Abstract
Linkage disequilibrium-based association mapping is a powerful tool for dissecting the genetic basis underlying complex traits. In this study, an association mapping panel consisting of 180 elite Upland cotton cultivars was constructed, evaluated in three locations across 2 years and genotyped using 228 SSRs to detect molecular markers associated with seed oil and protein contents. A total of 86 significant (á = 0.01) marker-trait associations were detected between 58 SSR markers and two seed quality traits in six environments. Fifteen SSR markers distributed on ten chromosomes (A3, A7, A9, A10, A12, A13, D2, D5, D6 and D9) and 12 across 9 chromosomes (A3, A7, A9, A10, A12, D2, D3, D5 and D9) associated with seed oil and protein contents, respectively, could be detected in more than one environment. Among the 18 SSR markers significantly associated with seed oil and/or protein contents, nine loci were associated with both seed traits simultaneously. The results of this study provide useful information for further understanding the genetic basis of cottonseed oil and protein traits, and they should facilitate future efforts to breed cotton containing seeds with high oil or high protein contents using MAS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gossypium hirsutum L., commonly referred to as Upland cotton, is an essential cash crop worldwide, which accounts for 95 % of the world’s cotton production (Zhang et al. 2008). Commercial seed cotton is composed of approximately 40 % lint and 60 % seed, which provides the most important natural fiber for the textile industry as well as seed nutrition for both humans and livestock. The importance of fiber quality has long been recognized due to the changing requirements of spinning technology, and considerable efforts have been devoted to improving fiber quality traits (Chen et al. 2011; Ashokkumar et al. 2014). By contrast, cottonseed is still considered to be a by-product of lint, and little emphasis has been placed on breeding for seed nutritional components (Wu et al. 2009). Cottonseed oil, which consists of approximately 70 % unsaturated and 30 % saturated fatty acids, can be refined to eliminate phenolic compounds, which can then directly be used for edible purposes (Lukonge et al. 2007), and it is also considered to be an important biofuel resource (Liu et al. 2009). Cottonseed protein is widely used to feed sheep, cattle and other ruminant livestock (Kohel et al. 1985; Coppock et al. 1987). If gossypol were eliminated from cottonseed protein, it would be fully edible, thereby providing a new, important source of nutrition, which would increase food security worldwide.
Cottonseed oil and protein contents are quantitative traits that are simultaneously affected by genetic and environmental factors during seed development; these traits often vary among different growing seasons, locations and years (Dani and Kohel 1989; Wu et al. 2010). Large-scale, repeated chemical testing during breeding is labor-intensive, costly and time-consuming, and it has proven to be unfeasible for effectively improving these two traits based simply on phenotypic selection (Wu et al. 2010; Ashokkumar and Ravikesavan 2011). Molecular markers tightly linked to target genes and/or QTLs can be used for marker-assisted selection (MAS), which markedly improves breeding efficiency (Xu and Crouch 2008; Ashokkumar and Ravikesavan 2011). In the past two decades, the availability of abundant molecular markers has made tagging QTLs harboring functional genes through family-based linkage mapping a routine process, and a large number of QTLs for agronomically important traits have been identified in cotton (Zhang et al. 2008; Said et al. 2013), including QTLs for cottonseed oil and protein (Song and Zhang 2007; An et al. 2010; Liu et al. 2012; Alfred et al. 2012; Yu et al. 2012; Liu et al. 2013). However, approximately 80 % of the QTLs identified by linkage mapping could not be confirmed in subsequent studies, and few have actually been applied in breeding programs (Lacape et al. 2010; Said et al. 2013). This may be due to the fact that most of the QTLs were population-specific, and the limited amount of recombination present in most populations used for linkage mapping makes it difficult to map QTLs at a high resolution, which has severely limited their application in breeding programs. Linkage disequilibrium (LD)-based association mapping (AM), which has the potential to exploit most recombination events that have occurred during the plant’s evolutionary history and to simultaneously evaluate the effects of many alleles of target loci, has become a powerful approach to dissecting complex traits in many crops (Zhu et al. 2008; Mackay et al. 2009). In cotton, AM had been used for QTL detection for fiber quality traits (Kantartzi and Stewart 2008; Abdurakhmonov et al. 2008, 2009; Zeng et al. 2009; Zhang et al. 2013; Cai et al. 2014), yield and its components (Zhang et al. 2013; Mei et al. 2013), disease resistance (Mei et al. 2014; Zhao et al. 2014) and salinity tolerance (Saeed et al. 2014). However, to date, to the best of our knowledge, no association-mapping study of seed oil and protein traits has been reported in cotton. In the present study, 180 elite Upland cotton cultivars and breeding lines were assembled into an AM panel, evaluated in three locations across 2 years and genotyped using 228 polymorphic SSR markers to perform marker-trait association analysis. The results provide useful information for further understanding the genetic basis of cottonseed oil and protein, and they should facilitate future efforts to breed cotton containing seeds with high oil or protein content by MAS.
Materials and methods
Plant materials
A total of 180 elite Upland cotton cultivars and breeding lines were selected from the AM population previously used in our laboratory (Mei et al. 2013) to construct a new panel. Among these accessions, 174 entries were developed in China (including 77, 64, 17 and 16 that were released in the Yellow River, Yangtze River, Northwest China and North China cotton growing regions, respectively) and six that were introduced from the U.S., including the genetic standard line TM-1. All accessions had been self-pollinated for more than eight generations. Detailed information about the 180 accessions is summarized in Table S1.
Measurements of seed oil and protein contents
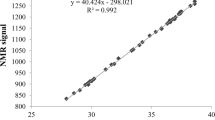
All 180 accessions were grown in the following three environments in 2011 and 2012: (1) Breeding Station at Nanyang Agricultural Research Institute, Nanyang, Henan, China (32°55′16″N, 112°34′07″E, designated NY); (2) Xinxiang Cotton and Wheat Research Institute, Xinxiang, Henan (35°09′34″N, 113°47′35″E, XX); and (3) Breeding Station at Biocentury Seed Company Limited, Korla, Xinjiang Uygur Autonomous Region (41°44′36″N, 86°07′40″E, XJ). The first location is in the Yangtze River cotton-growing region of China, the second is in the Yellow River region and the third is in the Northwestern inland region. A randomized complete block design with single row plot and two replications was used in each field trial. The sowing dates were from late March to early April in different years and locations. Seeds of 180 accessions were directly sown into the field in XJ, with 30 holes per row, a hole spacing of 10 and 40 cm between rows, and single plant was remained after seedling emergence. Seedling transplant were performed in NY and XX. Seedlings with 3–4 leaves were transplanted from seedbeds to the field, with 20 plants per row, a plant spacing of 30 and 80 cm between rows. Field management followed local practices. At the maturity stage, 25 fully opened bolls from each plot were hand harvested and ginned. Approximately 20 g cottonseed per sample was prepared and shell-intacted, and seed oil and protein contents were measured with the NIR (near infrared reflectance) spectrum method on a Foss NIRSystems 5000 (NIRSystems, Silver Spring, MD, USA) according to Huang et al. (2013).
SSR genotyping
Young leaves from each of the 180 accessions were collected and stored at −20 °C. Total genomic DNA was extracted from the leaf samples following the published method developed in our laboratory (Guo et al. 2007). In our previous study, a 360-accession panel was genotyped using 145 SSR markers. However, two of the 145 SSRs were not polymorphic in the new 180-accession panel. An additional 83 SSRs tightly linked to agronomically important traits (Qin et al. 2008; Mei et al. 2014) were then used to fingerprint the new panel. In total, the panel was genotyped using 228 markers (detailed information is summarized in Table S2) for marker-trait association analysis.
Data analysis
Summary statistics of genetic diversity were calculated using PowerMarker 3.25 software (Liu and Muse 2005). The Bayesian model-based program STRUCTURE 2.3 was used to infer the population structure with 66 unlinked and/or weakly linked markers (Pritchard et al. 2009; Mei et al. 2013). Both the burn-in period and the Markov Chain Monte Carlo replications were set at 100,000 using an admixture and allele frequency correlated model. Five run iterations were performed with the hypothetical number of subpopulations (k) ranging from 1 to 10. The correct estimation of k was performed by joining the probability [LnP(D)] output and the ad hoc statistic ∆k (Evanno et al. 2005). Based on the correct k, each accession was assigned to a subpopulation for which the membership value (Q value) was >0.5 (Pritchard et al. 2000), and the population structure matrix (Q) was generated for further marker-trait association mapping. SPAGeDi software was used to calculate the pair-wise relatedness coefficients (K, kinship matrix) with the negative value of kinship set to zero (Hardy and Vekemans 2002).
Statistical analysis of phenotype data from three locations across 2 years was performed using SAS 8.0 software (SAS Institute 1999). Analysis of variance (ANOVA) was calculated with PROC GLM based on the trait means for each line across six environments. Decomposition of variance components (genotype, location, year, replicate and the interactions among these factors) was evaluated using PROC VARCOMP.
The widely used mixed linear model (MLM) considering both Q and K implemented in the TASSEL software package was used to perform marker-trait association analysis, and the P value and R 2 of each association were determined (Yu et al. 2006; Bradbury et al. 2007).
Results
Genetic diversity, population structure and genetic relatedness
A total of 601 alleles were detected at 228 SSR loci. The allele number averaged 2.64 (ranging from 2 to 11), and approximately 86 % of the loci (198 of 228) yielded only two or three alleles (Table S2). The gene diversity, polymorphism information content (PIC) and heterozygosity values of the 228 loci averaged 0.37, 0.31 and 0.03, respectively, with ranges of 0.04–0.75, 0.04–0.72 and 0–0.22, respectively (Fig. 1).
Distribution of a gene diversity, b PIC and c heterozygosity of 228 SSR loci in 180 Upland cotton accessions. Data were calculated using PowerMarker 3.25 software (Liu and Muse 2005)
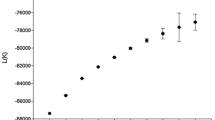
Model-based population structure analysis of the panel revealed that the LnP(D) value corresponding to each hypothetical k increased with increasing k value and did not exhibit a peak. By contrast, the ∆k value revealed a much higher likelihood at k = 2 than at k = 3–10 (Fig. 2), suggesting that the total population could be divided into two major subpopulations (Evanno et al. 2005), designated P1 and P2, respectively. The P1 group contained 56 accessions including 23 cultivars from the Yellow River cotton growing region, 13 from Northwest China, 12 from North China, six from the Yangtze River region and two from the United States. The P2 group consisted of 124 accessions including 58 lines from the Yangtze River region, 54 from the Yellow River region, four from Northwest China, four from the North China region and four from the United States (Table S1). The corresponding Q matrix at k = 2 was used for further marker-trait association analysis.
Model-based evaluation of population structure. a LnP(D) values for k from 1 to 10, b ∆k for k from 2 to 9. The LnP(D) of each hypothetical k continued to increase, and the ∆k values showed a much higher likelihood at k = 2 than at k = 3–10, suggesting that the total panel should be divided into two major subpopulations. LnP(D) values are mean values of five repeats estimated using STRUCTURE (Pritchard et al. 2009), and ∆k values were calculated according to Evanno et al. (2005)
A total of 86.98 % of the cotton accessions had kinship coefficient values less than 0.05, while 8.63 % had values ranging from 0.05 to 0.10, and the remaining 4.39 % showed varying degrees of genetic relatedness (Fig. 3). We constructed a K matrix for association analysis based on the results of relatedness analysis.
Distribution of pair-wise kinship coefficients among 180 Upland cotton accessions. Data were calculated using SPAGeDi (Hardy and Vekemans 2002) with 228 SSR markers
Variations in seed oil and protein contents
We measured the seed oil and protein contents of 180 Upland cotton accessions grown in three different locations across 2 years. Each trait varied widely, and ANOVA revealed that the genotype (G) and the interactions between genotype and environmental factors (G × E) were both highly significant (á = 0.001; Table 1), indicating that these two seed quality traits are strongly affected by the environment. The mean values of oil content of 180 seed samples from XJ, XX, and NY across 2 years were 31.76, 29.98 and 29.45 %, respectively, and highly significant differences (LSD, á = 0.01) were observed in the three locations. The mean values of protein content of 180 seed samples from NY and XX across 2 years were 49.19, 49.08 %, respectively, and they were not significantly different (LSD, á = 0.01) from each other. When compared with the mean value from XJ (45.61 %), highly significant differences were also observed in the former two and the latter one location. The mean coefficient of variance (CV) for oil and protein contents was 8.25 % and 5.31 %, respectively, demonstrating that there was a high degree of diversity in seed quality traits in the present panel. The absolute values of skewness and kurtosis for these two traits in most environments were less than 1, suggesting that the seed oil and protein contents in this panel approximately followed a normal distribution. High significant negative correlations (−0.95 to −0.98, á = 0.01, data not shown) were found between oil and protein contents in each of the six environments, which is consistent with the results of many other studies (Wu et al. 2009; Yu et al. 2012; Liu et al. 2013).
SSR markers associated with oil and protein contents
At the á = 0.01 (−lgP = 2) level, a total of 86 significant marker-trait associations were detected between 58 SSR markers and two seed quality traits in six environments. Among these associations, more than half (59 of 86) could be detected in only one environment. The proportion of phenotypic variation explained by markers ranged from 4.31 to 24.18 %, with an average of 10.11 % (Table S3). Significant marker-trait associations simultaneously detected in more than one environment are shown in Table 2. Fifteen SSR markers distributed on 10 chromosomes (A3, A7, A9, A10, A12, A13, D2, D5, D6 and D9) are associated with seed oil content, including two, two and 11 that could be detected in five, three and two environments, respectively. Twelve SSR markers across nine chromosomes (A3, A7, A9, A10, A12, D2, D3, D5 and D9) are associated with seed protein content, including one, four and seven that could be detected in five, three and two environments, respectively. Among the 18 related SSR markers, nine loci are significantly associated with the two seed traits simultaneously (Table 2).
Discussion
Genetic diversity and population structure of the AM panel
It is essential for genetic diversity to be present in the AM panel used for marker-trait AM studies (Flint-Garcia et al. 2005). Most Upland cotton cultivars developed in China were derived from a few germplasm resources introduced from abroad (Huang 2007). Therefore, it is especially critical to select samples that encompass genetic diversity as much as possible. Theoretically, a panel with a large number of accessions should best meet this requirement. However, working with large populations significantly increases phenotyping costs and can easily lead to errors due to differences in field conditions and management practices, especially for large-plant crops such as cotton, which could reduce the detection power of association analysis. The 180 accessions in the panel used in the current study were selected from our previously used panel (Mei et al. 2013) based on field performance and pair-wise genetic distances among the 356 accessions. The gene diversity and PIC values of the 228 loci averaged 0.37 and 0.31 with ranges of 0.04–0.75 and 0.04–0.72, respectively, and more than 86 % of the kinship coefficient values were less than 0.05. In spite of the fact that fewer accessions were used in the current study than in the previous study (180 vs. 356), the genetic diversities of both panels were similar (Mei et al. 2013).
Many crops have long, complex histories of domestication and breeding, including Upland cotton. Relatedness among entry samples can lead to population stratification, which can confound the results of AM (Price et al. 2006; Yu et al. 2006). Our model-based evaluation of the population structure of the 180 Upland cotton cultivars revealed that the population could be divided into two major subpopulations (Fig. 1). Group P1 contains almost all cultivars with early maturity and some cultivars with moderate maturity, while group P2 contains almost all cultivars with late maturity and some cultivars with moderate maturity. These results indicate that population stratification has occurred in the current AM panel, which should be considered in subsequent association analyses.
QTLs for cottonseed oil and protein contents detected by association mapping
In this study, marker-trait association analysis was performed with the optimal MLM model, which considers both population structure and relatedness, to detect SSR markers associated with seed oil and protein contents (Yu et al. 2006). A total of 86 highly significant (á = 0.01) associations were detected between 58 SSR markers and two seed quality traits (Table S3). Nonetheless, it is not easy to determine which significance level should be accepted. The use of stringent probability thresholds may reduce the danger of false positives but may pose the risk of rejecting true positives caused by setting the thresholds too high (Yan et al. 2011). If a more stringent Bonferroni-corrected threshold (P ≤ 0.05/228, −lgP ≥ 3.66) is adopted (Lander and Botstein 1989), only four associations were significant (Table S3). For the purpose of MAS, marker-trait associations should be environmentally stable and consistent. SSR markers with significant associations simultaneously detected in more than one environment could be considered candidate seed quality SSRs. Of these candidates, 15 SSR markers were associated with cottonseed oil content and 12 SSRs were associated with seed protein content. Among the 18 related SSR markers, nine loci were significantly associated with the two seed traits simultaneously (Table 2), which is also consistent with the fact that seed oil and protein are usually negatively correlated (Wu et al. 2009; Yu et al. 2012; Liu et al. 2013). The resulting stably associated markers, such as NAU845 and NAU1048, which were detected in almost all locations, should be quite useful for developing new cultivars with broad adaptability to different environments. Moreover, the materials used in this study are all cultivars and breeding lines with elite field performance, which can be directly utilized as parents in breeding programs.
The genetic basis of seed oil and protein contents appears to be complicated; as mentioned in several reports, additive, dominance and cytoplasmic effects all play important roles in their inheritance (Wu et al. 2010; Yu et al. 2012; Liu et al. 2013). The QTL mapping results of previous studies (Song and Zhang 2007; An et al. 2010; Liu et al. 2012; Alfred et al. 2012; Yu et al. 2012; Liu et al. 2013), as well as the current results, are too divergent to be compared. More research should be performed to better dissect the genetic architecture of the seed oil and protein traits in the future. Recently, two preliminary maps of the whole-genome scaffolds of G. raimondii (the putative diploid donor for tetraploid species) were separately released by two different groups (Paterson et al. 2012; Wang et al. 2012), which will facilitate tetraploid genome assembly. Moreover, true genome-wide AM will be realized in the near future through resequencing or other high-throughput genotyping technologies.
References
Abdurakhmonov IY, Kohel RJ, Yu JZ, Pepper AE, Abdullaev AA, Kushanov FN, Salakhutdinov IB, Buriev ZT, Saha S, Scheffler BE, Jenkins JN, Abdukarimov A (2008) Molecular diversity and association mapping of fiber quality traits in exotic G. hirsutum L. germplasm. Genomics 92:478–487
Abdurakhmonov IY, Saha S, Jenkins JN, Buriev ZT, Shermatov SE, Scheffler BE, Pepper AE, Yu JZ, Kohel RJ, Abdukarimov A (2009) Linkage disequilibrium based association mapping of fiber quality traits in G. hirsutum L. variety germplasm. Genetica 136:401–417
Alfred Q, Liu HY, Xu HM, Li JR, Wu JG, Zhu SJ, Shi CH (2012) Mapping of quantitative trait loci for oil content in cottonseed kernel. J Genet 91:289–295
An CF, Jenkins JN, Wu J, Guo Y, McCarty JC (2010) Use of fiber and fuzz mutants to detect QTL for yield components, seed, and fiber traits of upland cotton. Euphytica 172:21–34
Ashokkumar K, Ravikesavan R (2011) Conventional and molecular breeding approaches for seed oil and seed protein content improvement in cotton. Int Res J Plant Sci 2:37–45
Ashokkumar K, Kumar KS, Ravikesavan R (2014) An update on conventional and molecular breeding approaches for improving fiber quality traits in cotton: a review. Afr J Biotechnol 13:1097–1108
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Cai C, Ye W, Zhang T, Guo W (2014) Association analysis of fiber quality traits and exploration of elite alleles in Upland cotton cultivars/accessions (Gossypium hirsutum L.). J Integr Plant Biol 56:51–62
Chen X, Guo W, Zhang T (2011) Cotton omics in China. Plant Omics 4:278–287
Coppock CE, Lanham JK, Horner JL (1987) A review of the nutritive value and utilization of whole cottonseed, cottonseed meal and associated by-products by dairy cattle. Anim Feed Sci Technol 18:89–129
Dani RG, Kohel RJ (1989) Maternal effects and generation mean analysis of seed-oil content in cotton (Gossypium hirsutum L.). Theor Appl Genet 77:569–575
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Flint-Garcia SA, Thuillet AC, Yu J, Pressoir G, Romero SM, Mitchell SE, Doelley J, Goodman MM, Buckler ES (2005) Maize association population: a high-resolution platform for quantitative trait locus dissection. Plant J 44:1054–1064
Guo WZ, Cai C, Wang C, Han Z, Song X, Wang K, Niu X, Wang C, Lu K, Shi B, Zhang T (2007) A microsatellite-based, gene-rich linkage map reveals genome structure, function and evolution in Gossypium. Genetics 176:527–541
Hardy O, Vekemans X (2002) SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620
Huang ZK (ed) (2007) The cultivars and their pedigree of cotton in China. China Agriculture Press, Beijing
Huang Z, Sha S, Rong Z, Chen J, He Q, Khan DM, Zhu S (2013) Feasibility study of near infrared spectroscopy with variable selection for non-destructive determination of quality parameters in shell-intact cottonseed. Ind Crops Prod 43:654–660
Kantartzi SK, Stewart JMcD (2008) Association analysis of fibre traits in Gossypium arboreum accessions. Plant Breed 127:173–179
Kohel RJ, Glueck J, Rooney LW (1985) Comparison of cotton germplasm collections for seed-protein content. Crop Sci 25:961–963
Lacape JM, Llewellyn D, Jacobs J, Arioli T, Becker D, Calhoun S, Al-Ghazi Y, Liu S, Palaï O, Georges S, Giband M, de Assunçã H, Augusto Vianna Barroso P, Claverie M, Gawryziak G, Jean J, Vialle M, Viot C (2010) Meta-analysis of cotton fiber quality QTLs across diverse environments in a Gossypium hirsutum × G. barbadense RIL population. BMC Plant Biol 10:132
Lander ES, Botstein D (1989) Mapping mendelian factors underlying quantitative traits using RFLP linkage maps. Genetics 121:185–199
Liu KJ, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinfo Appl Note 9:2128–2129
Liu Q, Singh S, Chapman K, Green A (2009) Bridging traditional and molecular genetics in modifying cottonseed oil. In: Paterson AH (ed) Genetics and genomics of cotton. Springer, New York, pp 353–382
Liu HY, Quampah A, Chen JH, Li JR, Huang ZR, He QL, Shi CH, Zhu SJ (2012) QTL analysis for gossypol and protein contents in upland cottonseeds with two different genetic systems across environments. Euphytica 188:453–463
Liu X, Li J, Yu X, Shi Y, Jia F, Sun F, Liu A, Gong J, Shang H, Gong W, Wang T, Deng H, Yuan Y (2013) Identification of QTL for cottonseed oil and protein content in Upland cotton (Gossypium hirsutum L.) based on a RIL population. Mol Plant Breed 11:520–528
Lukonge E, Labuschagne MT, Hugo A (2007) The evaluation of oil and fatty acid composition in seed of cotton accessions from various countries. J Sci Food Agri 87:340–347
Mackay TFC, Stone EA, Ayroles JF (2009) The genetics of quantitative traits: challenges and prospects. Nat Rev Genet 10:565–577
Mei H, Zhu X, Zhang T (2013) Favorable QTL alleles for yield and its components identified by association mapping in Chinese Upland cotton cultivars. PLoS ONE 8(12):e82193
Mei H, Ai N, Zhang X, Ning Z, Zhang T (2014) QTLs conferring FOV 7 resistance detected by linkage and association mapping in Upland cotton. Euphytica 197:237–249
Paterson AH, Wendel JF, Gundlach H, Guo H, Jenkins J, Jin D, Llewellyn D, Showmaker KC, Shu S, Udall J, Yoo MJ, Byers R, Chen W, Doron-Faigenboim A, Duke MV, Gong L, Grimwood J, Grover C, Grupp K, Hu G, Lee TH, Li J, Lin L, Liu T, Marler BS, Page JT, Roberts AW, Romanel E, Sanders WS, Szadkowski E, Tan X, Tang H, Xu C, Wang J, Wang Z, Zhang D, Zhang L, Ashrafi H, Bedon F, Bowers JE, Brubaker CL, Chee PW, Das S, Gingle AR, Haigler CH, Harker D, Hoffmann LV, Hovav R, Jones DC, Lemke C, Mansoor S, ur Rahman M M, Rainville LN, Rambani A, Reddy UK, Rong JK, Saranga Y, Scheffler BE, Scheffler JA, Stelly DM, Triplett BA, Van Deynze A, Vaslin MF, Waghmare VN, Walford SA, Wright RJ, Zaki EA, Zhang T, Dennis ES, Mayer KF, Peterson DG, Rokhsar DS, Wang X, Schmutz J (2012) Repeated polyploidization of Gossypium genomes and the evolution of spinnable cotton fibres. Nature 492:423–427
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38:904–909
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Pritchard JK, Wen X, Falush D (2009) Documentation for structure software: Version 2.3 http://pritch.bsd.uchicago.edu/structure.html
Qin H, Guo W, Zhang Y, Zhang T (2008) QTL mapping of yield and fiber traits based on a four-way cross population in Gossypium hirsutum L. Theor Appl Genet 117:883–894
Saeed M, Guo W, Zhang T (2014) Association mapping for salinity tolerance in cotton (Gossypium hirsutum L.) germplasm from US and diverse regions of China. Aust J Crop Sci 8:338–346
Said J, Lin Z, Zhang X, Song M, Zhang J (2013) A comprehensive meta QTL analysis for fiber quality, yield, yield related and morphological traits, drought tolerance, and disease resistance in tetraploid cotton. BMC Genom 14:776
Song XL, Zhang TZ (2007) Identification of quantitative trait loci controlling seed physical and nutrient traits in cotton. Seed Sci Res 17:243–251
Wang K, Wang Z, Li F, Ye W, Wang J, Song G, Yue Z, Cong L, Shang H, Zhu S, Zou C, Li Q, Yuan Y, Lu C, Wei H, Gou C, Zheng Z, Yin Y, Zhang X, Liu K, Wang B, Song C, Shi N, Kohel RJ, Percy RG, Yu JZ, Zhu YX, Wang J, Yu S (2012) The draft genome of a diploid cotton Gossypium raimondii. Nat Genet 44:1098–1103
Wu J, Jenkins JN, McCarty JC, Thaxton P (2009) Seed trait evaluation of Gossypium barbadense L. chromosomes/arms in a G. hirsutum L. background. Euphytica 167:371–380
Wu J, McCarty JC, Jenkins JN (2010) Cotton chromosome substitution lines crossed with cultivars: genetic model evaluation and seed trait analyses. Theor Appl Genet 120:1473–1483
Xu Y, Crouch J (2008) Marker-assisted selection in plant breeding: from publications to practice. Crop Sci 48:391–407
Yan J, Warburton M, Crouch J (2011) Association mapping for enhancing maize (Zea mays L.) genetic improvement. Crop Sci 51:433–449
Yu J, Pressoir G, Briggs WH, Bi IV, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Yu J, Yu S, Fan S, Song M, Zhai H, Li X, Zhang J (2012) Mapping quantitative trait loci for cottonseed oil, protein and gossypol content in a Gossypium hirsutum × Gossypium barbadense backcross inbred line population. Euphytica 187:191–201
Zeng L, Meredith WR Jr, Gutierrez OA, Boykin DL (2009) Identification of associations between SSR markers and fibre traits in an exotic germplasm derived from multiple cross among Gossypium tetraploid species. Theor Appl Genet 119:93–103
Zhang HB, Li YN, Wang BH, Chee PW (2008) Recent advances in cotton genomics. Int J Plant Genomics ID 742304. doi: 10.1155/2008/742304
Zhang T, Qian N, Zhu X, Chen H, Wang S, Mei H, Zhang Y (2013) Variations and transmission of QTL alleles for yield and fiber qualities in Upland cotton cultivars developed in China. PLoS ONE 8(2):e57220
Zhao Y, Wang H, Chen W, Li Y (2014) Genetic structure, linkage disequilibrium and association mapping of Verticillium wilt resistance in elite cotton (Gossypium hirsutum L.) germplasm population. PLoS ONE 9(1):e86308
Zhu C, Gore M, Buckler ES, Yu J (2008) Status and prospects of association mapping in plants. Plant Genome 1:5–20
Acknowledgments
This work was financially supported in part by grants from the Major State Basic Research Development Program of China (973 Program) (2011CB109300), the National High Technology Research and Development Program of China (863 Program) (2011AA10A102) and the Priority Academic Program Development of Jiangsu Higher Education Institutions and 111 program.
Competing interests
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
Guizhen Liu and Hongxian Mei have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, G., Mei, H., Wang, S. et al. Association mapping of seed oil and protein contents in upland cotton. Euphytica 205, 637–645 (2015). https://doi.org/10.1007/s10681-015-1450-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-015-1450-z