Abstract
Antibiotic susceptibility test (AST) is an umbrella term for techniques to determine the susceptibility of bacteria to antibiotics. The antibiotic-resistant bacteria are a major threat to public health and a directed therapy based on accurate AST results is paramount in resistance control. Here we have briefly covered the progress of conventional, molecular, and automated AST tools and their limitations. Various aspects of microfluidic AST such as optical, electrochemical, colorimetric, and mechanical methods have been critically reviewed. We also address the future requirements of the microfluidic devices for AST. Cumulatively, we have outlined the overview of AST that can help to expand and improve the existing techniques and emphasize that microfluidics could be the future of AST and introduction of microtechnologies in AST will be extremely advantageous, especially for point-of-care testing.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The discovery of the penicillin, the first true antibiotic, is considered as the single most crucial step towards the war against microbial infections. Antibiotics are usually secondary metabolites of plants, fungi and microorganism with a potential to kill (bactericidal) or inhibit the growth (bacteriostatic) of the microorganism (Xue et al. 2018). With inappropriate use of the antibiotic, the bacteria have gained resistance towards several group of antibiotics. The antibiotic-resistant bacteria are a major threat to public health as a wide array of antibiotic resistance is reported in even diverse species of bacteria. Thus, the majority of them are difficult or impossible to detect. These bacteria have potential to induce severe clinical manifestations, and limited treatments are available (Blair et al. 2015). Moreover, it is also an obstacle to the development of novel drugs.
The antibiotic misuse/abuse leads to an extremely complex and high rate of mutation and evolution in the bacteria (Blair et al. 2015). The inappropriate prescription practices (Lansang et al. 1990), inadequate patient education (Lansang et al. 1990), unauthorized sale of antimicrobials (Saleh et al. 2015), limited diagnostic facilities (Ayukekbong et al. 2017), lack of appropriate functioning of drug regulatory authorities (Dua et al. 1994), and non-human use of antimicrobials such as in animals (Angulo et al. 2005; Petersen et al. 2002) are the major cause of the development of antibiotic resistance (Fig. 1). The rate of progression of antibiotic-resistant microbes is higher in the developing countries because of higher incidents of aforementioned malpractices (Ayukekbong et al. 2017; Blair et al. 2015; Van Boeckel et al. 2014).
Antibiotic susceptibility test (AST) is the most accepted approach for targeting the bacterial resistance and susceptibility (RamezanAli et al. 2012). AST is a comprehensive and reproducible practice to determine the potential antibiotics against the infectious bacteria (Jenkins and Schuetz, 2012). It is prominently used when the bacterial species under study have acquired resistance to commonly prescribed antibiotics (Jenkins and Schuetz 2012). ISO standard 20776 has categorized the infectious agent as “susceptible”, “intermediate”, or “resistant” to particular antibiotics (SIR) (Rodloff et al. 2008). These guidelines are helpful in the antibiotic prescription and personalized medicine, thus act as a formidable way to reduce the growing antibiotic resistance in bacteria, if followed properly.
The goal of the current review article is to present an overview of various aspects of conventional AST. Different classes of microfluidic AST tools such as optical, electrochemical, colorimetric and mechanical, have been presented with critical evaluation of their caveats (Table 1). Furthermore, the future perspective of microfluidic based AST is discussed.
Conventional AST methods
Both the phenotypic and genotypic methods have been developed to determine the antibiotic susceptibility of infectious bacteria. Several phenotypic techniques including disk diffusion, dilution (agar and broth), and E-test are well established. These methods are dependable, however, low sensitivity (more than 105 cells), complicated sample preparations, lack of automation and growth dependency are the major limitations (Balouiri et al. 2016). With the advancement, research shifted towards molecular detections such as polymerase-chain-reaction (PCR), matrix-assisted laser desorption/ionization- time of flight mass spectroscopy (MALDI-TOF MS), DNA arrays and chips. Molecular AST methods are generally attributed to rapid, direct, sensitive and specific detection of resistance genes, but need of specific assay for specific antibiotic, false negative due coincidental mutations, expensive reagents and equipment, and requirement of skilled personnel, are major drawbacks that diminish their clinical utility.
Also, automated machines like Phoenix, MicroScan, WalkAway, and Vitek came up to mitigate the limitations of conventional techniques (Pulido et al. 2013). These methods are attributed to the sensitive, rapid and automatic analysis of bacterial viability. Despite these advancements, routine laboratory maintenance, the requirement of trained operators, poor reproducibility, and the possibility of false-positive results are still an issue for clinical utility (Felmingham and Brown, 2001). Recently, several automated machines such as VITEK2, Sensititre ARIS 2 have been introduced, which facilitate sensitive and accurate evaluation of antibiotic susceptibility. These machines are more reliable and reproducible, but the inability to detect all clinically relevant bacteria against various antibiotics are downsides for wide acceptance (Karlowsky and Richter, 2015). In general, the prohibitive cost of equipment and consumables, the inability for single bacterial cell analysis and time-consuming procedure are the major drawbacks lying in the above-mentioned methods. These shortcomings have opened new vistas for microfluidics in AST.
Microfluidic AST methods
Microfluidics is an emerging technology that deals with the manipulation of a small number of fluids typically on an integrated circuit of micrometer-sized fluidic channels. Microfluidics is considered as a promising tool over the conventional technologies as it offers precise manipulation and control of micro-volume samples. Also, low cost, less sample volume, scalability, batch performance, multiplexing, and high throughput are its attractive features (Park et al. 2011). Major advancement on the microfluidic-based AST has been initiated after 2005. Researchers have shown that problem of conventional AST can be easily addressed by microfluidics as it offers automation and high-throughput analysis (Lee et al. 2017). Therefore, microfluidics can be expected as an attractive platform for rapid, sensitive, automated and accurate bio-sensing tool for clinical diagnosis. Based on the detection methods, microfluidic systems for AST can be categorized into optical, electrochemical, mechanical, and colorimetric systems. In the following, major features of microfluidic methods and their potential for the determination of antibiotic susceptibility and MIC are discussed.
Optical method
Recently, microfluidic observation through various microscopic techniques has become a mainstay for physiological and behavioral research (Obara et al. 2013). The Phase contrast and fluorescence microscopy with detection marker (fluorescence dyes and proteins) are the centerpiece to envisage the microfluidic antibiotic susceptibility (Skafte-Pedersen et al. 2012). In 2010, microfluidic channels with large surface-to-volume ratio and phase contrast microscopy were utilized to facilitate and monitor the rapid growth of Escherichia coli under the influence of antibiotics (Chen et al. 2010). In another study, antibiotic susceptibility was estimated by time-lapse photographs obtained from the phase contrast microscopy. In this case, the bacteria were cultured in a 3D microfluidic culture device. The images of growing bacteria under antibiotic stress were obtained and grayscale values of images were used to evaluate the susceptibility (Hou et al. 2014a, b) (Fig. 2). Bacterial species showing the resistance to wide range of antibiotics such as Pseudomonas aeruginosa pose another challenge as diagnostic cannot be relied on a single antibiotic, and thus become a heuristic process. Microfluidic approach provides automated and multiple detection process by the integration of fluidic circuits, and the AST of these strains performed in parallel with high precision by optical microfluidics has been successfully demonstrated (Matsumoto et al. 2016). The optical techniques are rapid but still rely on growth dynamic analysis. This approach can provide insufficient information about the behavior of bacteria towards a given antibiotic. To deliver a comprehensive bacterial behavioral profile, the growth phase study must be coupled with the morphological analysis. Therefore, recently in 2014, the microfluidic-based morphological survey has been carried out for 189 clinical samples along with CLSI recommended strains against variable antibiotics. The satisfactory AST results with a good agreement of 91.5% were obtained within 4 h (Choi et al. 2014). The overall time required for morphology inspection was found to be greater than growth analysis. The entire survey utilized the phase contrast microscopy for bacterial analysis.

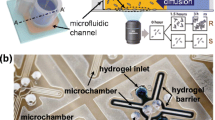
adapted with permission from (Hou et al. 2014a, b)}, II Colorimetric pH sensors {adapted with permission from (Tang et al. 2013)}, III electrochemical microfluidic device, bacteria trapped in confined microchannels for single cell AST {adapted with permission from (Lu et al. 2013)}, IV Mechanical, the asynchronous magnetic bead rotation (AMBR) method {adapted with permission from (Kinnunen et al. 2011)
A schematic presentation of various microfluidic AST methods, I optical, showing 3D microfluidic culture and device {
Apart from the phase contrast microscopy, fluorescence microscopy is often used for quantitative and qualitative analysis for AST. Bacterial strains are transformed with green fluorescent protein (GFP) labeled protein and emit fluorescence. These transformed bacteria are subjected to antibiotics. The resulting fluorescence is quantified using fluorescence microscope which is directly proportional to the number of viable cells present in the system. This property is widely exploited for AST (Lehtinen et al. 2004). Long-term time-lapse microscopy coupled with fluorescence property of GFP has been successfully utilized to study antibiotic resistance in Mycobacterium tuberculosis which is the root cause of tuberculosis. This study is vital as it is a rapidly mutating bacteria and multiple/extensive drug resistance (MDR/XDR) has already been reported (Ventola 2015). Recently, multiplexing capability of the microfluidic platform has been fully exploited, and the resistance acquired against ampicillin, cefalexin, chloramphenicol, and tetracycline in Escherichia coli has been successfully evaluated within 2–4 h by employing fluorescence-based microfluidic systems (Mohan et al. 2013a).
Fluorescent dyes such as SYTOX green is used for resistant analysis as it can permeate inside the dead bacteria only and produce the fluorescence signal when it binds to the DNA. The efficacy of SYTOX green for AST was well demonstrated on the assessment of Staphylococcus aureus which was associated with over 18,000 deaths in the United States in 2005 (Kalashnikov et al. 2012). After bacterial culture in antibiotic-containing media, SYTOX dye permeates inside the dead bacteria and binds with the DNA. SYTOX does not permeate in live bacteria. Thus, the susceptible bacteria can be differentiated by fluorescence emission. In another dye-based approach, resazurin has been added to the culture media. As resazurin is the fluorescent redox indicator, it is converted into resafurin by electron receptors used in the cellular metabolic activity, such as NADH and FADH. As the dye indicates viability, the fluoresces confirmed the resistant bacteria (Boedicker et al. 2008). Based on this method, MIC of ampicillin, chloramphenicol, and tetracycline against Escherichia coli and cefoxitin against Staphylococcus aureus had been accurately estimated (Boedicker et al. 2008).
Electrochemical method
Electrochemical sensors are another important bio-recognition tool. It exploits the electrochemical properties of target analytes. Electrochemical measurements relies on a specific combination of electrode and electrolyte which are targeted to capture specific signal (Rackus et al. 2015). A significant amount of work has been done on electrochemical biosensors in last decade which include several reports on antibiotic susceptibility test. Peitz et al. devised an automated and high throughput setup for single bacterial cell detection, which employed dielectrophoretic principle (DEP) to capture bacteria in microfluidic channels (Peitz and van Leeuwen, 2010). AST with Escherichia coli and polymyxin validated the performance of this platform. The obtained results strongly supported the implementation of an electrochemical sensor in the fields of diagnostics and biofilm investigations. DEP application is further extended for demonstrating a rapid detection of antibiotic susceptibility by using the DEP chip, where the crossover frequency (cof) change is recorded at different antibiotic concentration against Escherichia coli (Lu and Wong 2011) (Fig. 2). Quick estimation was the major advantage as the results could be estimated within 60 min. Electrochemical DNA biosensors are another key development in this class of biosensor. These Electrochemical DNA biosensors are not only limited to detect vitality, but they can also detect the bacteria on the basis of 16s rRNA. The DNA probe is stem-loop shaped, modified with thiol at its 5′ end and biotin at its 3′ end, was immobilized on a gold electrode. The probes were “closed” when the target was absent. The probe-target DNA hybridization induces the conformational changes to “open”, along with the biotin at its 3′ end binding with streptavidin–horseradish peroxidase (HRP), followed by quantification via electrochemically detecting the enzymatic product in the presence of substrate (Liu et al. 2011). This approach is used for AST with the potential of molecular-level characterization of bacteria in culture and also directly from the physiological samples such as urine (Halford et al. 2013).
Colorimetric method
The marked difference between the colorimetric and other microfluidic sensors is the output signal and affordability. Colorimetric based microfluidic devices are easiest to perceive as the results are visible to naked eyes. It provides rapid response, high sensitivity, simplicity in optical settings and multiplexing. The sensing device is generally composed of microfabricated units which include bacteria, antibiotic-containing growth media and pH indicator dyes (Mohan et al. 2013b; Tang et al. 2013) (Fig. 2). In AST, pH-sensitive dyes are a key element of the colorimetric detection. The pH of the growth media increases due to the growth of bacteria and subsequently, the color of the media changes. This change in color is observed either visually or with simple optical techniques. Both qualitative and quantitative results can be obtained accurately. Various bacteria and antibiotics have been investigated on the microfluidic platform for colorimetric antibiotic susceptibility sensing. A promising example of colorimetric AST was demonstrated using pH indicator dye, phenol red (Nordmann et al. 2012). In a self-loading microsystem, the bacteria are grown under the exposure of different concentration of antibiotics. As the bacteria consume the nutrient media, they metabolize and release acidic products, leading to pH and color changes. This color change can simply be visualized under the ambient light without the necessity of complex microscopy. Based on this method, MIC of vancomycin, tetracycline, and kanamycin against Enterococcus faecalis, Proteus mirabilis, Klebsiella pneumoniae, and Escherichia coli, have been determined. The obtained results revealed a good correlation with conventional method (Cira et al. 2012). The drug susceptibility testing microfluidic (DSTM) devices have been developed for testing Pseudomonas strain in which growth is detected through colorimetric methods using control bacteria (Matsumoto et al. 2016). The obtained morphological data represented the respectable efficiency with conventional broth dilution method. Later, a paper-based chromogenic microfluidic strategy has been developed for AST that showed the accuracy of 94% compared with the conventional technique. Chromogenic medium for Staphylococcus aureus, Escherichia coli, and Dung enterococcus and paper impregnated antibiotic were used to quantify the results. The whole protocol was completed within 15 h (Matsumoto et al. 2016; Nordmann et al. 2012). Recently, a prototype of conventional broth dilution method is transformed on the microfluidic platform for colorimetric AST analysis where wild-type (ATCC 29212) and vancomycin-resistant Enterococcus cells are tested using phenol red against five different vancomycin concentrations. A comparable performance is recorded with E-test within 24 h (Lee et al. 2017).
Mechanical method
One of the recently developed dimensions in the microfluidic AST is mechanical sensor. Mechanical method involves imposition of mechanical stress on bacteria and observing the change in physical properties like viscous drag or rotational rate. Bacteria immobilized on the magnetic beads were employed to study the AST (Li et al. 2014). This method is generally called asynchronous magnetic bead rotation (AMBR) which is composed of micro-droplets confining AMBR beads. These micro-droplets have conjugated bacteria and change the topographical property (volume or shape) and drag-coefficient of beads when these bacteria grow under antibiotic treatment. This change in drag-coefficient helps in discriminating the resistant bacteria from susceptible ones. The method enabled the rapid and simple approach for single cell susceptibility testing (Sinn et al. 2011) (Fig. 2). Modified approach was then formulated with label-free AMBR micro-viscometer instead of labeled AMBR (Sinn et al. 2012). In this concept, AMBR micro-viscometer was encapsulated in water–oil micelle containing uropathogenic Escherichia coli. The minute change in the viscosity with growth of bacteria in the medium was indicated within 20 min, providing rapid, sensitive and label-free approach for bacterial growth and drug susceptibility test. By applying this method, MIC of gentamicin was determined against Escherichia coli isolate within 100 min. This new system significantly reduced the AST time and complexity. Meanwhile, all the above-mentioned techniques suffer from the difficulty in imaging of single bacterial cell under the microscope. Later, self-assembled AMBR method has been introduced, which claimed the analysis of the cluster of self-assembled magnetic particles with increased sensitivity (Kinnunen et al. 2012). Self-assembled cluster of magnetic beads provided the direct implementation of AMBR biosensors on microfluidic platform with simple microscopic observations. When the effect of antibiotic is futile, the bacteria multiplies and slow down the rotation of the cluster. But if it is effective, the bacterial growth is inhibited, and the rotational period remains same. The rotation rate of the clusters is measured by microscope and observed proportional to the antibiotic susceptibility. Apart from the AMBR, the potential of fluid flow induced stress for testing the antibiotic resistance on microfluidic platform was also successfully demonstrated (Kalashnikov et al. 2012). Bacteria were immobilized on the microfluidic channels, then subjected to mechanical stress generated by flow of media along with the enzymatic stress by bactericidal agent lysostaphin. Resistant bacteria were successfully identified on the basis of cell viability by SYTOX dye, in 60 min (Kalashnikov et al. 2012, 2014).
Critical issues of microfluidics AST methods
Despite all the beneficial properties of microfluidic AST that enabled its rapid and accurate detection over conventional technologies, there are several limitations that make it difficult to implement it in clinical biology. In general, the selection of materials is limited by the required material properties such as electrical conductivity or optical transparency depending on types of microfluidic system. The polydimethylsiloxane (PDMS) is used regularly for microfluidic fabrications due to its optical transparency and chemical inertness, however, its structural flexibility makes it difficult to directly integrate other parts including metal electrodes and fluidic interconnects. While alternative plastic materials for mass production using simple fabrication methods, e.g. injection molding, have been applied in several researches, geometric resolution and complexity are still limited. Simple and versatile manufacturing will be one of the key enabling factors for future commercialization, and thus collaborative efforts from fabrication, material science and microfluidic experts should be done for the development of manufacturer-friendly devices. The variation in physical parameters like oxygen, CO2, pH, osmolarity and water vapor due to the permeability of PDMS may disturb the internal environment of culturing device (Firpo et al. 2015). The most profound effect is the change of pH which either kills the bacteria or gives false positive results. Complex and expertise-dependent fabrication of a microfluidic platforms are another major drawback for bacterial observation.
Lack of portability is a hurdle for producing the user friendly handheld device. Portability of a device can be divided into three major divisions, sample preparation, providing reaction condition, and detection. Microfluidics does offer simple sample preparation or direct application of unprocessed samples by utilizing high specificity assays and reaction conditions. However, output signal collection remains a major problem, for example, complex imaging system involves fluorescent detection instruments like external light source, microscope or microplate reader. Although massive and sophisticated optical and electrical instruments deliver accurate detection, the demands of routine maintenance, laboratory setup and storage might be their major impediments for clinical applications. The certain setup provides inadequate morphological information, whereas most can only display the viability status of bacteria. It cannot provide the dose-dependent behavioral information of several life phases of bacteria. Currently, a very few reports are present on morphological pattern analysis of bacteria. These issues should be addressed with immediate effect.
Future perspective
Evidently, microfluidic tools are the future of AST, especially for point-of-care testing. Current microfluidic devices have addressed many complications of conventional methods up to some extent by offering many advantages regarding time and complexity for susceptibility test. However, the rise of multidrug-resistant and extensively drug-resistant strains will remain as a matter of great challenge for permanency of micro-technologies. These challenges are not limited to microfluidics only but also with conventional methods that always compelled the governing organization (CLSI and FDA) to come up periodically with updated guidelines. Rapid mutations have become a prime challenge in commercializing the microfluidic device for AST and finding new conventional methods. Therefore, consideration of both the conventional and microfluidic methods is vital for technological advancement. Besides, considering both the genotypic and phenotypic traits with the single device might be useful in controlling resistance. Consequently, rapid, inexpensive, simple, safe, and compatible test methods remain a foremost critical feature that may improve workflow and quality of microfluidic AST techniques. Incorporating these features in a precise manner will be helpful in developing potential technology into the diagnostic world.
Colorimetric methods have potential that could substantially improve imaging-based experiments by allowing visual detection with automatic settings that might be important for diagnosis in developing countries. There is scarcity on colorimetric microfluidic AST, and extensive research is warranted. Real-time microscopy integrated with microfluidics could be another promising way to circumvent the intricacies of the current imaging techniques. Furthermore, advances in ubiquitous smartphone and its high-resolution camera technology have transformed the medical research by providing user-friendly applications for bacterial visualization and image-based quantification. Hence, such modulation on microfluidic AST would offer the possibilities to compete with an ever-changing mutation, leading to resistance in a bacterium in more convenient and portable form. Unavailability of reports on the morphological analysis of bacteria for AST is a major constraint for the development of sensitive and reliable conventional or microfluidic devices. As the resistance is developed by the mutation in the genes, there is a possibility that same mutation might induce a change in the morphology or growth dynamics. This knowledge gap restricts our understanding towards resistance mechanism that could be overcome by focusing on the morphological analysis of bacteria. Therefore, a comparative morphological survey is required to generate the complete understanding of resistance mechanism. A combined approach utilizing both phenotypic and genotypic parameters can overcome the key obstacle of false positive results.
There are few FDA approved commercial automated systems available for susceptibility test but are extremely costly and require routine maintenance for optimum performance. Microfluidics has a potential to replace the costly and sophisticated instruments, but the field is still underdeveloped for commercialization point of view. During the last ten years, microfluidic AST has not achieved much success in transferring conventional technologies to the miniaturized platform for clinical practitioner. Therefore, enormous scope and opportunities are available for research and development in microfluidic AST that could be an essential platform to target new challenges and ideas for suppressing antibacterial resistance.
The predictions are indicating a massive enhancement in antibiotic resistance. The appearance of resistance to the new disease is a challenge, but the reemergence of old diseases is equally threatening. Thus, point-of-care testing will put us in commanding position for early detection of these mutants. Microfluidic tools offer us the possibility to miniaturize the established conventional, molecular, and automated techniques.
Various nanoparticles (gold, silver, europium, and platinum, etc.) will be utilized to enhance the sensitivity and accuracy of AST. Cumulatively, all the branches will develop more accurate and sensitive tools for AST, and microfluidics, along with their developments, will translate these into microtechnologies.
Conclusion
In this review, we have outlined the overview of AST that can help to expand and improve the existing techniques. We have highlighted merits and demerits of conventional, automated and microfluidic susceptibility testing tools to aid in current understanding and get an overview of the requirement in antibiotic susceptibility tests. Over the past decades, advances in microfluidics have enabled to analyze rapid bacterial behavior with minimal manual errors and interruptions. However, it has shown its shortcomings. Looking forward, customer-centric approach with microfluidics will be needed to meet above listed limitations.
Availability of data and materials
Data sharing not applicable to this article as no datasets were generated or analyzed during the current study.
References
Angulo FJ, Collignon P, Wegener HC, Braam P, Butler CD (2005) The routine use of antibiotics to promote animal growth does little to benefit protein undernutrition in the developing world. Clin Infect Dis 41(7):1007–1013
Ayukekbong JA, Ntemgwa M, Atabe AN (2017) The threat of antimicrobial resistance in developing countries: causes and control strategies. Antimicrob Resist Infect Control 6(1):47
Balouiri M, Sadiki M, Ibnsouda SK (2016) Methods for in vitro evaluating antimicrobial activity: a review. J Pharm Anal 6(2):71–79
Blair JMA, Webber MA, Baylay AJ, Ogbolu DO, Piddock LJV (2015) Molecular mechanisms of antibiotic resistance. Nat Rev Microbiol 13(1):42–51
Boedicker JQ, Li L, Kline TR, Ismagilov RF (2008) Detecting bacteria and determining their susceptibility to antibiotics by stochastic confinement in nanoliter droplets using plug-based microfluidics. Lab Chip 8(8):1265–1272
Chen CH, Lu Y, Sin MLY, Mach KE, Zhang DD, Gau V, Liao JC, Wong PK (2010) Rapid antimicrobial susceptibility testing using high surface-to-volume ratio microchannels. Anal Chem 82(3):1012
Choi J, Yoo J, Lee M, Kim E-G, Lee JS, Lee S, Joo S, Song SH, Kim E-C, Lee JC (2014) A rapid antimicrobial susceptibility test based on single-cell morphological analysis. Sci Transl Med 6(267):267
Cira NJ, Ho JY, Dueck ME, Weibel DB (2012) A self-loading microfluidic device for determining the minimum inhibitory concentration of antibiotics. Lab Chip 12(6):1052–1059
Dua V, Kunin CM, White LV (1994) The use of antimicrobial drugs in Nagpur, India. A window on medical care in a developing country. Soc Sci Med 38(5):717–724
Felmingham D, Brown DF (2001) Instrumentation in antimicrobial susceptibility testing. J Antimicrob Chemother 48(suppl 1):81–85
Firpo G, Angeli E, Repetto L, Valbusa U (2015) Permeability thickness dependence of polydimethylsiloxane (PDMS) membranes. J Membr Sci 481:1–8
Golchin SA, Stratford J, Curry RJ, McFadden J (2012) A microfluidic system for long-term time-lapse microscopy studies of mycobacteria. Tuberculosis 92(6):489–496
Halford C, Gonzalez R, Campuzano S, Hu B, Babbitt JT, Liu J, Wang J, Churchill BM, Haake DA (2013) Rapid antimicrobial susceptibility testing by sensitive detection of precursor rRNA using a novel electrochemical biosensing platform. Antimicrob Agents Chemother 57(2):936–943
Hou Z, An Y, Hjort K, Hjort K, Sandegren L, Wu Z (2014a) Time lapse investigation of antibiotic susceptibility using a microfluidic linear gradient 3D culture device. Lab Chip 14(17):3409–3418
Hou Z, An Y, Hjort K, Hjort K, Sandegren L, Wu Z (2014b) Time lapse investigation of antibiotic susceptibility using a microfluidic linear gradient 3D culture device. Lab Chip 14(17):3409–3418
Jenkins SG, Schuetz AN (2012) Current concepts in laboratory testing to guide antimicrobial therapy. In: Mayo Clinic proceedings. Elsevier, pp 290–308
Kalashnikov M, Lee JC, Campbell J, Sharon A, Sauer-Budge AF (2012) A microfluidic platform for rapid, stress-induced antibiotic susceptibility testing of Staphylococcus aureus. Lab Chip 12(21):4523–4532
Kalashnikov M, Campbell J, Lee JC, Sharon A, Sauer-Budge AF (2014) Stress-induced antibiotic susceptibility testing on a chip. J Vis Exp 83:50282
Karlowsky JA, Richter SS (2015) Antimicrobial susceptibility testing systems*, manual of clinical microbiology, 11th edn. American Society of Microbiology, Washington, DC
Kinnunen P, Sinn I, McNaughton BH, Newton DW, Burns MA, Kopelman R (2011) Monitoring the growth and drug susceptibility of individual bacteria using asynchronous magnetic bead rotation sensors. Biosens Bioelectron 26(5):2751–2755
Kinnunen P, McNaughton BH, Albertson T, Sinn I, Mofakham S, Elbez R, Newton DW, Hunt A, Kopelman R (2012) Self-assembled magnetic bead biosensor for measuring bacterial growth and antimicrobial susceptibility testing. Small 8(16):2477–2482
Lansang MA, Lucas-Aquino R, Tupasi TE, Mina VS, Salazar LS, Juban N, Limjoco TT, Nisperos LE, Kunin CM (1990) Purchase of antibiotics without prescription in manila, the philippines. Inappropriate choices and doses. J clin epidemiol 43(1):61–67
Lee W-B, Fu C-Y, Chang W-H, You H-L, Wang C-H, Lee MS, Lee G-B (2017) A microfluidic device for antimicrobial susceptibility testing based on a broth dilution method. Biosens Bioelectron 87:669–678
Lehtinen J, Nuutila J, Lilius EM (2004) Green fluorescent protein–propidium iodide (GFP-PI) based assay for flow cytometric measurement of bacterial viability. Cytometry part A 60(2):165–172
Li Y, Burke DT, Kopelman R, Burns MA (2014) Asynchronous magnetic bead rotation (AMBR) microviscometer for label-free DNA analysis. Biosensors 4(1):76–89
Liu C, Zeng G-M, Tang L, Zhang Y, Li Y-P, Liu Y-Y, Li Z, Wu M-S, Luo J (2011) Electrochemical detection of Pseudomonas aeruginosa 16S rRNA using a biosensor based on immobilized stem–loop structured probe. Enzyme Microb Technol 49(3):266–271
Lu Y, Wong PK (2011) Single cell antimicrobial susceptibility testing using confined microchannels and electrokinetic loading. In: 15th International conference on miniaturized systems for chemistry and life sciences 2011, MicroTAS 2011
Lu Y, Gao J, Zhang DD, Gau V, Liao JC, Wong PK (2013) Single cell antimicrobial susceptibility testing by confined microchannels and electrokinetic loading. Anal Chem 85(8):3971–3976
Matsumoto Y, Sakakihara S, Grushnikov A, Kikuchi K, Noji H, Yamaguchi A, Iino R, Yagi Y, Nishino K (2016) A microfluidic channel method for rapid drug-susceptibility testing of Pseudomonas aeruginosa. PLoS ONE 11(2):e0148797
Mohan R, Mukherjee A, Sevgen SE, Sanpitakseree C, Lee J, Schroeder CM, Kenis PJA (2013a) A multiplexed microfluidic platform for rapid antibiotic susceptibility testing. Biosens Bioelectron 49:118–125
Mohan R, Mukherjee A, Sevgen SE, Sanpitakseree C, Lee J, Schroeder CM, Kenis PJA (2013b) A multiplexed microfluidic platform for rapid antibiotic susceptibility testing. Biosens Bioelectron 49:118–125
Mohan R, Sanpitakseree C, Desai AV, Sevgen SE, Schroeder CM, Kenis PJA (2015) A microfluidic approach to study the effect of bacterial interactions on antimicrobial susceptibility in polymicrobial cultures. RSC Advances 5(44):35211–35223
Nordmann P, Dortet L, Poirel L (2012) Rapid detection of extended-spectrum-β-lactamase-producing enterobacteriaceae. J Clin Microbiol 50(9):3016–3022
Obara B, Roberts MAJ, Armitage JP, Grau V (2013) Bacterial cell identification in differential interference contrast microscopy images. BMC Bioinform 14(1):134
Park S, Zhang Y, Lin S, Wang TH, Yang S (2011) Advances in microfluidic PCR for point-of-care infectious disease diagnostics. Biotechnol Adv 29(6):830–839
Peitz I, van Leeuwen R (2010) Single-cell bacteria growth monitoring by automated DEP-facilitated image analysis. Lab Chip 10(21):2944–2951
Petersen A, Andersen JS, Kaewmak T, Somsiri T, Dalsgaard A (2002) Impact of integrated fish farming on antimicrobial resistance in a pond environment. Appl Environ Microbiol 68(12):6036–6042
Pulido MR, García-Quintanilla M, Martín-Peña R, Cisneros JM, McConnell MJ (2013) Progress on the development of rapid methods for antimicrobial susceptibility testing. J Antimicrob Chemother 68(12):2710–2717
Rackus DG, Shamsi MH, Wheeler AR (2015) Electrochemistry, biosensors and microfluidics: a convergence of fields. Chem Soc Rev 44(15):5320–5340
Ramezan Ali A, Ali Mehrabi T, Mohammad Javad HS, Khadijeh M, Mahdi Ghorbananli Z (2012) A method for antibiotic susceptibility testing: applicable and accurate. Jundishapur J Microbiol 2012:341–345
Rodloff A, Bauer T, Ewig S, Kujath P, Müller E (2008) Susceptible, intermediate, and resistant—the intensity of antibiotic action. Deutsches Ärzteblatt Int 105(39):657–662
Saleh N, Awada S, Awwad R, Jibai S, Arfoul C, Zaiter L, Dib W, Salameh P (2015) Evaluation of antibiotic prescription in the Lebanese community: a pilot study. Infect Ecol Epidemiol. 5:27094. https://doi.org/10.3402/iee.v5.27094
Sinn I, Kinnunen P, Albertson T, McNaughton BH, Newton DW, Burns MA, Kopelman R (2011) Asynchronous magnetic bead rotation (AMBR) biosensor in microfluidic droplets for rapid bacterial growth and susceptibility measurements. Lab Chip 11(15):2604–2611
Sinn I, Albertson T, Kinnunen P, Breslauer DN, McNaughton BH, Burns MA, Kopelman R (2012) Asynchronous magnetic bead rotation microviscometer for rapid, sensitive, and label-free studies of bacterial growth and drug sensitivity. Anal Chem 84(12):5250–5256
Skafte-Pedersen P, Hemmingsen M, Sabourin D, Blaga FS, Bruus H, Dufva M (2012) A self-contained, programmable microfluidic cell culture system with real-time microscopy access. Biomed Microdevice 14(2):385–399
Tang Y, Zhen L, Liu J, Wu J (2013) Rapid antibiotic susceptibility testing in a microfluidic pH sensor. Anal Chem 85(5):2787–2794
Van Boeckel TP, Gandra S, Ashok A, Caudron Q, Grenfell BT, Levin SA, Laxminarayan R (2014) Global antibiotic consumption 2000 to 2010: an analysis of national pharmaceutical sales data. Lancet Infect Dis 14(8):742–750
Ventola CL (2015) The antibiotic resistance crisis: part 1: causes and threats. Pharm Ther 40(4):277–283
Webster TA, Sismaet HJ, Chan IPJ, Goluch ED (2015) Electrochemically monitoring the antibiotic susceptibility of Pseudomonas aeruginosa biofilms. Analyst 140(21):7195–7201
Xue Y, Wang M, Zhao P, Quan C, Li X, Wang L, Gao W, Li J, Zu X, Fu D, Feng S, Li P (2018) Gram-negative bacilli-derived peptide antibiotics developed since 2000. Biotechnol Lett 40:1271–1287
Funding
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (NRF-2015R1C1A1A01054762).
Author information
Authors and Affiliations
Contributions
ZK, MS, and SP wrote and edited the manuscript. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Khan, Z.A., Siddiqui, M.F. & Park, S. Progress in antibiotic susceptibility tests: a comparative review with special emphasis on microfluidic methods. Biotechnol Lett 41, 221–230 (2019). https://doi.org/10.1007/s10529-018-02638-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-018-02638-2