Abstract
Dextranase can hydrolyze dextran to low-molecular-weight polysaccharides, which have important medical applications. In the study, dextranase-producing strains were screened from various soil sources. The strain H6 was identified as Talaromyces pinophilus by a standard ITS rDNA analysis. Crude dextranase was purified by ammonium sulfate fractionation and Sepharose 6B chromatography, which resulted in a 6.69-fold increase in the specific activity and an 11.27% recovery. The enzyme was 58 kDa, lower than most dextranase, with an optimum temperature of 45 °C and an optimum pH of 6.0, and identified as an endodextranase. It was steady over a pH range from 3.0 to 10.0 and had reasonable thermal stability. The dextranase activity was increased by urea, which enhanced its activity to 115.35% and was conducive to clinical dextran production. Therefore, T. pinophilus H6 dextranase could show its superiority in practical applications.
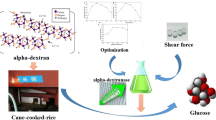
Graphical Abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Dextrans are high-molecular-weight (Mw) exopolysaccharides, which consist of α-d-(1 → 6) linked glucose units with various industrial applications [13, 27]. They distinctly vary in structures depending on the percentage and the manner of glucose branching [16]. The extensive branching of dextrans greatly affects their physicochemical properties in terms of viscosity and solubility [26]. With sucrose solution as substrate, dextran polymers are biosynthesized by dextransucrase (E.C. 2.4.1.5), which is mainly produced by Leuconostoc mesenteroides and Streptococcus lactis bacteria. Low-Mw fractions of dextrans, which are degraded by dextranase, have many important biotechnological applications. Different fractions of dextran have been used as blood volume expanders [14, 35], vaccines, and drug delivery vehicles [18, 23] in medical industries, additives in food and cosmetics, and for separating gels in studies [13]. Dextrans with relatively low Mw (6–8 kDa) can form complexes with iron, and these complexes have good therapeutic effects in the treatment of severe anemia [11]. Low-Mw dextran sulfate can replace heparin to aid in blood clotting and isomaltooligosaccharides have been used as prebiotics [9, 25]. At present, α-glucans are frequently industrially degraded by hydrolyzing the polymer with hydrochloric acid at high temperature, followed by separating the products from polymers and isomaltodextrins formed using an organic solvent. However, the hydrolysate containing a high concentration of chlorides causes serious adverse effects on products and leads to allergic reactions. In addition, this process facilitates the relatively broad Mw distribution because of the random disorder of acid hydrolysis. The yield of this method should also be further enhanced because of the original glucan loss. Therefore, enzymatic degradation of α-glucans is a green technology for industrial production.
Dextranase (α-1,6-d-glucan-6-glucanohydrolase; E.C. 3.2.1.11) is an inducible enzyme that catalyzes the endohydrolysis of the predominant α-1,6 glycosidic bonds of dextrans. Dextranases are extensively used in theoretical research and practical purposes. In addition to preparing dextran and its derivative, dextranase plays a crucial role in molasses and beverage processing in food industries [30], oral hygiene to prevent dental caries [7, 28], and in describing the structure of polysaccharides [21]. Dextrans are initially synthesized mainly by controlling the temperature and concentration of sucrose and dextransucrase [10], but this technique is restricted to industrial production because of the difficulties in separating and purifying dextrans. With dextranase being increasingly isolated from various sources, the enzymatic biocatalysis of the hydrolysis of α-glucans should be examined and developed. However, co-fermenting L. mesenteroides and Lipomyces starkeyi [14] is adverse to product purification because of the multiple substances in the mixed culture broth. Goulas et al. [9] prepared oligodextrans and isomaltooligosaccharides by combining dextransucrase and dextranase, but the products were not purified further. Moreover, dextranase with high catalytic ability and stability can improve the efficiency of α-glucan degradation and its potential commercial and industrial value. Thus, novel and various dextranases are worth further development.
Dextranases are present in various microorganisms, such as bacteria, mold, and yeast [13]. Dextranase produced by fungi has received much attention because of their high enzymatic activity and isomaltooligosaccharides products [15]. In the present study, Talaromyces pinophilus strain was separated from natural soil samples, and dextranase production conditions were optimized. This study was performed primarily to separate and purify T. pinophilus H6 dextranase from fermentation and measure its enzymatic characteristics. This study was the first to report the purification and characterization of a dextranase from isolated T. pinophilus strain. Then, the purified dextranase was used to degrade polymers with reasonable thermal stability. Moreover, the enzyme was beneficial for high-Mw dextran degradation, which could solve the broad Mw distribution and low yield. The glucan polymers degraded by T. pinophilus H6 dextranase can form products with controllable Mws (10–70 kDa) by controlling the contents of dextranase (5.00 U/mL) and dextrans (6%) and reaction time (150 min). The Mws of the products were 69,546, 40,203, 20,113, and 10,052 Da with a total yield of 82.60%. The glucan products were suitable for medical applications that require high purity and monodispersity. In addition, the productivity and yield were improved. The new bioprocess for α-glucan degradation facilitated its industrial production.
Materials and methods
Materials
Dextran T2000 (2000 kDa), Sephadex G150, G100, G15, and Sepharose 6B were Pharmacia products (Uppsala, Sweden). A series of dextran standards (dextran 5900, 3450, 2500, 1900, 1185, 655, 420, 275, 144, 47, 31, 20, and 3 kDa) for the calibration curves of Mw was obtained from APSC (Mentor, OH, USA). Dextran T20, T40, and T70 (with average Mws of 20, 40, and 70 kDa, respectively) were obtained from Jinyang (Shandong, China). The unstained protein Mw marker was obtained from TaKaRa (Dalian, China). Commercial dextranase was purchased from Sigma-Aldrich (St. Louis, MO, USA). All other chemicals were of analytical grade. Dextransucrase was produced from the engineered strain Escherichia coli BL21(DE3)/pET28-dexYG [36] constructed in our laboratory. The culture conditions and medium for enzyme production were similar to those in a previous report [8]. Dextran (7–8 kDa) was produced by the synergistic synthesis of the aforementioned dextransucrase and dextranase from Penicillium aculeatum F1001 produced in previous reports [8, 37].
Screening for dextranase-producing strains
Various soil samples were collected from different habitats (He Fei, Fu Yang, and Ma On Shan) in China. The microorganisms that can assimilate dextran were isolated and screened. The soil suspension was diluted and evenly spread on potato dextrose agar (PDA) plates. Fungi that grew on the PDA plates at 28 °C under aerobic conditions were cultured in the screening medium. The screening medium (pH 5.5) included (w/v) 1.5% dextran T2000, 0.5% NaNO3, 0.1% KCl, 0.1% K2HPO4·3H2O, 0.1% MgSO4·7H2O, 0.001% FeSO4·7H2O, and 1.8% agar powder. Colonies that assimilated dextrans were grown in the screening medium plates and cultured in the screening medium without agar powder. Dextran T2000 was degraded by the culture broth of the colonies, thereby producing hyaline halos in the plate. Among the dextranase-producing strains screened, strain H6 showed the highest dextranase activity in repetitive culture and was identified by strain morphology, cultural characteristics, and internal transcribed spacer (ITS) rDNA analysis.
Culture conditions for dextranase production
The fermentation medium of the strain H6 included (w/v) 1.5% dextran (7–8 kDa), 3% soluble starch, 2.5% KNO3, 0.1% KCl, 0.1% K2HPO4·3H2O, 0.1% MgSO4·7H2O, and 0.001% FeSO4·7H2O. The initial pH of the medium was pH 3.0–3.5. A single T. pinophilus H6 was suspended in the medium and cultivated on a shaker (220 rpm) at 33 °C for 8 days. The thalli of T. pinophilus H6 were separated by centrifugation for 30 min at 4 °C at 8000×g. The supernatant was stored at 4 °C.
Determination of protein concentration
Protein concentration (mg/mL) was measured by the Bradford method [3] using crystalline bovine serum albumin as the protein standard.
Dextranase activity assay
Dextranase activity was measured by determining the reducing sugar released during the enzyme–substrate reaction [24]. The diluted enzyme solution (4.0 mL) was incubated with 4.0 mL of substrate (3% dextran T70) solubilized in 20 mM acetate buffer (pH 6.0) at 45 °C for 30 min. One unit (1 U) of dextranase activity was defined as the quantity of enzyme that could hydrolyze dextran to produce reducing sugar equivalent to 1 μM glucose/min at standard assay conditions.
Purification of dextranase
The crude enzyme extract was subjected to ammonium sulfate fractionation from 40 to 70% saturation. The precipitate was collected by centrifugation at 8000×g for 20 min at 4 °C. The precipitate was then dissolved and dialyzed in 20 mM acetate buffer (pH 6.0). The enriched enzyme sample was eluted from a Sepharose 6B column (1.5 cm i.d. × 60 cm) with 20 mM acetate buffer (pH 6.0) at a flow rate of 15 mL/h. The absorbance of each fraction was determined at 280 nm to monitor the proteins during chromatographic separation. The fractions with high dextranase activity were gathered to measure dextranase activity and protein concentration according to the above description. Specific activity was calculated according to the above measurement results. The purity of the enzyme and its molecular mass were determined by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). Protein samples and unstained protein marker were separated using 10.0% polyacrylamide gel [20]. Following electrophoresis, the gel was stained with Coomassie blue G-250.
Effect of pH on enzyme activity and stability
Dextranase activity was determined at different pH values ranging from 3.0 to 10.0 (acetate buffer, pH 3.0–5.5; phosphate buffer, pH 6.0–8.0; Tris–HCl, pH 7.5–9.0; glycine-NaOH, pH 9.0–10.0). Relative activities were expressed as percentages of the maximum activity. The pH stability of the enzyme was evaluated by incubating the enzyme in reaction buffers at pH 3.0 to 10.0 at 25 °C for 1 h without any substrate. Dextran T70 was added to the enzyme to measure residual enzyme activity at standard assay conditions.
Effect of temperature on enzyme activity and stability
The effect of temperature on dextranase activity was evaluated by incubating the reaction mixtures at 20–75 °C and measuring the enzyme by the standard method. Relative activities were expressed as percentages of the maximum activity. The thermal stability of dextranase was evaluated by exposing the enzyme in 20 mM acetate buffer (pH 6.0) for 1 h at different temperatures without any substrate. Dextran T70 was added to the enzyme to measure the relative residual enzyme activity at standard assay conditions. The activity of non-heated enzyme was considered as 100%. To evaluate the temperature stability of dextranase, the enzyme was placed at −18, 4, or 45 °C at different intervals until the enzyme activity was reduced to half of its initial value.
Effects of metal ions and other reagents on dextranase activity
The effects of some metal ions and reagents [namely, K+ (KCl), NH4 + (NH4Cl), Mg2+ (MgSO4), Zn2+ (ZnSO4), Ca2+ (CaCl2), Mn2+ (MnCl2), Co2+ (CoCl2), Cu2+ (CuSO4), Ba2+ (BaCl2), Ni2+ (NiCl2), Al3+ (AlCl3), Fe3+ (FeCl3), urea, Tris, SDS, and EDTA] on dextranase activity were measured at standard assay conditions. Dextranase activity assayed without any compounds was considered as 100% to calculate relative activities with the aforementioned compounds.
Substrate specificity and analysis of final hydrolysis products
The rate of enzymatic hydrolysis of substrates with diverse glycosidic bonds was measured to evaluate the substrate specificity of dextranase. The enzyme was incubated at 45 °C for 30 min in 20 mM acetate buffer (pH 6.0) with different carbohydrates as substrates. Relative activities were expressed as percentages of the maximum activity. The final hydrolysis products of dextran T70 were analyzed by thin-layer chromatography (TLC) using a silica gel 60 plate developed in a solvent system consisting of 7:4:2:1 (v/v/v/v) n-butanol:isopropanol:acetic acid:water. Glucose, isomaltose, and isomaltotriose were used as the standards. A mixture of dextran T70 and heat-inactivated dextranase was the control sample. The carbohydrates were visualized on the plate by spraying a solution including diphenylamine (2 g), aniline (2 mL), and 85% phosphonic acid (10 mL) in 100 mL of acetone, followed by heating at 85 °C for 10 min. In addition, the mixture was treated as a sample and analyzed by HPLC (Waters, Milford, USA), in which a Luna 5 μm NH2 (4.6 mm × 25 cm, Phenomenex, USA) column was used. The chromatograph was operated with a flow rate of 1.0 mL/min at 40 °C and then connected to a Smart-line refractive index detector (Knauer, Berlin, Germany).
Enzyme kinetics
The initial velocity (ν) was determined to measure the kinetic constants using various concentrations (0.2–1.0% w/v) of dextran T100, T70, and T40 in 20 mM acetate buffer (pH 6.0) at 45 °C. The kinetic constants were calculated from Lineweaver–Burk plots [22].
Kinetic study of the Mw of α-glucan biocatalyzed by T. pinophilus H6 dextranase
Talaromyces pinophilus H6 dextranase was added to 100 mL of 6% dextran (Mw = 5046 kDa) in 20 mM acetate buffer (pH 6.0) at a final concentration of 5.0 U/mL. The reaction solution was incubated at 45 °C with mechanical stirring. Samples of the reaction mixture were obtained at intervals from 1 to 90 min, boiled for at least 5 min to stop the reaction, and diluted to 5 mg/mL. Subsequently, diluted samples were filtered through a mixed cellulose ester membrane (a filter membrane with 0.22 μm aperture and 25 mm diameter) and analyzed by HPLC (Waters, Milford, USA), in which TSK gel G4000XL (7.8 mm × 30 cm, TOSOH, HOSOL, Japan) and TSK gel G6000XL (7.8 mm × 30 cm, TOSOH, HOSOL, Japan) columns were used. The chromatograph was operated with a flow rate of 0.6 mL/min at 60 °C and then connected to a Smart-line refractive index detector (Knauer, Berlin, Germany). Calibration curves of the retention time and Mw were prepared for dextran standards (APSC, Mentor, Ohio, USA): 5900, 3450, 2500, 1900, 1185, 655, 420, 275, 144, 47, 31, 20, and 3 kDa dextran.
Enzymatic biocatalysis of α-glucan degradation
The substrate solution included 6% glucan polymer (Mw = 5046 kDa) in 20 mM acetate buffer (pH 6.0). T. pinophilus H6 dextranase was added to 1000 mL of the substrate solution at a final concentration of 5.0 U/mL. The reaction solution was incubated at 45 °C for 150 min with mechanical stirring and heated in a 90 °C water bath for 60 min to stop the reaction. Activated carbon was added to a final concentration of 5% (w/v) to decolorize the reaction solution, and the mixture was heated to microboiling for 10 min and filtered. The glucan products were separated by graded ethanol precipitation. The different concentrations (42, 49, 60, and 78%) of ethanol precipitated products with varying Mws. Water was added to the products to dissolve the precipitated dextrans and reprecipitated products with ethanol. The Mw of each glucan sample (5 mg/mL) was then determined by HPLC, as previously described.
The results reported are the means of at least three separate experiments.
Results and discussion
Screening and identification of dextranase-producing strains
Strain H6, which formed hyaline holes on the screening plate, was obtained. This strain grew under aerobic conditions and showed dextranase activity of 4.60 U/mL in the culture filtrate under the induction of dextran. The strain incubated on PDA plates at 28 °C initially exhibited white filamentous colonies, then gradually became yellow-green, and finally grew green or dark green spores. The colonies were large with flocculent felty surface and neat edge, and the reverse side turned dark red. The conidiophore grew broom-like whorled or solitary branches at the top, and the conidia were spherical or oval. The strain could be classified under the genus Penicillium because of its morphology. The phylogenetic tree of strain H6 is shown in Fig. 1. The ITS rDNA sequence of the strain closely matched that of T. pinophilus at 99% homology. Its GenBank accession number is KF751644. The strain was deposited in the China General Microbiological Culture Collection Center with the accession number of 9260. Dextranase production by T. pinophilus has not been reported. Thus, T. pinophilus H6 is a different strain for dextranase production.
Phylogenetic tree of H6 based on ITS rRNA gene sequences and the reference Talaromyces species. Evolutionary distances were calculated using MEGA 4.1, with 0.05 substitution per nucleotide. The numbers in parentheses represent the GenBank sequence accession numbers. The number at each branch point is the percentage supported by bootstrap
Purification of dextranase
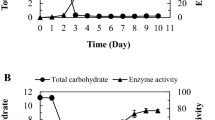
Dextranase from the supernatant was originally concentrated by ammonium sulfate from 40 to 70% saturation. The enzyme was eluted and separated effectively using Sepharose 6B chromatography (Fig. 2a). Fractions with high specific activity were collected from tubes 13–15. Tubes 13 and 14 were merged and applied to SDS-PAGE. Dextranase was purified to 6.69-fold and its yield was 11.27% (Table 1). The purified enzyme evidently showed high specific activity (14,894 U/mg), which was 5.35-fold higher than its counterpart reported in published methods [34] using a similar two-step purification. SDS-PAGE results showed that the Mw of purified dextranase was approximately 58 kDa (Fig. 2b), which suggested that T. pinophilus H6 dextranase should be a monomer. Only one band appeared in the image, reflecting the effectiveness of the purification process. Its Mw was similar to the counterparts of dextranase from Thermotoga lettingae [17], Chaetomium erraticum [31], Hypocrea lixii [34], and a recombinant Lipomyces starkeyi [5]. The known Mw of dextranase is generally from 40 kDa to 80 kDa. The minimum dextranase (26.5 kDa) is derived from Penicillium lilacinum [6], and the maximum one (175 kDa) is from Streptococcus sobrinus [2].
Purification of dextranase by Sepharose 6B chromatography. a The primary purified enzyme was purified by Sepharose 6B chromatography and gathered from tubes 13–15. b Pure dextranase collected from gel filtration chromatography with 10.0% SDS-PAGE. Lane M protein molecular mass marker; lane 1 culture broth; lane 2 ammonium sulfate precipitate; lane 3 purified dextranase in Sepharose 6B
Effects of pH and temperature on dextranase activity and enzyme stability
The effect of pH on dextranase activity is shown in Fig. 3a (closed square). The optimum pH for T. pinophilus H6 dextranase was 6.0, which was similar to those from other fungi [1, 29] and within the range of pH 4.5–6.5 reported for dextranases from filamentous fungi [32]. The results also implied that dextranase was active within the pH range of 4.0–7.0. Dextranase activity was extremely stable between pH 3.0 and 10.0 at 25 °C [Fig. 3a (open square)].
Effect of pH (a) and temperature (b) on the activity (closed square) and stability (open square) of dextranase from T. pinophilus H6. The reactions consisting of 3% dextran T70, dextranase, and buffer were performed by incubation for 30 min. The effect of pH on enzyme activity was determined at 45 °C in acetate buffer, phosphate buffer, Tris–HCl, and glycine–NaOH. The pH stability was evaluated by incubating the enzyme in reaction buffers at pH 3.0 to 10.0 at 25 °C for 1 h. Dextran T70 was added to the enzyme to measure residual enzyme activity at standard assay conditions. The effect of temperature was determined at 20–75 °C in 20 mM acetate buffer (pH 6.0). The thermal stability was evaluated by incubating the enzyme in 20 mM acetate buffer (pH 6.0) at 20–75 °C after 1 h incubation. For the effect of pH and temperature on dextranase activity and stability, the values are shown as percentages of the maximum activities, which were taken to be 100%
The effect of temperature on dextranase activity is shown in Fig. 3b. The optimal temperature of T. pinophilus H6 dextranase was 45 °C, and the enzyme evidently showed activity from 30 to 65 °C, with peak relative activity of no less than 75% (Fig. 3b). The thermal stability of dextranase revealed that the enzyme retained almost 100% residual activity after storage at 45 °C (pH 6.0) for 10 h. The half-life (t 1/2) of H6 dextranase was 19 days at 45 °C (data not shown). Moreover, dextranase was stable at −18 and 4 °C (for 56 days) in 20 mM acetate buffer (pH 6.0). Table 2 summarizes the main characteristics and functions of several microbial dextranases. It is obvious that the characteristics are different due to various sources [19]. The purified dextranase from T. pinophilus H6 has high specific activity (14,894 U/mg) and reasonable thermal stability (keeps high activity for 456 h at optimum temperature), showing priority to dextranases from other sources. Also, compared with other dextranases, our dextranase has lower pH sensitivity (exerts satisfactory catalysis ability from pH 3 to 10). Additionally, it can avoid the compound action of enzymolysis and acidolysis/alkaline hydrolysis and it would be easier to gain homogeneous dextran than the dextranases with heat and acid/alkaline catalysis conditions [5, 17, 26].
Effects of metal ions and reagents on dextranase activity
The effects of several compounds on dextranase activity are presented in Table 3. Dextranase activity increased from 100% (without addition of compounds) to 115.35 and 114.80% in the presence of urea at 1 mM and SDS at 10 mM, respectively. Additionally, the enzyme was found to be inactivated completely by Cu2+ (with sulfate) at 10 mM. Consistent with the results from Das and Dutta [6], dextranase activity was strongly inhibited by Zn2+ (with sulfate). Dextranase activity was also slightly inhibited by Co2+, Ni2+, Fe3+ (with chloride), and EDTA. Other tested metal ions, such as K+, NH4 +, Ca2+, Ba2+ (with chloride), and Mg2+ (with sulfate), did not show perceptible influences.
Substrate specificity and final hydrolysis products of T. pinophilus dextranase
Dextranase activities in the catalyzed hydrolysis of substrates with diverse glycosidic bonds were measured to evaluate the substrate specificity of dextranase (Table 4). The enzyme revealed high specificity to dextran primarily composed of α-1,6 glucosidic bond, which was similar to other dextranases [1, 7]. The optimum substrate was dextran T70, and the optimum substrate concentration was 3%. Dextranase was less effective in the hydrolysis of other carbohydrates containing α-1,6 glucosidic bond, such as Sephadex. It had no activity toward β-cyclodextrin, chitin, Sepharose 6B, chitosan, and sucrose. In addition, the activity toward soluble starch, which included most α-1,4 glucosidic bonds and a few α-1,6 bonds, was relatively low. This phenomenon is more evident than that of H. lixii F1002 dextranase [34], which indicated that the enzyme had more capacity for hydrolyzing discontinuous α-1,6 glucosidic bonds in soluble starch. We chose dextrans (1000–4000 kDa) as the substrate, and the dextran molecular weight showed a rapid downward trend during the first 3 h and turned to a slow downward trend after that, showing a more prominent degradation effect of dextran than many dextranases from other sources [12, 17, 26]. As illustrated by supplementary Table 1 and Table 4, it is obvious that our dextranase shows high affinity to and catalysis efficiency for high-molecular-weight dextran and is preferred for the hydrolysis of high-molecular-weight dextran. Thus, our dextranase hydrolyzes high-molecular-weight dextran under different conditions and low-molecular-weight dextran can be obtained by fractional collection, which is beneficial to gain various dextrans and its derivatives that are applicable to medicine, e.g. iron-dextrin [11] for anemia, dextran sulfate [25] for anticoagulation and direct use for the prevention and treatment of dental caries [28]. In addition, the culture broth of T. pinophilus H6 also exhibited high activity with laminarin consisting of β-1,3 linkages. Further studies on this discovery are in progress. The final products of the optimum substrate hydrolysis catalyzed by T. pinophilus dextranase were isomaltose, isomaltotriose, and some isomaltooligosaccharides (Fig. 4), which were similar to the counterparts of dextranase from C. erraticum [30]. The enzyme did not show any transglucosylation products by TLC and HPLC (data not shown), suggesting that dextran T70 hydrolysis is the main reaction and the enzyme is of the endotype. The dextranase from T. pinophilus has a mode of action similar to that of the dextranases from T. lettingae [17], H. lixii [34], and Bacillus licheniformis [38].
Michaelis constant (K m) values
The initial rate of hydrolysis (the liberation velocity of the reducing sugar) of dextran was measured to evaluate the ν value for the breakage of glucosidic bonds. The kinetic constants calculated from Lineweaver–Burk plots are summarized in Table 5. Dextranase had different K m values for dextrans of various molecular masses (Dextran T100, T70, and T40, respectively). The results suggested that the enzyme was attractive to high-Mw dextrans. This behavior was similar to those of P. aculeatum dextranase [8], L. starkeyi dextranase [33], and H. lixii F1002 dextranase [34]. This property of the enzyme was superior to that of Streptomyces sp. NK458 dextranase [26], further suggesting the suitability of the strain for industrial applications.
Kinetic study of the Mw of α-glucan biocatalyzed by T. pinophilus dextranase
The dextran Mw (data not shown) indicated that the minimum Mw obtained within 90 min of the hydrolysis reaction was 18,032 Da. In addition, the Mw decreased faster from 1 to 20 min than that from 25 to 90 min, because the enzyme was more attractive to α-glucans with higher Mw. This phenomenon was inferred to be associated with the kinetic constants of the enzyme. The hydrolysis products were obtained in a relatively short time because T. pinophilus H6 dextranase has a larger ν value than H. lixii F1002 dextranase [34].
Enzymatic biocatalysis of α-glucan degradation
The Mw values of the glucans obtained were 69,546, 40,203, 20,113, and 10,052 Da, corresponding to elution times of 12.958, 13.523, 14.237, and 14.952 min, respectively. The yield of each type was 13.17, 39.35, 18.21, and 11.87%. The separated products had high purities and monodispersities (data not shown) because of the unique hydrolytic affinity and high dextranase activity of T. pinophilus H6 dextranase. Therefore, the enzyme was efficient in producing α-glucans of specified Mws from glucan polymers by controlling the concentrations of dextranase and substrates, as well as the reaction time.
Conclusions
In summary, a novel dextranase from T. pinophilus H6 was purified and characterized. The purified enzyme had high specific activity (14,894 U/mg) and reasonable thermal stability. Additionally, it was significantly stable over a broad pH range from pH 3.0 to 10.0. The enzyme showed good performance in hydrolyzing high-Mw glucan, which can solve broad Mw distribution and low yield. Therefore, dextranase with unusual enzymatic properties can be a potential biocatalyst to degrade α-glucans in industry.
References
Abdel-Naby MA, Ismail AS, Abdel-Fattah AM, Abdel-Fattah AF (1999) Preparation and some properties of immobilized Penicillium funiculosum 258 dextranase. Process Biochem 34(4):391–398
Barret JF, Barret TA, Curtiss R (1987) Purification and partial characterization of the multicomponent dextranase complex of Streptococcus sobrinus and cloning of the dextranase gene. Infect Immun 55(3):792–802
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Cai RH, Lu MS, Fang YW, Jiao YL, Zhu Q, Liu ZP, Wang SJ (2014) Ann Microbiol 64:147–155
Chen L, Zhou X, Fa W, Zhang Y (2008) Expression, purification and characterization of a recombinant Lipomyces starkey dextranase in Pichia pastoris. Protein Expres Purif 58(1):87–93
Das DK, Dutta SK (1996) Purification, biochemical characterization and mode of action of an extracellular endo-dextranase from the culture filtrate of Penicillium lilacinum. Int J Biochem Cell Biol 28:107–113
Finnegan PM, Brumbley SM, O’Shea MG, Nevalainen H, Bergquist PL (2004) Isolation and characterization of genes encoding thermoactive and thermostable dextranases from two thermo tolerant soil bacteria. Curr Microbiol 49(5):327–333
Gan WW, Zhang HB, Zhang YQ, Hu XQ (2014) Biosynthesis of oligodextrans with different Mw by synergistic catalysis of dextransucrase and dextranase. Carbohydrate Polym 112:387–395
Goulas AK, Fisher DA, Grimble GK, Grandison AS, Rastall RA (2004) Synthesis of isomaltooligosaccharides and oligodextrans by the combined use of dextransucrase and dextranase. Enzyme Microb Technol 35:327–338
Hayacibara MF, Koo F, Smith AMV, Kopec LK, Scott-Anne K, Cury JA, Bowen WH (2004) The influence of mutanase and dextranase on the production and structure of glucans synthesized by streptococcal glucosyltransferases. Carbohyd Res 339(12):2127–2137
Hussain I, Bhoyroo J, Butcher A, Koch TA, He A, Bregman DB (2013) Direct comparison of the safety and efficacy of ferric carboxymaltose versus iron dextran in patients with iron deficiency anemia. Anemia 169107:1–11
Jiao YL, Wang SJ, Lv MS, Jiao BH, Li WJ, Fang YW, Liu S (2014) Characterization of a marine-derived dextranase and its application to the prevention of dental caries. J Ind Microbiol Biotechnol 41:17–26
Khalikova E, Susi P, Korpela T (2005) Microbial dextran-hydrolyzing enzymes: fundamentals and applications. Microbiol Mol Biol R 69:306–325
Kim D, Day DF (1994) A new process for the production of clinical dextran by mixed-culture fermentation of Lipomyces starkeyi and Leuconostoc mesenteroides. Enzyme Microb Tech 16:844–848
Kim D, Robyt JF, Lee SY, Lee JH, Kim YM (2003) Dextran molecular size and degree of branching as a function of sucrose concentration, pH, and temperature of reaction of Leuconostoc mesenteroides B-512FMCM dextransucrase. Carbohyd Res 338:1183–1189
Kim YM, Kimura A, Kim D (2011) Novel quantitative method for the degree of branching in dextran. Food Sci Biotechnol 20(2):537–541
Kim YM, Kim D (2010) Characterization of novel thermostable dextranase from Thermotoga lettingae TMO. Appl Microbiol Biotechnol 85(3):581–587
Kulthida K, Pranee I, Emmanuelle M, Alain D (2012) Enzymatically degradable nanoparticles of dextran esters as potential drug delivery systems. Carbohyd Polym 88:875–881
Kang C, Yu XW, Xu Y (2014) Purification and characterization of a prolyl endopeptidase isolated from Aspergillus oryzae. J Ind Microbiol Biotechnol 41:49–55
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227(5259):680–685
Larsson AM, Andersson R, Stahlberg J, Kenne L, Jones TA (2003) Dextranase from Penicillium minioluteum: reaction course, crystal structure, and product complex. Structure 11(9):1111–1121
Lineweaver H, Burk D (1934) The determination of enzyme dissociation constants. J Am Chem Soc 56:658–666
Mehvar R (2000) Dextrans for targeted and sustained delivery of therapeutic and imaging agents. J Control Release 69:1–25
Nelson N (1944) A photometric adaptation of the Somogyi method for the determination of glucose. J Biol Chem 153:375–380
Olcer Z, Tanriseven A (2010) Co-immobilization of dextransucrase and dextranase in alginate. Process Biochem 45:1645–1651
Purushe S, Prakash D, Nawani N, Dhakephalkar P, Kapadnis B (2012) Biocatalytic potential of an alkalophilic and thermophilic dextranase as a remedial measure for dextran removal during sugar manufacture. Bioresource Technol 115:2–7
Li RH, Zhang HB, Hu XQ, Gan WW, Li QP (2016) An efficiently sustainable dextran-based flocculant: synthesis, characterization and flocculation. Chemosphere 159:342–350
Singleton V, Horn J, Bucke C, Adlard M (2001) A new polarimetric method for the analysis of dextran and sucrose. Int Sugar J 103(1230):251–254
Sugiura M, Ito A, Ogiso T, Kato K, Asano H (1973) Purification of dextranase from Penicillium funiculosum and its enzymatic properties. B B A 309(2):357–362
Thitaram SN, Chung CH, Day DF, Hinton A, Bailey JS, Siragusa GR (2005) Isomaltooligosaccharide increases cecal Bifidobacterium population in young broiler chickens. Poultry Sci 84(7):998–1003
Virgen-Ortíza JJ, Ibarra-Junqueraa V, Escalante-Minakataa P, Ornelas-Pazb J de J,Osuna-Castroc JA, González-Potes A (2015) Kinetics and thermodynamic of the purified dextranase from Chaetomium erraticum. J Mol Catal B-Enzym 122:80–86
Walker GJ (1978) Dextrans. Biochem Carbohyd 11(16):75–126
Webb E, Spencer-Martins I (1983) Extracellular endodextranase from the yeast Lipomyces starkeyi. Can J Microbiol 29(9):1092–1095
Wu DT, Zhang HB, Huang LJ, Hu XQ (2011) Purification and characterization of extracellular dextranase from a novel producer, Hypocrea lixii F1002, and its use in oligodextran production. Process Biochem 46:1942–1950
Zdolsek HJ, Vecfors M, Lindahl TL, Tornquist T, Bortnik P, Hahn RG (2011) Hydroxyethyl starches and dextran during hip replacement surgery: effects on blood volume and coagulation. Acta Anaesth Scand 55(6):677–685
Zhang HB, Hu YJ, Zhu CB (2008) Cloning, sequencing and expression of a dextransucrase gene (dexYG) from Leuconostoc mesenteroides. Biotechnol Lett 30(8):1441–1446
Zhang HB, Wu DT, Huang LJ, Hu XQ, Wang X (2011) Purification, characterization of an extracellular dextranase from an isolated Penicillium sp. Acta Microbiol Sin 51(4):495–503
Zohra RR, Aman A, Ansari A, Haider MS, Qader SAU (2015) Purification, characterization and end product analysis of dextran degrading endodextranase from Bacillus licheniformis KIBGE-IB25. Int J Biol Macromol 78:243–248
Acknowledgements
The study was supported by the National Natural Science Foundation of China (Grant No. 81573399).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, YQ., Li, RH., Zhang, HB. et al. Purification, characterization, and application of a thermostable dextranase from Talaromyces pinophilus . J Ind Microbiol Biotechnol 44, 317–327 (2017). https://doi.org/10.1007/s10295-016-1886-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-016-1886-8