Abstract
Human health is threatened by the spread of antimicrobial resistance and resulting infections. One reason for the resistance spread is the treatment with inappropriate and ineffective antibiotics because standard antimicrobial susceptibility testing methods are time-consuming and laborious. To reduce the antimicrobial susceptibility detection time, minimize treatments with empirical broad-spectrum antibiotics, and thereby combat the further spread of antimicrobial resistance, faster and point-of-care methods are needed. This requires many different research approaches. Microfluidic systems for antimicrobial susceptibility testing offer the possibility to reduce the detection time, as small sample and reagent volumes can be used and the detection of single cells is possible. In some cases, the aim is to use human samples without pretreatment or pre-cultivation. This chapter first provides an overview of conventional detection methods. It then presents the potential of and various current approaches in microfluidics. The focus is on microfluidic methods for phenotypic antimicrobial susceptibility testing.
Graphical Abstract

Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Antibiotic resistance test
- Antibiotic susceptibility tests (AST)
- Antimicrobial resistance
- Microfluidics
- Point-of-care systems
1 Introduction
According to the WHO, the spread of antimicrobial-resistant (AMR) infections is a leading global threat to human health. Today, about 700,000 people worldwide die annually from infections with multi-resistant pathogens [1]. Current forecasts predict ten million deaths worldwide in 2050 if the current situation is not changed [1]. This number would exceed the number of cancer-related deaths. The spread of AMR threatens the ability of modern medicine to use well-established procedures such as complex surgery, organ transplants, or chemotherapy in its fight against disease [2]. As an example for the spread of AMR, Fig. 1 shows the increase in antibiotic resistance of the most common sepsis-causing bacteria in Europe in the past few years.
Escherichia coli resistant to third-generation cephalosporins across Europe in (a) 2009 and (b) 2019 (adapted from [3])
One reason for the emergence of multi-resistant bacteria strains is the unnecessary use of antibiotics, as it is a common practice in animal fattening [4]. The increase in AMR can also be attributed to the long detection times and the resultant early non-specific therapy typical in the treatment of humans infected with such pathogens. Conventional detection methods require about 48–72 h after sample collection until antibiotic resistance is confirmed [5, 6]. However, the mortality risk of infected people increases the longer the patients do not receive appropriate therapy (7% per hour) [5]. For this reason, empirical broad-spectrum antibiotic treatments are already used in cases of suspected infection with AMR pathogens, especially in sepsis [7,8,9]. In such cases, this problem is exacerbated by the use of reserve antibiotics in treatment, although they may not be strictly necessary or may not have the desired effects. As a consequence, these reserve antibiotics may also lose their effectiveness.
Thus, the challenge is to extend access to antibiotics while at the same time limiting inappropriate use, in particular of broad-spectrum antibiotics and reserve antibiotics [10]. Rapid antibiotic susceptibility tests (AST) can be essential for the correct and economically efficient use of antibiotics. Besides preventing AMR development by adequate drug prescription, rapid resistance diagnostics can help to initiate appropriate antibiotic treatment promptly. Thus, rapid AST improves therapy outcomes and saves lives [11].
2 Antimicrobial Susceptibility Testing (AST)
The methods currently available for AST can be divided into genotypic and phenotypic methods [4]. Figure 2 shows an overview of phenotypic and genotypic AST methods and their assay times. Genotypic methods examine the genetic nature of the bacteria. However, one limitation of these methods is that they cannot detect unknown resistance genes [12]. For known genotypes associated with antibiotic resistance, the result is available after about 90 min. Since only known mechanisms of resistance can be proven, this may lead to false-negative results and thus to the wrong medication for the patient. When testing uncultured blood, false-positive results can be caused by even the slightest contamination of the sample if unrelated DNA is detected [13]. The disadvantages of genotypic tests make phenotypic assays more suitable for AST. These culture-based methods, such as broth microdilution or disk diffusion, are the gold standard AST methods.
Phenotypic methods are used to investigate the growth behavior of bacterial strains in the presence of different antibiotics [4]. This method also allows a direct assessment of whether an antibiotic has stopped bacterial growth. Thus, a decision can be made about the optimal therapeutic measures. For conventional phenotypic AST, the patient sample has to be isolated and then pre-cultured, which takes between 24 and 48 h [14, 15]. In the case of broth microdilution, the growth of the bacterial culture in the presence of various antibiotic concentrations is determined by measuring the optical density (OD) [16]. This phenotypic method can also be used to determine the minimum antibiotic concentration which prevents bacterial growth, otherwise referred to as minimum inhibitory concentration (MIC) [16]. False-positive results of the phenotypic AST may be caused when, for example, bacteria form thread-like structures in the presence of antibiotics without dividing [14]. A further disadvantage of this phenotypic method is the low sensitivity of about 107 colony-forming units (CFU) per mL [16]. In addition, phenotypic methods require inoculum sizes of ~5 × 105 CFU/ml [17]. The associated lead time for cultivation is another disadvantage [14]. The advantage, however, is that resistances can be determined directly and without prior knowledge of the resistance mechanisms.
3 Microfluidic AST
New phenotypic AST based on microfluidics have been developed in recent years to shorten the detection time and increase sensitivity [20]. With microfluidic techniques, the assay time of AST can be reduced to 1–3 h [21].
Microfluidic systems have the advantage of [4, 16, 22]:
-
small sample and reagent volumes,
-
detection of single cells,
-
combining several sample processing steps,
-
potentially accelerating biochemical reactions.
Thus, microfluidic systems can achieve high sensitivity and allow for automation and high-throughput analysis [23]. The use of small volumes at the single-cell level can also prevent cross-contamination [16].
As a basis for this novel AST, standard principles used in microfluidics were developed further. Microfluidic channel systems are therefore widely used [24]. Different detection methods are used to measure physiological or biochemical changes during or after bacterial growth [22]. In the following, selected microfluidic AST methods are described and classified according to their detection principles.
3.1 Optical Detection
3.1.1 Single-Cell Imaging
Single-cell imaging describes a group of microscopy techniques that allow imaging and single-cell detection. It is used to study cell dynamics and usually based on fluorescent molecules. In the case of AST, single-cell imaging is used to monitor bacterial growth with different antibiotic treatments. For this purpose, the bacterial cells are immobilized in various microfluidic systems. [21, 25, 26].
Baltekin et al. use the mother-machine design, a multichannel system widely used in microfluidics to test for urinary tract infections (shown in Fig. 3a). The microfluidic chip is made of a PDMS base structured by soft lithography with microchannels and closed with a cover glass bonded to the PDMS [21]. The system consists of a feeding channel through which the bacteria are first loaded, and which then ensures the culture medium supply. Several parallel growth channels branch off from this feeding channel. During loading, bacteria enter the growth channels. These so-called mother cells are where the subsequent bacterial growth originates. A sample with a low bacteria content of 104 CFU/ml can be used for loading (shown in Fig. 3b) [21]. The culture medium supply is achieved via a 300 nm gap [21]. The growth in each microchannel is detected separately by taking phase-contrast images using a microscope [21]. The average growth rate is calculated across all channels. Antimicrobial resistance can be detected by determining the growth rate after treatment with different antibiotics. With this system, the detection time of bacterial growth including loading can be reduced to 30 min [21].
Microchannel system for immobilization (a) Schematic illustration of the channel system (red: captured cells); (b) Bacterial loading of different density cell cultures at different points in time; (c) Phase contrast image of E.coli (lighter regions) in the system using a 20× objective (adapted with permission from [21])
A similar system was developed by Li et al. [27], which also permits the sorting of bacteria of different sizes from polymicrobial clinical samples by applying different pressures and incubating them separately. Lu et al. also use a microchannel system to immobilize the bacteria. It consists of 68 parallel channels with one inlet and one outlet. By integrating two microelectrodes in the PDMS system, the bacteria can be positioned in the microchannels. A sample with a bacterial content of 105 CFU/ml can be used for loading [28].
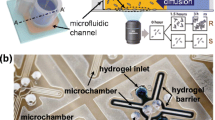
Choi et al. use a microfluidic agarose channel system for immobilization and bacterial cell incubation. The microfluidic chip consists of a centered inlet for cell loading and six channels running outwards in a star arrangement for bacterial incubation. The microfluidic chip is composed of a PDMS base structured by soft lithography with microchannels and closed with a cover PDMS coated by bonding. When loaded through the inlet, an agarose-bacteria solution is distributed evenly over all six channels and cures there [24]. At the six side-branched channels, different antibiotics or antibiotic concentrations can be added to the culture medium. By monitoring the individual bacterial growth and calculating the area occupied by bacteria in a microfluidic channel, antimicrobial susceptibility can be determined [24]. An automatic analysis can be performed using image-based single-cell morphological analysis (SCMA). Here, morphological changes in individual bacterial cells are automatically analyzed and categorized under different antimicrobial conditions [29]. Thus, antimicrobial sensitivity can be determined by SCMA in less than 4 h [29].
3.1.2 Time-Lapse Microscopy
Using a combination of a liquid bacterial culture and agarose, alternative microfluidic approaches can achieve an assay time between 2.5 and 4 h [30,31,32]. Here, bacterial density is determined by grayscale intensity changes in the images (from black to white) resulting from bacterial growth [30].
Kim et al. use a microfluidic system of two parallel PDMS channels sealed with a cover glass (shown in Fig. 4) [32]. An agarose-bacteria solution is fed in between the two parallel microchannels (see Fig. 4a). The medium flows through one channel and medium, the antibiotic through the other. Due to the different antibiotic concentrations of the two parallel channels, the antibiotic molecules diffuse into the agarose-bacteria solution (shown in Fig. 4b). A concentration gradient forms within 30 min. As a result of treatment under antibiotic gradients, changes in local bacterial growth can be observed [32]. Dose-effect diagrams can be reconstructed from the gray intensity changes in the images (see Fig. 4c). 2 × 108 CFU/ml was used. The changes in local bacterial growth were determined after 6 h.
Microchannel system with agarose-bacteria solution: (a) Schematic illustration of antibiotic treatment using diffusion process; (b) Experimental process using the microfluidic chip: (1) Agarose-bacteria solution is introduced. (2) The antibiotic concentration gradient is formed (30 min). (3) Changes in local bacteria growth are observed; (c) Analyzing gray intensity changes of images along with linearized antibiotic concentrations (reproduced from [32], with permission of AIP Publishing)
3.1.3 Interferometry
Other approaches use interferometry to measure bacterial growth [33, 34]. The approach of Busche et al. is shown in Fig. 5a. Here an optofluidic microchip is used, which is fabricated by etching nanochannels into a thin silicon oxide layer and then closed by anodic bonding of a structured glass. The nanochannels, connected by a nanogap, create an optical grating (shown in Fig. 5b). This combines a nanofluidic bacterial cell trap with an optical asymmetric grating [34]. The channel system consists of detection channels and reference channels, which can be pressurized and supplied with bacteria and a culture medium via bypass channels. The bacteria are first loaded into the detection channels and immobilized at the nanogap by bypass cross-flow [35]. Then culture medium flows through all channels. The refractive index in the detection channels is changed by bacterial growth. Using a collimated laser beam, a change in the intensity distribution can be measured, as shown in Fig. 5c [36]. By treating the bacteria with and without antibiotics, the effect of the antibiotic on bacterial growth can be determined (shown in Fig. 5d).
Optofluidic chip for bacterial growth detection. Schematic illustrations: (a) Detection method; (b) Microchannel system as an optical grating (green: detection channels, yellow: reference channels; red: collimated laser beam); (c) Intensity distribution during bacterial growth; (d) Diffraction signal during bacterial growth with and without antibiotics (adapted from [34])
3.1.4 Fluorescence Imaging
Fluorescence imaging is the visualization of fluorescence dyes or proteins as markers for molecular processes or structures [4]. Using a green fluorescent protein (GFP) requires genetic modification of the bacteria, so this detection method is not suitable for the analysis of clinical samples [38, 39]. GFP is suitable for real-time observation of bacterial growth due to the high correlation between fluorescence intensity and cell density.
Mohan et al. have developed a channel/chamber system made from PDMS that is filled using negative pressure (shown in Fig. 6a) [37]. One antibiotic channel and one bacterial channel lead to a set of eight chambers (2.4 nl each), as shown in Fig. 6b [39]. When all chambers are filled, a mixing valve is used to achieve a homogeneous mixture within the culture chambers (shown in Fig. 6c, d). The bacterial count can be determined by fluorescence imaging [37], which requires the bacteria used to express GFP. This system is suitable as a multiplex microfluidic system since parallel tests can be carried out with high throughput. In the research work presented in this paragraph, a 48-well array was used, in which 12 different antibiotics or antibiotic concentrations can be tested in parallel. This takes less time than a conventional phenotypic test, as there is no need for pre-cultivation [37]. For the MIC determination of polymicrobial samples, initial cell counts of ~100 to 300 cells were set [39]. Bacterial growth was observed over 16 h. A decision regarding antibiotic susceptibility including the determination of MIC could be made after 2–4 h [39].
Multiplex microfluidic AST system with bacteria immobilization in nanoliter arrays: (a) Entire array with inlet ports for bacteria and antibiotics and vacuum ports for filling using negative pressure; (b) Close-up of the 48 wells; each well can be loaded with bacteria concentration (red) or antibiotics (green); (c) Set of eight chambers during filling; (d) Set of eight chambers during mixing (reproduced with permission from [39])
In droplet microfluidics, discrete microdroplets (volumes from pL to mL) are generated and analyzed in an immiscible phase [40]. Each microdroplet is separated by a liquid–liquid interface, usually stabilized by amphiphilic surfactants.
In AST, the droplets serve as incubators for the encapsulated bacterial cells, allowing both antibiotics and viability dyes to be added. The fluorescent dye resazurin is used to demonstrate the viability of the bacteria treated with antibiotics [41,42,43]. It is a metabolism marker. Resazurin reacts irreversibly to resofurin, which has strong fluorescent properties, through the cellular reaction potential [42]. This reaction occurs at a rate that is proportionate to the aerobic respiration of the cells in a bacterial culture [42].
Derzsi et al. have developed a passive-dilution platform generating droplets using five different pipettes. They measured the fluorescence intensity of resofurin after a 4 h droplet incubation. With different concentrations of antibiotics, they were able to determine the MIC [44].
3.1.5 Relative Optical Density
Liu et al. use high-throughput screening of antibiotic-resistant bacteria in picodroplets [45]. The microfluidic system is based on PDMS structured with microchannels using soft lithography [45]. The detection is carried out using a system for light scattering without bacteria labeling [45]. This method is used to study the relative optical density of droplets. It allows distinguishing between droplets with growth and those with inhibited growth/resistance. Bacterial concentrations of 105 CFU/ml have been measured. In other approaches, the droplet microfluidics was used to treat bacteria with combinations of antibiotics [40, 47].
Another approach combines optical density measurement with hydrodynamic immobilization in traps [46]. The system consists of cup-shaped structure arrangements that are used as mechanical barriers to capture bacteria flowing in a microchannel. The structure dimensions are designed to trap exactly one bacterial cell. Based on this, the bacterial growth rate under the influence of antibiotics can be determined at a bacterial concentration of 107 CFU/ml [46].
3.2 Electrical Detection
3.2.1 Measuring the Electrical Resistance Change
Yang et al. [48] use a microfluidic channel system as reported in [21, 27] to capture and incubate the bacteria in microchannels (2 μm × 2 μm with a gap of 800 nm) [48]. In contrast to the systems mentioned earlier, the detection is completely electrical. As the bacteria grow, the electrical resistance in the channel increases while the electrical current decreases. The change in the bacteria population is proportionate to the electrical resistance. Thus, bacterial growth can be tested directly with antibiotic treatment. With this microfluidic system, small urine samples can be tested without pretreatment [48].
3.2.2 Electrochemical Detection
Besant et al. use a system of 2.5 nl wells in which microbeads with a diameter of 5 μm are immobilized in stacks and thus serve as a bacterial filter [49]. For detection, an electrochemical approach is used. By adding resazurin, a shrinking of a redox-active molecule is reached, which is directly related to the number of metabolically active bacteria. Antibiotic-resistant bacteria reduce resazurin to resorufin, while antibiotic-sensitive bacteria do not reduce it [49]. The two molecular states can be distinguished by measurement using electrodes integrated into the well. A minimum cell concentration of 105 CFU/ml can be used [49]. In this phenotypic approach, growth is measured over 100 min in the presence of an antibiotic [49].
3.3 Biochemical Detection
One common marker to measure the metabolic activities of bacteria is the adenosine triphosphate (ATP). Using this marker, the efficient emission photons (550–570 nm; of which the luciferin substrate is catalyzed by luciferase in the presence of ATP) and oxygen is measured [50]. This measurement process requires sensitive microplate readers, able to capture the photons. The detection method using ATP is known as ATP bioluminescence.
Dong et al. have developed a multilayer system consisting of a culture layer with 384 reaction chambers and culture medium veins made of polystyrene; a membrane layer made of fiberglass membrane filter; and a sample layer with sample and air veins made of polystyrene (shown in Fig. 7a). The system is covered by two glass layers. On the reaction chamber side, the fiberglass membrane filter is coated with antibodies for bacteria immobilization. Figure 7b shows the standard process for using this system. The metabolism of a bacterial concentration generates an ATP bioluminescence signal. The antimicrobial effect of an antibiotic can be determined directly from a urine sample after 3–6 h if the bacterial concentration is above 1,000 CFU/ml [51].
Microfluidic AST using ATP bioluminescence for detection and antibody binding for bacteria immobilization: (a) Multilayer system design; (b) Schematic illustration of the standard AST process: (1) the urine sample is loaded into the sample veins, and the bacteria cells are captured by antibodies; (2) unbound bacterial cells are removed; (3) the bound cells are encapsulated by calcium alginate gel; (4) the alginate solution is washed away; (5) paraffin oil is introduced to isolate each reaction chamber; the bound cells (6.0) can then be reproduced, (6.2) inhibited by series of antibiotics, or (6.1) inhibited by a single antibiotic; (7) the cells are quantified using ATP bioluminescence in a microplate reader. (Adapted with permission from [51]. Copyright 2015 American Chemical Society)
Dong et al. also use antibody binding to membranes for label-free optical detection [52]. However, antibodies are not available for all bacteria.
Further biochemical approaches use Raman spectroscopy of biomarkers [53, 54], DNA samples [55,56,57], or RNA markers [58, 59]. These are genotypic tests.
3.4 Mechanical Detection
Mechanical detection methods are based, for example, on a mechanical cantilever signal to determine antimicrobial susceptibility. For immobilization, the cantilever surface is functionalized with bacteria-specific receptors such as antibodies. The binding results in either a cantilever deflection or a resonance frequency change [60,61,62].
Other cantilever-based systems use cantilevers with embedded microchannels, also known as suspended microchannel resonators [62, 63]. Etayash et al. use a bi-material cantilever with a 50-pL volume microchannel, which can provide three signals (adsorbed mass, adsorption stress, and mid-infrared spectroscopy of the adsorbates; shown in Fig. 8a, b, e) [62]. For the immobilization of the bacteria, the microfluidic channel surfaces are functionalized with chemical or physical receptors capturing the bacteria selectively, as shown in Fig. 8c, d. The bacteria accumulation within the microchannel leads to a change in the cantilever resonance frequency and a cantilever deflection (shown in Fig. 8f, g), where the cantilever resonance frequency is caused by the absorbed mass and the cantilever deflection is caused by the adsorption stress. Additional irradiation with infrared light stimulates the captured bacteria and causes deflection proportionate to the infrared absorption of the bacteria. Thus, a nanomechanical infrared spectrum can be generated for selective identification of dead and living bacteria. Bacteria that are treated with antibiotics show a significant nanomechanical reaction [62]. Cell concentrations between 102 and 105 CFU/ml were tested [62]. In initial experiments, when captured E. coli were treated with antibiotics and exposed, a deflection of the cantilever and a shift in resonance frequency were observed after 30 min.
Bi-material cantilever with embedded microchannel: (a) Schematic illustration of the system. The cantilever is made of silicon nitride coated with gold, which is fixed on a silicon substrate. The microchannel surface is functionalized with specific receptors. Additionally, the cantilever is irradiated with infrared light. (b) Cross-section of the inlet at the bottom of the system using scanning electron microscopy. (c) Cross-section of the cantilever with the embedded microfluidic channel. (d) Fluorescent image of the microchannel with immobilized bacteria. (e) Scanning electron microscopy image of the cantilever tip. (f) Nanomechanical cantilever deflection resulting from infrared absorption of the bacteria. (g) Change in the resonance frequency due to adsorption of bacteria in the microchannel. (h) The wavelength at which the bacteria absorb infrared light depends on the nanomechanical cantilever deflection resulting from irradiation with a certain range of infrared light. (Reprinted with permission from [62])
4 Conclusion
This chapter provides a review of different research approaches for microfluidic AST, which are mainly used to detect antibiotic susceptibility based on bacterial cultures (phenotypic). Basic microfluidic concepts were further developed to provide suitable solutions for microfluidic AST. The different microfluidic systems found in the literature can be classified by their detection methods: optical, electrical, biochemical, and mechanical. Table 1 lists the detection and immobilization methods covered in this chapter.
In all microfluidic systems mentioned, the bacterial cells are physically trapped, and their mobility is thus restricted. The nutrients/culture medium required for growth are supplied via channels or by diffusion.
It seems preferable to use systems that allow on-chip cultivation, as it requires no time-consuming pre-cultivation. However, one fundamental difficulty is the cell loading at very low bacterial concentrations, which can impede the accumulation of a sufficient number of bacterial cells. A solution for loading uncultivated and therefore low-concentration samples into a microfluidic system for analysis is an on-chip filtration of the bacteria in small volumes to increase the concentration.
The systems used for optical detection are transparent and often consist of PDMS or PDMS and glass. While PDMS are easy to fabricate and air permeable, they cannot be used to test small MIC values of antibiotics, as small molecules and drugs are absorbed during testing [64,65,66].
Some detection methods seem complex and expensive, so they are only partially suitable for use as AST. This is particularly the case with some optical detection methods. Another issue arises when using dead-end systems, which tend to clog.
In conclusion, several microfluidic AST have been shown to provide faster results than conventional AST (<7 h) and thus support clinical decisions on the correct use of antibiotics. The analysis of morphology and the number of single cells by minimizing the incubation area significantly improve the resolution. Reducing the analysis size, in turn reducing the number of samples and reagents used, allows the integration of several parallel tests on one microfluidic device. Due to their size and the intended ease of use, microfluidic approaches for novel AST can contribute to the development of point-of-care systems. However, steps still need to be taken to develop microfluidic AST, to provide a cost-effective, portable, accurate, and time-efficient device.
Microfluidic devices can also simplify and accelerate sample pretreatment (cleaning and isolation) for the subsequent AST. Besides the use as microfluidic AST, microfluidic devices can also be used for antibiotic development or research on the mechanisms of AMR development [45].
References
World Health Organization (2016) United Nations meeting on antimicrobial resistance. Bull World Health Organ 94(9):638–639. https://doi.org/10.2471/BLT.16.020916
Friedman ND, Temkin E, Carmeli Y (2016) The negative impact of antibiotic resistance. Clin Microbiol Infect 22(5):416–422. https://doi.org/10.1016/j.cmi.2015.12.002
European Centre for Disease Prevention and Control (2017) Surveillance atlas of infectious diseases. https://www.ecdc.europa.eu/en/surveillance-atlas-infectious-diseases. Accessed 3 Nov 2020
Khan ZA, Siddiqui MF, Park S (2019) Progress in antibiotic susceptibility tests: a comparative review with special emphasis on microfluidic methods. Biotechnol Lett 41(2):221–230. https://doi.org/10.1007/s10529-018-02638-2
Stürenburg E, Junker R (2009) Point-of-care testing in microbiology: the advantages and disadvantages of immunochromatographic test strips. Dtsch Arztebl Int 106(4):48–54. https://doi.org/10.3238/arztebl.2009.0048
Mach KE, Mohan R, Baron EJ, Shih M-C, Gau V, Wong PK, Liao JC (2011) A biosensor platform for rapid antimicrobial susceptibility testing directly from clinical samples. J Urol 185(1):148–153. https://doi.org/10.1016/j.juro.2010.09.022
Ferrer R, Martin-Loeches I, Phillips G, Osborn TM, Townsend S, Dellinger RP, Artigas A, Schorr C, Levy MM (2014) Empiric antibiotic treatment reduces mortality in severe sepsis and septic shock from the first hour: results from a guideline-based performance improvement program. Crit Care Med 42(8):1749–1755. https://doi.org/10.1097/CCM.0000000000000330
Karam G, Chastre J, Wilcox MH, Vincent J-L (2016) Antibiotic strategies in the era of multidrug resistance. Crit Care 20(1):136. https://doi.org/10.1186/s13054-016-1320-7
Kollef MH (2008) Broad-spectrum antimicrobials and the treatment of serious bacterial infections: getting it right up front. Clin Infect Dis 47(Suppl 1):S3–S13. https://doi.org/10.1086/590061
Laxminarayan R, Matsoso P, Pant S, Brower C, Røttingen J-A, Klugman K, Davies S (2016) Access to effective antimicrobials: a worldwide challenge. Lancet 387(10014):168–175. https://doi.org/10.1016/S0140-6736(15)00474-2
Bhattacharyya RP, Bandyopadhyay N, Ma P, Son SS, Liu J, He LL, Wu L, Khafizov R, Boykin R, Cerqueira GC, Pironti A, Rudy RF, Patel MM, Yang R, Skerry J, Nazarian E, Musser KA, Taylor J, Pierce VM, Earl AM, Cosimi LA, Shoresh N, Beechem J, Livny J, Hung DT (2019) Simultaneous detection of genotype and phenotype enables rapid and accurate antibiotic susceptibility determination. Nat Med 25(12):1858–1864. https://doi.org/10.1038/s41591-019-0650-9
Hou HW, Bhattacharyya RP, Hung DT, Han J (2015) Direct detection and drug-resistance profiling of bacteremias using inertial microfluidics. Lab Chip 15(10):2297–2307. https://doi.org/10.1039/c5lc00311c
Khan ZA, Siddiqui MF, Park S (2019) Current and emerging methods of antibiotic susceptibility testing. Diagnostics (Basel) 9(2). https://doi.org/10.3390/diagnostics9020049
Murray C, Adeyiga O, Owsley K, Di Carlo D (2015) Research highlights: microfluidic analysis of antimicrobial susceptibility. Lab Chip 15(5):1226–1229. https://doi.org/10.1039/c5lc90017d
Garcia-Prats JA, Cooper TR, Schneider VF, Stager CE, Hansen TN (2000) Rapid detection of microorganisms in blood cultures of newborn infants utilizing an automated blood culture system. Pediatrics 105(3 Pt 1):523–527. https://doi.org/10.1542/peds.105.3.523
Zhang K, Qin S, Wu S, Liang Y, Li J (2020) Microfluidic systems for rapid antibiotic susceptibility tests (ASTs) at the single-cell level. Chem Sci 11(25):6352–6361. https://doi.org/10.1039/D0SC01353F
Smith KP, Kirby JE (2018) The inoculum effect in the era of multidrug resistance: minor differences in inoculum have dramatic effect on MIC determination. Antimicrob Agents Chemother 62(8). https://doi.org/10.1128/AAC.00433-18
van Belkum A, Burnham C-AD, Rossen JWA, Mallard F, Rochas O, Dunne WM (2020) Innovative and rapid antimicrobial susceptibility testing systems. Nat Rev Microbiol 18(5):299–311. https://doi.org/10.1038/s41579-020-0327-x
Vasala A, Hytönen VP, Laitinen OH (2020) Modern tools for rapid diagnostics of antimicrobial resistance. Front Cell Infect Microbiol 10:308. https://doi.org/10.3389/fcimb.2020.00308
Mishra P, Mishra KP, Singh D, Ganju L, Kumar B, Singh SB (2018) Advances in rapid detection and antimicrobial susceptibility tests: a review. Def Life Sci J 4(1):12–20. https://doi.org/10.14429/dlsj.4.12572
Baltekin Ö, Boucharin A, Tano E, Andersson DI, Elf J (2017) Antibiotic susceptibility testing in less than 30 min using direct single-cell imaging. Proc Natl Acad Sci U S A 114(34):9170–9175. https://doi.org/10.1073/pnas.1708558114
S-u H, Zhang X (2020) Microfluidics as an emerging platform for tackling antimicrobial resistance (AMR): a review. Curr Anal Chem 16(1):41–51. https://doi.org/10.2174/1573411015666181224145845
Lee W-B, Fu C-Y, Chang W-H, You H-L, Wang C-H, Lee MS, Lee G-B (2017) A microfluidic device for antimicrobial susceptibility testing based on a broth dilution method. Biosens Bioelectron 87:669–678. https://doi.org/10.1016/j.bios.2016.09.008
Choi J, Jung Y-G, Kim J, Kim S, Jung Y, Na H, Kwon S (2013) Rapid antibiotic susceptibility testing by tracking single cell growth in a microfluidic agarose channel system. Lab Chip 13(2):280–287. https://doi.org/10.1039/c2lc41055a
Hiratsuka T, Komatsu N (2019) Single-cell live imaging. Methods Mol Biol 1979:409–421. https://doi.org/10.1007/978-1-4939-9240-9_24
Mullassery D, Horton CA, Wood CD, White MRH (2008) Single live-cell imaging for systems biology. Essays Biochem 45:121–133. https://doi.org/10.1042/BSE0450121
Li H, Torab P, Mach KE, Surrette C, England MR, Craft DW, Thomas NJ, Liao JC, Puleo C, Wong PK (2019) Adaptable microfluidic system for single-cell pathogen classification and antimicrobial susceptibility testing. Proc Natl Acad Sci U S A 116(21):10270–10279. https://doi.org/10.1073/pnas.1819569116
Lu Y, Gao J, Zhang DD, Gau V, Liao JC, Wong PK (2013) Single cell antimicrobial susceptibility testing by confined microchannels and electrokinetic loading. Anal Chem 85(8):3971–3976. https://doi.org/10.1021/ac4004248
Choi J, Yoo J, Lee M, Kim E-G, Lee JS, Lee S, Joo S, Song SH, Kim E-C, Lee JC, Kim HC, Jung Y-G, Kwon S (2014) A rapid antimicrobial susceptibility test based on single-cell morphological analysis. Sci Transl Med 6(267):267ra174. https://doi.org/10.1126/scitranslmed.3009650
Hou Z, An Y, Hjort K, Hjort K, Sandegren L, Wu Z (2014) Time lapse investigation of antibiotic susceptibility using a microfluidic linear gradient 3D culture device. Lab Chip 14(17):3409–3418. https://doi.org/10.1039/C4LC00451E
Li B, Qiu Y, Glidle A, McIlvenna D, Luo Q, Cooper J, Shi H-C, Yin H (2014) Gradient microfluidics enables rapid bacterial growth inhibition testing. Anal Chem 86(6):3131–3137. https://doi.org/10.1021/ac5001306
Kim S, Lee S, Kim J-K, Chung HJ, Jeon JS (2019) Microfluidic-based observation of local bacterial density under antimicrobial concentration gradient for rapid antibiotic susceptibility testing. Biomicrofluidics 13(1):14108. https://doi.org/10.1063/1.5066558
Leonard H, Halachmi S, Ben-Dov N, Nativ O, Segal E (2017) Unraveling antimicrobial susceptibility of bacterial networks on micropillar architectures using intrinsic phase-shift spectroscopy. ACS Nano 11(6):6167–6177. https://doi.org/10.1021/acsnano.7b02217
Busche JF, Möller S, Klein A-K, Stehr M, Purr F, Bassu M, Burg TP, Dietzel A (2020) Nanofluidic immobilization and growth detection of Escherichia coli in a Chip for antibiotic susceptibility testing. Biosensors (Basel) 10(10). https://doi.org/10.3390/bios10100135
Busche JF, Möller S, Stehr M, Dietzel A (2019) Cross-flow filtration of Escherichia coli at a Nanofluidic gap for fast immobilization and antibiotic susceptibility testing. Micromachines (Basel) 10(10). https://doi.org/10.3390/mi10100691
Purr F, Bassu M, Lowe RD, Thürmann B, Dietzel A, Burg TP (2017) Asymmetric nanofluidic grating detector for differential refractive index measurement and biosensing. Lab Chip 17(24):4265–4272. https://doi.org/10.1039/c7lc00929a
Mohan R, Mukherjee A, Sevgen SE, Sanpitakseree C, Lee J, Schroeder CM, Kenis PJA (2013) A multiplexed microfluidic platform for rapid antibiotic susceptibility testing. Biosens Bioelectron 49:118–125. https://doi.org/10.1016/j.bios.2013.04.046
Kim S, Masum F, Jeon JS (2019) Recent developments of Chip-based phenotypic antibiotic susceptibility testing. Biochip J 13(1):43–52. https://doi.org/10.1007/s13206-019-3109-7
Mohan R, Sanpitakseree C, Desai AV, Sevgen SE, Schroeder CM, Kenis PJA (2015) A microfluidic approach to study the effect of bacterial interactions on antimicrobial susceptibility in polymicrobial cultures. RSC Adv 5(44):35211–35223. https://doi.org/10.1039/C5RA04092B
Kaminski TS, Scheler O, Garstecki P (2016) Droplet microfluidics for microbiology: techniques, applications and challenges. Lab Chip 16(12):2168–2187. https://doi.org/10.1039/c6lc00367b
Boedicker JQ, Li L, Kline TR, Ismagilov RF (2008) Detecting bacteria and determining their susceptibility to antibiotics by stochastic confinement in nanoliter droplets using plug-based microfluidics. Lab Chip 8(8):1265–1272. https://doi.org/10.1039/b804911d
Avesar J, Rosenfeld D, Truman-Rosentsvit M, Ben-Arye T, Geffen Y, Bercovici M, Levenberg S (2017) Rapid phenotypic antimicrobial susceptibility testing using nanoliter arrays. Proc Natl Acad Sci U S A 114(29):E5787–E5795. https://doi.org/10.1073/pnas.1703736114
Churski K, Kaminski TS, Jakiela S, Kamysz W, Baranska-Rybak W, Weibel DB, Garstecki P (2012) Rapid screening of antibiotic toxicity in an automated microdroplet system. Lab Chip 12(9):1629–1637. https://doi.org/10.1039/c2lc21284f
Derzsi L, Kaminski TS, Garstecki P (2016) Antibiograms in five pipetting steps: precise dilution assays in sub-microliter volumes with a conventional pipette. Lab Chip 16(5):893–901. https://doi.org/10.1039/C5LC01151E
Liu X, Painter RE, Enesa K, Holmes D, Whyte G, Garlisi CG, Monsma FJ, Rehak M, Craig FF, Smith CA (2016) High-throughput screening of antibiotic-resistant bacteria in picodroplets. Lab Chip 16(9):1636–1643. https://doi.org/10.1039/c6lc00180g
Pitruzzello G, Thorpe S, Johnson S, Evans A, Gadêlha H, Krauss TF (2019) Multiparameter antibiotic resistance detection based on hydrodynamic trapping of individual E. coli. Lab Chip 19(8):1417–1426. https://doi.org/10.1039/C8LC01397G
Shang L, Cheng Y, Zhao Y (2017) Emerging droplet microfluidics. Chem Rev 117(12):7964–8040. https://doi.org/10.1021/acs.chemrev.6b00848
Yang Y, Gupta K, Ekinci KL (2020) All-electrical monitoring of bacterial antibiotic susceptibility in a microfluidic device. Proc Natl Acad Sci U S A 117(20):10639–10644. https://doi.org/10.1073/pnas.1922172117
Besant JD, Sargent EH, Kelley SO (2015) Rapid electrochemical phenotypic profiling of antibiotic-resistant bacteria. Lab Chip 15(13):2799–2807. https://doi.org/10.1039/c5lc00375j
Mirasoli M, Guardigli M, Michelini E, Roda A (2014) Recent advancements in chemical luminescence-based lab-on-chip and microfluidic platforms for bioanalysis. J Pharm Biomed Anal 87:36–52. https://doi.org/10.1016/j.jpba.2013.07.008
Dong T, Zhao X (2015) Rapid identification and susceptibility testing of uropathogenic microbes via immunosorbent ATP-bioluminescence assay on a microfluidic simulator for antibiotic therapy. Anal Chem 87(4):2410–2418. https://doi.org/10.1021/ac504428t
Rostova E, Ben Adiba C, Dietler G, Sekatskii SK (2016) Kinetics of antibody binding to membranes of living Bacteria measured by a photonic crystal-based biosensor. Biosensors (Basel) 6(4). https://doi.org/10.3390/bios6040052
Liu C-Y, Han Y-Y, Shih P-H, Lian W-N, Wang H-H, Lin C-H, Hsueh P-R, Wang J-K, Wang Y-L (2016) Rapid bacterial antibiotic susceptibility test based on simple surface-enhanced Raman spectroscopic biomarkers. Sci Rep 6:23375. https://doi.org/10.1038/srep23375
Han Y-Y, Lin Y-C, Cheng W-C, Lin Y-T, Teng L-J, Wang J-K, Wang Y-L (2020) Rapid antibiotic susceptibility testing of bacteria from patients' blood via assaying bacterial metabolic response with surface-enhanced Raman spectroscopy. Sci Rep 10(1):12538. https://doi.org/10.1038/s41598-020-68855-w
Rolain JM, Mallet MN, Fournier PE, Raoult D (2004) Real-time PCR for universal antibiotic susceptibility testing. J Antimicrob Chemother 54(2):538–541. https://doi.org/10.1093/jac/dkh324
Schoepp NG, Khorosheva EM, Schlappi TS, Curtis MS, Humphries RM, Hindler JA, Ismagilov RF (2016) Digital quantification of DNA replication and chromosome segregation enables determination of antimicrobial susceptibility after only 15 minutes of antibiotic exposure. Angew Chem Int Ed Engl 55(33):9557–9561. https://doi.org/10.1002/anie.201602763
Hindson CM, Chevillet JR, Briggs HA, Gallichotte EN, Ruf IK, Hindson BJ, Vessella RL, Tewari M (2013) Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat Methods 10(10):1003–1005. https://doi.org/10.1038/nmeth.2633
Khazaei T, Barlow JT, Schoepp NG, Ismagilov RF (2018) RNA markers enable phenotypic test of antibiotic susceptibility in Neisseria gonorrhoeae after 10 minutes of ciprofloxacin exposure. Sci Rep 8(1):11606. https://doi.org/10.1038/s41598-018-29707-w
Halford C, Gonzalez R, Campuzano S, Hu B, Babbitt JT, Liu J, Wang J, Churchill BM, Haake DA (2013) Rapid antimicrobial susceptibility testing by sensitive detection of precursor rRNA using a novel electrochemical biosensing platform. Antimicrob Agents Chemother 57(2):936–943. https://doi.org/10.1128/AAC.00615-12
Mader A, Gruber K, Castelli R, Hermann BA, Seeberger PH, Rädler JO, Leisner M (2012) Discrimination of Escherichia coli strains using glycan cantilever array sensors. Nano Lett 12(1):420–423. https://doi.org/10.1021/nl203736u
Wang J, Morton MJ, Elliott CT, Karoonuthaisiri N, Segatori L, Biswal SL (2014) Rapid detection of pathogenic bacteria and screening of phage-derived peptides using microcantilevers. Anal Chem 86(3):1671–1678. https://doi.org/10.1021/ac403437x
Etayash H, Khan MFR, Kaur K, Thundat T (2016) Microfluidic cantilever detects bacteria and measures their susceptibility to antibiotics in small confined volumes. Nat Commun 7:12947. https://doi.org/10.1038/ncomms12947
Godin M, Delgado FF, Son S, Grover WH, Bryan AK, Tzur A, Jorgensen P, Payer K, Grossman AD, Kirschner MW, Manalis SR (2010) Using buoyant mass to measure the growth of single cells. Nat Methods 7(5):387–390. https://doi.org/10.1038/nmeth.1452
van Meer BJ, de Vries H, Firth KSA, van Weerd J, Tertoolen LGJ, Karperien HBJ, Jonkheijm P, Denning C, IJzerman AP, Mummery CL (2017) Small molecule absorption by PDMS in the context of drug response bioassays. Biochem Biophys Res Commun 482(2):323–328. https://doi.org/10.1016/j.bbrc.2016.11.062
Toepke MW, Beebe DJ (2006) PDMS absorption of small molecules and consequences in microfluidic applications. Lab Chip 6(12):1484–1486. https://doi.org/10.1039/B612140C
Shirure VS, George SC (2017) Design considerations to minimize the impact of drug absorption in polymer-based organ-on-a-chip platforms. Lab Chip 17(4):681–690. https://doi.org/10.1039/C6LC01401A
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Klein, AK., Dietzel, A. (2021). Microfluidic Systems for Antimicrobial Susceptibility Testing. In: Bahnemann, J., Grünberger, A. (eds) Microfluidics in Biotechnology. Advances in Biochemical Engineering/Biotechnology, vol 179. Springer, Cham. https://doi.org/10.1007/10_2021_164
Download citation
DOI: https://doi.org/10.1007/10_2021_164
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-04187-7
Online ISBN: 978-3-031-04188-4
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)