Abstract
JAK2 and JAK3 are non-receptor protein tyrosine kinases implicated in B-cell- and T-cell-mediated diseases. Both enzymes work via different pathways but are involved in the pathogenesis of common lymphoid-derived diseases. Hence, targeting both Janus kinases together can be a potential strategy for the treatment of these diseases. In the present study, two separate pharmacophore-based 3D-QSAR models ADRR.92 (\(Q_{\mathrm{test}}^{2} 0.663, R^{2}_{\mathrm{train}} 0.849\), F value 219.3) for JAK2 and ADDRR.142 (\(Q^{2}_{\mathrm{test}}0.655, R_{\mathrm{train}}^{2}\) 0.869, F value 206.9) for JAK3 were developed. These models were employed for the screening of a PHASE database of approximately 1.5 million compounds; subsequently, the retrieved hits were screened employing docking simulations with JAK2 and JAK3 proteins. Finally, ADME properties of screened dual inhibitors displaying essential interactions with both proteins were calculated to filter candidates with poor pharmacokinetic profiles. These candidates could serve as novel therapeutic agents in the treatment of lymphoid-related diseases.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
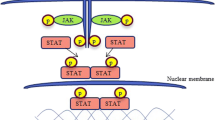
Janus kinases are non-receptor protein tyrosine kinases which mediate the signaling of various cytokines pathways, and anomalous regulation of these pathways can lead to various cancerous conditions. For the prevention or control of above-mentioned diseases, inhibition of these cytokine pathways is necessary either directly or by the inhibiting Janus kinases that participate in cytokine-mediating signaling as secondary messengers. JAK2 is involved in multi cytokine pathways along with other JAK members, but its role in IL-6-mediated signaling is significant in various physiological processes such as bone metabolism, acute phase response, hematopoiesis, and B-cell differentiation [1]. Imbalance in IL-6-mediated signaling can increase B-cell differentiation which further leads to lymphoid-derived diseases. JAK3 also plays a vital role in lymphoid development via IL-2 pathway that regulates the activity of distinct lymphoid cell populations including T and B lymphocytes and natural killer (NK) cells [2]. The imbalance in IL-2-mediated signaling also contributes in the pathogenesis of lymphoid-derived diseases. Thus, the development of dual JAK2/3 inhibitors could be efficacious strategy to treat a variety of lymphoid-derived diseases that are dependent on the JAK2 and JAK3 signaling cascade. A drug in clinical trials, AG-490, which is dual JAK2/3 inhibitor, effectively blocks uncontrolled B-cell growth in the patients suffering from acute lymphoblastic leukemia by inhibiting the abnormal constitutive activation of JAK2 detected within these cells [3]. It also blocks the IL-2-mediated cell growth of phytohemagglutinin (PHA) or antigen-specific-activated human T cells through inactivation of JAK3 and STAT5 signaling pathway [4].
In the present study, two separate pharmacophore models were developed for JAK2 and JAK3 enzymes; consequently, the models were employed for the screening of a PHASE molecular database of 1.5 million molecules. The combined approach of pharmacophore-based virtual screening (PBVS) and structure-based virtual screening (SBVS) was utilized in the identification of new dual inhibitory agents for JAK2 and JAK3. We used AG490 as reference for our study so as to get final hits having potency and activity closer to this drug.
Computational methods
Molecular modeling—data selection and processing
Two datasets consisting of 252 JAK2 and 211 JAK3 inhibitor compounds mentioned in Table S1 and S2 (supplementary data), respectively, were selected from the literature [5–23]. The biological activities of both the datasets extended (IC\(_{50}\) values) over a wide range 0.00009–29.000 and 0.0006–51.300 \(\upmu \)M, respectively. The reported \(\hbox {IC}_{50}\) values were converted into respective \(\hbox {pIC}_{50 }\)values by taking the negative logarithm of the \(\hbox {IC}_{50}\) values (\(-\)log\(\hbox {IC}_{50})\) and were subsequently utilized for the development of pharmacophore model. Maestro, an integrated visualization interface for all Schrödinger software, was used to sketch all compounds [24], and these molecules were optimized using Ligprep [25] employing the OPLS_2005 force field.
Pharmacophore generation
PHASE, a high-performance program module of Schrödinger for ligand-based drug design, was used to generate pharmacophore models [26]. Since most of the studied ligands were flexible, all possible conformations for a ligand were generated using mixed Monte Carlo minimization method (MCMM)/low mode docking (LMOD) approach. For conformation sampling, energy window of 20 kcal/mol was employed to increase the chances of finding the representative conformer close to the bound structure. After conformation generation, ligands were assigned as actives and inactives by giving an appropriate activity threshold value. The activity threshold value was selected on the basis of dataset activity distribution. Using the “create site” option in PHASE, molecules were assigned different pharmacophoric features. PHASE provides six pharmacophoric features: hydrogen bond acceptor (HBA) (A), hydrogen bond donor (HBD) (D), hydrophobic (H), negatively charged (N), positively charged (P), and aromatic ring (R) features. The conformational space of active molecules was utilized for the generation of common pharmacophore hypotheses (CPH), which groups together similar pharmacophores according to their intersite distances. The resulting hypotheses obtained were scored and ranked on the basis of scoring parameters, i.e., survival and survival minus inactive (\(S-I\)) score. The scoring was done to identify the best candidate hypothesis, which provided an overall ranking of all the hypotheses. The hypotheses so generated were then clustered, and a representative model from each cluster was selected on the basis of the highest \(S-I\) score. The selected models were thereafter utilized for the alignment of non-model molecules which were subsequently intended for the development of 3D-QSAR model.
3D-QSAR development
Three-dimensional quantitative structure–activity relationship (QSAR) is the computational method used in the development of relationship between independent (structural components) and dependent variables (biological activity) to obtain a reliable statistical model for prediction of the activities of new molecules [27, 28]. QSAR modeling was carried out using the selected hypotheses by randomly dividing the dataset into training set and test set on the basis of proper variation of activity. For the generation of 3D-QSAR models, all training set molecules aligned over the common pharmacophoric sites were placed into regular 1 Å cubic grids. Each cube was allocated 0 or 1 “bits” to account the different types of atomic features in training set molecules that occupy the cube. Each occupied cube gives rise to one or more volume bits, where a separate bit is allocated for each different categories of atom that occupy the cubes. A large pool of binary values (0 and 1) was formed for the dataset molecules that were treated as independent variable for the generation of QSAR models. The best QSAR model was selected on the basis of high value of \(Q^{2}_{\mathrm{test}}\) (correlation of prediction for test set) and \(R^{2}_{\mathrm{train}}\) (correlation of prediction for training set). The 5 and 7 component (PLS factor) models with good statistics were obtained for the dataset of JAK2 and JAK3, respectively, whereas the maximum number of PLS factors in each model can be 1/5 of the total number of training set molecules. Further increase in the number of PLS factors did not improve the model statistics or predictive ability.
Since pharmacophore model is theoretical model, it is necessary to analyze whether or not the developed model is able to predict the active compounds. Thus, the developed models need to be validated before going for further implementation. The validation of the models was carried out by different validation methods including both internal and external validation. For the validation of the generated QSAR model for its external predictive ability and reliability, test set prediction using Pearson-r was examined. Pearson coefficient of correlation determines the predictive reliability of the generated model for external dataset molecules that have not been considered for the development of model i.e., test set molecules. In addition, the validation for the external predictive ability of the generated model further validated by calculating a set of parameters i.e., \(R_{\mathrm{o}}^{2}\) or \({R_{\mathrm{o}}^{\prime }}^{2}\) close to \(R^{2}, [(R^{2}-R_{\mathrm{o}}^{2})/R^{2}] < 0.1\) or \([R^{2}-{R_{\mathrm{o}}^{\prime }}^{2}/R^{2}] < 0.1\), and the corresponding 0.85 \(\le k \le \) 1.15 or 0.85 \(\le k^{\prime } \le \) 1.15 [29].
The other validation methods include applicability domain (APD) calculation that determines the resemblance between training and test set compounds and Y-randomization which analyze the robustness of the generated model. The APD calculation was carried out using “canvas” program in which similarity measurements were determined on the basis of Euclidean distances between all pairs of training and test set compounds. In this validation parameter, the similarity of test set molecules must reside within the threshold (APD) otherwise selected model is considered to be unreliable for the prediction of new compounds.
\(\langle d \rangle \) new average, \(\sigma \) is standard deviation of training set molecules with distances lower than previously calculated average value, and Z is an empirical cutoff with default value of 0.5.
Y-randomization is determined by scrambling the activity data of training set molecules in random manner and to generate different training sets from the original training set. Thereafter, for the random sets, value of \(R^{2}\) was determined and the average value so obtained was reported as \(R_{\mathrm{scramble}}^{2}\). This \(R_{\mathrm{scramble}}^{2}\) was calculated using PHASE module. The value of \(R_{\mathrm{scramble}}^{2}\) should be less than the \(\hbox {R}_{\mathrm{train}}^{2}\) of the original selected model.
Virtual screening
Virtual screening (VS) was performed in order to identify those structures among conformers database which are most likely to bind to a drug target. VS was carried out systematically using integral use of two techniques classified as SBVS and ligand-based virtual screening (LBVS). The database screening using pharmacophore model is a method of LBVS to screen millions of multi-conformational compounds to retrieve structurally diverse hits. To overcome the drawback of LBVS, i.e., lacking the ability to identify false hits, combination of SBVS and LBVS is used as integrated approach in drug discovery protocol.
A PHASE database of 1.5 million molecules was employed for the screening with the selected JAK2 and JAK3 pharmacophore models for the identification of new hits. These hit compounds contain the structural features that overlap the selected model. The hit molecules were further docked using Glide [30, 31] in the JAK2 and JAK3 proteins to remove the false negative and positive hits. Some parameters related to oral bioavailability and pharmacokinetic profile of the designed molecules were computationally calculated using Qikprop [32]. These calculated parameters were finally used for filtering and ranking the large number of screened molecules. Qikprop is based on the principle of Lipinski’s rule of five. According to this rule, poor absorption is expected if molecular weight (MW) \(>\) 500, partition coefficient (logP) \(>\) 5, HBDs \(>\) 5, and HBAs \(>\) 10. The ADME parameters include partition coefficient (\(\textit{QP}log\textit{P}_{\mathrm{o}/\mathrm{w}})\), water solubility (QPlogS), cell permeability (QPPCaco), percentage human oral absorption, etc.
Finally, ligands were sampled for post-processing with Prime/MM–GBSA which predicts the binding energy of set of ligands and a single receptor. The MM–GBSA binding energy (\(\Delta G_{\mathrm{bind}})\) is estimated in kcal/mol using equation:
where ER:EL is prime energy of the optimized complex, EL is prime energy of optimized free ligand, and ER is prime energy of optimized free receptor.
Results and discussion
Pharmacophore generation and 3D QSAR
A pharmacophore modeling study for phenylaminopyrimidine JAK2 inhibitors has been reported in 2011 considering 44 molecules [33]. In the present study, diverse dataset of 252 inhibitors of JAK2 including the above-mentioned series was utilized for the study. For the generation of pharmacophore model, the molecules were divided into active, inactive, and moderately active molecules. For JAK2 inhibitors, threshold value was 9.35 for active ligand and 5.10 for inactive ligand. For this, a total of 16 molecules were considered active, 14 molecules as inactives, and rest were considered moderately active. In the case of JAK3 inhibitors, threshold value was taken 8.20 for actives and 5.00 for inactives that contain total of 16 molecules active and 15 molecules as inactive. The conformational space of each molecule was then sampled using MCMM/LMOD algorithm. A maximum of 1,000 conformers were generated for each molecule within energy window of 20 kcal/mol and root mean square deviation (RMSD) value of 1 Å to remove unnecessary conformers. For the generation of pharmacophore models, the software was restricted to explore minimum of 4 and maximum of 5 sites for both JAK2 and JAK3 inhibitors and these models were restricted to match 16 of 16 and 12 of 16 active molecules, respectively. A total of 17 hypotheses were obtained for JAK2 and 13 for JAK3 inhibitors and were subsequently ranked on the basis of survival score and survival minus inactive (\(S-I\)) score that correspond to score active and score inactive, respectively. The hypotheses so generated had very similar active and inactive score; therefore, in order to avoid selection of similar kind of hypothesis, all the generated hypotheses were clustered and a representative of each cluster was selected on the basis of highest \(S-I\) score. Thus, a total of 5 hypotheses were selected each for JAK2 and JAK3 belonging to different cluster and the statistical parameters of these models are reported in the Table 1.
The 3D-QSAR models for both the datasets were generated by dividing the dataset molecules into test and training set molecules considering uniform variation of biological activity of the molecules given in Table 2. The best hypothesis of JAK2 was ADRR.92, indicating that JAK2 inhibitors have one hydrogen bond acceptor (A), one HBD (D), and two ring aromatic (R) features. This hypothesis was selected on the basis of highest \(Q_{\mathrm{test}}^{2}\) value i.e., 0.663 and also showed high \(R_{\mathrm{train}}^{2}\) (0.849) and F value (219.3). The large value of F (Fisher test value) indicated a statistically significant regression model. Similarly, best hypothesis of JAK3 (ADDRR.142) was selected which showed high \(Q^{2}_{\mathrm{test}}\) (0.655) and also showed high \(R^{2}_{\mathrm{train}}\) (0.869) and F value (206.9). The ADDRR.142 comprised five features including one HBA (A), two HBD (D), and two ring aromatic (R) features. The selected models of both JAK2 and JAK3 also rendered the good predictive power over other models. The statistical results of generated QSAR models are mentioned in Table 3. The spatial arrangement of features along with their distance present in four (ADRR.92) and five (ADDRR.142) featured pharmacophore models of JAK2 and JAK3 and its mapping over their corresponding highest active molecules is shown in the Fig. 1a–d. The correlation graphs obtained between experimental and the predicted activity of training set molecules and test set molecules obtained from the best models are displayed in Fig. 2a–d. The high values of Pearson-r for test set molecules i.e., 0.837 (JAK2) and 0.816 (JAK3) described the high external predictive ability of these models. The best models also showed acceptable values of \(k\), 1.007; \(k^{\prime }\), 0.984; \(R_{\mathrm{o}}^{2}, 0.995, {R_{\mathrm{o}}^{\prime }}^{2}\), 0.989 for JAK2 and \(k\), 0.991; \(k^{\prime }\), 1.000; \(R_{\mathrm{o}}^{2}\), 0.993, \({R_{\mathrm{o}}^{\prime }}^{2}\), 1.000 for JAK3, confirming the prediction reliability of both selected models. The values of calculated APD of test set molecules were also observed to be within the range of calculated APD of training set molecules indicating the reliability of models for the prediction of new compounds (Table 4). In Y-randomization, the selected 3D-QSAR models of JAK2 and JAK3 exhibited lower values of \(R_{\mathrm{scramble}}^{2}\), i.e., 0.638 and 0.684, respectively, as compared to corresponding original values of \(R_{\mathrm{train}}^{2}\), confirming the trueness of the selected models. Three features, i.e., HBA (A), HBD (D), and ring aromatic (R) of the selected model of JAK2 inhibitors are similar to that of already published pharmacophore model.
Virtual screening
In our integrated VS protocol, 1.5 million PHASE database molecules were screened through the validated pharmacophore model of JAK2 and 1,000 molecules were retrieved due to restriction of maximum output molecules. These 1,000 molecules were subsequently screened through JAK3 model that retrieved 436 molecules. These 436 molecules contained the structural features of both pharmacophore models but could have different conformations that might not properly interact with JAK2 and JAK3 proteins. To avoid the selection of false negative molecules that did not interact properly with proteins and false positive molecules that show the interactions but not favourable, docking analysis was carried out. A number of 3D structures of both JAK2 and JAK3 in complex with different ligands are available in PDB data bank. For the docking analysis, suitable structures were selected on the basis of cross-docking experiments (Fig 3).
Protein selection
Cross-docking is the process of extraction of all ligands from their crystal structures and then redocked those ligands in each crystal protein individually. The redocked ligands are then aligned over their actual crystal ligands, and deviation between the two ligands is assessed as RMSD. The average RMSD of all redocked ligands in each protein was determined to signify the quality of crystal protein. The lower RMSD value represents the ability of the crystal protein to dock the molecules more accurately. Among 28 crystal structures available for JAK2 with different ligands, six (PDB ID: 3KRR, 3E62, 3E63, 3E64, 3UGC, and 4FVQ) were selected on the basis of good resolution \(<\)2 Å. From these, 3UGC showed the lowest RMSD value [34–37]. On the other hand, eight crystal structures of JAK3 (PDB ID: 1YVJ, 4HVD, 3PJC, 4HVI, 4HVH, 4HVG, 3LXK, and 3LXL) were available at PDB site [15, 38–40]. Among them, 4HVD was selected for further docking processes on the basis of cross-docking experiments. The result of cross-docking analysis is mentioned in Table 5. Thus, crystal proteins 3UGC for JAK2 and 4HVD for JAK3 were selected for further docking analysis of hits retrieved after VS.
Investigation of important amino acid residues of JAK2 and JAK3 for their inhibition
Not all amino acid residues present in active site of JAK2 and JAK3 are equally important for their inhibition. Hence, relative importance of active site amino acid residues for inhibition of both enzymes was determined using docking analysis of highest active molecule. Additionally, a dual inhibitor of JAK3, compound AG490, which is under clinical development, was also subjected to docking analysis. The interactions shown by highest active molecule of JAK2 and JAK3 and dual inhibitor AG490 with active site amino acid residues of these enzymes are mentioned in Table 6. From the above docking analysis it was concluded that amino acid residues Glu930 and Leu932 are crucial for JAK2 inhibitory activity, whereas Glu903 and Leu905 amino acid residues are important for JAK3 inhibitory activity. Moreover, available crystal structures of JAK2 and JAK3 were visually analyzed for determination of important amino acid residues for inhibition of these enzymes. The amino acid residues found important after visual inspection were complementary to obtained docking results. The interactions shown by crystal structures of JAK2 and JAK3 are mentioned in Table 5.
Docking, ADME, and energy calculation
Docking was carried out to increase the power of pharmacophore-based screening and to differentiate between active and inactive ligands. The 436 retrieved database candidates from VS were docked into the active site of JAK2 protein. Among them, 204 molecules showing the important inhibitory interactions with amino acid residues Glu930 and Leu932 were selected and subsequently docked with JAK3 protein. After visual inspection, 69 compounds were extracted on the basis of interaction of these candidates with important amino acid residues Glu903 and Leu905. These 69 molecules thus obtained possess dual inhibitory activity against JAK2 and JAK3 and showed interaction with both types of proteins.
Finally to filter the molecules with poor pharmacokinetic profiles, ADME properties were calculated. Incorporation of ADME predictions as a part of the drug development process can generate lead compounds that are more likely to exhibit satisfactory ADME performances during clinical trials. A total of 24 candidates out of 69 were filtered out on the basis of pharmacokinetic parameters thereby indicating their potential to act as drug-like molecules. The best 24 dual JAK2/JAK3 inhibitors were further subjected to Prime MM–GBSA for the calculation of binding free energy. The MM–GBSA binding free energy of all ligands along with its pharmacokinetic parameters of the final 24 hits is given in Table 7. The Prime MM–GBSA binding free energy ranging \(-\)59.545 to \(-\)98.069 (JAK2) and \(-\)54.544 to \(-\)77.280 (JAK3) and glide docking score \(-\)12.405 to \(-\)6.414 (JAK2) and \(-\)6.029 to \(-\)10.224 (JAK3) of the selected 24 molecules are comparable to the binding free energy of clinical trial drug AG-490 suggesting good binding affinity with enzymes. The MM–GBSA scores of the selected molecules were also found to be comparable to those of the co-crystal ligands with MM–GBSA scores for JAK2 and JAK3 ligands being \(-\)152.081 and \(-\)79.942, respectively. Interestingly, it was found that basic structural motifs of two hits (Phase ID 0855481 and 0855521) among 24 have been filed as patent by Almirall S.A. as dual JAK2/3 inhibitor [41]. Thus, these 24 candidates could be better drug candidates for targeting JAK2 and JAK3 enzymes. The interaction details along with glide g-score of all 24 final hits are mentioned in Table 8.
Docking results of one of the hit compounds, i.e., compound 0288703, that showed good dual inhibitory activity are displayed in Fig. 4. The compound 0288703 shows two hydrogen bonding interactions with the JAK2 protein (Fig. 4a). The hydroxyl group and the carbonyl group of the compound interact with carbonyl group of Glu 930 and amino group of Leu 932, respectively. This compound also interacts with JAK3 protein giving five hydrogen bonding interactions (Fig. 4b). The hydroxyl group of the compound forms two hydrogen bonds with Leu 905; the oxygen atom and hydrogen atom of this hydroxyl group give interaction with amino group and carbonyl group of Leu 905, respectively. The nitrogen of pyridine nucleus of the compound shows interaction with NH of the side chain of Arg 911, the carbonyl group of the compound interacts with SH of Cys903, and the NH of the compound interacts with Leu828.
Conclusion
Multi-target potential of the drug molecules is considered as beneficial for the treatment of multi-pathway diseases. Lymphoid-derived diseases originate from multiple pathways; thus, its intervention with molecules having multi-target potential would serve as better therapy. Two well-known kinases JAK2 and JAK3 reported in the literature are involved in the progression of lymphoid-derived diseases. Thus, our study aimed to design some new molecules describing inhibitory potential for both JAK2 and JAK3.
Two ligand-based pharmacophore models were generated for the dataset of inhibitor molecules of JAK2 and JAK3 to dig out the essential structural features required for inhibition of both enzymes which are helpful for screening of novel molecules having inhibitory activity against both enzymes. The selected models as shown by the correlation statistics and predictive statistics are very much significant to draw explicit inferences. Finally, 24 potential hits with good pharmacokinetic profile and predicted activity were identified by calculating ADME properties and docking analysis, respectively. This study provides a set of guidelines that will greatly help in designing the novel and more potent JAK2/JAK3 dual inhibitors.
References
Heinrich PC, Behrmann I, Müller-Newen G, Schaper F, Graeve L (1998) Interleukin-6-type cytokine signalling through the gp130/Jak/STAT pathway. Biochem J 334:297–314
Smith KA (1988) Interleukin-2: inception, impact, and implications. Science 240:1169–1176. doi:10.1126/science.3131876
Meydan N, Grunberger T, Dadi H, Shahar M, Arpaia E, Lapidot Z, Leeder JS, Freedman M, Cohen A, Gazit A, Levitzki A, Roifman CM (1996) Inhibition of acute lymphoblastic leukaemia by a Jak-2 inhibitor. Nature 379:645–648. doi:10.1038/379645a0
Kirken RA, Erwin RA, Taub D, Murphy WJ, Behbod F, Wang L, Pericle F, Farrar WL (1999) Tyrphostin AG-490 inhibits cytokine-mediated JAK3/STAT5a/b signal transduction and cellular proliferation of antigen-activated human T cells. J Leukoc Biol 65:891–899
Weinberg LR, Albom MS, Angeles TS, Breslin HJ, Gingrich DE, Huang Z, Lisko JG, Mason JL, Milkiewicz KL, Thieu TV (2011) 2,7-Pyrrolo[2,1-f][1,2,4]triazines as JAK2 inhibitors: modification of target structure to minimize reactive metabolite formation. Bioorg Med Chem Lett 21:7325–7330. doi:10.1016/j.bmcl.2011.10.032
Zificsak CA, Gingrich DE, Breslin HJ, Dunn DD, Milkiewicz KL, Theroff JP, Thieu TV, Underiner TL, Weinberg LR, Aimone LD (2012) Optimization of a novel kinase inhibitor scaffold for the dual inhibition of JAK2 and FAK kinases. Bioorg Med Chem Lett 22:133–137. doi:10.1016/j.bmcl.2011.11.049
Lim J, Taoka B, Otte RD, Spencer K, Dinsmore CJ, Altman MD, Chan G, Rosenstein C, Sharma S, Su H-P (2011) Discovery of 1-amino-5H-pyrido[4,3-b]indol-4-carboxamide inhibitors of Janus kinase 2 (JAK2) for the treatment of myeloproliferative disorders. J Med Chem 54:7334–7349. doi:10.1021/jm200909u
Siu T, Kozina ES, Jung J, Rosenstein C, Mathur A, Altman MD, Chan G, Xu L, Bachman E, Mo J-R (2010) The discovery of tricyclic pyridone JAK2 inhibitors. Part 1: hit to lead. Bioorg Med Chem Lett 20:7421–7425. doi:10.1016/j.bmcl.2010.10.031
Harikrishnan LS, Kamau MG, Wan H, Inghrim JA, Zimmermann K, Sang X, Mastalerz HA, Johnson WL, Zhang G, Lombardo LJ (2011) Pyrrolo[1,2-f]triazines as JAK2 inhibitors: achieving potency and selectivity for JAK2 over JAK3. Bioorg Med Chem Lett 21:1425–1428. doi:10.1016/j.bmcl.2011.01.022
Gerspacher M, Furet P, Pissot-Soldermann C, Gaul C, Holzer P, Vangrevelinghe E, Lang M, Erdmann D, Radimerski T, Regnier CH (2010) 2-Amino-aryl-7-aryl-benzoxazoles as potent, selective and orally available JAK2 inhibitors. Bioorg Med Chem Lett 20:1724–1727. doi:10.1016/j.bmcl.2010.01.069
Pissot-Soldermann C, Gerspacher M, Furet P, Gaul C, Holzer P, McCarthy C, Radimerski T, Regnier CH, Baffert F, Drueckes P (2010) Discovery and SAR of potent, orally available 2,8-diaryl-quinoxalines as a new class of JAK2 inhibitors. Bioorg Med Chem Lett 20:2609–2613. doi:10.1016/j.bmcl.2010.02.056
Malerich JP, Lam JS, Hart B, Fine RM, Klebansky B, Tanga MJ, D’Andrea A (2010) Diamino-1,2,4-triazole derivatives are selective inhibitors of TYK2 and JAK1 over JAK2 and JAK3. Bioorg Med Chem Lett 20:7454–7457. doi:10.1016/j.bmcl.2010.10.026
Schenkel LB, Huang X, Cheng A, Deak HL, Doherty E, Emkey R, Gu Y, Gunaydin H, Kim JL, Lee J (2011) Discovery of potent and highly selective thienopyridine Janus kinase 2 inhibitors. J Med Chem 54:8440–8450. doi:10.1021/jm200911r
Thoma G, Nuninger F, Falchetto R, Hermes E, Tavares GA, Vangrevelinghe E, Zerwes HG (2010) Identification of a potent Janus kinase 3 inhibitor with high selectivity within the Janus kinase family. J Med Chem 54:284–288. doi:10.1021/jm101157q
Burns CJ, Bourke DG, Andrau L, Bu X, Charman SA, Donohue AC, Fantino E, Farrugia M, Feutrill JT, Joffe M (2009) Phenylaminopyrimidines as inhibitors of Janus kinases (JAKs). Bioorg Med Chem Lett 19:5887–5892. doi:10.1016/j.bmcl.2009.08.071
Derenzini E, Lemoine M, Buglio D, Katayama H, Ji Y, Davis R, Sen S, Younes A (2011) The JAK inhibitor AZD1480 regulates proliferation and immunity in Hodgkin lymphoma. Blood Cancer J 1:e46. doi:10.1038/bcj.2011.46
Pardanani A, Lasho T, Smith G, Burns C, Fantino E, Tefferi A (2009) CYT387, a selective JAK1/JAK2 inhibitor: in vitro assessment of kinase selectivity and preclinical studies using cell lines and primary cells from polycythemia vera patients. Leukemia 23:1441–1445. doi:10.1038/leu.2009.50
Quintás-Cardama A, Vaddi K, Liu P, Manshouri T, Li J, Scherle PA, Caulder E, Wen X, Li Y, Waeltz P, Rupar M, Burn T, Lo Y, Kelley J, Covington M, Shepard S, Rodgers JD, Haley P, Kantarjian H, Fridman JS, Verstovsek S (2010) Preclinical characterization of the selective JAK1/2 inhibitor INCB018424: therapeutic implications for the treatment of myeloproliferative neoplasms. Blood 115:3109–3117. doi:10.1182/blood-2009-04-214957
Wilson LJ, Malaviya R, Yang C, Argentieri R, Wang B, Chen X, Murray WV, Cavender D (2009) Synthetic staurosporines via a ring closing metathesis strategy as potent JAK3 inhibitors and modulators of allergic responses. Bioorg Med Chem Lett 19:3333–3338. doi:10.1016/j.bmcl.2009.04.039
Chen JJ, Thakur KD, Clark MP, Laughlin SK, George KM, Bookland RG, Davis JR, Cabrera EJ, Easwaran V, De B (2006) Development of pyrimidine-based inhibitors of Janus tyrosine kinase 3. Bioorg Med Chem Lett 16:5633–5638. doi:10.1016/j.bmcl.2006.08.022
Clark MP, George KM, Bookland RG, Chen J, Laughlin SK, Thakur KD, Lee W, Davis JR, Cabrera EJ, Brugel TA (2007) Development of new pyrrolopyrimidine-based inhibitors of Janus kinase 3 (JAK3). Bioorg Med Chem Lett 17:1250–1253. doi:10.1016/j.bmcl.2006.12.018
Adams C, Aldous DJ, Amendola S, Bamborough P, Bright C, Crowe S, Eastwood P, Fenton G, Foster M, Harrison TK (2003) Mapping the kinase domain of Janus kinase 3. Bioorg Med Chem Lett 13:3105–3110. doi:10.1016/S0960-894X(03)00657-7
Cole AG, Bohnstedt AC, Paradkar V, Kingsbury C, Quintero JG, Park H, Lu Y, You M, Neagu I, Diller DJ (2009) 2-Benzimidazolyl-9-(chroman-4-yl)-purinone derivatives as JAK3 inhibitors. Bioorg Med Chem Lett 19:6788–6792. doi:10.1016/j.bmcl.2009.09.080
Maestro, version 9.3, User Manual, Schrödinger, LLC, New York (2012)
Ligprep, version 2.5, User Manual, Schrödinger, LLC, New York (2012)
PHASE, version 3.4, Schrödinger, LLC, New York (2012)
Dixon SL, Smondyrev AM, Knoll EH, Rao SN, Shaw DE, Friesner RA (2006) PHASE: a new engine for pharmacophore perception, 3D QSAR model development, and 3D database screening.1. Methodology and preliminary results. J Comput Aided Mol Des 20:647–671. doi:10.1007/s10822-006-9087-6
Verma J, Khedkar VM, Coutinho EC (2010) 3D-QSAR in drug design: a review. Curr Top Med Chem 10:95–115. doi:10.2174/156802610790232260
Golbraikh A, Tropsha A (2002) Beware of q2!. J Mol Graph Model 20:269–276. doi:10.1016/S1093-3263(01)00123-1
Glide, version 5.8, User Manual, Schrödinger, LLC, New York (2012)
Friesner RA, Banks J, Murphy RB, Halgren TA, Klicic JJ, Mainz DT, Repasky MP, Knoll EH, Shaw DE, Shelley M, Perry JK, Francis P, Shenkin PS (2004) Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J Med Chem 47:1739–1749. doi:10.1021/jm0306430
QikProp, version 3.5, User Manual, Schrödinger, LLC, New York, NY (2012)
Singh D, Karthikeyan M, Kirubakaran P, Nagamani S (2011) Pharmacophore filtering and 3D-QSAR in the discovery of new JAK2 inhibitors. J Mol Graph Model 30:186–197. doi:10.1016/j.jmgm.2011.07.004
Baffert F, Régnier CH, De Pover A, Pissot-Soldermann C, Tavares GA, Blasco F, Brueggen J, Chène P, Drueckes P, Erdmann D (2010) Potent and selective inhibition of polycythemia by the quinoxaline JAK2 inhibitor NVP-BSK805. Mol Cancer Ther 9:1945–1955. doi:10.1158/1535-7163
Antonysamy S, Hirst G, Park F, Sprengeler P, Stappenbeck F, Steensma R, Wilson M, Wong M (2009) Fragment-based discovery of JAK-2 inhibitors. Bioorg Med Chem Lett 19:279–282. doi:10.1016/j.bmcl.2008
Andraos R, Qian Z, Bonenfant D, Rubert J, Vangrevelinghe E, Scheufler C, Marque F, Régnier CH, De Pover A, Ryckelynck H (2012) Modulation of activation-loop phosphorylation by JAK inhibitors is binding mode dependent. Cancer Discov 2:512–523. doi:10.1158/2159-8290
Bandaranayake RM, Ungureanu D, Shan Y, Shaw DE, Silvennoinen O, Hubbard SR (2012) Crystal structures of the JAK2 pseudokinase domain and the pathogenic mutant V617F. Nat Struct Mol Biol 19:754–759. doi:10.1038/nsmb.2348
Boggon TJ, Li Y, Manley PW, Eck MJ (2005) Crystal structure of the Jak3 kinase domain in complex with a staurosporine analog. Blood 106:996–1002. doi:10.1182/blood-2005-02-0707
Soth M, Hermann JC, Yee C, Alam M, Barnett JW, Berry P, Browner MF, Frank K, Frauchiger S, Harris S (2012) 3-Amido pyrrolopyrazine JAK kinase inhibitors: development of a JAK3 vs JAK1 selective inhibitor and evaluation in cellular and in vivo models. J Med Chem 56:345–356. doi:10.1021/jm301646k
Chrencik JE, Patny A, Leung IK, Korniski B, Emmons TL, Hall T, Weinberg RA, Gormley JA, Williams JM, Day JE (2010) Structural and thermodynamic characterization of the TYK2 and JAK3 kinase domains in complex with CP-690550 and CMP-6. J Mol Biol 400:413–433. doi:10.1016/j.jmb.2010.05.020
Dymock BW, See CS (2013) Inhibitors of JAK2 and JAK3: an update on the patent literature 2010–2012. Expert Opin Ther Pat 23:449–501. doi:10.1517/13543776.2013.765862
Acknowledgments
Authors would like to thank Dr. Ravikumar Muttineni (Application Scientist), Er. Anirban Banerjee (IT Consultant), and Mr. Raghu Rangaswamy from Schrödinger, Bangalore, for their constant scientific and technical support to handle Schrödinger software and work smoothly. Authors also thank University Grant Commission, New Delhi, for providing the financial support; Grant No. 37-324/2009(SR).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jasuja, H., Chadha, N., Kaur, M. et al. Dual inhibitors of Janus kinase 2 and 3 (JAK2/3): designing by pharmacophore- and docking-based virtual screening approach. Mol Divers 18, 253–267 (2014). https://doi.org/10.1007/s11030-013-9497-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-013-9497-z