraints and templates. The output including the confidence score (C-score), the template modeling score (TM-score), the root mean square deviation (RMSD), the number of decoys, and the cluster density was provided by the server for quantitative assessments of the predictions (models).
C-score is an estimate of the confidence of structure prediction. In a typical range of [−5, 2], the higher value of C-score indicates the better quality and the C-score value \(-\)1.5 indicates the correct topology of predicted model. RMSD is an average distance of all residue pairs in the predicted structure and the native structure, ranging between 1 and 2 Å for high-resolution models and between 2 and 5 Å for medium resolution models. Since RMSD might be influenced by a local error, TM-score which also measures the structural similarity between two structures is proposed to solve this problem. TM-score <0.17 means random predictions and TM-score >0.5 means correct topology for all sizes of proteins [72]. The cluster density is the number of structure decoys (low temperature replicas) at a unit of space in the SPICKER cluster and a higher cluster density means a better quality model.
In addition to the above parameters, the stereochemical quality of the final model was checked using PROCHECK [73]. Our structure in PDB format was uploaded to PDBsum to produce its Ramachandran plot of the phi–psi torsion angles for all residues in the structure, except those at the chain termini. Because glycine residues are not restricted to any particular region of the plot, they are separately identified by triangles. Based on an analysis of 118 structures of resolution of at least 2.0 Å and R-factor no greater than 20.0, a good quality model would be expected to have over 90 % in the most favored regions [A,B,L] (also called core regions).
Molecular docking
Ligand and protein preparation
Both ligands and protein were prepared to be used for docking. The 2D structures of the ligands were built in ChemBioDrawUltra [34] if not available and subsequently energy minimized in MOE [29]. The best P-gp homology model in complex with ligand was protonated, tethered, and minimized by the LigX tool in MOE [29] and the ligand was subsequently removed. The binding site of the target protein was also predicted by the I-TASSER server based on the similar binding site of template protein.
Docking
Potential P-gp inhibitors which were previously identified before by the QSAR models from two virtual screening databases (in-house and drug-bank) were docked into the ligand-binding pocket of P-gp homology model using the FlexX package in LeadIT 2.0.2 [74] to provide insights into molecular recognition via protein–ligand interactions. In this process, the triangle matching algorithm was chosen for the place base fragment, while the maximum numbers of solutions per iteration and per fragmentation were defined as 1000 and 200, respectively. Compounds having both good predicted pIC\(_{50}\) values and docking scores were more likely to be the hits for P-gp inhibitory activity.
Results and discussion
2D-QSAR models
A total of 1628 MOE and PaDEL descriptors were initially computed for the whole dataset of 499 compounds. Among them, five descriptors (Kier1, Kier2, Kier3, apol, and bpol) which were determined as duplicates were eliminated. All 400 training and internal test set compounds with no missing attributes were used for selecting features. The feature reduction in RapidMiner, including raw filter by the Remove Useless Attributes and Remove Correlated Attributes operators, and GA-based optimization by the Optimize Selection (Evolutionary) operator resulted in 383 attributes. This number of attributes which was still abundant for the model development was continuously reduced to 89 attributes with number of folds \(\ge \)10 % by the BestFirst algorithm in Weka. However, only the 34 most relevant attributes with number of folds \(\ge \)80 % (Supplementary Material 1, SM1-Table 1) were chosen to generate the machine learning models. All parameters in the variable selection process were set to the default.
Out of the selected attributes, the ones computed by PaDEL were in the majority compared with the ones computed by MOE (27/7) to represent for the whole dataset. MOE descriptors belonged to three classes: Adjacency and Distance Matrix Descriptors (2), Atom Counts and Bond Counts (1), and Partial Charge Descriptors (4), while PaDEL descriptors belonged to 10 classes: Autocorrelation (4); Barysz matrix (4); Burden modified eigenvalues (9); Atom type electrotopological state (4); Information content (1); Longest aliphatic chain (1); Molecular linear free energy relation (1); Ring count (1); Topological charge (1) and Topological distance matrix (1).
As mentioned above, the Numeric Predictor node with integrated machine learning algorithms was put in a simple stream in Clementine for automated modeling of the relationship between chemical structure (descriptors) and P-gp inhibitory activity (a numeric range target), using 300 training set compounds derived from the diverse and random partitions. Based on the 34 chosen descriptors, all six candidate models as mentioned in the machine learning methods (Neural Network, C&R Tree, CHAID, Regression, Generalized Linear and SVM) were created by default to predict these continuous numeric outcomes for each type of division (Supplementary Material 1, SM1-Table 2). Having the same results in both separation cases, only one (Regression) of the two models (Regression and Generalized Linear) was chosen for further analysis.
The generated models were compared together using the squared correlation coefficient (\({R}^{2})\) which indicates a stronger relationship when being closer to 1. In the diverse partition, the \({R}^{2}\) values in the training set were less than those in the test set in cases of the Regression, SVM, and Neural Network models and vice versa in cases of the CHAID and C&R Tree models. In the random partition, the \({R}^{2}\) value in the training set was less than that in the test set in case of the Neural Network model and vice versa in cases of the CHAID, C&R Tree, Regression, and SVM models. Furthermore, although the \({R}^{2}\) values of all models in the diverse division were approximate to those in the random division in case of the training set, most of these values in the diverse division were considerably greater than those in the random division in case of the test set. The obtained results demonstrated the diverse training set was more appropriate than the random training set to be used for the development of machine learning models.
Based on the \({R}^{2}\) values, the CHAID and C&R Tree models ranked best in both diverse and random training sets (85.80, 85.91, 82.56, and 84.00 %, respectively), while two pairs of models SVM, C&R Tree and Neural Network, C&R Tree performed best in the diverse and random internal tests, respectively (81.21, 81.20, 74.14, and 73.69 %, respectively). The greatest difference in \({R}^{2}\) between the training and internal test sets was found in case of CHAID with the random partition. Conversely, there was no remarkable difference in this metric between the diverse and random training sets as well as the diverse and random internal test sets in case of Neural Network. To avoid such limitations of individual models, the solution of combining predictions from multiple models was put forward in our study.
Apart from Generalized Linear, the remaining models were combined into a single aggregated model using the Ensemble node. To compare each of the individual models with the Ensemble model, the option “Filter out fields generated by ensembled models” was not selected. The ensemble scores for this range target were generated by averaging the scores for the individual models. One frequent benefit of this approach is the ability to yield more accurate predictions than can be gained from any one model [46]. Indeed, the Ensemble model with its aggregated predictions gained the high performance in the diverse training set (\({R}^{2} \approx 0.84, {Q}_{\mathrm{LOO}}^2 \approx 0.70, \left| {{R}^{2}-\hbox {Q}_{\mathrm{LOO}}^2 } \right| \approx 0.14, {r}_\mathrm{m}^2 \approx 0.80, {r}_\mathrm{m}^{{\prime }2} \approx 0.64, \overline{{r}_\mathrm{m}^2 }\approx 0.72\), and \(\Delta {r}_\mathrm{m}^2 \approx 0.16\)), while the individual models failed at least one of the mentioned validation criteria, viz \({Q}_{\mathrm{LOO}}^2 \) and \(\left| {{R}^{2}-{Q}_{\mathrm{LOO}}^2 } \right| \) in the CHAID and C&R Tree models; \(\overline{{r}_\mathrm{m}^2 }\) in the Regression, SVM, and Neural Network models, and \(\Delta {r}_\mathrm{m}^2 \) in the Regression model (Table 1). All six models had \(\left| {{R}^{2}-{R}_{\mathrm{Yi}}^2 } \right| \ge 0.2\) and met the criterion of \({R}_\mathrm{p}^2 (\ge \)0.5) in the y-randomization process (Table 1). Despite being not quite as well as the individual models in some concrete conditions, the combination of multiple models performed as the best in overall. The Ensemble model showed itself as robust one which is likely to perform better when applied to other datasets in general terms, without having to dig deeply into the specifics of any one model.
The QSAR models were subsequently validated by the internal and external test sets. For the internal test set, the Ensemble model continued performing as the best with \({Q}_{\mathrm{F}1}^2 \approx 0.83, {Q}_{\mathrm{F}2}^2 \approx 0.83, {Q}_{\mathrm{F}3}^2 \approx 0.81, {r}_\mathrm{m}^2 \approx 0.80, {r}_{\mathrm{m}}^{{\prime }2} \approx 0.64, \overline{{r}_\mathrm{m}^2 } \approx 0.72, \Delta {r}_\mathrm{m}^2 \approx 0.16\) and CCC \(\approx \) 0.90 (Table 2). For the external test set, the best model of all was surprisingly SVM whose predictions showed the square value of the correlation coefficient (\({R}^{2} \approx 0. 74\)) only higher than that of Neural Network (\({R}^{2} \approx 0.73\)) on the diverse training set (Table 1). In this case, the Ensemble model showed its performance that was comparable to the best model SVM with \({Q}_{\mathrm{F}1}^2 \approx 0.83, {Q}_{\mathrm{F}2}^2 \approx 0.82, {Q}_{\mathrm{F}3}^2 \approx 0.83, {r}_{\mathrm{m}}^2 \approx 0.82, {r}_\mathrm{m}^{{\prime }2} \approx 0.67, \overline{{r}_\mathrm{m}^2} \approx 0.74, \Delta {r}_{\mathrm{m}}^2 \approx 0.16\), and CCC \(\approx 0.90\) (Table 3). These results proved the benefits of combining multiple models for handling diverse problems in biological data modeling.
Gramatica and Chirico [63] in their quantitative analysis revealed the importance of evaluating the scatter plot of experimental and predicted data to discover unacceptable QSAR models hidden by good statistical values. The plots of regression of predicted \(\mathrm{pIC}_{50}\) values by observed \(\mathrm{pIC}_{50}\) values on P-gp for the Ensemble model are displayed with error bars in Fig. 2 in cases of the training set (a), the internal test set (b), and the external test set (c). The points clustered along the diagonal visually indicated a good correlation between observed and predicted values for all the datasets. In other words, the generated Ensemble model was a good fit for these data.
For decades, the ligand-based approaches (SAR, 2D-QSAR, 3D-QSAR, and pharmacophore modeling) have been mainly employed for the discovery and design of novel and potential P-gp inhibitors when the structure of this protein has still not been resolved [26]. The molecular mechanisms of MDR modulation in general and P-gp inhibition in particular not being yet fully understood until now have been challenging QSAR studies of P-gp inhibitors [75]. Apart from binary classifiers for predicting P-gp inhibition (inhibitor: 1; noninhibitor: 0), many other 2D-QSAR studies [25, 76–86] treating P-gp inhibitory activity as a range target (e.g., \(\mathrm{pIC}_{50}, {pK}_{\mathrm{d}})\) were also published in the past and summarized in Table 4. Using different methods from simple (multiple linear regression) to complex (neural networks, support vector machine, hybrid), these studies showed satisfactory statistical results and contributed to determine structural features necessary for P-gp- mediated MDR reversal. Nevertheless, the applicability of the reported models to other datasets is questionable because of their limited numbers of investigated molecules.
In our study, the AD determination process resulted in 7 outliers (C89, C288, C381, C411, C422, C433, and C472) among 300 diverse training set compounds; 1/100 internal test set compounds (C358); and 2/99 external test set compounds (C40, C407) residing outside the AD. Without excluding these compounds, two individual models (Regression, SVM) and the Ensemble model were categorized as “good” and three individual models (CHAID, C&R Tree, Neural Network) were categorized as “moderate” based on the estimation of MAE-based metrics after removing 5 % external test set objects with high residual values (Table 5; Supplementary Material 2, SM2-MAE). The problems related to classical \({R}^{2}\)-based metrics and CCC (their values which are influenced by the range and the distribution of response values around the training/test set mean of the test set compounds) may lead to the wrong conclusion on the model acceptability [65]. By determining the allowable error limit using the training set response range, the MAE-based criteria suggested by Roy et al. [65] help to examine the predictive error values. The determined AD and the checked MAE-based criteria in this study showed the internal and external test set compounds lied in both chemical and response domains of the training set. In other words, the “good” predictability of our QSAR models was asserted with higher confidence by the combination of conventional and new validation methods.
Based on the same experimental assay, cell line, and cytotoxic agent, our data from different sources that are relatively comparable were merged to increase the database size and ensure sufficient structural diversity for the model development. Strict validation criteria also helped to confirm the high quality of the developed QSAR models, especially the Ensemble model which averaged the scores of the remaining models. From obtained results, this aggregated model is expected as a promising in silico tool which could be subjected to high throughput screening to yield rapid and accurate predictions for P-gp inhibitory potency of candidate drugs. Additionally, there was a good agreement between the present work and the previously published works in important molecular physicochemical properties for P-gp modulation such as the surface area, the lipophilicity, the longest aliphatic chain, and the positive charge (Table 4) that were represented by PEOE_VSA_FHYD, PEOE_VSA_FPNEG, PEOE_VSA_FPPOS, LipoaffinityIndex, nAtomLAC, and PEOE_RPC+, respectively, in our work. This information may help to orient the synthesis through suitable structural modifications resulting in activity enhancement.
Homology models
As a result of molecular modeling by the automated I-TASSER server, four P-gp homology models were generated for molecular docking in the next step. Four proteins whose PDB ID codes are 3g61A [10], 4m1mA [87], 4f4cA [88], and 3g5uA [10] were used by this server as the top threading templates. The quality estimation is shown in Table 6, with only the first model (model 1) having TM-score and RMSD values predicted. Since the correlation of C-score and quality of lower-ranked models were much weaker than that for the first model, the absolute quality (TM-score and RMSD) of the lower rank models could not be meaningfully estimated. However, the relative quality of lower rank models could be predicted based on the relative rank and their C-score information [68].
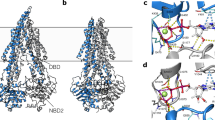
With its best quality according to I-TASSER indicators (C-score = 0.58, TM-score = 0.79, RMSD = 8.1, number of decoys = 1850, and cluster density = 0.1667), the model 1 which was built from the chain A of P-gp in mus musculus/house mouse (3g61) [10] was subsequently used for generating Ramachandran plot. In black-and-white, this plot showed the most favorable regions in the darkest gray and the less favorable regions in progressively lighter tones (Fig. 3). The statistics indicated that among 1150 nonglycine and nonproline residues, 78.7 % (905 residues), 16.4 % (189 residues), 3.0 % (35 residues), and 1.8 % (21 residues) were, respectively, found in the most favored regions [A,B,L]; the additional allowed regions [a,b,l,p]; the generously allowed regions [\(\sim {a},\sim {b},\sim {l},\sim {p}\)]; and the disallowed regions [XX]. Below 90 % of residues in the core regions as expected could be attributed to the resolution of the structure on which the homology model had been based (4.35 Å). For docking in virtual screening, its binding pocket was also predicted using the same template protein but from the chain B with the ligand 0JZ (cyclic-tris-(R)-valineselenazole/QZ59-RRR) (Fig. 4). This result was completely in accordance with the information revealed by Chang et al. [10] about the internal cavity of P-gp responsible for polyspecific drug binding.
Ramachandran plot of the best P-gp homology model, in which the most favored regions, the additional allowed regions, the generously allowed regions and the disallowed regions were labeled [A,B,L]; [a,b,l,p]; [\(\sim \)a,\(\sim \)b,\(\sim \)l,\(\sim \)p] and [XX], respectively. The darker area represented the more favorable phi–psi combination
In the past, many efforts were made for homology modeling of human P-gp from the resolved structures of bacterial pumps such as MsbA [89], Sav1866 [90, 91], and BtuCD [92, 93] to enable structure-based drug design [26]. However, to obtain a high- quality homology model, the template protein must be of high resolution and have a high sequence identity with the target protein [26]. This explains why the crystallization and structure determination of mammalian P-gp had great expectations. The X-ray structure of mouse P-gp in complex with 0JZ which was used in this study has the resolution of 4.35 Å, 87 % sequence identity to human P-gp and \(\approx 100\,\%\) identity of residues in the binding cavity excepting mSer725/hAla729 [10, 94]. Although this structure has a register shift of one amino acid (Tyr303/Ile302), it seems to be a suitable template to generate homology models of human P-gp for docking rather than new and corrected crystal structures of mouse P-gp (PDB ID: 4KSB, 4KSC, and 4KSD) which did not have bound ligands [95], since a reference ligand (0JZ in this case) is necessary to make the binding site prediction in I-TASSER and to exactly specify this region for docking-based virtual screening in LeadIT.
In silico screening
Bioactivity prediction The generated QSAR models were applied on two libraries of 87 in-house chalcones and 6874 drug-bank compounds to make predictions for their bioactivity (\(\hbox {IC}_{50}\) or \(\hbox {pIC}_{50}\)) on the multidrug resistance efflux pump P-gp, aiming to identify potential inhibitors with predicted \(\hbox {IC}_{50}\) values \(\le 15 \upmu {M}\) (threshold value) [96]. Based on 34 descriptors provided in Supplementary Material 1, SM1-Table 1, the compounds having no missing values were quickly searched for the unusuals in combination with 499 compounds of the whole dataset used for training and validating QSAR models, using the Anomaly Detection node in Clementine. The percentage of most anomalous compounds was set to 1 %. In addition, a principle component analysis (PCA) of all these compounds was also conducted with the maximum number of five components.
After the preliminary screening, 73 potential anomalies were detected from 6763 drug-bank compounds with no missing attributes (Supplementary Material 2, SM2-AD). Based on two first principal components, the distribution of the compounds belonging to different databases is presented in Supplementary Material 1, SM1-Fig. 1. The first scatter plot (Supplementary Material 1, SM1-Fig. 1a) showed the rationality of eliminating anomalous compounds which somewhat separated from the others in two-dimensional space. While the second scatter plot (Supplementary Material 1, SM1-Fig. 1b) showed the dispersion of 87 in-house chalcones within the chemical space of the database of 499 compounds for the development of prediction models, the third scatter plot (Supplementary Material 1, SM1-Fig. 1c) showed many points scattered outside this area. In other words, the QSAR models might be succeeded in bioactivity prediction for chalcones in the in-house database but are likely to inaccurately predict for some compounds in the DrugBank database.
The detailed results of bioactivity prediction on two mentioned databases by all generated QSAR models could be found in the Supplementary Material 2 (SM2-BAP). The prediction process by the best model (Ensemble) resulted in 22/87 in-house chalcones and 2374/6690 drug-bank compounds having their predicted \(\hbox {IC}_{50}\) values less than or equal to the threshold value of \(\hbox {15}\, \upmu \mathrm{M}\) to be considered as P-gp inhibitors. Based on the substituents, these chalcones could be categorized into five groups: 2\(^{\prime }\)-hydroxy, 4\(^{\prime }\)-bromo, methoxy, amino, and heterocyclic ring A (Supplementary Material 1, SM1-Table 3). Among 44 drug-bank compounds with their predicted \(\hbox {pIC}_{50}\) values \(\ge \)7 (Supplementary Material 1, SM1-Table 4), Amiodarone, Tacrolimus, Indinavir, Ritonavir, Cyclosporine, and Saquinavir are well-known inhibitors of P-gp according to the U.S. Food and Drug Administration [97].
All 22 hit chalcones with their predicted activity values were taken into account for the structure–activity relationship (SAR) hypotheses of P-gp inhibition: (i) the presence of ortho-hydroxyl group on the ring A of chalcone scaffold makes the bioactivity decrease (F7 and F29), while the para-halogen group substitution (bromo) on this ring leads to a contrary result (F18 and F14); (ii) the replacement of methoxy group by dimethyl amino group at the para position on the ring B helps to increase the inhibitory activity (F4 and F18); (iii) the heterocyclic ring A is not necessary for good inhibitors (F18 and F65); (iv) methoxy substituents at the ortho and para positions on the ring A (F29, F33, and F36) have a positive influence on the effect of inhibitors; and (v) the inhibitory potency is enhanced by halogen group (cloro) at the ortho position (F44 and F45) or methoxy group at the ortho or meta position (F4, F5 and F7) on the ring B (ortho substitution is better: F35, F36 and F37).
Molecular docking A set of 22 in-house chalcones and 44 drug-bank compounds derived from the in silico prediction was applied for docking to determine their possible binding modes and affinities with the best homology model of P-gp, using FlexX in LeadIT. In addition, docking study was also implemented with 65 remaining chalcones, Reserpine, Tariquidar, and Elacridar (small molecule inhibitors of P-gp) [22] for comparison purpose. With no flips, protonation, and torsions, the binding site of protein was defined by choosing the ligand in the predicted binding site (0JZ) as a reference to include amino acids within radius of 6.5 Å. A total of 120 small molecules including 87 chalcone derivatives and 33 drug-bank compounds were successfully docked into the active site of receptor. However, the docking algorithm failed on 11 other drug-bank compounds because of their complex and bulky structures. The docking results could be found in detail in the Supplementary Material 2 (SM2-MD).
With docking scores ranging from −23.853 to −10.828 kJ/mol, the in-house chalcones showed good binding abilities into the internal cavity of P-gp homology model. The consensus of prediction and docking results suggested F45 as the most promising chalcone for inhibiting P-gp function (Supplementary Material 1, SM1-Table 3). Despite of having predicted \(\hbox {IC}_{50}\) values >15 \(\upmu \)M, F57, F53 and F55 with docking scores of −23.853, −21.750, and −21.483 kJ/mol, respectively, were the derivatives possessing the highest binding affinities for P-gp out of all 87 chalcones. Higher docking scores in general displayed weaker binding abilities of the drug-bank compounds in comparison with the chalcone derivatives, although they had significantly lower predicted \(\hbox {IC}_{50}\) values (Supplementary Material 1, SM1-Table 4). Apart from the compounds unable to be docked, there were 7/33 drug-bank compounds having positive docking scores. As well as F45, some drug-bank compounds such as DB01578, DB0278, DB02009, DB04378, and DB01232 stood out for both prediction and docking results. Compared with docking scores of Reserpine, Tariquidar, and Elacridar which were −3.465, −13.262, and −22.858 kJ/mol, respectively, hit compounds derived from the DrugBank database (e.g., DB01578, DB02785, DB02009, and DB04378) and our private database (e.g., F44 and F45) emerged as potential candidates to modulate P-gp-mediated drug efflux and reverse MDR in cancer cells.
Docking poses of three in-house chalcones having the highest predicted \(\hbox {pIC}_{50}\) or the lowest predicted \(\hbox {IC}_{50}\) values (F45, F17, and F44); three in-house chalcones having the lowest docking scores (F57, F53, and F55); three drug-bank compounds having the highest predicted \(\hbox {pIC}_{50}\) or the lowest predicted \(\hbox {IC}_{50}\) values (DB01118, DB01578, and DB01362); three drug-bank compounds having the lowest docking scores (DB01232, DB02785, and DB01578) and three P-gp inhibitors (Reserpine, Tariquidar, and Elacridar) are illustrated in Fig. 5 (F45, DB01578) and Supplementary Material 1, SM1-Fig.F2 (other ligands). These images indicated key residues responsible for receptor–ligand-binding interactions (hydrogen bonding, arene-cation, and hydrophobic), which were His61, Gln946, Tyr950, Leu65, Phe194, and Gln195 in case of chalcones and Gln347, Phe343, Phe194, Gln195, Ser344, and Gln946 in case of drug-bank compounds. Amino acids Phe194, Gln195 in TM3 and Gln946, Tyr950 in TM11 reaffirmed the drug binding pocket located at the TM3/TM11 interface which was indicated before in the work of Chiba et al. [14]. Other methods such as photoaffinity labeling, site-directed mutagenesis, thiol probes and co-crystal structural studies of P-gp in the presence of mentioned ligands could be used to validate residues identified by our docking study.
Conclusions
In this work, we employed multiple different machine learning methods on a large and diverse database of compounds whose bioactivity values (\(\hbox {IC}_{50})\) are comparable to develop QSAR models for ligand-based design of P-gp inhibitors. The results of internal and external validation processes with strict validation criteria proved the predictivity of these models, especially the Ensemble model. In addition, high-quality homology models of P-gp were also generated for structure-based design. Based on these in silico tools, virtual screening was applied on two databases, one from in-house source and one from Drug Bank to discover new P-gp-mediated MDR reverters. Finally, structural optimization for more potent and safer drug candidates is required before further experimental investigations.
Supplementary material
The supplementary material associated with this article is presented in one PDF file (additional tables and figures) and one XLSX file (databases, external validation, anomaly detection, bioactivity prediction, and molecular docking results).
References
Teodori E, Dei S, Martelli C, Scapecchi S, Gualtieri F (2006) The functions and structure of ABC transporters: implications for the design of new inhibitors of Pgp and MRP1 to control multidrug resistance (MDR). Curr Drug Targets 7:893–909. doi:10.2174/138945006777709520
Zhou SF (2008) Structure, function and regulation of P-glycoprotein and its clinical relevance in drug disposition. Xenobiotica 38:802–832. doi:10.1080/00498250701867889
Sharom FJ (2006) Multidrug resistance protein: P-glycoprotein. In: Drug transporters. Wiley, New York, pp 223–262. doi:10.1002/9780470140505.ch10
Glaeser H, Fromm MF, König J (2008) Transporters and drugs–an overview. In: Antitargets. Wiley-VCH Verlag GmbH & Co. KGaA, pp 341–366. doi:10.1002/9783527621460.ch15
Thai KM, Ngo TD, Tran TD, Le MT (2013) Pharmacophore modeling for antitargets. Curr Top Med Chem 13:1002–1014. doi:10.2174/1568026611313090004
Vaz RJ, Klabunde T (2008) A personal foreword. In: Antitargets. Wiley-VCH Verlag GmbH & Co. KGaA, pp XXI–XXIV. doi:10.1002/9783527621460.fmatter
Lehne G (2000) P-glycoprotein as a drug target in the treatment of multidrug resistant cancer. Curr Drug Targets 1:85–99. doi:10.2174/1389450003349443
Binkhathlan Z, Lavasanifar A (2013) P-glycoprotein inhibition as a therapeutic approach for overcoming multidrug resistance in cancer: current status and future perspectives. Curr Cancer Drug Targets 13:326–346. doi:10.2174/15680096113139990076
Ronchi E, Sanfilippo O, Di Fronzo G, Bani MR, Della Torre G, Catania S, Silvestrini R (1989) Detection of the 170 kDa P-glycoprotein in neoplastic and normal tissues. Tumori 75:542–546
Aller SG, Yu J, Ward A, Weng Y, Chittaboina S, Zhuo R, Harrell PM, Trinh YT, Zhang Q, Urbatsch IL, Chang G (2009) Structure of P-glycoprotein reveals a molecular basis for poly-specific drug binding. Science 323:1718–1722. doi:10.1126/science.1168750
Shintre CA, Pike AC, Li Q, Kim JI, Barr AJ, Goubin S, Shrestha L, Yang J, Berridge G, Ross J, Stansfeld PJ, Sansom MS, Edwards AM, Bountra C, Marsden BD, von Delft F, Bullock AN, Gileadi O, Burgess-Brown NA, Carpenter EP (2013) Structures of ABCB10, a human ATP-binding cassette transporter in apo- and nucleotide-bound states. Proc Natl Acad Sci USA 110:9710–9715. doi:10.1073/pnas.1217042110
Montanari F, Ecker GF (2015) Prediction of drug-ABC-transporter interaction—recent advances and future challenges. Adv Drug Deliv Rev 86:17–26. doi:10.1016/j.addr.2015.03.001
Chufan EE, Kapoor K, Sim HM, Singh S, Talele TT, Durell SR, Ambudkar SV (2013) Multiple transport-active binding sites are available for a single substrate on human P-glycoprotein (ABCB1). PloS One 8:e82463. doi:10.1371/journal.pone.0082463
Chiba P, Mihalek I, Ecker GF, Kopp S, Lichtarge O (2006) Role of transmembrane domain/transmembrane domain interfaces of P-glycoprotein (ABCB1) in solute transport. Convergent information from photoaffinity labeling, site directed mutagenesis and in silico importance prediction. Curr Med Chem 13:793–805. doi:10.2174/092986706776055607
Palmeira A, Sousa E, Vasconcelos MH, Pinto MM (2012) Three decades of P-gp inhibitors: skimming through several generations and scaffolds. Curr Med Chem 19:1946–2025. doi:10.2174/092986712800167392
Szakacs G, Paterson JK, Ludwig JA, Booth-Genthe C, Gottesman MM (2006) Targeting multidrug resistance in cancer. Nat Rev Drug Discov 5:219–234. doi:10.1038/nrd1984
Saneja A, Khare V, Alam N, Dubey RD, Gupta PN (2014) Advances in P-glycoprotein-based approaches for delivering anticancer drugs: pharmacokinetic perspective and clinical relevance. Expert Opin Drug Deliv 11:121–138. doi:10.1517/17425247.2014.865014
Saneja A, Dubey RD, Alam N, Khare V, Gupta PN (2014) Co-formulation of P-glycoprotein Substrate and inhibitor in nanocarriers: an emerging strategy for cancer chemotherapy. Curr Cancer Drug Targets 14:419–433. doi:10.2174/1568009614666140407112034
Callaghan R, Luk F, Bebawy M (2014) Inhibition of the multidrug resistance P-glycoprotein: time for a change of strategy? Drug Metab Dispos 42:623–631. doi:10.1124/dmd.113.056176
Zhang S, Morris ME (2003) Effects of the flavonoids biochanin A, morin, phloretin, and silymarin on P-glycoprotein-mediated transport. J Pharmacol Exp Ther 304:1258–1267. doi:10.1124/jpet.102.044412
Bansal T, Jaggi M, Khar RK, Talegaonkar S (2009) Emerging significance of flavonoids as P-glycoprotein inhibitors in cancer chemotherapy. J Pharm Pharm Sci 12:46–78. doi:10.18433/J3RC77
Srivalli KMR, Lakshmi PK (2012) Overview of P-glycoprotein inhibitors: a rational outlook. Braz J Pharm Sci 48:353–367. doi:10.1590/s1984-82502012000300002
Sak K (2014) Cytotoxicity of dietary flavonoids on different human cancer types. Pharmacogn Rev 8:122–146. doi:10.4103/0973-7847.134247
Ferreira A, Pousinho S, Fortuna A, Falcão A, Alves G (2015) Flavonoid compounds as reversal agents of the P-glycoprotein-mediated multidrug resistance: biology, chemistry and pharmacology. Phytochem Rev 14:233–272. doi:10.1007/s11101-014-9358-0
Parveen Z, Brunhofer G, Jabeen I, Erker T, Chiba P, Ecker GF (2014) Synthesis, biological evaluation and 3D-QSAR studies of new chalcone derivatives as inhibitors of human P-glycoprotein. Bioorg Med Chem 22:2311–2319. doi:10.1016/j.bmc.2014.02.005
Palmeira A, Sousa E, Vasconcelos MH, Pinto M, Fernandes MX (2012) Structure and ligand-based design of P-glycoprotein inhibitors: a historical perspective. Curr Pharm Des 18:4197–4214. doi:10.2174/138161212802430530
Zdrazil B, Pinto M, Vasanthanathan P, Williams AJ, Balderud LZ, Engkvist O, Chichester C, Hersey A, Overington JP, Ecker GF (2012) Annotating human P-glycoprotein bioassay data. mol inform 31:599–609. doi:10.1002/minf.201200059
Bento AP, Gaulton A, Hersey A, Bellis LJ, Chambers J, Davies M, Kruger FA, Light Y, Mak L, McGlinchey S, Nowotka M, Papadatos G, Santos R, Overington JP (2014) The ChEMBL bioactivity database: an update. Nucleic Acids Res 42:D1083–1090. doi:10.1093/nar/gkt1031
MOE. 2008.10 edn. Chemical Computing Group Inc., 1010 Sherbrooke St. W, Suite 910, Montreal, Quebec, Canada H3A 2R7. http://www.chemcomp.com/
Wishart DS, Knox C, Guo AC, Shrivastava S, Hassanali M, Stothard P, Chang Z, Woolsey J (2006) DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res 34:D668–672. doi:10.1093/nar/gkj067
Wishart DS, Knox C, Guo AC, Cheng D, Shrivastava S, Tzur D, Gautam B, Hassanali M (2008) DrugBank: a knowledgebase for drugs, drug actions and drug targets. Nucleic Acids Res 36:D901–906. doi:10.1093/nar/gkm958
Knox C, Law V, Jewison T, Liu P, Ly S, Frolkis A, Pon A, Banco K, Mak C, Neveu V, Djoumbou Y, Eisner R, Guo AC, Wishart DS (2011) DrugBank 3.0: a comprehensive resource for ’omics’ research on drugs. Nucleic Acids Res 39:D1035–1041. doi:10.1093/nar/gkq1126
Law V, Knox C, Djoumbou Y, Jewison T, Guo AC, Liu Y, Maciejewski A, Arndt D, Wilson M, Neveu V, Tang A, Gabriel G, Ly C, Adamjee S, Dame ZT, Han B, Zhou Y, Wishart DS (2014) DrugBank 4.0: shedding new light on drug metabolism. Nucleic Acids Res 42:D1091–1097. doi:10.1093/nar/gkt1068
ChemBioDrawUltra. 12.0 edn. PerkinElmer, CambridgeSoft. http://www.cambridgesoft.com/
Zhang J, Huan J (2010) Comparison of chemical descriptors for protein-chemical interaction prediction. Int J Comput Biosci. doi:10.2316/Journal.210.2010.1.210-1010
Yap CW (2011) PaDEL-descriptor: an open source software to calculate molecular descriptors and fingerprints. J Comput Chem 32:1466–1474. doi:10.1002/jcc.21707
Demel MA, Janecek AGK, Thai K-M, Ecker GF, Gansterer WN (2008) Predictive QSAR Models for polyspecific drug targets: the importance of feature selection. Curr Comput Aided Drug Des 4:91–110. doi:10.2174/157340908784533256
RapidMiner. 5.3.008 edn. Rapid-I and contributors, Stockumer Str. 475, 44227 Dortmund, Germany. http://rapidminer.com/
WEKA. 3.7.9 edn. The University of Waikato, Hamilton, New Zealand. http://www.cs.waikato.ac.nz/ml/weka/
Jaworska J, Nikolova-Jeliazkova N, Aldenberg T (2005) QSAR applicabilty domain estimation by projection of the training set descriptor space: a review. Altern Lab Anim 33:445–459
OECD (2014) Guidance Document on the Validation of (Quantitative) Structure-Activity Relationship [(Q)SAR] Models. OECD Publishing, OECD Series on Testing and Assessment. doi:10.1787/9789264085442-en
Netzeva TI, Worth A, Aldenberg T, Benigni R, Cronin MT, Gramatica P, Jaworska JS, Kahn S, Klopman G, Marchant CA, Myatt G, Nikolova-Jeliazkova N, Patlewicz GY, Perkins R, Roberts D, Schultz T, Stanton DW, van de Sandt JJ, Tong W, Veith G, Yang C (2005) Current status of methods for defining the applicability domain of (quantitative) structure-activity relationships. The report and recommendations of ECVAM Workshop 52. Altern Lab Anim 33:155–173
Roy K, Kar S, Ambure P (2015) On a simple approach for determining applicability domain of QSAR models. Chemometr Intell Lab 145:22–29. doi:10.1016/j.chemolab.2015.04.013
Roy K, Kar S (2015) Importance of applicability domain of QSAR models. In: Kunal R (ed) Quantitative Structure-activity relationships in drug design, predictive toxicology, and risk assessment. IGI Global, Hershey, pp 180–211. doi:10.4018/978-1-4666-8136-1.ch005
Dobchev DA, Pillai GG, Karelson M (2014) In silico machine learning methods in drug development. Curr Top Med Chem 14:1913–1922. doi:10.2174/1568026614666140929124203
Clementine. 12.0 edn. SPSS Inc., 233 South Wacker Drive, 11th Floor, Chicago, IL 60606-6307, USA. http://www.spss.com/
Witten IH, Frank E, Hall MA (2011) Data mining : practical machine learning tools and techniques. Morgan Kaufmann series in data management systems, 3rd edn. Morgan Kaufmann, Burlington
Bengio Y, Grandvalet Y (2004) No unbiased estimator of the variance of K-fold cross-validation. J Mach Learn Res 5:1089-1105. www.jmlr.org/papers/v5/grandvalet04a.html
Pratim Roy P, Paul S, Mitra I, Roy K (2009) On two novel parameters for validation of predictive QSAR models. Molecules 14:1660–1701. doi:10.3390/molecules14051660
Golbraikh A, Tropsha A (2002) Beware of q2!. J Mol Graph Model 20:269–276. doi:10.1016/S1093-3263(01)00123-1
Tropsha A, Gramatica P, Gombar VK (2003) The importance of being earnest: validation is the absolute essential for successful application and interpretation of QSPR models. QSAR Comb Sci 22:69–77. doi:10.1002/qsar.200390007
Gramatica P (2007) Principles of QSAR models validation: internal and external. QSAR Comb Sci 26:694–701. doi:10.1002/qsar.200610151
Tropsha A (2010) Best practices for QSAR model development, validation, and exploitation. Mol Inform 29:476–488. doi:10.1002/minf.201000061
Shi LM, Fang H, Tong W, Wu J, Perkins R, Blair RM, Branham WS, Dial SL, Moland CL, Sheehan DM (2001) QSAR models using a large diverse set of estrogens. J Chem Inf Comput Sci 41:186–195. doi:10.1021/ci000066d
Schuurmann G, Ebert RU, Chen J, Wang B, Kuhne R (2008) External validation and prediction employing the predictive squared correlation coefficient test set activity mean vs training set activity mean. J Chem Inf Model 48:2140–2145. doi:10.1021/ci800253u
Consonni V, Ballabio D, Todeschini R (2009) Comments on the definition of the Q2 parameter for QSAR validation. J Chem Inf Model 49:1669–1678. doi:10.1021/ci900115y
Consonni V, Ballabio D, Todeschini R (2010) Evaluation of model predictive ability by external validation techniques. J Chemom 24:194–201. doi:10.1002/cem.1290
Roy PP, Roy K (2008) On some aspects of variable selection for partial least squares regression models. QSAR Comb Sci 27:302–313. doi:10.1002/qsar.200710043
Ojha PK, Mitra I, Das RN, Roy K (2011) Further exploring rm2 metrics for validation of QSPR models. Chemometr Intell Lab 107:194–205. doi:10.1016/j.chemolab.2011.03.011
Ojha PK, Roy K (2011) Comparative QSARs for antimalarial endochins: importance of descriptor-thinning and noise reduction prior to feature selection. Chemometr Intell Lab 109:146–161. doi:10.1016/j.chemolab.2011.08.007
Lin LI (1989) A concordance correlation coefficient to evaluate reproducibility. Biometrics 45:255–268. doi:10.2307/2532051
Chirico N, Gramatica P (2011) Real external predictivity of QSAR models: how to evaluate it? Comparison of different validation criteria and proposal of using the concordance correlation coefficient. J Chem Inf Model 51:2320–2335. doi:10.1021/ci200211n
Chirico N, Gramatica P (2012) Real external predictivity of QSAR models. Part 2. New intercomparable thresholds for different validation criteria and the need for scatter plot inspection. J Chem Inf Model 52:2044–2058. doi:10.1021/ci300084j
Roy K, Mitra I, Kar S, Ojha PK, Das RN, Kabir H (2011) Comparative studies on some metrics for external validation of QSPR models. J Chem Inf Model 52:396–408. doi:10.1021/ci200520g
Roy K, Das RN, Ambure P, Aher RB (2016) Be aware of error measures. Further studies on validation of predictive QSAR models. Chemometr Intell Lab 152:18–33. doi:10.1016/j.chemolab.2016.01.008
Ravna AW, Sylte I (2012) Homology modeling of transporter proteins (carriers and ion channels). Methods Mol Biol 857:281–299. doi:10.1007/978-1-61779-588-6_12
Zhang Y (2008) I-TASSER server for protein 3D structure prediction. BMC Bioinform 9:40. doi:10.1186/1471-2105-9-40
Roy A, Kucukural A, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5:725–738. doi:10.1038/nprot.2010.5
Wu S, Skolnick J, Zhang Y (2007) Ab initio modeling of small proteins by iterative TASSER simulations. BMC Biol 5:17. doi:10.1186/1741-7007-5-17
Zhang Y (2009) I-TASSER: fully automated protein structure prediction in CASP8. Proteins 77(Suppl 9):100–113. doi:10.1002/prot.22588
Multidrug resistance protein 1 [Homo sapiens] - NCBI Reference Sequence: NP_000918.2. NCBI. http://www.ncbi.nlm.nih.gov/protein/42741659?report=fasta. Accessed 22 Sept 2015
Xu J, Zhang Y (2010) How significant is a protein structure similarity with TM-score = 0.5? Bioinformatics 26:889–895. doi:10.1093/bioinformatics/btq066
Laskowski RA, Macarthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Cryst 26:283–291. doi:10.1107/S0021889892009944
LeadIT. 2.0.2 edn. BioSolveIT GmbH, An der Ziegelei 79, 53757 St. Augustin, Germany. http://www.biosolveit.de/
Baguley BC (2010) Multiple drug resistance mechanisms in cancer. Mol Biotechnol 46:308–316. doi:10.1007/s12033-010-9321-2
Litman T, Zeuthen T, Skovsgaard T, Stein WD (1997) Structure-activity relationships of P-glycoprotein interacting drugs: kinetic characterization of their effects on ATPase activity. Biochim Biophys Acta 1361:159–168. doi:10.1016/S0925-4439(97)00026-4
Österberg T, Norinder U (2000) Theoretical calculation and prediction of P-glycoprotein-interacting drugs using MolSurf parametrization and PLS statistics. Eur J Pharm Sci 10:295–303. doi:10.1016/S0928-0987(00)00077-4
Dearden JC, Al-Noobi A, Scott AC, Thomson SA (2003) QSAR studies on P-glycoprotein-regulated multidrug resistance and on its reversal by phenothiazines. SAR QSAR Environ Res 14:447–454. doi:10.1080/10629360310001624024
Wang RB, Kuo CL, Lien LL, Lien EJ (2003) Structure-activity relationship: analyses of p-glycoprotein substrates and inhibitors. J Clin Pharm Ther 28:203–228. doi:10.1046/j.1365-2710.2003.00487.x
Kupsáková I, Rybár A, Dočolomanský P, Drobná Z, Stein U, Walther W, BarančıÃk M, Breier A (2004) Reversal of P-glycoprotein mediated vincristineresistance of L1210/VCR cells by analogues of pentoxifylline: A QSARstudy. Eur J Pharm Sci 21:283–293. doi:10.1016/j.ejps.2003.10.019
Wang Y-H, Li Y, Yang S-L, Yang L (2005) An in silico approach for screening flavonoids as P-glycoprotein inhibitors based on a Bayesian-regularized neural network. J Comput Aided Mol Des 19:137–147. doi:10.1007/s10822-005-3321-5
Wu J, Li X, Cheng W, Xie Q, Liu Y, Zhao C (2009) Quantitative Structure activity relationship (QSAR) approach to multiple drug resistance (MDR) modulators based on combined hybrid system. QSAR Comb Sci 28:969–978. doi:10.1002/qsar.200860134
Sousa IJ, Ferreira M-JU, Molnár J, Fernandes MX (2013) QSAR studies of macrocyclic diterpenes with P-glycoprotein inhibitory activity. Eur J Pharm Sci 48:542–553. doi:10.1016/j.ejps.2012.11.012
Jabeen I, Wetwitayaklung P, Chiba P, Pastor M, Ecker GF (2013) 2D- and 3D-QSAR studies of a series of benzopyranes and benzopyrano[3,4b][1,4]-oxazines as inhibitors of the multidrug transporter P-glycoprotein. J Comput Aided Mol Des 27:161–171. doi:10.1007/s10822-013-9635-9
Vazquez RN, Camargo AB, Marchevsky EJ, Luco JM (2014) Molecular factors influencing the affinity of flavonoid compounds on P-glycoprotein efflux transporter. Curr Comput Aided Drug Des 10:250–258. doi:10.2174/157340991003150302231140
Shen J, Cui Y, Gu J, Li Y, Li L (2014) A genetic algorithm- back propagation artificial neural network model to quantify the affinity of flavonoids toward P-glycoprotein. Comb Chem High Throughput Screen 17:162–172. doi:10.2174/1386207311301010002
Li J, Jaimes KF, Aller SG (2014) Refined structures of mouse P-glycoprotein. Protein Sci 23:34–46. doi:10.1002/pro.2387
Jin MS, Oldham ML, Zhang Q, Chen J (2012) Crystal structure of the multidrug transporter P-glycoprotein from Caenorhabditis elegans. Nature 490:566–569. doi:10.1038/nature11448
Ward A, Reyes CL, Yu J, Roth CB, Chang G (2007) Flexibility in the ABC transporter MsbA: alternating access with a twist. Proc Natl Acad Sci USA 104:19005–19010. doi:10.1073/pnas.0709388104
Dawson RJ, Locher KP (2006) Structure of a bacterial multidrug ABC transporter. Nature 443:180–185. doi:10.1038/nature05155
Dawson RJ, Locher KP (2007) Structure of the multidrug ABC transporter Sav 1866 from Staphylococcus aureus in complex with AMP-PNP. FEBS Lett 581:935–938. doi:10.1016/j.febslet.2007.01.073
Locher KP, Lee AT, Rees DC (2002) The E. coli BtuCD structure: a framework for ABC transporter architecture and mechanism. Science 296:1091–1098. doi:10.1126/science.1071142
Hvorup RN, Goetz BA, Niederer M, Hollenstein K, Perozo E, Locher KP (2007) Asymmetry in the structure of the ABC transporter-binding protein complex BtuCD-BtuF. Science 317:1387–1390. doi:10.1126/science.1145950
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. doi:10.1093/nar/25.17.3389
Ward AB, Szewczyk P, Grimard V, Lee CW, Martinez L, Doshi R, Caya A, Villaluz M, Pardon E, Cregger C, Swartz DJ, Falson PG, Urbatsch IL, Govaerts C, Steyaert J, Chang G (2013) Structures of P-glycoprotein reveal its conformational flexibility and an epitope on the nucleotide-binding domain. Proc Natl Acad Sci USA 110:13386–13391. doi:10.1073/pnas.1309275110
Rautio J, Humphreys JE, Webster LO, Balakrishnan A, Keogh JP, Kunta JR, Serabjit-Singh CJ, Polli JW (2006) In vitro p-glycoprotein inhibition assays for assessment of clinical drug interaction potential of new drug candidates: a recommendation for probe substrates. Drug Metab Dispos 34:786–792. doi:10.1124/dmd.105.008615
Drug Development and Drug Interactions: Table of Substrates, Inhibitors and Inducers. U.S. Food and Drug Administration. http://www.fda.gov/Drugs/DevelopmentApprovalProcess/DevelopmentResources/DrugInteractionsLabeling/ucm093664.htm. Accessed 22 Sept 2015
Acknowledgments
This research is funded by Vietnam National Foundation for Science and Technology Development (NAFOSTED) under Grant number 106-YS.05-2015.31 to Khac-Minh Thai.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors confirm that this article content has no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ngo, TD., Tran, TD., Le, MT. et al. Computational predictive models for P-glycoprotein inhibition of in-house chalcone derivatives and drug-bank compounds. Mol Divers 20, 945–961 (2016). https://doi.org/10.1007/s11030-016-9688-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-016-9688-5