Abstract
Fusarium wilt is a worldwide disease that affects cotton production. Molecular markers tightly linked to resistance genes can be used for marker-assisted and/or genomic selection. We performed both family-based linkage mapping and population-based association mapping (AM) to detect quantitative trait loci (QTLs) conferring resistance against Fusarium oxysporum f. sp. vasinfectum race 7 (FOV 7) in Upland cotton. To identify QTLs underlying FOV 7 resistance by linkage mapping, three Upland cotton cultivars/lines, Xuzhou 142, Yumian 21 and Shang 9901, were used to obtain the composite cross population, designated as Xuzhou 142/Yumian 21//Xuzhou 142/Shang 9901. A linkage map containing 185 simple sequence repeat loci and 40 linkage groups was constructed with an average distance of 7.5 cM between adjacent markers. Seven QTLs were detected by linkage mapping, explaining 2.9–6.6 % of the total phenotypic variance. We also performed marker–trait AM with the MLM model (Q + K) in a panel composed of 356 Upland cotton cultivars. In total, 27 loci were significantly associated with FOV 7 resistance at the α = 0.01 level (−log10 P ≥ 2), which were distributed on 16 chromosomes and explained 1.48–12.99 % of phenotypic variation. Three of the 7 QTLs identified by linkage mapping could be detected in AM. We identified the favorable allele for each of the 27 associated loci and investigated the number of favorable alleles in each accession. The results should increase our understanding of the genetic basis of FOV resistance and facilitate future resistance breeding in Upland cotton.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fusarium wilt (FW), caused by Fusarium oxysporum f. sp. vasinfectum (FOV), is a widespread disease that causes huge losses in cotton (Gossypium ssp.) production worldwide (Ma 2007; Davis et al. 2006; Ulloa et al. 2006; Constable et al. 2007). FOV is a soil-inhabiting fungus that generally invades seedlings through the root tips, subsequently spreading along the vascular system, which leads to FW symptoms including leaf wilt, necrosis, chlorosis, stunted growth, vascular discoloration, leaf abscission and plant death (Rodríguez-Gálvez and Mendgen 1995). Once established in soil, FOV can survive in the field for several years as Chlamydia spores, even in the absence of a host, and is nearly impossible to eliminate. Although both chemical controls and cultural practices have been employed in protecting plants from damage, the most effective and efficient control should be provided through host resistance (Ma 2007; Ulloa et al. 2006; Constable et al. 2007). In the past several years, eight races (race 1–8) of FOV had been indentified worldwide that use both cotton and non-cotton differential hosts (Davis et al. 2006). Three FOV races (race 3, 7 and 8) had been found in China; race 7 possesses the highest virulence and is the most widely distributed race (Chen et al. 1985; Sun et al. 1999). Recently, DNA-based techniques were employed in conjunction with pathogenicity tests to validate these races and to test new isolates (Skovgaard et al. 2001; Kim et al. 2005; Holmes et al. 2009), and highly virulent isolates of FOV were identified in Australia (Wang et al. 2004) and the United States (Kim et al. 2005; Holmes et al. 2009). Host resistance to FOV races has been widely evaluated in cotton germplasm under both field nursery and greenhouse conditions, and many highly resistant cotton cultivars and breeding lines have been developed through conventional breeding (Ma et al. 2002; Ulloa et al. 2006, 2009). However, little is known about the mechanism and genetic basis of FOV resistance. Some early classical genetic studies have suggested that the inheritance of FOV resistance in cotton is determined by a single gene (Smith and Dick 1960; Netzer et al. 1985; Feng et al. 1996), while other studies have suggested that FOV resistance is controlled by multiple genes (Kappelman 1971; Jiao 1985; Wang et al. 1989).
Large-scale resistance evaluations in breeding programs are time-consuming and labor-intensive, and it is not easy to obtain the ideal genotype simply through phenotypic selection. The cultivar development process has been slow in the face of the emergence of new, highly virulent FOV isolates (Ulloa et al. 2006; Constable et al. 2007). Molecular markers tightly linked to the target genes and QTLs can be used for marker-assisted selection (MAS) and/or genomic selection (GS), thus improving breeding efficiency (Xu and Crouch 2008; Jannink et al. 2010). In the past two decades, the availability of abundant molecular markers has made tagging QTLs harboring functional genes through family-based linkage mapping a routine process (Mackay et al. 2009), and a large number of QTLs for agronomically important traits have been identified in cotton (Zhang et al. 2008; Chen et al. 2011), including QTLs for FOV resistance (Wang and Roberts 2006; Chen et al. 2008; Wang et al. 2009, 2010; Ulloa et al. 2011, 2013; Lopez-Lavalle et al. 2012). However, approximately 80 % of the QTLs identified by linkage mapping could not be confirmed in subsequent studies, and few have actually been applied in breeding programs (Lacape et al. 2010; Rong et al. 2007). This may be due to the fact that most of the QTLs were population-specific, and the limited recombination present in most populations used for linkage mapping makes it difficult to map QTLs at a high resolution, which has severely limited their application in breeding programs.
Linkage disequilibrium (LD) based association mapping (AM), which has the potential to exploit most recombination events that have occurred in the evolutionary history of a plant and to simultaneously evaluate the effects of many alleles of target traits, has become a powerful approach to dissecting complex traits in many crops (Mackay et al. 2009; Zhu et al. 2008), including cotton (Kantartzi and Stewart 2008; Abdurakhmonov et al. 2008, 2009; Zeng et al. 2009; Zhang et al. 2013). However, several inherent constraints to natural populations have limited the successful use of association mapping in plants, such as genetic relatedness and rare alleles (Gupta et al. 2005; Myles et al. 2009). Genetic relatedness among individuals can result in spurious marker-trait associations (Gupta et al. 2005). Several statistical strategies have been developed to account for issues related to population structure and relatedness (Price et al. 2006; Yu et al. 2006; Yu et al. 2008). Given that the number of individuals with a specific genotype is quite small, the effect of rare alleles on mapping can go far beyond the effect of small population sizes (Myles et al. 2009). However, family-based linkage mapping can make use of alleles that occur at low frequencies in natural populations by designing crosses to create artificial populations with inflated frequencies of those alleles. Therefore, joint linkage and association mapping was recommended as an alternative approach to overcome some of the inherent limitations of both linkage and association mapping (Gupta et al. 2005; Myles et al. 2009), and this technique has proven to be a powerful approach to detecting QTLs underlying complex traits (Nemri et al. 2010; Lu et al. 2010; Brachi et al. 2010; Korir et al. 2013; Cadic et al. 2013).
In the present study, we performed joint family-based linkage mapping and population-based association mapping to detect QTLs underlying FOV 7 resistance in Upland cotton. In addition, we identified a set of favorable alleles and representative germplasm resources. The results of this study should provide useful information for further understanding the genetic basis of FOV resistance and for facilitating future resistance breeding by GS in Upland cotton.
Materials and methods
Plant materials
A composite cross population (CP) with three parents was developed for linkage map construction and QTL mapping. Three Upland cotton cultivars, Xuzhou 142, Yumian 21 and Shang 9901, were chosen as parents according to many years of FOV 7 resistance evaluation. Xuzhou 142, an obsolete cultivar selected from STV 2B in the 1970s with large boll size and high lint percentage that is severely infested by FOV 7, was selected as the susceptible parent. Yumian 21 was released in 1999 and is currently used as a resistant control in national cotton regional trials due to its extremely high resistance to FOV 7. The cultivar Shang 9901 is an anonymous breeding line with high yield potential and moderately high resistance to FOV 7. Two single crosses (Xuzhou 142/Yumian 21 and Xuzhou 142/Shang 9901, SF1) were made in Hainan, China in the winter of 2008. The two SF1 were double-crossed to develop the CP (Xuzhou 142/Yumian 21//Xuzhou 142/Shang 9901, DF1) at Jiangpu breeding station at Nanjing Agricultural University (NAU), Nanjing, China during the summer of 2009. The DF1 seeds were planted in Hainan in the winter of 2009 to collect leaves for DNA extraction and to produce DF1:2 families for resistance evaluation. A total of 241 DF1 were self-pollinated, but only 239 individuals produced seeds enough for FW resistance evaluation. Meanwhile, a total of 356 representative Upland cotton cultivars and breeding lines were selected from the cotton germplasm collection in our laboratory and at the Cotton Research Institute, Chinese Academy of Agricultural Sciences (CRI-CAAS) and assembled to construct an association mapping panel. All accessions were self-pollinated for more than six generations; detailed information about the accessions is summarized in Table S1.
Resistance evaluation
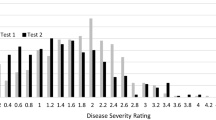
Accessions of the association panel were tested in both the greenhouse and field nursery, while the CP progenies were only tested in the greenhouse due to limited seed production. In both assays, the cultivars Yumian 21 and Simian 3 were treated as the resistant and susceptible control, respectively. The 356 entries of the association panel were tested in the greenhouse of NAU by artificially inoculating the plants with FOV 7 in the winter of 2009, and the 239 DF1:2 families of the CP (for evaluating resistance of DF1 individuals) were tested in the same way in the winter of 2010. The FOV 7 isolate was provided by the Institute of Plant Protection, Jiangsu Academy of Agriculture Sciences (PPI-JAAS), China. To produce the inoculum, boiled wheat seeds were mixed with sand in a 3:1 ratio and sterilized at 120 °C for 2 h in an autoclave. The mixture was inoculated with pre-cultured FOV pathogen on a clean bench, incubated at 25 °C for 10 days and air-dried for use. The culture soil was sterilized at 160 °C for 2 h in a drying oven and mixed with 2 % inoculum (by weight) to produce culture medium. The medium was loaded into paper cups (~250 g per cup) and irrigated prior to sowing. A randomized complete block design with two replications was carried out. Each replication comprised 15 cups; six to eight acid-delinted seeds were sown into each cup and only two seedlings remained after emergence; so about 30 plants per genotype were tested in each replication. The room temperature was 18 to 20 °C at night and 25 to 28 °C in the daytime. Symptoms appeared 18 days after planting and leaf damage was measured every 5 days. The susceptibility of each plant was scored according to China’s national standard for FW evaluation (Ma 2007). The disease grades of 0, 1, 2, 3 and 4 for symptoms were scored as follows: 0 indicates healthy, with no disease symptoms; 1 indicates <25.0 % of the leaf surface exhibited disease symptoms; 2 indicates 25.1–50.0 % of the leaf surface exhibited disease symptoms or plants were slightly dwarfed in stature; 3 indicates 50.1–75.0 % of the leaf surface exhibited disease symptoms or plants obviously dwarfed in stature; and 4 indicates >75.0 % of the leaf surface exhibited disease symptoms or plants completely defoliated or died. The disease index (DI) of each accession was calculated using the following formula:
where DI is the FW disease index of each accession, x is the disease grade of 0–4, y is the number of plants with corresponding disease grade, and α is the total number of investigated plants of each accession. F test and multiple comparisons (LSD) for DIs of three parents and 239 DF1:2 families were performed using the software SPSS 13.0. The additive (A) and dominance (D) effects of FOV resistance were estimated in the two single crosses according to the methods described by Kong (2006).
In the field trial, cotton seeds of the 356 entries were sown directly into an FW nursery at Shihezi Cotton Research Institute, Xinjiang Autonomous Region, China, in April, 2010. The nursery had been artificially inoculated with FOV 7 for many years and was heavily and evenly infested by the pathogen. A randomized complete block design with two replications was performed; each plot included a single 4 m row with a 15 cm plant-to-plant distance and a 60 cm row-to-row distance. 20–25 plants for each accession were assayed at the seedling stage, in early June, ~35 days after planting. The disease grades of individual plants were scored and the DI of each accession was calculated with methods described above. The mean DI value of two replications for each accession was used for QTL detection.
SSR genotyping
Young leaves from three parents, two SF1 and 241 DF1 individuals of the CP population and 356 accessions of the association panel were collected and stored at −20 °C. Total genomic DNA was extracted from the samples as described by Guo et al. (2007). The three parents, Xuzhou 142, Yumian 21 and Shang 9901, were first screened with 6,300 pairs of SSR primers to search for polymorphism. Polymorphic markers were then used to genotype two SF1 and 241 DF1 individuals. Individual genotypes were scored according to the method described in the JoinMap 4.0 manual (Van Ooijen and Voorrips 2006), and Chi square tests of goodness-of-fit were performed on segregation data for all loci to determine their agreement with the expected ratios. To fingerprint the entries of the association panel, 381 pairs of SSR primers evenly distributed in the tetraploid cotton genome (one marker per 10 cM) were selected according to a dense genetic linkage map constructed in our laboratory (Guo et al. 2007). The procedure for PCR-amplification and product analysis followed our published methods (Zhang et al. 2002; Zhao et al. 2012a).
Linkage map construction and QTL mapping
A CP with three homozygous parents (P1/P2//P1/P3) is analogous to a 4WC population (P1/P2//P3/P4) and can be regarded as a 4WC population with a common parent, P1. The methods of linkage analysis and QTL mapping in a full-sib family of outbreeding species can be applied to a four-way cross (4WC) population without any modification (Qin et al. 2008), and therefore, the methods can be extended to the CP with three parents (Zhang et al. 2012). The software JoinMap 4.0 (Van Ooijen and Voorrips 2006) was employed to construct the linkage map. Log-of-odds (LOD) scores ≥4.0 were used to determine linkage groups. The Kosambi map function was used to convert recombination frequencies to map distances. Previously chromosome-anchored SSR markers (Guo et al. 2007; Zhao et al. 2012b) were used to assign the linkage groups to chromosomes. QTL analyses were conducted on DI data using MapQTL 5.0 (Van Ooijen 2004) with the Multiple-QTL model (MQM). The significance thresholds for LOD scores were calculated by permutation tests in MapQTL 5.0, with a genome-wide significance level of α = 0.05, n = 1,000 as a significant QTL and a linkage group-wide significance level of α = 0.05, n = 1,000 as a suggestive QTL (Van Ooijen 1999). The determination of a QTL depended on the highest peak LOD and the directions of additive effects. The graphic representation of the linkage groups and QTLs was created by MapChart 2.2 (Voorrips 2002). QTL nomenclature was adopted using the method developed in rice (McCouch et al. 1997).
Marker-trait association mapping
Genetic diversity in 356 accessions was evaluated using the software PowerMarker 3.25 (Liu and Muse 2005). The Bayesian model-based program STRUCTURE 2.3 was used to infer the population structure with 66 unlinked and/or weakly linked SSR markers (Pritchard et al. 2009). Based on the correct k, each accession was assigned into a subpopulation for which the membership value (Q value) was >0.5 (Pritchard et al. 2000), and the population structure matrix (Q) was generated for further marker-trait association mapping. The software SPAGeDi was used to calculate the pair-wise relatedness coefficients (K, kinship matrix) to estimate the genetic relatedness among individuals with the negative value of kinship set as zero (Hardy and Vekemans 2002). The mixed linear model (MLM) considering both Q and K implemented in the TASSEL software package was used to perform marker-trait association, and the P value and R 2 of each association were determined (Yu et al. 2006; Bradbury et al. 2007).
Favorable allele identification
Based on the results of linkage and association mapping, QTL alleles of loci significantly associated with FW resistance were further analyzed. The phenotypic allele effect was estimated through comparisons between the average phenotypic value over accessions with the specific allele and that of all accessions:
where a i is the phenotypic effect of the ith allele; x ij is the phenotypic value over the jth material with the ith allele; n i is the number of materials with the ith allele; N k is the phenotypic value over all accessions and n k is the number of all accessions. If a i > 0, the allele is determined to be positive, and a i < 0 corresponds to a negative allele. The favorable allele was then identified according to breeding objectives of the target trait (Zhang et al. 2013).
Results
FW susceptibility of assayed accessions
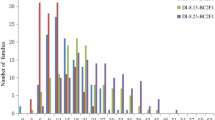
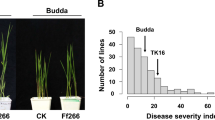
The FW disease indices of three parents, two SF1 crosses and 239 DF1:2 families of the CP were shown in Table 1 and Table 2. Highly significant differences were observed between the S (Xuzhou 142) and R (Yumian 21 and Shang 9901) parents and 239 DF1:2 families by F test. The additive (A) and dominance (D) effects of FOV resistance were estimated through classical quantitative genetics methods (Kong 2006) and were shown in Table 1. The dominant and additive effect ratios in Xuzhou 142/Yumian 21 and Xuzhou 142/Shang 9901 were 0.57 and 0.60 (Table 1), respectively, which shows that partially dominant action of the FOV resistance gene occurred in both single crosses. The DIs of 239 DF1:2 families followed a normal distribution, and transgressive segregations were found (Table 2), which suggests that FW resistance in Upland cotton may be controlled by multiple genes. The FW indices of 356 accessions of the association panel were scored in both the greenhouse and field nursery; summary statistics were shown in Table 2. The DIs of 356 entries in the greenhouse and field nursery evaluations averaged 33.15 and 20.97, with ranges of 1.56–66.67 and 0–97.70, respectively. The DIs obtained in the greenhouse followed a normal distribution, while those obtained in the field nursery did not. This result may be due to the fact that some of the entries escaped from pathogen infection under field nursery conditions. Nonetheless, a significant positive correlation (r = 0.424, P ≤ 0.01) was found between the two measurements.
QTLs detected through linkage mapping
A total of 6,300 SSR primer pairs were used to screen the polymorphisms among Xuzhou 142, Yumian 21 and Shang 9901. Of these primer pairs, 216 (3.5 %) showed polymorphism and amplified 219 loci when they were used to genotype the 241 CP individuals. A Chi square test for goodness of fit was used to assess Mendelian segregation ratios, including 1:1, 1:2:1, and 1:1:1:1 in CP, and showed that 28 loci significantly deviated from the expected segregation ratios (P ≤ 0.05), accounting for 12.79 % of the total. A linkage map with 185 loci and 40 groups was obtained when 191 loci were used to construct the linkage groups (Table S3), leaving six loci unlinked. The map spanned 1,378.8 cM, with an average distance of 7.5 cM between adjacent markers, accounting for approximately 37.6 % of the entire tetraploid cotton genome (Guo et al. 2007; Zhao et al. 2012b). Thirty-nine of 40 linkage groups were assigned to 20 chromosomes, and one could not be assigned. Nineteen linkage groups were assigned to the A-subgenome, containing 80 loci and spanning 623.7 cM, with an average distance of 7.8 cM. Twenty-one linkage groups were assigned to the D-subgenome, containing 103 loci and spanning 748.3 cM, with an average distance of 7.3 cM (Table S3).
QTL analyses were performed with MQM implemented in MapQTL 5.0 (Van Ooijen 2004). A total of seven suggestive QTLs with 2.9–6.6 % of the total phenotypic variance explained were detected; these QTLs were localized to six chromosomes (Fig. 1). A summary of the QTLs, including the chromosome position, LOD score, percentage of phenotypic variance explained by the QTL (PVE), additive effects of a 1 (additive effect of the first single cross) and a 2 (that of the second single cross) and overall dominance effect d was shown in Table 3. The a 1 values of four QTLs were positive (qFW-A3-1, qFW-A12-1, qFW-D3-1 and qFW-D5-1, with values of 2.29, 1.48, 2.19 and 0.98, respectively), which indicates that these alleles, from the resistant parent Yumian 21, led to an increase in FOV 7 resistance (decreasing DI). Four QTLs with positive a 2 values (qFW-A12-1, qFW-D3-1, qFW-D5-1 and qFW-D8-1 with 1.48, 2.19, 2.58 and 2.22, respectively) were detected, which suggests that the other resistant parent, 9901, contributed favorable alleles on these loci. Furthermore, the effects of alleles from the two resistant parents on qFW-A12-1 and qFW-D3-1 were equal; they were maybe the same alleles or different alleles with same effects. Interestingly, the susceptible parent Xuzhou 142 actually contributed favorable alleles on locus qFW-D13-1 and qFW-D13-2 (with negative a 1 and a 2 values). In addition, a single-marker analysis was also conducted among DIs and 34 non-located markers using the Kruskal–Wallis model (Van Ooijen 2004). Four of the 34 markers were significantly associated with FW resistance (K** to K***). According to our saturated linkage map, NAU3695, NAU3519 and Gh401 were assigned to chromosome A11, A12 and D13, respectively (Guo et al. 2007; Zhao et al. 2012a, b), while NAU2666 could not be assigned to any chromosome.
QTLs identified by association mapping
Of the 381 SSR markers evenly distributed across the tetraploid cotton genome, 145 were polymorphic in the 356 accessions, and a total of 415 alleles were detected (Table S2). The allele number, gene diversity and PIC values of the 145 loci averaged 2.86, 0.32 and 0.27, with ranges of 2–9, 0.01–0.73 and 0.27–0.68, respectively (Table S2). Model-based evaluation of the population structure of the 356 accessions showed that the LnP(D) value corresponding to each hypothetical k kept increasing along with the k value and did not show any peak or platform, while the Δk value showed a much higher likelihood at k = 2 than at k = 3–10 (Figure S1), which suggests that the panel should be divided into two major subpopulations. The corresponding Q-matrix was used for subsequent association mapping. Of the kinship coefficient values, 86.85 % were less than 0.05, 8.56 % had a range of 0.05–0.10 and the remaining 4.59 % showed various degrees of genetic relatedness (data not shown). Based on the relatedness among accessions, a K matrix was constructed.
Marker–trait association mapping was performed with the MLM model (Q + K) implemented in TASSEL software. A total of 35 markers were significantly associated with FOV 7 resistance at the α = 0.05 level (−log10 P ≥ 1.30) and were distributed on 18 chromosomes (Table S4), while at the α = 0.01 level (−log10 P ≥ 2.0), a total of 27 loci were significant and were localized to 16 chromosomes (Table 4). Of these loci, 23 were detected under field nursery conditions, 10 were detected in the greenhouse and six were detected under both conditions. The proportion of phenotypic variation explained by the markers ranged from 1.48 to 12.99 %, with an average of 4.21 %.
Favorable QTL alleles and their representative materials
The phenotypic effects of each QTL allele of the 27 associated loci were estimated in both the greenhouse and field nursery evaluations according to the method mentioned above, and therefore, favorable alleles for FOV 7 resistance were identified (Table 5). The phenotypic effects of the 27 loci in the greenhouse and field nursery averaged −1.31 and −3.45, with ranges of −3.81 to −0.01 and −14.88 to −0.20, respectively. Among the favorable alleles, NAU934-2 had the most negative phenotypic effect in the greenhouse assay and was able to decrease FW DI by 3.8, while NAU6966-3 had the most negative phenotypic effect in the field nursery and was able to decrease FW DI by 14.88. A wide variation of the favorable alleles processed in the 356 accessions was observed, with an average of 14.84 and a range of 7–21 (Table S1). Pearson correlation analysis between the number of favorable alleles and FW DI was carried out, and highly negative significance was found in both the greenhouse (r = −0.344, P < 0.001) and the field nursery (r = −0.488, P < 0.001) evaluations. The top-5 resistant accessions for each favorable allele were listed as representative materials in Table 5.
Discussion
QTLs for FOV 7 resistance identified by linkage and association mapping
FOV 7 is the most widely distributed FOV race in China (Chen et al. 1985; Sun et al. 1999). Molecular tagging and favorable allele mining of FOV 7 resistance QTLs/genes will be highly beneficial for cotton improvement. In previous studies, Chen et al. (2008) detected four QTLs conferring FOV 7 resistance using an interspecific cross (G. hirsutum 98134 × G. barbadense Xinhai 14) and located these QTLs to Chr 3 (A3), 15 (D1), 23 (D9) and 26 (D12), respectively. Wang et al. (2009) detected four QTLs associated with FOV 7 resistance in Upland cotton and located these QTLs to four chromosomes [Chr 7 (A7), 15 (D1), 17 (D3) and 23 (D9)], with a major QTL on Chr 17 (D3) that explained 52.5–60.9 % of the total phenotypic variation. In a subsequent report, Wang et al. (2010) detected five QTLs associated with FW 7 resistance in two intraspecific populations, with one located on Chr 15 (D1) and four on Chr 2 (A2) or 17 (D3), and indicated that the latter four QTLs were linked to JESPR304 or CIR 35; it is possible that these QTLs were the same as the major QTL previously detected (Wang et al. 2009). In addition, the authors also detected two major QTLs on Chr 9 (A9) and Chr 12 (A12) or 26 (D12) in another interspecific population. In the current study, we detected 7 suggested QTLs conferring FOV 7 resistance using linkage mapping, and we localized the QTLs to six chromosomes (A3, A12, D3, D5, D8 and D13; Fig. 1; Table 3), which greatly exceeded that detected in previous studies (Chen et al. 2008; Wang et al. 2009, 2010). The results showed that using a composite CP with more than two parents is more powerful for QTL detection (Qin et al. 2008; Zhang et al. 2012) than the populations used in previous studies, although it is not easy to compare QTL mapping results from different studies due to divergent plant materials and markers used. Three (qFW-A3-1, qFW-A12-1, qFW-D3-1) of the 7 QTLs were similar in map position to that detected in previous studies (Chen et al. 2008; Wang et al. 2009, 2010), while the LOD scores (1.22–3.45) and PVEs (2.9–6.6 %) were somewhat lower in the current study. This result may be due to the fact that resistance evaluation was carried out only at the seedling stage, when the susceptible plants had not time enough to develop severe symptoms. Therefore, resistance should be evaluated at more than one plant developmental stages under greenhouse conditions.
Association mapping can be affected by many factors, such as population structure, relatedness among accessions, small sample size and low frequency of specific alleles; these factors may increase the detection of false positive associations (Gupta et al. 2005; Yu et al. 2006). In this study, AM was performed with a moderately large-sized panel (356 accessions) using the optimal model of MLM, which considers both population structure and relatedness, to detect SSR markers associated with FOV 7 resistance. Nonetheless, it is still not easy to determine which significance level is the most appropriate. The use of stringent probability thresholds reduces the danger of false positives but poses the risk of rejecting true positives caused by setting the thresholds too high (Yan et al. 2011). In the present study, 35 loci were significantly associated with FOV 7 resistance at the α = 0.05 level (Table S4), and 27 loci were significant at the α = 0.01 level (Table 4). If a more stringent threshold by the Bonferroni correction (P ≤ 0.05/145, −log10 P ≥ 3.46) is adopted (Lander and Botstein 1989), only 10 associations were significant. Five (qFW-A3-1, qFW-A12-1, qFW-D3-1, qFW-D5-1 and qFW-D8-1) of the 7 QTLs identified by linkage mapping could be detected by association mapping at the α = 0.05 level (Table S4), while only three QTLs (qFW-D3-1, qFW-D5-1 and qFW-D8-1) could be detected at the α = 0.01 level (Table 4). The marker JESPR304, which is closely linked with a major QTL [Chr 17 (D3)] (Wang et al. 2009, 2010), was indeed significantly (−log10 P = 7.09) associated with FOV 7 resistance in our association mapping study, while this locus should be located on Chr 2 (A2) (Guo et al. 2007; Zhao et al. 2012a, b; Ulloa et al. 2013). Unfortunately, JESPR304 did not amplify polymorphic bands among the three parents of our CP.
Inheritance and genetic basis of FOV resistance in cotton
As FW has long been and will continue to be an important disease that affects cotton production, most studies of this disease have been performed to unravel the genetics of FW resistance. In early classical genetic studies, the inheritance of FOV resistance in cotton was considered to be controlled by a single gene (Smith and Dick 1960; Netzer et al. 1985; Feng et al. 1996) or multiple genes (Kappelman 1971; Jiao 1985; Wang et al. 1989) with partially dominant effects. In the current study, partial dominance was also observed in the two single crosses, Xuzhou 142/Yumian 21 and Xuzhou 142/Shang 9901, with dominant-additive ratios of 0.57 and 0.60, respectively (Table 1).
Wang and Roberts (2006) identified a major resistance gene (Fov1) against FOV 1 in G. barbadense cv. Pima-S7 and indicated that one or more minor genes in Acala NemX could delay wilt symptoms. Recently, Ulloa et al. (2011) performed QTL mapping in an F2 (Pima-S7 × Acala NemX) and a recombinant inbred line (RIL; G. hirsutum TM-1 × G. barbadense Pima 3-79) population. The authors detected six QTLs (Fov1-C06, Fov1-C08, Fov1-C111, Fov1-C112, Fov1-C16 and Fov1-C19) conferring FOV 1 resistance and confirmed a major QTL (Fov1-C16) in different genetic backgrounds. Lopez-Lavalle et al. (2012) used an intraspecific cross between the Upland cotton MCU-5 and Siokra 1-4 to detected QTLs conferring resistance to Australian FOV races and found that MCU-5 resistance is complex, with 3 QTLs identified in F3 and 8 ones in F4. The QTLs were localized to chromosomes A6, D4 and D6. Ulloa et al. (2013) investigated three intraspecific (G. hirsutum × G. hirsutum L. and G. barbadense × G. barbadense L.), five interspecific (G. hirsutum × G. barbadense) and one RIL population in four greenhouse and two field experiments and identified a set of 11 SSR markers across six linkage groups/chromosomes (3, 6, 8, 14, 17 and 25) associated with FOV race 4 resistance. Integrating information mentioned above with the QTL mapping results for FOV 7 in our study,, it suggests that the inheritance of FOV resistance in cotton should be far more complex than has been elucidated to date. Interestingly, the map positions of some QTLs conferring resistance against different FOV races coincided, suggesting that these genes may play the same or similar roles in the process of different host-pathogen and/or environment interactions.
Favorable alleles and their potential use in breeding programs
Host resistance has always been considered to be an effective way to manage plant diseases. Although significant progress in the production of FOV-resistant cotton has been made by conventional phenotype-based selection, this process has been slow and inefficient in the face of emergence of new, highly virulent isolates of FOV in Australia and the United States (Wang et al. 2004; Kim et al. 2005). The development of effective markers for FOV resistance offers the promise of increasing the selective gain while ensuring sustainable targeted breeding progress. In the current study, favorable alleles for each of the 27 associated loci were identified, and the number of favorable alleles in each of 356 accessions were also investigated (Table 5, Table S1). Pearson correlation analysis showed that there was highly negative significance between DIs and the number of favorable alleles. While great differences in the resistance of cotton genotypes against FOV races have been demonstrated in many evaluations worldwide, a genotype with a high level of resistance against all FOV races has not been found. Given that FW is the result of interactions among the host, pathogen and environment, many genes must be involved in this process. The accumulation of resistance alleles, even with relatively minor individual effects, will result in higher levels of resistance. The Upland cotton cultivars from China and Africa have shown more resistance to Australian FOV races (Constable et al. 2007). Therefore, the favorable alleles and their typical germplasm resources indentified in this study should have great potential for developing highly resistant Upland cotton cultivars in future molecular breeding programs.
References
Abdurakhmonov IY, Kohel RJ, Yu JZ, Pepper AE, Abdullaev AA, Kushanov FN, Salakhutdinov IB, Buriev ZT, Saha S, Scheffler BE, Jenkins JN, Abdukarimov A (2008) Molecular diversity and association mapping of fiber quality traits in exotic G. hirsutum L. germplasm. Genomics 92:478–487
Abdurakhmonov IY, Saha S, Jenkins JN, Buriev ZT, Shermatov SE, Scheffler BE, Pepper AE, Yu JZ, Kohel RJ, Abdukarimov A (2009) Linkage disequilibrium based association mapping of fiber quality traits in G. hirsutum L. variety germplasm. Genetica 136:401–417
Brachi B, Faure N, Horton M, Flahauw E, Vazquez A, Nordborg M, Bergelson J, Cuguen J, Roux F (2010) Linkage and association mapping of Arabidopsis thaliana flowering time in nature. PLoS Genet 6:e1000940
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Cadic E, Coque M, Vear F, Grezes-Besset B, Pauquet J, Piquemal J, Lippi Y, Blanchard P, Romestant M, Pouilly N, Rengel D, Gouzy J, Langlade N, Mangin B, Vincourt P (2013) Combined linkage and association mapping of flowering time in Sunflower (Helianthus annuus L.). Theor Appl Genet. doi:10.1007/s00122-013-2056-2
Chen Q, Ji X, Sun W (1985) Identification of cotton-wilt Fusarium races in China. Scientia Agricultura Sinica 6:1–6
Chen X, Ge F, Wang D, Shi W, Xu L, Wu W, Huang Q (2008) Mapping of QTLs conferring resistance to Fusarium wilt in cotton. Mol Plant Breed 6:1127–1133
Chen X, Guo W, Zhang T (2011) Cotton omics in China. Plant Omics 4:278–287
Constable GA, Reid PE, Stiller WN (2007) Breeding for resistance to a new strain of Fusarium wilt in Australia, In: Stephens M (ed) Proceedings of the World Cotton Research Conference-4, Sept 10–14, Lubbock
Davis RM, Colyer PD, Rothrock CS, Kochman JK (2006) Fusarium wilt of cotton: population diversity and implications for management. Plant Dis 90:692–703
Feng C, Zhang J, Liu J, Guo J, Sun J (1996) Inheritance of resistance to Fusarium wilt in several Chinese upland cotton varieties. Acta Agronomica Sinica 22:550–554
Guo WZ, Cai C, Wang C, Han Z, Song X, Wang K, Niu X, Wang C, Lu K, Shi B, Zhang TZ (2007) A microsatellite-based, gene-rich linkage map reveals genome structure, function and evolution in Gossypium. Genetics 176:527–541
Gupta PK, Rustgi S, Kulwal PL (2005) Linkage disequilibrium and association studies in higher plants: present status and future prospects. Plant Mol Biol 57:461–485
Hardy O, Vekemans X (2002) SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620
Holmes EA, Bennett RS, Spurgeon DW, Colyer PD, Davis RM (2009) New genotypes of Fusarium oxysporum f. sp. vasinfectum from the Southeastern United States. Plant Dis 93:1298–1304
Jannink JL, Lorenz AJ, Iwata H (2010) Genomic selection in plant breeding: from theory to practice. Brief Funct Genomics 9:166–177
Jiao B (1985) Primary research of general combining ability and heritability of resistance to Fusarium wilt and Verticillium wilt in upland cotton. Acta Agronomica Sinica 11:267–273
Kantartzi SK, Stewart McD J (2008) Association analysis of fibre traits in Gossypium arboreum accessions. Plant Breed 127:173–179
Kappelman AJ (1971) Inheritance of resistance to Fusarium wilt in cotton. Crop Sci 11:672–674
Kim Y, Hutmacher RB, Davis RM (2005) Characterization of California isolates of Fusarium oxysporum f. sp. vasinfectum. Plant Dis 89:366–372
Kong FL (ed) (2006) Quantitative genetics in plants. China Agricultural University Press, Beijing, pp 15–16
Korir PC, Zhang J, Wu K, Zhao T, Gai J (2013) Association mapping combined with linkage analysis for aluminum tolerance among soybean cultivars released in Yellow and Changjiang River Valleys in China. Theor Appl Genet. doi:10.1007/s00122-013-2082-0
Lacape JM, Llewellyn D, Jacobs J, Arioli T, Becker D, Calhoun S, Al-Ghazi Y, Liu S, Palaï O, Georges S, Giband M, de Assunçã H, Barroso PA, Claverie M, Gawryziak G, Jean J, Vialle M, Viot C (2010) Meta-analysis of cotton fiber quality QTLs across diverse environments in a Gossypium hirsutum × G. barbadense RIL population. BMC Plant Biol 10:132
Lander ES, Botstein D (1989) Mapping mendelian factors underlying quantitative traits using RFLP linkage maps. Genetics 121:185–199
Liu KJ, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinfo Appl Note 9:2128–2129
Lopez-Lavalle LAB, Gillespie VJ, Tate WA, Ellis MH, Stiller WN, Llewenllyn L, Wilson IW (2012) Molecular mapping of a new source of Fusarium wilt resistance in tetraploid cotton (Gossypium hirsutum L.). Mol Breed 30:1181–1191
Lu Y, Zhang S, Shah T, Xie C, Hao Z, Li X, Farkhari M, Ribaut J-M, Cao M, Rong T, Xu Y (2010) Joint linkage–linkage disequilibrium mapping is a powerful approach to detecting quantitative trait loci underlying drought tolerance in maize. Proc Natl Acad Sci USA 107:19585–19590
Ma C (ed) (2007) Researches on Fusarium and Verticillium wilt in cotton. China Agriculture Press, Beijing, pp 10–19; pp 303–305
Ma C, Jian G, Zheng C (2002) The advances in cotton breeding resistance to Fusarium and Verticillium wilts in China during past fifty years. Scientia Agricultura Sinica 35:508–513
Mackay TFC, Stone EA, Ayroles JF (2009) The genetics of quantitative traits: challenges and prospects. Nat Rev Genet 10:565–577
McCouch SR, Cho YG, Yano PE, Blinstrub M, Morishima H, Kinoshita T (1997) Report on QTL nomenclature. Rice Genet Newslet 14:11–13
Myles S, Peiffer J, Brown PJ, Ersoz ES, Zhang Z, Costich DE, Buckler ES (2009) Association mapping: critical consideration shift from genotype to experimental design. Plant Cell 21:2194–2202
Nemri A, Atwell S, Tarone AM, Huang YS, Zhao K, Studholme DJ, Nordborg M, Jones JDG (2010) Genome-wide survey of Arabidopsis natural variation in downy mildew resistance using combined association and linkage mapping. Proc Natl Acad Sci USA 107:10302–10307
Netzer D, Tal Y, Marani A, Weintall C (1985) Resistance of interspecific cotton hybrids (Gossypium hirsutum × Gossypium barbadense containing Gossypium harknessii cytoplasm) to fusarium wilt. Plant Dis 69:312–313
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38:904–909
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Pritchard JK, Wen X, Falush D (2009) Documentation for structure software: Version 2.3 http://pritchardlab.stanford.edu/structure.html
Qin H, Guo W, Zhang Y, Zhang T (2008) QTL mapping of yield and fiber traits based on a four-way cross population in Gossypium hirsutum L. Theor Appl Genet 117:883–894
Rodríguez-Gálvez E, Mendgen K (1995) The infection process of Fusarium oxysporum in cotton root tips. Protoplasma 189:61–72
Rong J, Feltus FA, Waghmare VN, Pierce GJ, Chee PW, Draye X, Saranga Y, Wright RJ, Wilkins TA, May OL, Smith CW, Gannaway JR, Wendel JF, Paterson AH (2007) Meta-analysis of polyploid cotton QTL shows unequal contributions of subgenomes to a complex network of genes and gene clusters implicated in lint fiber development. Genetics 176:2577–2588
Skovgaard K, Nirenberg HI, O’Donnell K, Rosendahl S (2001) Evolution of Fusarium oxysporum f. sp vasinfectum races inferred from multigene genealogies. Phytopathology 91:1231–1237
Smith AL, Dick JB (1960) Inheritance of resistance to Fusarium wilt in upland cotton and sea island cotton as complicated by nematodes under field conditions. Phytopathology 50:44–48
Sun W, Jian G, Chen Q (1999) Study on monitoring of physiological race of cotton Fusarium wilt in China. Scientia Agricultura Sinica 32:51–57
Ulloa M, Hutmacher RB, Davis RM, Wright SD, Percy R, Marsh B (2006) Breeding for Fusarium wilt race 4 resistance in cotton under field and greenhouse conditions. J Cotton Sci 10:114–127
Ulloa M, Percy R, Zhang J, Hutmacher RB, Wright SD, Davis RM (2009) Registration of four Pima cotton germplasm lines having good levels of Fusarium wilt race 4 resistance with moderate yields and good fibers. J Plant Regist 3:198–202
Ulloa M, Wang C, Hutmacher RB, Wright SD, Davis RM, Saski CA, Roberts PA (2011) Mapping Fusarium wilt race 1 resistance genes in cotton by inheritance, QTL and sequencing composition. Mol Genet Genomics 286:21–36
Ulloa M, Hutmacher RB, Roberts PA, Wright SD, Nichols RL, Davis RM (2013) Inheritance and QTL mapping of Fusarium wilt race 4 resistance in cotton. Theor Appl Genet 126:1405–1418
Van Ooijen J (1999) LOD significance thresholds for QTL analysis in experimental populations of diploid species. Heredity 83:613–624
Van Ooijen JW (2004) MapQTL 5, software for the mapping of quantitative trait loci in experimental populations. Kyazma BV, Wageningen
Van Ooijen JW, Voorrips RE (2006) JoinMap 4.0: software for the calculation of genetic linkage maps. Plant Research International, Wageningen
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Wang C, Roberts PA (2006) A Fusarium wilt resistance gene in Gossypium barbadense and its effect on root-knot nematode-wilt disease complex. Phytopathology 96:727–734
Wang Z, Ma Z, Qu J (1989) Analysis of gene effect of resistance to Fusarium and Verticillum wilt in upland cotton. J Heibei Agric Univ 12:21–25
Wang B, Brubaker CL, Burdon JJ (2004) Fusarium species and Fusarium wilt pathogens associated with native Gossypium populations in Australia. Mycol Res 108:35–44
Wang P, Su L, Qin L, Hu B, Guo W, Zhang T (2009) Identification and molecular mapping of a Fusarium wilt resistant gene in upland cotton. Theor Appl Genet 119:733–739
Wang P, Shi L, Su L, Hu B (2010) Quantitative trait loci for resistance against Fusarium wilt based on three cotton F2 populations. Agric Sci China 9:1799–1806
Xu Y, Crouch J (2008) Marker-assisted selection in plant breeding: from publications to practice. Crop Sci 48:391–407
Yan J, Warburton M, Crouch J (2011) Association mapping for enhancing maize (Zea mays L.) genetic improvement. Crop Sci 51:433–449
Yu J, Pressoir G, Briggs WH, Bi IV, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Yu J, Holland JB, McMullen MD, Buckler ES (2008) Genetic design and statistical power of nested association mapping in maize. Genetics 178:539–551
Zeng L, Meredith WR Jr, Gutierrez OA, Boykin DL (2009) Identification of associations between SSR markers and fibre traits in an exotic germplasm derived from multiple cross among Gossypium tetraploid species. Theor Appl Genet 119:93–103
Zhang J, Guo W, Zhang T (2002) Molecular linkage map of allotetraploid cotton (Gossypium hirsutum L. × Gossypium barbadense L.) with a haploid population. Theor Appl Genet 105:1166–1174
Zhang HB, Li Y, Wang B, Chee PW (2008) Recent advances in cotton genomics. Int J Plant Genomics. doi:10.1155/2008/742304
Zhang K, Zhang J, Ma J, Tang S, Liu D, Teng Z, Liu D, Zhang Z (2012) Genetic mapping and quantitative trait locus analysis of fiber quality traits using a three-parent composite population in upland cotton (Gossypium hirsutum L.). Mol Breed 29:335–348
Zhang T, Qian N, Zhu X, Chen H, Wang S, Mei H, Zhang Y (2013) Variations and transmission of QTL alleles for yield and fiber qualities in Upland cotton cultivars developed in China. PLoS ONE 8:e57220
Zhao L, Cai CP, Mei HX, Guo WZ (2012a) Screening of microsatellite loci for identifying genome barcoding of cotton cultivars. ACTA Agronomica Sinica 38:1810–1817
Zhao L, Lv Y, Cai C, Tong X, Chen X, Chen X, Zhang W, Du H, Guo X, Guo W (2012b) Toward allotetraploid cotton genome assembly: integration of a high-density molecular genetic linkage map with DNA sequence information. BMC Genom 13:539
Zhu C, Gore M, Buckler ES, Yu J (2008) Status and prospects of association mapping in plants. Plant Genome 1:5–20
Acknowledgments
We thank professor Xiongming Du, Cotton Research Institute, Chinese Academy of Agriculture Sciences, for providing part of the cotton cultivars used in this research. This work was financially supported in part by Grants from the Major State Basic Research Development Program of China (973 Program) (2011CB109300), the National High Technology Research and Development Program of China (863 Program) (2011AA10A102) and the Priority Academic Program Development of Jiangsu Higher Education Institutions and 111 program.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mei, H., Ai, N., Zhang, X. et al. QTLs conferring FOV 7 resistance detected by linkage and association mapping in Upland cotton. Euphytica 197, 237–249 (2014). https://doi.org/10.1007/s10681-014-1063-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-014-1063-y