Abstract
A simple yet effective dynamic bead-based microarray is necessary for multiplexed high-throughput screening applications in the fields of biology and chemistry. This paper introduces a microfluidic-based dynamic microbead array system using pneumatically driven elastomeric valves integrated with a microchannel in a single polydimethylsiloxane (PDMS) layer that performs the following functions: single-microbead arraying with loading and trapping efficiencies of 100 %, sequential microbead release for selective retrieval of microbeads of interest, and rapid microarray resettability (<1 s). The key feature is the utilization of an elastomeric membrane as a valve for trapping and releasing single microbeads; this membrane is deformable depending on the applied pneumatic pressure, thereby simply providing a dual trap-and-release function. We propose an effective single-microbead-trapping mechanism based on a dynamic flow-change network and a mathematical model as the design criterion of a trapping site. A sequential microbead release technique via a multistep “release-retrap-and-repeat” method was developed for the selective retrieval of trapped microbeads with a simple configuration consisting of a single PDMS layer and a simple macro-to-micro connection. The proposed dynamic microbead array could be a powerful tool for high-throughput multiplex bead-based drug screening or disease diagnosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Microarray-based techniques have been developed to meet the demand for multiplexed high-throughput screening for the investigation of numerous biomolecules, chemicals, and the interactions among them. Depending on the sample arraying method, microarrays can be categorized into two types: static and dynamic. With advancements in particle synthesis technology, the dynamic microarray composed of heterogeneous microcarriers such as microbeads with surfaces that are coated with various molecules or their compounds (e.g., amino acids, peptide) is emerging as a powerful tool for biomedical applications such as drug screening and disease diagnosis (Nolan and Mandy 2006; Skelley et al. 2009); in contrast, static microarrays consist of samples patterned on a static solid support (Robinson et al. 2002; Steinert et al. 2009; Malainou et al. 2012). Because of their large surface-to-volume ratio, microbeads have a higher capacity to bind molecules, thereby improving the detection limit and reaction time compared to static arrays with planar surfaces (Verpoorte 2003). In addition, they can easily be transported, mixed, and separated for various purposes (e.g., multiplexed assays).
However, to fully exploit its advantages, a dynamic microarray system should offer the following essential functions: (1) high trapping efficiency of single microbeads as well as high loading efficiency to observe the binding specificity of surface molecules of each microbead in real time and over the long term under continuous flow conditions; (2) selective retrieval to identify molecules of interest on the surfaces of microbeads or additional uses of these microbeads; and (3) microarray resettability to allow the reuse of a device and efficient data collection by reducing device fabrication costs (e.g., time and labor costs).
Several research groups have reported microfluidic-based dynamic microarray systems that are capable of handling small volumes of liquids and that can therefore accurately manipulate microsized particles and reduce sample and reagent consumption. Microfluidic-based dynamic microarrays can be classified as those using active and passive methods depending on the types of applied forces. Active methods, including electric, magnetic, and acoustic types, require a medium (i.e., a microelectrode) to generate the force while allowing precise manipulation (i.e., trap-and-release) of individual microparticles. In addition, the connection pads, which link the microelectrodes with the external power systems, should be integrated into the chip, and they will become larger numbers as the number of manipulated microparticles increases (Voldman et al. 2002; Zhu et al. 2012). Therefore, in active methods, chip design and fabrication are complex procedures (Choi et al. 2000; Wang and Zhe 2011). Furthermore, electric and magnetic methods strongly rely on the type (i.e., electromagnetic properties) of the particle that can be used.
In passive methods, microparticle-trapping mechanisms are based on dynamic changes in the flow field before and after particle trapping, and therefore, they require no external control systems except for a fluid delivery system. When semipermeable structures are used as microparticle traps, the trapping stream is diverted to prevent the additional inflow of sequential particles once a trapping site is filled with a single particle (Carlo et al. 2006). In this manner, infused microparticles can be immobilized in designated trapping sites for convenient detection under continuous flow conditions. For further investigation (e.g., off-chip cell lysis or peptide identification) of microparticles of interest, a combined method using a laser-induced microbubble technique was developed to realize a device with a selective release function, with the trapping of single microparticles achieved using the passive method (Tan and Takeuchi 2007, 2008). In the microbubble technique, the connection pads do not need to be integrated because energy is directly supplied from a laser to each microelectrode (i.e., to generate a microbubble); however, simple device fabrication by using only the PDMS replica molding that is generally adopted in the passive method is not possible, because an additional patterned electrode layer is required. To overcome this disadvantage and the impractical resetting process (i.e., release of one microbead at a time), the same research group integrated pneumatically driven valves instead of the electrode into the previous microarray device (Iwai and Takeuchi 2009). By adequately controlling the hydraulic resistance toward a trap depending on the ON/OFF state of the valve, selective trapping and releasing of microbeads can be realized. However, this system requires multiple sample injection processes and a number of valves to control each trap individually for multiplexed screening applications (i.e., to manipulate different types of microbeads). Recently, some research groups have proposed resettable dynamic microarrays to rapidly retrieve a large number of microparticles with an extremely simple device configuration and operation (Sochol et al. 2010, 2012; Iwai et al. 2011). These systems utilize the asymmetrical property of the flow field depending on the flow direction. In the normal direction, introduced microbeads were trapped, and when the flow was reversed, trapped microbeads were released. However, this system offered no selective retrieval function, and thus, an additional device is required for particle sorting (e.g., a flow-cytometry-based sorting device).
In this regard, we have previously proposed a microfluidic-based hydrodynamic trap-and-release technique for single microparticles that utilizes the deformable characteristics of an elastomeric membrane integrated in a single layer with a microchannel (Kim et al. 2012). However, this technique must be expanded to a microarray format for practical applications such as high-throughput screening. Although some research groups have introduced dynamic microarrays using the deformable elastomeric membrane as a removable trap, they have focused on manipulating microparticles at the group level (Shao et al. 2011; Fu et al. 2011; Jin et al. 2011) or on clustering different types of single microbeads for particle–particle interaction study (Tonooka et al. 2012). As mentioned previously, a dynamic microarray system that can perform these essential functions without the aid of the electrode layer has not yet been realized.
In this study, we introduce a dynamic microarray system with pneumatically driven elastomeric valves to integrate the essential functions into a single chip, while retaining the advantages (i.e., simple micro-to-macro connection and replica molding) of the passive method. First, to realize a microbead array with high loading (i.e., to minimize the loss of infused microbeads) and trapping efficiencies, we used a mathematical model that depends on the microparticle size to develop a trapping mechanism in order to effectively utilize the dynamic change in hydraulic resistance after microbead trapping. Next, a resettable function was realized by restoring the original state (i.e., no deflection) of the deformed valve so that it could be used as a microbead trap without the need for flow direction tuning or the application of additional force. Finally, to selectively retrieve the microbeads of interest, we introduced the first sequential release technique for trapped microbeads using a mechanism that retrapped the released microbeads by continuously switching between the ON and OFF states of the valve.
2 Concept and design criterion
2.1 Dynamic single-microbead array with pneumatic valves
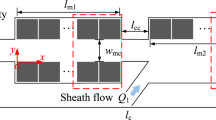
Figure 1a shows a schematic view of the microfluidic traps for trapping and selectively retrieving single microbeads using pneumatic valves. It consists of three modes: trapping, releasing, and retrapping. In each trap, single microbeads are trapped based on the dynamic change in the hydraulic resistance (i.e., before and after microbead trapping) as a function of the cross-sectional shape of a microchannel. Briefly, depending on the relationship between the center position of a microbead and the flow pattern [i.e., a virtual width (W v)], the path of the introduced microbeads is determined (Yamada et al. 2007). First, to regulate the position of microbeads and to ensure the trapping of single microbeads, all introduced microbeads are aligned along one side wall using the buffer flow. In addition, according to the design configuration, the alignment of microbeads is maintained in the main channel through the symmetrical flow of each trapping site under low-Reynolds-number flows with negligible inertial force. Next, when the microbead radius exceeds the virtual width of an empty trap, the microbead (Fig. 1b, red) flows along the trapping stream and toward the trap. Once the trap is filled, a subsequent microbead bypasses the occupied trap because its virtual width is larger than the microbead radius owing to the increased hydraulic resistance toward the trap, and it enters the next empty trap. In this manner, traps are sequentially filled with single microbeads. The release and retrapping modes allow us to perform resettable and selective retrieval functions using pneumatically driven elastomeric valves with one external valve controller (i.e., to create a simple macro-to-micro connection), as shown in Fig. 1c, d. In the releasing mode, microbeads can be released easily by switching the valve off, making our dynamic microarray resettable. By switching the valve on again after a short time, the released microbeads can be retrapped in the next trap, allowing the sequential release of trapped microbeads. This feature facilitates the selective retrieval of the microbeads of interest by switching the valve a certain number of times depending on the initial location of the microbeads in the microarray.
Schematic top views of a portion of a dynamic microarray device integrated with pneumatic valves and with three operation modes. a Introduced microbeads are aligned along one side wall using the buffer flow, and then, microbeads are immobilized sequentially in each trap depending on the relationship between the virtual width (W v) and the microbead radius. b Trapping mode: once a trap under the “ON” valve state is filled with a single microbead (yellow), the virtual width (long and short dashed lines) increases to bypass the subsequent microbead (red) due to an increase in the hydraulic resistance toward the trap. Hence, the subsequent microbead (red) is trapped at the next empty trap. c, d Releasing and retrapping mode: by switching off the pneumatic valve, trapped microbeads can easily be released and then retrapped at the next trap if the valves are switched on again after a short time. In this manner, trapped microbeads can be released sequentially toward an outlet by controlling the number of times the valve is switched (color figure online)
2.2 Theoretical analysis
To efficiently determine the design dimensions depending on the size of an applied microbead without experimental trial and error and thus reducing the time and labor costs, we developed a mathematical model that describes the relationship between the virtual width and the design dimensions with the following assumptions: (1) the flow is pressure-driven, steady, and laminar; (2) the fluid is an incompressible Newtonian one; and (3) the velocity profile is a two-dimensional parabola, as follows:
where μ is the fluid viscosity, dp/dx is the pressure gradient between channel ends, W M is the main channel width, and 0 ≤ y ≤ W M [note that the velocity profile would be assumed to be a two-dimensional parabola if the aspect ratio of the channel height (H) to width (W M) is larger than ~1] (Lee et al. 2006).
Depending on our single-microbead-trapping mechanism, the virtual width (W v), which denotes the width of the virtual stream entering the bypass channel, should be satisfied by the following two inequations (see Fig. 2):
where W v,b and W v,a are virtual widths before and after single-microbead trapping, respectively, and r is the microbead radius.
Based on the law of mass conservation, the flow rate of the virtual stream in the main channel should be the same as that (Q B) into the bypass channel. The virtual width can then be expressed as follows:
where Q B is the flow rate into the bypass channel and Q M, the total flow rate [details of these calculations are given in Online Resource 1 eqns. (S1)–(S3)].
According to the Hagen–Poiseuille law, which is analogous to Ohm’s law, the fluidic circuit can be interpreted as an electric circuit. The ratio Q B/Q M can then be replaced by the ratio of hydraulic resistances, R T/(R T + R B), where R T and R B are the hydraulic resistances of the trap and the bypass channel, respectively. By substituting inequation (2) into Eq. (3), the ratio R T /(R T + R B ) should be satisfied with the following two inequations:
where subscripts b and a denote states before and after single-microbead trapping, respectively. Here, the hydraulic resistance can be defined by the design dimensions as a function of the cross-sectional shape of the microchannel. The hydraulic resistance of the bypass channel of a rectangular shape is given by
where f Re is the laminar friction constant, L B is the bypass channel length, D h is the hydraulic diameter, and A B is the channel cross-sectional area [two definitions that correspond to f Re and D h are given in Online Resource 1 eqns. (S4) and (S5), respectively]. Next, the hydraulic resistance of a trap of an arbitrary shape is given by
where L T is the trap length, P T is the perimeter of the trap, and A T is the cross-sectional area of the trap. To calculate the perimeter and the cross-sectional area of the trap, the deflection curve of the pneumatic valve should be defined. In our previous study, we assumed that the valve deflection curve follows a double sine function.18 The validation of that assumption was demonstrated experimentally [the definition of the valve deflection curve is given in Online Resource 1 eqns. (S7) and (S8)].
By substituting Eqs. (5) and (6) into inequation (4), we can obtain a mathematical expression as the design criterion for achieving single-microbead trapping. To satisfy inequation (4) under certain values of the microbead radius (r) and channel width (W M), the ratio R T/(R T + R B) can be varied by the bypass channel length (L B) because the other geometrical variables [i.e., f Re, D h, A, and P, which are functions of the channel width and height, and the trap length (L T)] depend on an applied microbead size (i.e., to effectively alter the width of the virtual stream according to the microbead trapping). For a microbead with a diameter of 21 μm under a valve central deflection (D V,c) of 10 μm, the bypass channel length (L B) should be ~300–1500 μm, as follows:
Table 1 lists the design dimensions and theoretical analysis results according to different bypass channel lengths. Four different devices were designed to demonstrate the validation of the proposed mathematical model. In addition, from Eq. (6), R T (i.e., P T, A T ) could be different based on the application of pneumatic pressure as an operation parameter, leading to valve deflection. According to the valve central deflection (D V,c), the results of the theoretical analysis of the virtual width are given in Online Resource 1 Table S1.
3 Experimental
3.1 Device fabrication
The dynamic microarray device was fabricated using a standard soft lithography technique (Duffy et al. 1998). The device is composed of a glass slide and a single PDMS (Sylgard 184, Dow Corning Corp.) layer in which the microchannels are connected to two inlets (one for supplying a microbead suspension and the other for buffer flow) and an outlet, a pressure port (for supplying a pneumatic pressure into the pneumatic channel), and pneumatic valves (see Online Resource 1 Fig. S1).
The PDMS layer was fabricated via replica molding, and the mold was prepared by the photolithographic patterning of a negative photoresist (KMPR 1035, Microchem. Corp.). To characterize the effect of the PDMS mixture ratio and baking time as process parameters on the pneumatic valve deflection, 10:1, 12:1, and 14:1 mixtures of PDMS prepolymer and curing agent were cured on a hot plate for 10 min at 100 °C, and three 12:1 PDMS mixtures were baked for 10, 30, and 60 min, respectively. Cured PDMS slabs were punched for the macro-to-micro connection and bonded with the glass slide after O2 plasma treatment of 45 W for 35 s under 500 mtorr O2 pressure.
3.2 Preparation of a polystyrene suspension
A polystyrene suspension (Φ = 20.9 ± 0.6 μm, coefficient of variation (CV) = 6.7 %, Thermal Fisher Scientific Inc.) mixed with phosphate-buffered saline (PBS) solution containing 0.4 % (v/v) of Tween 20 (Sigma–Aldrich) was used in this study. Tween 20 was added to minimize the adhesion force between the PDMS channel walls (including the pneumatic valves) and the polystyrene beads in order to allow the easy release of the trapped microbeads. The polystyrene bead concentration was 2 × 106 beads per ml. A PBS solution containing 0.4 % (v/v) of Tween 20 was used as the buffer fluid.
3.3 Experimental setup
The experimental setup consists of the dynamic microarray device, a fluid delivery system, and an external solenoid valve system to precisely control the pneumatic valve (see Online Resource 1 Fig. S1). Two syringe pumps (KD Scientific Inc.) were used to supply the polystyrene suspension and the buffer fluid into the device, respectively. The external solenoid valve system is composed of a pneumatic pressure supply and control parts. The pneumatic pressure was supplied using a diaphragm pump (Hargraves Technology Corp.), an electro-pneumatic regulator (ITV series, SMC Corp.), and a pressure monitor. To electrically switch the pressure applied to the pneumatic channel to either positive or ambient, a solenoid valve (LHD series, Lee products Ltd.) and LabView software (National Instruments) with a data acquisition device (NI DAQ USB-6251) were used. All experimental images and movies were obtained using a color CCD camera (WV-CP244, Panasonic Corp.) or a high-speed CCD camera (NR4-S3, Integrated Design Tools Inc.) mounted on an upright microscope (Mitutoyo Corp.).
4 Results and discussion
4.1 Pneumatic valve deflection according to applied pressure with different PDMS mixture ratios and baking times
The pneumatic valve deflection can be altered by varying the design dimensions (i.e., valve thickness, length, and width), operation conditions (i.e., amplitude of applied pneumatic pressure), and fabrication conditions (e.g., PDMS mixture ratio and baking temperature and time). Owing to our trapping mechanism, the design dimensions such as the valve length and width are dominated by the applied microbead size.
Therefore, to estimate the valve flexibility, devices made with different PDMS mixture ratios and baking times were tested. Figure 3a shows the degree of valve deflection as the applied pressure is increased, leading to the reduction in the gap distance (W G) of the trap. The gap distance must be less than the applied microbead diameter (i.e., ~21 μm) to be used as a trap; otherwise, all introduced microbeads would pass through. Figure 3b shows the effect of the mixture ratio on the valve deflection. As the mixture ratio is increased from 10:1 to 14:1, resulting in a reduction in the PDMS’s Young’s modulus (Fuard et al. 2008), the gap distance decreases under the same value of the applied pressure. In particular, the gap distance varies dramatically according to the applied pressure when the ratio is 14:1. Depending on our theoretical analysis, microbeads will flow along the bypassing stream if the gap distance becomes less than~14.5 μm in the case of device C with L B of 500 μm because the virtual width would be larger than the microbead radius. Hence, the gap distance should be ~14.5–21 μm.
Effects of valve pressurization on the valve deflection [i.e., gap distance of the trap (W G)] with different PDMS mixture ratios and baking times. a As the pneumatic pressure applied to the valve is increased, the gap distance of the trap decreases (with a PDMS mixture ratio of 12:1 and baking time of 10 min). This shows that to work as a trap, a pressure greater than a certain value should be applied against a microbead of a given size. b Effect of a PDMS mixture ratio with a constant PDMS baking time of 10 min on the gap distance; the valve stiffness varies with the PDMS mixture ratio (10:1, 12:1, and 14:1). The dotted line indicates the microbead diameter line (i.e., 21 μm). The dashed line indicates the minimum value (14.5 μm) required for microbead trapping based on the theoretical analysis result with respect to device C (i.e., having bypass channel length (LB) of 500 μm); otherwise, all introduced microbeads will bypass all traps. c Effect of the PDMS baking time with a constant PDMS mixture ratio of 12:1 on the gap distance: as the baking time is increased (10, 30, and 60 min), the gap distance against the applied pressure increases owing to increased valve stiffness caused by the additional polymerization of a PDMS prepolymer. The gap distance against the applied pressure was measured using the measure tool in Adobe PhotoShop CS5. Each data point is obtained from 10 valves. Error bars (i.e., standard deviation) are included in data points
To characterize the effect of PDMS baking time on the gap distance, three devices with a constant mixture ratio of 12:1, which were cured for 10, 30, and 60 min at 100 °C, were examined. It is well known that as the baking time is increased, the PDMS prepolymer can become additionally cross-linked, thereby becoming progressively stiffer. Figure 3c shows the results of this measurement. As the baking time is decreased, more data points are included in the trapping section, providing a wide range of operation conditions for trapping the microbead [note that a baking time of 10 min was the minimum time required for polymerization of the PDMS prepolymer].
Thus, all devices used for microbead-trapping and retrapping experiments were manufactured with a PDMS mixture ratio of 12:1 and baked for 10 min.
4.2 Trapping test of single microbeads with four different devices
Four microarray devices (devices A–D) with different L B values were tested to demonstrate our proposed trapping mechanism and to validate the mathematical model. To align the introduced microbeads along the side wall, both sample and buffer solutions were infused at a rate of 50 μL/h during all trapping experiments. Two superimposed high-speed images, shown in Fig. 4a, b, illustrate the bypassing and trapping modes using devices A and B, respectively.
Superimposed high-speed images and a single-microbead array image demonstrating microbead-trapping experiments. At a frame rate of 1000 fps, each trapping process (a–c) was captured and the cut images were created from two frames. Sample and buffer flow rates were 50 μL/h. a Bypassing mode: a superimposed high-speed image shows the bypass mode using device A without the application of pneumatic pressure. b Trapping mode: the trapping process of a single microbead using device B with an application of 150 kPa pressure is demonstrated, and it is observed that the microbead alignment was maintained at all positions in front of each trapping site. c Under an applied pressure of 210 kPa, a relatively small microbead bypassed traps, although the traps were empty. In empty traps, the distance between each microbead track, corresponding to two frames, decreases. This shows the change in dynamic flow according to microbead trapping. d Using device B under 210 kPa pressure, a single-microbead array with a trapping efficiency of 100 % proves the effectiveness of our trapping mechanism (N = 60)
According to the results of our theoretical analysis, in device A, the virtual width would be ~12.8 μm with no valve deflection [note that the virtual width increases with valve deflection]. Based on inequation (2), the introduced microbeads never go to the trap, although a portion of the trapping stream is larger than that of the bypassing stream because the microbeads are aligned to the side wall. Figure 4b shows the successful trapping of a single microbead under an applied pressure of 150 kPa using the device B. During the bypassing of each microbead-filled trap, the microbead alignment in the main channel was always maintained by the trapping stream functioning as the buffer flow, thereby preventing additional inflows of subsequent microbeads. Moreover, even in the absence of buffer flow, other traps aside from the first trap were occupied by single microbeads owing to the self-alignment of microbeads by the trapping stream. In device B, the trapping of the relatively small microbeads that arise from the polydispersity of the microbeads occasionally failed (Fig. 4c). From inequation (7), the experimental result corroborates and therefore validates the mathematical model, because device B with an L B of 300 μm would be under bypassing or trapping modes by the size variation of applied microbeads. The differences in the distances (i.e., velocity) of the microbead tracks according to the trapped microbead, as shown in Fig. 4c, demonstrate the dynamic change of the flow field.
The trapping efficiency of single microbeads was checked using an array of 60 traps. Our dynamic microarray with pneumatic valves provided a trapping efficiency of 100 % without vacancies, as shown in Fig. 4d. Single microbeads were carried sequentially to each trap from an upstream position (see Online Resource 2). Indeed, microbead clogging was not observed in the microchannel. This high performance results from the following two features: (1) geometry-induced self-alignment of microbeads and (2) consistency among flow patterns of each trapping site. Therefore, the position of the microbeads and the virtual width as the main parameters with respect to single-microbead trapping are under the same condition in each trapping site.
The loading and trapping efficiencies were evaluated in order to quantify the performance of our dynamic microarray device by observing the trapping process. Table 2 lists the various modes (bypassing, passing, sliding, and trapping) and loading/trapping efficiencies of the four different devices according to the applied pressure. As mentioned above, device A was operated only in the bypassing mode regardless of the amplitude of the applied pressure (0–210 kPa), whereas the other devices have three modes. Under pressures of 0 and 60 kPa, the pneumatic valves did not work as traps because the gap distances are larger than the microbead diameter, and therefore, all infused microbeads were thrown to the outlet along the main channel. When pressures of 90 and 120 kPa were applied, microbeads occasionally slipped on traps and were captured at the next trap that was filled with a single microbead, resulting in multiple trapping or the clogging of the microchannel. This may be caused by a number of factors, including the size distribution of microbeads, a slight difference in the deflections of each valve, and distortion of the valves caused by the pushing of the microbeads. At pressures of 150, 180, and 210 kPa, the microarrays were composed of a single microbead per trap without vacancies. Moreover, the infused microbeads were sequentially loaded without any loss of microbeads in the case of devices C and D (note that device B showed less loading efficiency owing to the size variation of microbeads, which was mentioned previously). Online Resources 3 and 4 show the sequential trapping process with trapping and loading efficiencies of 100 % for devices C and D. It appears that 60 traps are slightly inadequate for quantitative evaluation; however, obtaining many trapping experiment results using the 60 traps under various experimental conditions can provide a reasonable demonstration of high loading/trapping performances.
4.3 Sequential release function in retrapping mode
A sequential release function in a retrapping mode is key for selectively retrieving trapped microbeads using just one pneumatic controller (for example, for a simple macro-to-micro connection) by taking advantage of the deformation of pneumatic valves, because the individual control of each valve is very inefficient in an array owing to the increased number of external pressure controllers and pressure ports depending on the number of trapping sites.
Retrapping of released microbeads at later traps can be achieved by precisely controlling the ON/OFF states and the opening time of pneumatic valves depending on the main flow rate (Q M). High-speed images, as shown in Fig. 5a–e, reveal the sequential retrapping processes in device B. Here, the main flow rate and the valve OFF duration were 10 μL/h and 20 ms, respectively. Indeed, the release of the trapped microbead occurred simultaneously with the valve being switched off, and the microbead was retrapped after an extremely short time at the next trap when the valve deflection was restored.
Sequential images of a retrapping process using device B with a main flow rate of 10 μL/h, applied pressure of 150 kPa, and OFF duration of 20 ms. a–e show a trapped microbead immediately before switching to the OFF state, being released during the OFF valve state, as positive pressure is applied, in the ON valve state, and when it is retrapped at the next trap by restoring the valve deflection
The data in Table 3 indicate the relationship between the valve OFF duration and the main flow rate for successful retrapping in an array format. As the flow rate was increased, thus increasing the velocity of the released microbeads, the valve OFF duration was decreased to prevent the released microbeads from passing by the next trap so that they would be retrapped at a later trap. Otherwise, we would lose location information for each microbead based on the switching number of the valve. Thus, successful retrapping of released microbeads should be achieved without defects in the array. Figure 6a–d shows microbeads shifting downstream depending on the switching number. We used high-speed photography to confirm that all of the released microbeads were retrapped at subsequent traps during continuous valve switching when the main flow rate and valve ON/OFF durations were 20 μL/h and 300/10 ms, respectively (see Online Resource 5). Indeed, two microbeads in blue circles, as shown in Fig. 6a, d, were identical, i.e., the microbead immobilized at the first trap moved to the last trap (i.e., 60th) after the valve was switched 59 times.
Sequential high-speed images of the sequential release of trapped microbeads in an array (N = 60). All trapped microbeads were released sequentially by continuously switching between ON and OFF states with constant durations of 300 (for ON state) and 10 (for OFF state) ms. Images were captured at 300 Hz under a main flow rate of 20 μL/h. Red arrows indicate empty traps. a The circle (blue) and the square (green) indicate microbeads trapped in the 1st and 60th traps, respectively. b Each microbead shifted one step (i.e., one trap) downstream with each valve switching. Indeed, only the microbead immobilized at the last trap (60th) was released. c During 30 switches, microbeads that were initially immobilized in traps 31–60 were released in reverse order (i.e., from the 60th to the 31st trap). Here, with each additional switch, the microbead initially trapped at the 30th trap can be released and then retrieved. In this manner, all microbeads can be selectively retrieved depending on their initial locations. d After 59 switches, the microbead in the last trapping site (i.e., indicated by the blue circle) was identical to the microbead indicated by the blue circle in a (color figure online)
We have demonstrated that microbeads of interest can be retrieved by controlling the valve switching number depending on the initial location of the microbeads. In addition, our microarray can easily be reset in a short time (less than approximately 1 s) when the valve is switched OFF (see Online Resource 6). Furthermore, by using this resettable function, unwanted particles (e.g., dust and microbead clogs in the sliding mode) can be removed easily (data not shown).
5 Conclusion
In this study, we have introduced and demonstrated a simple yet effective dynamic microarray integrated with pneumatic valves. In a single-layer PDMS chip, we realized the following three essential features: a single-microbead array with 100 % loading and trapping efficiencies, a sequential release function for selective retrieval of trapped microbeads, and device resettability. Hence, the proposed system overcomes the limited function of passive methods as well as the complexity of active systems.
For quantitative assays, the single-microbead array technique with high trapping efficiency could reduce the effort required to filter out defects from experimental results. Moreover, the sequential trapping method with high loading efficiency will be useful in working with rare or expensive microparticles.
Because our trapping method is velocity-independent, it can also be applied for trapping biological cells, which can be deformed by fluid shear stress. In our previous study, we confirmed the feasibility of trapping a single cell using a pneumatic valve. For quantitative single-cell analysis, establishing a single-cell microarray with high trapping efficiency is necessary as a vital step. However, doing so is a big challenge because biological cells have a wide size distribution even within an isogenic group (Frimat et al. 2011). Our experimental results (Fig. 4c) show that with our trapping mechanism, the trapping of microbeads was dependent on the microbead size; only microbeads larger than the virtual width were trapped, allowing the trapping stream to be diverted to prevent additional inflows of subsequent microbeads. In this regard, our trapping mechanism would allow a highly efficient single-cell microarray.
Furthermore, the proposed mathematical model for the microfluidic-based hydrodynamic trapping of a single microparticle can be utilized to determine design dimensions depending on the sizes and size distribution of the applied particles, thereby optimizing the design parameters and minimizing the time and labor costs incurred by an experimental trial-and-error or simulation approach. However, the minimum particle size that can be applied using our method would be limited to ~10 μm owing to the intrinsic characteristics of this contact-based trapping method (i.e., fabrication difficulty of the pneumatic valve with thinner membrane and non-specific binding problem based on scaling laws).
We have proved the availability of the sequential release technique for selectively retrieving microbeads using pneumatic valves operated with just one controller. Therefore, our proposed dynamic microarray with essential functions could be parallelized to include more trapping sites to permit its use in multiplexed high-throughput screening applications such as drug screening and disease diagnosis, while retaining a simple system configuration using just one pneumatic controller.
Furthermore, the sequential release technique using a controlled immobilization time may be utilized as a microfluidic-based conveyor belt for microbeads. For example, microbeads could be sequentially synthesized, immobilized, stimulated, observed, and retrieved on a single chip if each trapping site has various functions. In particular, the selective retrieval function would be useful in applications where detection events of the trapped beads have to be further investigated (e.g., to identify the sequence of a peptide on the surface of a microbead of interest in the application of one bead one compound combinatorial libraries). In this regard, we believe that the proposed integrated single-layer dynamic microarray device could be a powerful and useful tool for bead-based applications.
6 Supporting information available
Derivation of the relationship between the virtual width and the ratio Q B /Q M; a formula to calculate the hydraulic resistance of the trap; theoretical analysis results (Online Resource 1); and five video files including trapping mode (Online Resources 2–4), retrapping mode (Online Resource 5), and device resettability (Online Resource 6) in an array format.
References
Carlo DD, Wu LY, Lee LP (2006) Dynamic single cell culture array. Lab Chip 6:1445–1449
Choi JW, Ahn CH, Bhansali S, Henderson HT (2000) A new magnetic bead-based, filterless bio-separator with planar electromagnet surfaces for integrated bio-detection systems. Sens Actuators B 68:34–39
Duffy DC, McDonald JC, Schueller OJA, Whitesides GM (1998) Rapid prototyping of microfluidic systems in poly(dimethylsiloxane). Anal Chem 70:4974–4984
Frimat JP, Becker M, Chiang YY, Marggraf U, Janasek D, Hengstler JG, Franzke J, West J (2011) A microfluidic array with cellular valving for single cell co-culture. Lab Chip 11:231–237
Fu Z, Shao G, Wang J, Lu D, Wang W, Lin Y (2011) Microfabricated renewable beads-trapping/releasing flow cell for rapid antigen-antibody reaction in chemiluminescent immunoassay. Anal Chem 83:2685–2690
Fuard D, Chevolleau TT, Decossas S, Tracqui P, Schiavone P (2008) Optimization of poly-di-methyl-siloxane (PDMS) substrates for studying cellular adhesion and motility. Microelectron Eng 85:1289–1293
Iwai K and Takeuchi S (2009) A dynamic microarray with pneumatic valves for selective trapping and releasing of microbeads. Proceedings of IEEE 22nd International Conference on Micro Electro Mechanical Systems, Sorrento
Iwai K, Tan WH, Ishihara H, Takeuchi S (2011) A resettable dynamic microarray device. Biomed Microdevices 13:1089–1094
Jin HJ, Cho YH, Gu JM, Kim J, Oh YS (2011) A multicellular spheroid formation and extraction chip using removable cell trapping barriers. Lab Chip 11:115–119
Kim H, Lee S, Kim J (2012) Hydrodynamic trap-and-release of single particles using dual-function elastomeric valves: design, fabrication, and characterization. Microfluid Nanofluidcs 13:835–844
Lee GB, Chang CC, Huang SB, Yang RJ (2006) The hydrodynamic focusing effect inside rectangular microchannels. J Micromech Microeng 16:1024–1032
Malainou A, Petrou PS, Kakabakos SE, Gogolides E, Tserepi A (2012) Creating highly dense and uniform protein and DNA microarrays through photolithography and plasma modification of glass substrates. Biosens Bioelectron 34:273–281
Nolan JP, Mandy F (2006) Multiplexed and microparticle-based analyses: quantitative tools for the large-scale analysis of biological systems. Cytometry A 69A:318–325
Robinson WH et al (2002) Autoantigen microarrays for multiplex characterization of autoantibody responses. Nat Med 8:295–301
Shao G, Cai Z, Wang J, Wang W and Lin Y (2011) A pneumatic actuated microfluidic beads-trapping device. Proceedings of SPIE7929: Microfluidics, BioMEMS, and Medical Microsystems IX (SPIE 2011), San Francisco, California
Skelley AM, Kirak O, Suh H, Jaenisch R, Voldman J (2009) Microfluidic control of cell pairing and fusion. Nat Methods 6:147–152
Sochol RD, Iwai K, Higa AT, Lo JC, Zhou E, Lo L, Luong C, Dueck M, Li S, Lee LP and Lin L (2010) A resettable high-density microfluidic cell trapping system. Proceedings of 14th International Conference on Miniaturized Systems for Chemistry and Life Sciences (MicroTAS 2010), Groningen
Sochol RD, Dueck ME, Li S, Lee LP, Lin L (2012) Hydrodynamic resettability for a microfluidic particulate-based arraying system. Lab Chip 12:5051–5056
Steinert CP, Kalkandjiev K, Zengerle R (2009) TopSpot® Vario: a novel microarrayer system for highly flexible and highly parallel picoliter dispensing. Biomed Microdevices 11:755–761
Tan WH, Takeuchi S (2007) A trap-and-release integrated microfluidic system for dynamic microarray applications. Proc Natl Acad Sci 104:1146–1151
Tan WH, Takeuchi S (2008) Dynamic microarray system with gentle retrieval mechanism for cell-encapsulating hydrogel beads. Lab Chip 8:259–266
Tonooka T, Teshima T, Takeuchi S (2012) Clustering triple microbeads in a dynamic microarray for timing-controllable bead-based reactions. Microfluid Nanofluidcs. doi:10.1007/s10404-012-1111-7
Verpoorte E (2003) Beads and chips: new recipes for analysis. Lab Chip 3:60N–68N
Voldman J, Gray ML, Toner M, Schmidt MA (2002) A microfabrication-based dynamic array cytometer. Anal Chem 74:3984–3990
Wang Z, Zhe J (2011) Recent advances in particle and droplet manipulation for lab-on-a-chip devices based on surface acoustic waves. Lab Chip 11:1280–1285
Yamada M, Kano K, Tsuda Y, Kobayashi J, Yamato M, Seki M, Okano T (2007) Microfluidic devices for size-dependent separation of liver cells. Biomed Microdevices 9:637–645
Zhu Z, Frey O, Ottoz DS, Rudolf F, Hierlemann A (2012) Microfluidic single-cell cultivation chip with controllable immobilization and selective release of yeast cells. Lab Chip 12:906–915
Acknowledgments
This work was supported by the National Research Foundation of Korea (NRF) Grant funded by the Korea government (MEST) (No. 2011-0030075) and Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education, Science and Technology (2012R1A1A2006305). We would like to express our appreciation to Prof. Jehyun Baek and Prof. Dongsung Kim, POSTECH for their support on our experimental setup to perform this work.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 2 (WMV 4694 kb)
Supplementary material 3 (WMV 3607 kb)
Supplementary material 4 (WMV 4085 kb)
Supplementary material 5 (WMV 3288 kb)
Supplementary material 6 (WMV 1863 kb)
Rights and permissions
About this article
Cite this article
Kim, H., Kim, J. A microfluidic-based dynamic microarray system with single-layer pneumatic valves for immobilization and selective retrieval of single microbeads. Microfluid Nanofluid 16, 623–633 (2014). https://doi.org/10.1007/s10404-013-1267-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10404-013-1267-9