Abstract
XPO5 (Exp5, Exportin-5) is a transporter protein mainly mediating pre-microRNAs’ nuclear export. Recent studies have demonstrated that XPO5 may play crucial roles in a few of cancers. However, little is known about XPO5 in hepatocellular carcinoma (HCC). In the present study, we elucidated the expression of XPO5 by quantitative real-time PCR (qRT-PCR) and immunohistochemical staining in HCC samples and conducted several functional analyses to address its effects on HCC development. The results demonstrated that both mRNA and protein levels of XPO5 were downregulated in HCC tissues compared to adjacent non-cancerous livers. Ectopic expression of XPO5 significantly suppressed cell proliferation, colony formation, growth in soft agar, and tumorigenicity in nude mice, whereas knockdown of XPO5 by RNA inference showed opposite phenotypes. Moreover, XPO5 knockdown promoted HCC cell migration and decreased the expression of E-cadherin and p53. Additionally, after treatment with DAC and TSA, the mRNA level of XPO5 was upregulated in HCC cells tested, implicating that epigenetic modulation may be involved in the transcription of XPO5. Collectively, our findings suggest that XPO5 functions as a potential tumor suppressor in the development and progression of HCC as well as a promising molecular target for HCC therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC) is the most common primary malignancy of hepatocytes, which serves as the third leading cause of cancer-related death worldwide [1]. Surgical resection or liver transplantation is the mainstay of treatment for early HCC patients; however, the majority of patients present at an advanced stage. Accordingly, several other treatment modalities including chemotherapy and radiotherapy are used to treat HCC but their effects are still not satisfactory. So, it is urgent to identify more effective diagnostic markers and therapeutic strategies associated with the initiation or progression of HCC [2].
Growing evidences have shown that the genes in processing pathway of microRNA (miRNA) are involved in the regulation of a large variety of human cancer types [3]. Alteration of these genes such as DGCR8, Dicer, XPO5, AGO2, and TRBP by genomic mutations, aberrant expression, or other means could significantly affect cancer initiation, progression, and metastasis [4–6]. Among them, XPO5 is a member of the karyopherin β family that uses the GTPase Ran to control cargo association. It is mainly responsible for exporting precursor microRNAs (pre-miRNAs) through the nuclear membrane to the cytoplasm [7]. A previous study indicates that the XPO5-inactivating mutations in its C-terminal have been found in a subset of human cancers with microsatellite instability, including colon, stomach, and endometrium [8]. This genetic defect traps pre-miRNAs in the nucleus, decreases miRNA processing, and reduces the target repression of mature miRNAs, thereby leading to cancer development. In addition, a single-nucleotide polymorphism (SNP, rs11077) of the XPO5 gene in its 3′ UTR region could increase the risk of renal cell carcinoma [9], could have association with survival of advanced small-cell lung cancer patients [10], and also could predict independently worse survival for HCC patients [11]. Notably, there have been implications that XPO5 is upregulated in urothelial carcinoma of the bladder and breast cancer [12, 13]. Inhibition of XPO5 induction during cell cycle results in delayed G1/S transition and suppresses cell growth [14]. Therefore, the roles of XPO5 in different cancer cells seem to be conflicting.
In the present study, we evaluated the expression of XPO5 in HCC tissues and conducted several in vitro and in vivo assays to characterize the exact function of XPO5 in HCC cell growth and migration. Our final data suggest that XPO5 is frequently downregulated in cancer samples and this downregulation may contribute to HCC.
Materials and methods
Tissue specimens and cell culture
Thirty seven pairs of human HCC and adjacent non-tumor tissues were obtained from patients who underwent surgical liver resection and signed an informed consent in Shanghai East Hospital, Tongji University, China. The diagnoses of these liver cancer samples were verified by pathologists. The samples were frozen in liquid nitrogen and stored at −80 °C until processed. The human HCC tissue array (HLiv-HCC150CS-01) was purchased from Shanghai Outdo Biotech Corporation, China, which contains 75 pairs of HCC samples. The immunohistochemical staining was performed according to the commercial protocol (Outdo Biotech). XPO5 expression was assessed using an anti-XPO5 antibody (Cat#ab57491, Abcam, USA) at a dilution of 1:100. The use of all above tissue materials for research was approved by the Ethics Committee of Shanghai East Hospital. HCC cell lines were taken from our laboratory stocks. All of the cells were maintained in Dulbecco’s Modified Eagle’s Medium (DMEM) supplemented with 10 % fetal bovine serum (FBS) (Gibco, USA) and antibiotics (50 U/ml penicillin and 50 µg/ml streptomycin) (Gibco, USA) at 37 °C in a 5 % CO2 humidified incubator.
RNA interference and plasmid construction
The specific small interference RNA (siRNA) against XPO5 was chemically synthesized by GenePharma, Shanghai, China, which has been described previously [7]. The siXPO5 sequence was as follows: sense 5′-GAUGCUCUGUCUCGAAUUGUATTdTdT-3′. The irrelevant nucleotides not targeting any annotated human genes were used as a negative control (si-NC): sense 5′-UUCUCCGAACGUGUCACGUdTdT-3′. The full-length cDNA of XPO5 is a gift from Dr. Bryan R. Cullen, Duke University Medical Center. The ORF of XPO5 was amplified and then subcloned into pcDNA3.1, generating pcDNA3.1-XPO5. Accurate reading frame insertion was verified by DNA sequencing. The lentivirus-overexpressing XPO5 (LV-XPO5, 1 × 108 TU/ml) and knocking down XPO5 (LV-shXPO5, 1 × 108 TU/ml) were packaged and purchased from Tuzhu Biotech and GenePharma, Shanghai, respectively, using the above corresponding sequences. HCC cells were infected with lentivirus at 10–20 MOI, and pools of infected cells were selected with 3 μg/ml puromycin.
Cell transfection
Cell transfection with plasmids or siRNAs was conducted using Lipofectamine 2000 (Invitrogen, USA) in accordance with the manufacturer’s instructions.
Cell proliferation and colony formation assay
Cell proliferation was measured using the Cell Counting Kit-8 (CCK-8; Dojindo, Kumamoto, Japan) as previously described [15]. For colony formation, HCC cells (1 × 104 per 6-cm plate) transfected with empty vector or pcNDA3.1-XPO5 were selected in appropriate medium with the addition of 0.6–0.8 mg/ml G418 (Life Technologies, USA). After 3 weeks, the colonies stained with crystal violet were photographed and counted.
Soft agar colony formation assay
HCC cells were suspended in the media containing 0.4 % agar and overlaid on 1 % agar in 24-well plates (1000 cells/well). After 2–4 weeks, colonies were counted and photographed. All experiments were independently repeated at least three times.
Western blot analysis
Total protein extracts of HCC cells were prepared as described previously [16]. Samples were boiled for 5 min, subjected to electrophoresis in SDS-PAGE, and transferred onto a polyvinylidene difluoride (PVDF) membrane. The membrane was blocked in PBS with 5 % nonfat milk for 2 h at room temperature and then incubated with the primary antibody overnight at 4 °C. Incubation with the secondary antibody was performed for 1 h at room temperature. Anti-E-cadherin (#14472, Cell Signaling Technology, USA), XPO5, P53, and β-actin (sc-66885, sc-126, sc-47778, Santa Cruz Biotechnology, CA, USA) antibodies were used in this study. Detection of proteins was achieved using the Odyssey Infrared Imaging System (Li-COR, USA).
Cell migration assay
Migration assays were performed using 24-well transwell chambers (8 μm pore size, BD biosciences, USA) according to the instructions of the manufacturer. Briefly, serum-starved cells were trypsinized and counted. 5 × 104 of HCC cells in 200 μl serum-free medium were placed into the upper chamber. The lower chamber was filled with 500 μl of 10 % FBS–DMEM. After 24–48 h, cells that migrated to the lower chamber were fixed and stained with 0.5 % crystal violet. Cells in at least five random microscopic fields (magnification ×100) were counted and photographed. All experiments were performed in duplicate and repeated three times.
Tumor implantation study
One group of 4–5-week-old nude mice (strain BALB/c nude; SLAC Laboratories Animal, Shanghai, China) received subcutaneous injections with the following amount of HCC cells: 1.5 × 106 of Sk-Hep-1 cells stably expressing XPO5 or empty vector, 1 × 106 of MHCC-LM3 cells stably silencing XPO5 expression, or control shNC. Tumor growth was measured weekly for 4–5 weeks and the isolated tumors were weighed after mice were sacrificed. All animal handling and experimental procedures were approved by the Ethics Committee of the Shanghai East Hospital, Tongji University.
RNA extraction and quantitative real-time PCR
HCC cells were treated with decitabine (DAC, 1 μM, Selleck, USA) for 48 h or trichostatin A (TSA, 0.3 μM, Selleck, USA) for 24 h. Cells treated with DMSO were used as a control. Total RNA was extracted from HCC cells or HCC samples using TRIZOL reagent (Invitrogen, USA) and was reverse transcribed into cDNA using M-MLV reverse transcriptase kit (Promega, USA). Quantitative RT-PCR was performed with Thermal Cycler Dice Real Time System (TaKaRa, Japan) and SYBR Green I reagent (TaKaRa, Japan). The mRNA level of XPO5 was normalized by β-actin prior to comparative analysis using \(2^{-{\Delta}Ct}\) method. The following primer sets were used: For XPO5, forward: 5-CACAACAAGGAGAGGTGATGAG-3; reverse: 5-AAGGTGAGAAGACGGAACAGAG-3. For β-actin, forward: 5-AGAGCCTCGCCTTTGCCGATCC-3; reverse: 5-CTGGGCCTCGTCGCCCACATA-3. All reactions were performed independently at least three times in triplicate.
Statistical analysis
All statistical analyses were performed with Student’s t test using GraphPad Prism 5 software. P values of less than 0.05 were considered statistically significant.
Results
XPO5 is downregulated in HCC samples
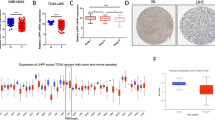
Quantitative real-time PCR (qRT-PCR) was performed to detect XPO5 expression in 37 pairs of HCC specimens. The results showed that XPO5 expression in HCC samples was much lower than that in the corresponding adjacent non-cancerous tissues (Fig. 1a). Among them, 17/37 (45.9 %) of HCC samples have at least a 1.5-fold decrease. Furthermore, immunohistochemistry for XPO5 expression was performed on additional 75 pairs of HCC samples in a tissue microarray. 32/75 (42.7 %) of cancerous tissues displayed weak staining compared with their normal counterparts. Two pairs of representative tissue images of XPO5 staining are shown in Fig. 1b. These results indicated that both mRNA and protein level of XPO5 were downregulated in HCC, implicating that XPO5 may play tumor-suppressive roles in HCC development. In addition, we also explored the expression of XPO5 in a series of HCC cell lines by qRT-PCR (Fig. 1c). Based on the data, we employed Huh7, HepG2, or Sk-Hep-1 cells for XPO5 overexpression and MHCC-LM3 and PLC/PRF/5 cells for XPO5 silencing in the following functional assays.
Expression pattern of XPO5 in HCC samples and cell lines. a XPO5 expression was measured in 37 pairs of HCC and non-HCC tissues by qRT-PCR. Relative transcript levels were normalized by ΔCt method using β-actin as an internal reference. N non-cancerous tissue, C cancerous tissue. **P < 0.01. b Representative pictures showing XPO5 expression in two pairs of human HCC samples. Magnification ×200. c Relative mRNA level of XPO5 was evaluated in HCC cell lines by qRT-PCR, where β-actin was used as an internal control
XPO5 overexpression inhibits and knockdown promotes HCC cell growth
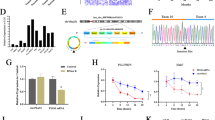
To elucidate the effect of XPO5 on cell growth, pcDNA3.1-XPO5 plasmid was transiently transfected into Hhu7 cells and specific siRNAs against XPO5 were transiently transfected into MHCC-LM3 and PLC/PRF/5 cells. Western blot results confirmed that XPO5 was successfully overexpressed or silenced in these cells (Fig. 2a). The effect of XPO5 on cell growth was evaluated through CCK8 method in the three HCC cell lines above. Figure 2b–d shows that overexpression of XPO5 inhibits Huh7 cell growth and knockdown of XPO5 promotes MHCC-LM3 and PLC/PRF/5 cell growth.
Effect of XPO5 expression on cell proliferation. a Ectopic expression of XPO5 in Huh7 cells and silencing expression by siXPO5 in MHCC-LM3 and PLC/PRF/5 cells were analyzed by western blotting. b Cell viability assays were performed to explore the effect of XPO5 overexpression on cell growth in Huh7 cells. c and d Cell viability assays were performed to explore the effect of XPO5 knockdown on cell growth in MHCC-LM3 and PLC/PRF/5 cells. The bars represent the standard deviation of independent experiments conducted in triplicate (mean ± SD). *P < 0.05; **P < 0.01
XPO5 overexpression reduces and knockdown enhances colony formation of HCC cells
Next, colony formation assays were carried out to further examine the suppression of XPO5 on cell growth. As expected, overexpression of XPO5 in Huh7 and HepG2 cells weakened colony formation on the normal culture plates, where the colonies were smaller and less than those with control vector (Fig. 3a). Furthermore, we also detected the growth ability of HCC cells influenced by XPO5 expression in soft agar. This anchorage-independent growth manner reflects the cancer cell malignancy and metastatic potential. As shown in Fig. 3b, knockdown of XPO5 in MHCC-LM3 cells increased colony formation and overexpression of XPO5 in Sk-Hep-1 cells decreased colony formation when cells were allowed to grow in soft agar.
Effect of XPO5 expression on colony formation of HCC cells. a Ectopic expression of XPO5 inhibited colony formation of Huh7 and HepG2 cells on normal plates. b XPO5 knockdown in MHCC-LM3 cells enhanced and XPO5 overexpression in Sk-Hep-1 cells decreased cell growth ability in soft agar, where si-NC and empty vector were used as controls. The histogram shows the mean and standard deviation of three independent experiments. *P < 0.05; **P < 0.01
XPO5 plays a suppressive role in xenograft tumor growth
To address whether XPO5 expression could play a role in xenograft tumor growth, HCC cells were subcutaneously injected into two flanks of the same nude mice (n = 6). Figure 4a shows the images and weights of tumors derived from Sk-Hep-1 cells stably expressing XPO5 and control cells, indicating that XPO5 overexpression significantly suppressed xenograft tumor growth. Conversely, the tumors derived from MHCC-LM3 cells stably knocking down XPO5 were larger in size and more in weight than those from the corresponding control cells (Fig. 4b). These results strongly suggested that XPO5 suppresses tumorigenicity of HCC cells in nude mice.
XPO5 plays suppressive roles in tumor growth of subcutaneous xenograft. a Overexpression of XPO5 in Sk-Hep-1 cells suppressed xenograft tumor growth in nude mice (n = 6), where those with empty vector were used as a control. b Tumor growth was also observed in nude mice injected with MHCC-LM3 cells in which XPO5 was stably knocked down (n = 6). Nude mice were sacrificed at the end of experiment, and tumors were excised and weighed
Knockdown of XPO5 facilitates HCC cell migration
We next elucidated the role of XPO5 in cell migration by transwell chamber assay. As shown in Fig. 5, a larger number of PLC/PRF/5 cells migrated to the underlayer of membrane in XPO5 knockdown group compared with control group. In the meantime, similar results were observed in MHCC-LM3 cells (Fig. 5), suggesting that XPO5 suppresses migration of HCC cells.
XPO5 knockdown promotes cell migration. PLC/PRF/5 and MHCC-LM3 cells were stably infected with lentivirus-silencing XPO5 or control (LV-shXPO5 and LV-shNC), and then these cells were suffered to determine migration ability through transwell chambers. The histogram shows the mean and standard deviation of three independent experiments. *P < 0.05; **P < 0.01
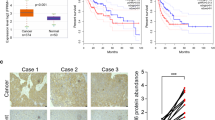
Knockdown of XPO5 results in downregulation of E-cadherin and p53
To explore the mechanism by which XPO5 suppresses HCC cell growth and migration, a series of cancer proliferation- and metastasis-related factors were detected by western blot analysis. Figure 6a indicates that E-cadherin and p53, two well-documented tumor suppressors, were reduced in XPO5 knockdown cells, suggesting that the two factors may be involved in the XPO5-mediated suppression of HCC development. Due to the potential mesenchymal phenotype of MHCC-LM3 cells, the epithelial marker E-cadherin was not detected by western blot. In addition, DAC (decitabine, a DNA demethylation agent) and TSA (trichostatin A, a histone deacetylase inhibitor) were used to test whether XPO5 expression is regulated epigenetically. As shown in Fig. 6b, XPO5 expression level was significantly increased after treatment with DAC or TSA in both Sk-Hep-1 and MHCC-LM3 cells, implying that DNA methylation and histone deacetylation inhibit XPO5 expression.
Discussion
Nucleocytoplasmic transport plays important roles in a series of cellular processes, dysregulation of which often results in abnormal cell growth, apoptosis, differentiation, and even cell transformation to tumor [17, 18]. Here we report that a nucleocytoplasmic transporter, the pre-miRNA export machinery XPO5 is involved in the suppression of HCC and is downregulated in HCC samples for the first time.
Cell viability and colony formation assays demonstrated that XPO5 upregulation or downregulation significantly affected HCC cell growth and malignancy. Moreover, the results from subcutaneous xenograft tumor growth model provided the in vivo evidence that XPO5 plays tumor-suppressive roles in HCC cells, in line with the result from the in vitro assays. Importantly, XPO5 was also identified as a tumor suppressor in colorectal cancer cells when similar experiments were employed [8]. We also performed cell migration assay by transwell chambers. Our data indicated that XPO5 may mediate the suppression of cell mobility in vitro. Although there is no report on the role of XPO5 in cell migration before, a study on Dicer has provided some insights into the miRNA processing pathway involved in the regulation of epithelial-to-mesenchymal transition (EMT) [19]. As for the mechanisms by which XPO5 suppresses cell growth and migration, we found that E-cadherin and p53 could act as the downstream effector of XPO5 since the western blot results showed downregulation of the two factors with XPO5 knockdown. In addition, after treatment of DAC and TSA, XPO5 is increased in mRNA level, implying that XPO5 is under control epigenetically. A previous report has shown that the promoter region of XPO5 is suffered with DNA methylation and this alteration in the methylation pattern of XPO5 promoter could be a risk biomarker in breast cancer [13].
As known, miRNAs are short noncoding RNAs that are involved in a wide range of biological and pathological processes through regulating target gene expression [20]. Currently, a great number of miRNAs have been identified as oncogenes or tumor suppressors in various cancer types. By modulating oncogenic and tumor suppressor pathways, they could, in principle, control human cancer development, progression, or metastasis [21, 22]. A report reveals that the global downregulation of mature miRNAs is emerging as a common feature of human cancers [23]. There has also been evidence that global change in miRNA level was caused by expression alteration and gene mutation in those miRNA processing machineries, such as Dicer, TRBP, and XPO5 [5, 8]. In terms of XPO5, studies have shown that the expression alteration or mutation of XPO5 gene has a significant effect on global miRNA level since XPO5 acts as the determiner of pre-miRNAs transporting from nucleus to cytoplasm [8]. Besides, XPO5 could also recognize and export a few types of structured RNAs, including adenovirus VA1 RNA, tRNA, and the signal recognition particle (SRP) RNA [24–26], along with certain other proteins, such as STAU2, ILF3, JAZ, and ADAR1 [27–29]. XPO5 seems to play different roles in different cell types, which is possibly due to these complex potential cargoes in tumor. Further studies should be performed to classify which cargoes are induced to Dys translocation by XPO5 alteration in HCC or in other cancers in future.
In summary, this is the first time the function and expression of XPO5 in HCC has been explored. Our findings support the notion that XPO5 functions as a tumor suppressor in tumor. XPO5 may serve as a potential target for the therapy of HCC and also help gain some insights into the conflicting functions of XPO5 in different cancer cell types.
References
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D (2011) Global cancer statistics. CA Cancer J Clin 61:69–90. doi:10.3322/caac.20107
Thorgeirsson SS, Grisham JW (2002) Molecular pathogenesis of human hepatocellular carcinoma. Nat Genet 31:339–346. doi:10.1038/ng0802-339
Ohtsuka M, Ling H, Doki Y, Mori M, Calin GA (2015) MicroRNA processing and human cancer. J Clin Med 4:1651–1667. doi:10.3390/jcm4081651
Cheng N, Li Y, Han ZG (2013) Argonaute2 promotes tumor metastasis by way of up-regulating focal adhesion kinase expression in hepatocellular carcinoma. Hepatology 57:1906–1918. doi:10.1002/hep.26202
Melo SA, Ropero S, Moutinho C, Aaltonen LA, Yamamoto H, Calin GA, Rossi S, Fernandez AF, Carneiro F, Oliveira C, Ferreira B, Liu CG, Villanueva A, Capella G, Schwartz S Jr, Shiekhattar R, Esteller M (2009) A TARBP2 mutation in human cancer impairs microRNA processing and DICER1 function. Nat Genet 41:365–370. doi:10.1038/ng.317
Merritt WM, Lin YG, Han LY, Kamat AA, Spannuth WA, Schmandt R, Urbauer D, Pennacchio LA, Cheng JF, Nick AM, Deavers MT, Mourad-Zeidan A, Wang H, Mueller P, Lenburg ME, Gray JW, Mok S, Birrer MJ, Lopez-Berestein G, Coleman RL, Bar-Eli M, Sood AK (2008) Dicer, Drosha, and outcomes in patients with ovarian cancer. N Engl J Med 359:2641–2650. doi:10.1056/NEJMoa0803785
Yi R, Qin Y, Macara IG, Cullen BR (2003) Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev 17:3011–3016. doi:10.1101/gad.1158803
Melo SA, Moutinho C, Ropero S, Calin GA, Rossi S, Spizzo R, Fernandez AF, Davalos V, Villanueva A, Montoya G, Yamamoto H, Schwartz S Jr, Esteller M (2010) A genetic defect in exportin-5 traps precursor microRNAs in the nucleus of cancer cells. Cancer Cell 18:303–315. doi:10.1016/j.ccr.2010.09.007
Horikawa Y, Wood CG, Yang H, Zhao H, Ye Y, Gu J, Lin J, Habuchi T, Wu X (2008) Single nucleotide polymorphisms of microRNA machinery genes modify the risk of renal cell carcinoma. Clin Cancer Res 14:7956–7962. doi:10.1158/1078-0432.CCR-08-1199
Ding C, Li C, Wang H, Li B, Guo Z (2013) A miR-SNP of the XPO5 gene is associated with advanced non-small-cell lung cancer. Onco Targets Ther 6:877–881. doi:10.2147/OTT.S48284
Liu S, An J, Lin J, Liu Y, Bao L, Zhang W, Zhao JJ (2014) Single nucleotide polymorphisms of microRNA processing machinery genes and outcome of hepatocellular carcinoma. PLoS ONE 9:e92791. doi:10.1371/journal.pone.0092791
Han Y, Liu Y, Gui Y, Cai Z (2013) Inducing cell proliferation inhibition and apoptosis via silencing Dicer, Drosha, and Exportin 5 in urothelial carcinoma of the bladder. J Surg Oncol 107:201–205. doi:10.1002/jso.23214
Leaderer D, Hoffman AE, Zheng T, Fu A, Weidhaas J, Paranjape T, Zhu Y (2011) Genetic and epigenetic association studies suggest a role of microRNA biogenesis gene exportin-5 (XPO5) in breast tumorigenesis. Int J Mol Epidemiol Genet 2:9–18
Iwasaki YW, Kiga K, Kayo H, Fukuda-Yuzawa Y, Weise J, Inada T, Tomita M, Ishihama Y, Fukao T (2013) Global microRNA elevation by inducible Exportin 5 regulates cell cycle entry. RNA 19:490–497. doi:10.1261/rna.036608.112
Huang J, Zheng DL, Qin FS, Cheng N, Chen H, Wan BB, Wang YP, Xiao HS, Han ZG (2010) Genetic and epigenetic silencing of SCARA5 may contribute to human hepatocellular carcinoma by activating FAK signaling. J Clin Invest 120:223–241. doi:10.1172/JCI38012
Zhou B, Li Y, Deng Q, Wang H, Wang Y, Cai B, Han ZG (2013) SRPK1 contributes to malignancy of hepatocellular carcinoma through a possible mechanism involving PI3K/Akt. Mol Cell Biochem 379:191–199. doi:10.1007/s11010-013-1641-7
Kuusisto HV, Wagstaff KM, Alvisi G, Roth DM, Jans DA (2012) Global enhancement of nuclear localization-dependent nuclear transport in transformed cells. FASEB J 26:1181–1193. doi:10.1096/fj.11-191585
Poon IK, Jans DA (2005) Regulation of nuclear transport: central role in development and transformation? Traffic 6:173–186. doi:10.1111/j.1600-0854.2005.00268.x
Martello G, Rosato A, Ferrari F, Manfrin A, Cordenonsi M, Dupont S, Enzo E, Guzzardo V, Rondina M, Spruce T, Parenti AR, Daidone MG, Bicciato S, Piccolo S (2010) A microRNA targeting dicer for metastasis control. Cell 141:1195–1207. doi:10.1016/j.cell.2010.05.017
Wahid F, Shehzad A, Khan T, Kim YY (2010) MicroRNAs: synthesis, mechanism, function, and recent clinical trials. Biochim Biophys Acta 1803:1231–1243. doi:10.1016/j.bbamcr.2010.06.013
Medina PP, Slack FJ (2008) MicroRNAs and cancer: an overview. Cell Cycle 7:2485–2492
Orellana EA, Kasinski AL (2015) MicroRNAs in cancer: a historical perspective on the path from discovery to therapy. Cancers (Basel) 7:1388–1405. doi:10.3390/cancers7030842
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, Golub TR (2005) MicroRNA expression profiles classify human cancers. Nature 435:834–838. doi:10.1038/nature03702
Gwizdek C, Ossareh-Nazari B, Brownawell AM, Doglio A, Bertrand E, Macara IG, Dargemont C (2003) Exportin-5 mediates nuclear export of minihelix-containing RNAs. J Biol Chem 278:5505–5508. doi:10.1074/jbc.C200668200
Calado A, Treichel N, Muller EC, Otto A, Kutay U (2002) Exportin-5-mediated nuclear export of eukaryotic elongation factor 1A and tRNA. EMBO J 21:6216–6224
Takeiwa T, Taniguchi I, Ohno M (2015) Exportin-5 mediates nuclear export of SRP RNA in vertebrates. Genes Cells 20:281–291. doi:10.1111/gtc.12218
Chen T, Brownawell AM, Macara IG (2004) Nucleocytoplasmic shuttling of JAZ, a new cargo protein for exportin-5. Mol Cell Biol 24:6608–6619. doi:10.1128/MCB.24.15.6608-6619.2004
Macchi P, Brownawell AM, Grunewald B, DesGroseillers L, Macara IG, Kiebler MA (2004) The brain-specific double-stranded RNA-binding protein Staufen2: nucleolar accumulation and isoform-specific exportin-5-dependent export. J Biol Chem 279:31440–31444. doi:10.1074/jbc.C400226200
Fritz J, Strehblow A, Taschner A, Schopoff S, Pasierbek P, Jantsch MF (2009) RNA-regulated interaction of transportin-1 and exportin-5 with the double-stranded RNA-binding domain regulates nucleocytoplasmic shuttling of ADAR1. Mol Cell Biol 29:1487–1497. doi:10.1128/MCB.01519-08
Acknowledgments
We gratefully thank Dr. Bryan R. Cullen and Dr. Manel Esteller for kindly providing the plasmids expressing XPO5. This work was supported by grants from the National Natural Science Foundation of China (81302064 and 81472576), the Shanghai Natural Science Foundation of China (13ZR1434000), and the Key Disciplines Group Construction Project of Pudong Health Bureau of Shanghai (PWZxq2014-04).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of Interest
The authors declare that no conflicts of interest exist.
Additional information
Yandong Li and Xiao Wang contributed equally to this work.
Rights and permissions
About this article
Cite this article
Li, Y., Wang, X., He, B. et al. Downregulation and tumor-suppressive role of XPO5 in hepatocellular carcinoma. Mol Cell Biochem 415, 197–205 (2016). https://doi.org/10.1007/s11010-016-2692-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-016-2692-3