Abstract
Cells migrating in clusters play a significant role in a number of biological processes such as embryogenesis, wound healing, and tumor metastasis during cancer progression. A variety of environmental and biochemical factors can influence the collective migration of cells with differing degrees of cell autonomy and inter-cellular coupling strength. For example, weakly coupled cells can move collectively under the influence of contact guidance from neighboring cells or the environment. Alternatively strongly coupled cells might follow one or more leader cells to move as a single cohesive unit. Additionally, chemical and mechanical signaling between these cells may alter the degree of coupling and determine effective cluster sizes. Being able to understand this collective cell migration process is critical in the prediction and manipulation of outcomes of key biological processes. Here we focus on understanding how various environmental and cellular factors influence small clusters of cells migrating collectively within a 3D fibrous matrix. We combine existing knowledge of single-cell migration in 2D and 3D environments, prior experimental observations of cell–cell interactions and collective migration, and a newly developed stochastic model of cell migration in 3D matrices, to simulate the migration of cell clusters in different physiologically relevant environments. Our results show that based on the extracellular environment and the strength of cell–cell mechanical coupling, two distinct optimal approaches to driving collective cell migration emerge. The ability to effectively employ these two distinct migration strategies might be critical for cells to collectively migrate through the heterogeneous tissue environments within the body.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Collective cell migration is vital for physiological processes such as tissue growth, morphogenesis, wound healing, and cancer metastasis (Arima et al. 2011; Munjal and Lecuit 2014; Alexander et al. 2008; Gillitzer and Goebeler 2001). In the first three processes mentioned above, cells move collectively within sheets or at the boundary of a 2D sheet, and the factors that drive these collective phenomena have been widely described and studied (Petitjean et al. 2010; Sepúlveda et al. 2013; Vedula et al. 2012; Garcia et al. 2015; Palamidessi et al. 2019). Primarily, guiding forces in these cases come from neighboring non-motile cells providing contact guidance, cell shape and polarity changes driven by inter-cellular tension, forces from cell mitosis or cell death, and chemotactic and durotactic signaling from the environment (DuChez et al. 2019; Lo et al. 2000; Arrieumerlou and Meyer 2005; Parker et al. 2002; Cai et al. 2014; Mansury et al. 2002; Robertson-Tessi et al. 2015). However, these scenarios are different from those encountered during collective cell migration in 3D matrix like environments as observed during cancer metastasis. In the case of metastasis, collective cell migration occurs either as long finger-like protrusions emanating from the main tumor or as small migrating clusters of cells that separate from the main tumor (Lambert et al. 2017). This collective dissemination and migration of tumor cells enhances successful seeding at a secondary tumor site (Wang et al. 2016). Here, we aim to understand the factors that govern and promote the migration of small cell clusters in 3D matrices. Metastatic cancer cells migrate through extracellular matrices (ECMs), which can exhibit a range of different properties depending on the tissue type (Wang et al. 2016; Nasrollahi and Pathak 2016). ECM properties that play a significant role in cell migration are stiffness (Wu et al. 2014; Plotnikov et al. 2012), fiber density (Carey et al. 2012), fiber alignment (Fraley et al. 2015), ECM porosity (Ilina et al. 2011), bond density (Burgess et al. 2000), and chemical signaling (Carey et al. 2012; Fraley et al. 2015). In addition, cell properties also affect migration, such as proteolytic ability (Levental et al. 2009), cell stiffness (Lange and Fabry 2013), cell mechanoactivity (Bosgraaf and Van Haastert 2009), and expression levels of cell–cell and cell–ECM adhesion receptors (Gallant et al. 2005). Additionally, in vivo and in vitro studies have shown that collectives are led by phenotypically different “leader” cells with “follower” cells trailing behind (Friedl and Mayor 2017). Leader cells exhibit characteristics that are mesenchymal-like—softer, fluidized cells with high mechanical activity (Wolf et al. 2007; Cross et al. 2008), whereas follower cells resemble an epithelial phenotype—stiffer cells with low mechanical activity (Saez et al. 2005). The transition of cells from an epithelial to a mesenchymal phenotype (EMT) is often regarded as the proximate cause of cancer metastasis. During EMT, downregulation of adheren junctions decreases cell–cell adhesion, which when coupled with increased mechanoactivity drives cell migration (Nasrollahi and Pathak 2016). EMT is a dynamic bidirectional process that does not always run to completion (Lambert et al. 2017). As such, clusters of metastatic cancer cells may experience phenotypically dynamic states, where followers transition into leaders as directed by extracellular and inter-cellular signaling. For example, followers transition to leaders when bound to ECM integrins in the front and cell cadherins in the rear (Kato et al. 2014). Furthermore, the presence of cancer-associated fibroblasts (CAFs) promotes transitioning into a leader phenotype (An et al. 2013). CAFs increase collective cell migration by realigning local tumor environments with tube-like pathways of highly aligned fibers (Gaggioli et al. 2007). Alternatively, only partial EMT of all cells might result in the absence of well-defined leader cells (Bronsert et al. 2014).
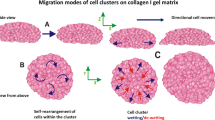
We investigate how cell clusters with or without defined leader cells migrate within different ECM environments using stochastic simulations of cell–cell and cell–ECM interactions. Our approach builds on existing in silico models (Frascoli et al. 2013; Mousavi et al. 2014; Drasdo and Hoehme 2012; Chen and Zou 2017) by adding a number of unique features as detailed below. We employ a 3D computational model that simulates long-term (> 48-h real time) cell–matrix and cell–cell interactions to track collective migration in a fiber-based 3D environment. Factors affecting migration such as bond density, fiber direction, fiber density, cell mechanoactivity, pseudopod protrusion frequency, protrusion length, active contractility, proteolytic activity, transmission of active forces and biochemical signals from leader cells to follower cells, and passive adhesive and elastic forces between cells are some of the key tunable parameters within the model. Using this model, we determine optimal migration environments by modulating the fiber density, fiber alignment, adhesion strength, and cluster size parameters. Additionally, we test two distinct cluster migration scenarios—(1) a defined leader phenotype that is maintained for the duration of the simulation (48 h) drives the cluster, while the rest of the cells are purely follower cells (Fig. 1a and Video S1), and (2) peripheral cells in contact with the matrix elements can easily switch in and out of leading phenotype (Fig. 1b and Video S2). We find that the leader and follower dynamics are an important feature for collective cell migration. While the exact mechanics of cell migration are still not well known, evidence for both these scenarios of defined one or two leaders driving small clusters (Bianco et al. 2007) or undefined leadership such as the phenotypic variability for cells in collectives undergoing EMT, embryonic development, and migration through dynamic ECM environments can be found in the literature (Friedl and Mayor 2017; Ewald et al. 2008; Jakobsson et al. 2010).
Cartoon showing two collective cell migration types. a Cluster cell migration with a defined leader where a single cell maintains the leader phenotype (blue cell) for the duration of migration. The path it migrates is traced out by the purple line. b Switching leader scenario where the cells can switch between the leader and the follower phenotype depending on their migration phase. Only one cell can have the leader phenotype at a given point in time, but the cell does not maintain this phenotype. First leader traces the purple path while the second leader traces the green path and so on
2 Methods
The proposed mathematical migration model was developed from our previous single-cell migration model (Yeoman and Katira 2018) and an altered multi-cell model proposed by Palsson (2008). Here we summarize the salient features of the combined model.
In Yeoman and Katira (2018), we presented a simulation setup to predict cell migration in 3D matrix environments as a function of cell and ECM mechanical properties such as adhesion strength, active contractility, mechanoactivity, matrix fiber density, fiber diameter distribution, fiber alignment, matrix stiffness, and the presence of chemotactic signaling. This setup is as follows: (1) Cells migrate by extending pseudopods along the length of the matrix fibers it is interacting with (Kim et al. 2015). (2) The fiber along which a new pseudopod extends is selected based on its proximity to the preferred direction of motion as determined by cell polarity and case cell shape. (3) The cell shape, which varies between elongated and rounded, is determined as a function of the alignment between the fibers the cells are in contact with, cell–matrix adhesion strength, contractile force exerted by the cell, and the stiffness of the matrix (Ahmadzadeh et al. 2017). (4) The fibers in this simulation setup are generated on an as-needed basis, stochastically, using the fiber density distribution, fiber diameter distribution, and fiber alignment angle distribution of the simulated matrix. This allows for a rapid simulation of cell–fiber interactions with a handful of fibers it is currently interacting with and allows for long-term cell migration predictions within 3D environments. (5) Gradients in fiber density or stiffness can be introduced by changing the local averages of the distributions from which new fiber and cross-links along these fibers are stochastically generated. Additional details on the single-cell stochastic migration model can be found in Yeoman and Katira (2018).
We modify this existing setup to allow for cell–cell interactions and the altered dynamics of leader and follower cells. Additionally, we calculate our cell shape differently from that described above, with shape dependent on the whether a cell is leading or following. The steps of the algorithm can be seen in the model summary Fig. 2. The starting point for this algorithm is the ECM fiber generation. Fibers are stochastically generated as needed and are populated with a random distribution of binding sites. The cells then react to the number of binding sites on the fiber they are extending a pseudopod along by entering one of three phases: retraction, outgrowth, or contraction. The cells are initialized in the retraction phase and can enter the other phases depending on the number of bonds between the pseudopod tip and fiber. During outgrowth, the cell extends its pseudopod by an incremental distance each time step, and the number of bonds between pseudopod tip and fiber is counted. If the number of bonds is above a maximum threshold, then the cell will enter contraction; if the number of bonds is below a minimum threshold, then the cell will enter retraction. Alternatively, cells will switch from outgrowth to retraction phase if the pseudopod has been extending for a certain stochastically determined time. Fiber cross-links are randomly distributed along the length of a fiber when it is stochastically generated. If a fiber cross-link is reached, then the cell is likely to continue outgrowth along the obtuse angle between the current fiber and a new stochastically generated fiber. During retraction, the current pseudopod shrinks, while a new pseudopod undergoes outgrowth. The new pseudopod may grow along the existing fiber in the reverse direction with a 20% chance or grow along a new fiber stochastically generated with an 80% chance. In the collective cell migration model, all cells can enter the outgrowth and retraction phases. However, only leader cells can enter the contraction phase if the growing pseudopod encounters enough binding sites. Cells are assigned as followers when another cell in the cluster has become a leader. Determination of the leader cell phenotype is described later in this section. During the contraction phase, an active force, \({\mathbf {F}}_i^{\mathrm{act}}\), is generated along the pseudopod vector. The pseudopod contracts in length, dragging the cell center forward under the action of the active force. Active migration force is a function of the number of adhesions between the pseudopod and the fiber it is attached to, the matrix stiffness, and cell contractility (Yeoman and Katira 2018). As the cell center moves forward, the pseudopod length decreases, and when the pseudopod length reaches zero (or a minimum threshold), the cell enters retraction.
2.1 Modeling cell–cell interactions
Cell–cell interaction forces, section A in Fig. 2, are determined by calculating the distance from one cell center to every other cell center, or \(r_{ij}\). If \(r_{ij}\le d_{\mathrm{break}}\) then the cells are considered to be in contact and passive forces will be calculated between cells i and j. A 2D representation of the adhesive and compressive passive forces can be seen in Fig. 3. The equations used to compute the passive forces are (Palsson 2008):
The scalar passive force, \(F_{ij}^{\mathrm{pass}}\), is continuous at \(x=0\), and, depending on \(x_{ij}\), the passive force is either positive or negative. \(F_{\mathrm{compressive}}\) is the passive force pushing two cells away from each other, and \(F_{\mathrm{adhesive}}\) pulls cells toward one another. \(F_{\mathrm{comp}}\) is a compressive force constant, \(\chi\) is an orientation factor, \(x_{ij}\) is an adjusted cell–cell distance factor. \(F_{\mathrm{adh}}\) is an adhesive force constant, \(x_0\) and \(v_0\) are constants for continuity, and \(\lambda\) is a strength constant.
The orientation factor is solved with:
\(r_{\mathrm{cell}}\) is the average cell radius, \(d_i\) and \(d_j\) is the distance from cell center to cell membrane along the \({\mathbf {r}}_{ij}\) unit vector.
The calculation for \(x_{ij}\) is:
\(d_{ij}\) is the distance from cell membrane to cell membrane along the \({\mathbf {r}}_{ij}\) unit vector, \({\hbox {min}}_{\mathrm{dist}}\) is a value derived from the minimum radius from packing deformed incompressible ellipsoids into a fixed space and that comes out to \({\hbox {min}}_{\mathrm{dist}}\approx -\,0.1*r_{\mathrm{cell}}\).
The calculation for the constants \(x_0\) and \(v_0\) is:
The following equation determines the passive force vector with:
The magnitude of the force is multiplied by the unit vector from the center of cell i to cell j, and \({\mathbf {F}}_{ij}^{\mathrm{pass}}\) is the passive force between cell i and cell j.
The net force per cell, \({\mathbf {F}}_i^{\mathrm{net}}\), is calculated here:
\({\mathbf {F}}_i^{\mathrm{act}}\) is the force vector generated during the contraction phase of a cell.
This force is determined with, \(l_{0,i}\), the protrusion length of a cell’s extending pseudopod, \(F_0\), the maximum contractile force in the pseudopod, and is multiplied by the extending pseudopod’s unit vector. \({\mathbf {p}}_i\) is the vector for cell i’s extending pseudopod. The contractile force, \(F_0\), is calculated here:
where \(n_{\mathrm{b}}\) is the number of bonds between the pseudopod and ECM fiber, and \(n_{\frac{1}{2}}\), is the cell–ECM bond density at which the generated force is half of \(F_{\mathrm{max}}\).
The drag force per cell, \({\mathbf {F}}_i^{\mathrm{D}}\), is based on the cell–cell common surface area and cell–matrix common surface area, and is calculated here:
The first term on the right is the drag force from cell–bond interactions of the ECM, and the second term on the right is the drag force from cells moving past one another. The constant \(\mu _{\mathrm{s}}\) is the viscosity coefficient for cell–matrix interactions, \(\mu _{\mathrm{c}}\) is the viscosity coefficient for cell–cell interactions, A is the total surface area of the cell, \(A_{is}\) and \(A_{ij}\) are the common surface areas between the cell and the matrix, and cell and cell, respectively. \(j\in {N(i)}\) denotes that all neighbors of i are j, so the summation is only for cells in contact of cell i. \({\mathbf {v}}_i\) and \({\mathbf {v}}_j\) are the vector velocities of cells i and j, respectively.
The calculation for common surface area between cells is:
sf is a surface factor and was left at 1 for simplicity, found in Palsson (2001).
The velocity of each cell is determined from the following sets of equations:
After expanding out the drag force equation for each cell, the values for each drag coefficient can be grouped into either \({\mathrm {SE}}_{ii}\) or \({\mathrm {SN}}_{ij}\), with one being the friction surrounding cell i or the friction between cells i and j, respectively. Matrix division is used to solve for the velocity. The proposed model for \({\mathrm {SE}}_{ij}\) substitutes in \(f_{\mathrm{b}}\) and \(f_{\mathrm{v}}\) for the frictional calculation in Palsson (2008) because using a friction coefficient based on the number of bonds, rather than a viscosity-based coefficient from cells moving past ECM seems more appealing when considering that the bond information is readily available in the model. \(f_{\mathrm{b}}\) is an adjusted frictional component from Yeoman and Katira (2018) with the removal of an exponential factor. \(n_{\mathrm{br}}^i\) is the number of bonds made between a cell and ECM fibers, \(k_{\mathrm{ECM}}\) is an ECM spring constant, \(k_i\) is the cell–matrix bond stiffness, and \(k_{\mathrm{off}}\) is a cell–matrix dissociation rate. \(f_{\mathrm{v}}\) is a viscous frictional component, \(\eta\) is the ECM viscosity, and \(k_{\mathrm{prime}}\) is a drag adjustment factor.
The change in position \({\mathbf {r}}_i\) of the cell center of cell i is then simply obtained by:
where \({{\Delta } t}\) is the simulation time step. (Optimal time step of 2 s determined previously in Yeoman and Katira (2018) is used.)
2.2 Quantifying migration
The algorithm shown in Fig. 2 and the equations described above are used to calculate the position of every cell at every time step. The overall center of mass of the cluster is tracked and its trajectory is analyzed using different approaches as it migrates (Dickinson and Tranquillo 1993). The four main characteristics evaluated are mean squared displacement (MSD), cluster speed, persistence length, and lifetime. MSD is obtained between non-overlapping points at specific time intervals along the trajectory. We fit this MSD \(\langle R^2\rangle\) vs time interval data to find the motility coefficient, \(\mu\), and exponent, \(\alpha\) (Yeoman and Katira 2018). Using the motility coefficient and the exponent, an effective MSD of that particular cluster over the cluster’s lifetime \(T_L\) is back calculated using:
This is repeated for 10 instances of cluster migration, with every combination of environmental and cell–cell adhesion parameter tested. Sample MSD for cluster trajectories are shown in Supplementary Figures S1 and S2.
Cluster speed is calculated by averaging the instantaneous velocity of each cell in the cluster at each time step. We use cluster speed to show us how quickly a cluster is migrating through the ECM. Lifetime is the simulated time that it takes for a single cell to break away from the cluster. We use this as a metric for determining how well a cluster stays together in certain ECM conditions. Persistence length indicates the distance over which a cluster maintains its directionality of migration. Persistence length \(L_{\mathrm{P}}\) was calculated by fitting MSD vs contour length data to Eq. 19 using a nonlinear least squares regression:
The maximum contour length L used was 10 \(\upmu {\hbox {m}}\). High persistence values indicate that the clusters are migrating with little change in a particular direction of travel, while low values tell us that the cluster changes the direction often.
2.3 Leader scenarios
Clusters of cells are driven by two different scenarios—defined single leader and switching leaders. For a defined leader, the same cell continues as the leader throughout the simulation, while the remaining cells are labeled as followers. For the switching leader scenario, there is a set of rules in place to select the leader during each time step. Any cell can be the leader, but only one leader can be active at a time. The leader cell is selected when a cell enters contraction before any other cell in the cluster. If two or more cells enter contraction during a time step, then leadership is randomized. If the current leader and another cell enter contraction during a time step, then the current leader keeps leadership. Followers are able to enter retraction and outgrowth, but cannot enter contraction. Followers also take on a spherical shape, while leaders have an ellipsoidal front and spherical rear.
2.4 Generation of the 2-phase optimal migration plots
Figure 5a–d shows a heat map generated by a thin-plate spline fit to the data in Supplementary Figures S3 and S4. However, only the migration strategy with the higher MSD is plotted on the heat map to show which behavior dominates in any given region, with blue representing cluster migration strategy with a defined leader and red representing cluster migration strategy with switching leaders.
2.5 Model validation
The model parameters are obtained from previously published experimental and theoretical studies (Table 1). The model equations are also based on the previously published theoretical work. The model predicts cluster cell migration speeds on the order of 1–50 \(\upmu {\hbox {m}}/{\mathrm {h}}\) , and clusters migrate distances on the order of a few hundred micrometers without breaking. These values seem reasonable (Friedl et al. 1995; Carmona-Fontaine et al. 2008; Cai et al. 2016). Additionally, the model predicts biphasic relationship for MSD, speed, and persistence length as a function of fiber density, which is consistently observed in typical single-cell migration studies (Burgess et al. 2000; Zaman et al. 2006; Palecek et al. 1997; DiMilla et al. 1991; Gaudet et al. 2003). As shown in Fig. 4 and Supplementary Figures S3 and S4, our model predicts these biphasic trends for migration with both, defined leader and switching leader strategies. Beyond this, it has been hard to find quantitative cluster cell migration data in the literature to validate other key predictions of our model.
Effect of fiber density on MSD, cluster lifetime, cluster speed, and persistence length for various cluster sizes for a defined leader cluster migration strategy. a MSD as a function of fiber density. b Cluster lifetime as a function of fiber density. c Cluster speed as a function of fiber density. d Persistence as a function of fiber density. Blue line is a 3-cell cluster, green line is a 5-cell cluster, and red line is a 10-cell cluster. Fiber density was increased linearly from \(0.5\times 10^{-3}\) to \(2\times 10^{-3}\,{\mathrm {fibers}}/\upmu {\hbox {m}}^3\). Cell–cell adhesion was set to moderate (50 nN cell–cell dissociation force). Ten simulations for each scenario were run at each fiber density and run for 48 h of simulated time or until cluster dissociation. Error bars represent ± SEM
3 Results
The primary result of our model is that for different levels of cell–cell adhesion, fiber alignment, cluster size, and either of the two cluster migration scenarios described above, cell migration distance shows a biphasic relationship with fiber density (Figs. S3 and S4). There seems to be an optimal ECM density ideal for cluster cell migration for every different combination of environmental and cellular conditions. This is along the lines of the biphasic relationship reported previously between migration distance and ECM density for single-cell migration. More importantly, our results show that for cluster cell migration, there are unique environmental and cellular conditions where one of the two migration strategies clearly outperforms the other (Fig. 5). (1) Cluster cell migration with a defined leader cell (migration scenario 1, blue region in Fig. 5) is ideal for clusters with high cell–cell adhesion migrating in high fiber density environments and preferable fiber alignment. (2) Cluster cell migration where the cells switch between leader and follower phenotypes (migration scenario 2, red region in Fig. 5) is ideal for clusters with low-to-intermediate cell–cell adhesion migrating in low fiber density environments.
Heat maps derived from thin-plate spline fit to MSD collected at 3 cell–cell adhesion levels and 5 fiber density levels. a Five-cell cluster with no fiber alignment. b Five-cell cluster with fiber alignment. c Ten-cell cluster with no fiber alignment. d Ten-cell cluster with high fiber alignment. Red regions are where clusters with switching leader phenotype have the highest MSD; blue regions are where clusters with a defined leader have the highest MSD
Analyzing the results in further detail, when inter-cellular adhesion is low, corresponding to low cadherin expression (with a cell–cell separation force of \(\sim\) 25 nN) (Chu et al. 2004), the MSD is significantly higher for clusters able to switch between leaders (cluster migration scenario 2) in both aligned and unaligned low-density matrices (Fig. 6a, red and orange lines). The radar plots in Fig. 6b, d, f show how cluster speed, persistence, and lifetime are also affected by the fiber density and cell–cell adhesion strength. When cell–cell adhesion is low in low fiber density environments, cluster lifetime and persistence are similar for both leading scenarios in aligned and unaligned matrices (Fig. 6b). Persistence is about twice as high for clusters with a defined leader in aligned matrices, but the higher cluster speed for clusters with no defined leader is greater by more than a factor of 4, leading to a higher migratory efficiency (Fig. 6b). In less dense environments, the ability to switch between leaders allows the cluster to overcome the sparsity of binding sites and probe a larger region of space. The cluster can more quickly find fibers with sufficient binding sites for force generation and displacement. Interestingly, the ability to switch between leaders also helps redistribute the migration forces evenly between the loosely bound cells, allowing the cells to stay clustered together even in high-density environments. We attribute this to a herding effect, which can be enhanced by increasing the number of cells in the cluster, thereby increasing the lifetime of the cluster (Fig. S5A). On the other hand, when there is single leader driving the cluster of loosely bound cells in high-density environments, the leader cell is more likely to break off quickly from the main cluster due to a buildup of migration forces between the cell–cell interface of the leader and follower cells (Fig. S5B). (Figure 6d shows the difference in lifetimes of loosely bound clusters migrating in high-density environments for both migration scenarios.)
As inter-cellular adhesion increases, cell migration with a single defined leader becomes more advantageous in denser matrices (Fig. 6c dark and light blue lines). Cluster speed is higher for clusters with undefined leaders for the same reasons as in less dense matrices, but the persistence decreases due to an increased probability of changing direction as the leaders switch between peripheral cells (Fig. 6d). A single leader in these cases allows for a more persistent migration, while the higher adhesion between the cells allows for cluster to hold up against the increased migration forces.
Effect of fiber density and cell–cell adhesion on MSD. a MSD for low cell–cell adhesion. b Radar plot showing values of persistence length, cluster lifetime, and cluster speed at low fiber density and low cell–cell adhesion. c MSD for mid cell–cell adhesion. d Radar plot at high fiber density and low adhesion. e MSD for high cell–cell adhesion. f Radar plot for high fiber density and high cell–cell adhesion. Blue toned lines are clusters with a defined leader; red toned lines are clusters with an undefined leader. Solid lines are in an unaligned ECM and dashed in an aligned ECM. Fiber density was increased linearly from \(0.5 \times 10^{-3}\) to \(2 \times 10^{-3}\) \({\mathrm {fibers}}/\upmu {\hbox {m}}^3\). Ten simulations for each scenario were run at each fiber density and run for 48 h of simulated time or until a single cell dissociated from the cluster. Error bars represent ± SEM
When the cell–cell separation force exceeds 50 nN, cluster migration driven by a single defined leader becomes even more advantageous in matrices with high fiber density (Fig. 6e). Single leader clusters migrate more persistently, especially with highly aligned fibers, through matrices with high fiber density and therefore have a higher MSD (Fig. 6f). When the cluster’s leader is undefined, the cluster’s migration direction will change every time the leading cell switches, thus reducing the migratory efficiency (Fig. 6f). Interestingly, cluster migration driven by switching leaders loses its edge in keeping the cluster together in high-density environments as well (Fig. 6f). Because all the peripheral cells can generate migratory forces, the likelihood that one of them generates strong enough forces to rip it apart from its neighbors goes up as compared to the case where only one leader cell is dragging the cluster behind it.
The single leader migration scenario is more suited to take advantage of fiber alignment, especially for high adhesion strength clusters in high-density environments (Fig. 6a, e). This is again because having a single leader allows the cluster to migrate more persistently along aligned fibers (Fig. 6b, d, f). However, at intermediate and low adhesion strengths, the fiber alignment in high-density environments stretches out the cluster more along the persistent migration path, straining the contact between the leader and the follower cells. This increases the likelihood of the cluster disintegrating, lowering the cluster lifetime and overall migration distance (Fig. 6c).
Overall, the smaller five-cell clusters have a higher MSD than the larger 10-cell clusters because they experience less drag as they migrate through the ECM. Cluster speed increases with increased cell–cell adhesion because the cluster becomes more compact as the trailing cells migrate closer to the leading cell. Regardless of cluster size, clusters with an undefined leader have a greater migratory efficiency in regions of low fiber density at any cell–cell adhesion strength, in both aligned and unaligned ECMs. Having a defined leader is favored when both adhesion and fiber density are high, and this type of migration is enhanced when the fibers are aligned.
4 Discussion
Clusters that are driven by a defined leader cell (scenario 1) are akin to animal foraging behaviors commonly seen in certain bees, ants, and fish species (Reebs 2000; Couzin et al. 2005; Sumpter 2006), where a single or a few leaders act as catalysts for coordinating directionality. A minority of defined leaders within a collective can enhance group movement in a single direction. However, this requires signal transmission (chemical, mechanical or otherwise) from the leader(s) to the rest of the collective and can become less effective as the overall size of the collective grows. Alternatively, clusters driven by switching leaders between all peripheral cells (scenario 2) are akin to herding that can give rise to self-organization in flocks of starlings or schools of fish (Sumpter 2006; Cavagna et al. 2010; Goodenough et al. 2017). Neighbor mimicking and distributed decision making are two behavior patterns reminiscent of herding. Neighbor mimicking helps align the motion of individuals, leading to more cohesive moments that helps maintain the integrity of mass-migrating groups (Buhl et al. 2006). Distributed decision making on the other hand can allow the collective to evaluate and choose from alternatives to increase migration efficiency (Mallon 2001).
To our knowledge, this study is the first to show that under certain environmental conditions, leading- and herding-like behaviors that are similar to the self-organizing, active systems seen throughout the animal kingdom can differentially dominate and govern optimal strategies in relatively small collectives of cells migrating through a 3D ECM. In extracellular matrices where fibrous proteins are scarce, for clusters with the ability to switch between leaders, multiple cells can probe the environment to overcome the scarcity of binding sites. By sharing the role of finding a sufficient number of binding sites for displacement, the cluster spends less time searching than a cluster with a single leader. In fiber dense environments, the more strategic behavior is dependent on cell–cell adhesion strength and the force required to separate two cells. For clusters with low cell–cell adhesion, herding occurs when leadership can switch between cells. The peripheral cells in these clusters nudge their neighbors toward the cluster’s center, generating compressive forces between neighboring cells. This helps align their motion to their nearest neighbor, leading to more cohesive movements that help maintain the cluster stability and extend the cluster lifetime. The advantages of defined leadership only become apparent when cell–cell adhesion is high enough, and motion of the leader can be transmitted farther along the follower cells effectively. High adhesion greatly improves the migratory efficiency in dense ECM by allowing a single leader to maintain its directional persistence for longer, especially in environments with high fiber alignment.
Our results show that collective cell migration is possible for significant distances, even when cells only weakly couple with each other. Under these conditions where cluster dissociation and single-cell migration would be expected, collective migration can be maintained by herding-like behavior. Because small clusters of cancer cells are more likely to establish secondary tumor sites, herding and self-organization of migrating cancer cells may increase metastatic potential in unfavorable environmental conditions. Furthermore, collective cell migration is also possible and may be beneficial in low fiber density environments. In these environments, cells in a cluster are together able to probe a larger volume of the ECM for binding sites than an individual cell, and therefore, decision making for movement is faster.
The results presented here are based on cells displaying a predefined migratory phenotype. For example, in a single defined leader scenario, the leader cell may be a cell that has undergone EMT and pulls along a group of cells that have remained epithelial, whereas if all cells have undergone partial EMT, they can switch between different leaders. However, the model presented here also provides a platform to examine how genotypic and phenotypic changes can alter individual and collective cell migration and examine whether these changes are specifically targeted to promote any particular migration scenario to suit a particular outcome. Studies have shown that genetic regulators can activate partial EMT and collective cell migration during metastases in Drosophila intestinal tumors (Campbell et al. 2017). Cells are known to downregulate cell–cell adhesion proteins during EMT (Nasrollahi and Pathak 2016), so the extent of this regulation may be important for the metastatic potential of a tumor depending on the properties of the extracellular environment around the tumor (VanderVorst et al. 2019). Other recent experimental studies have also shown that the matrix architectural context can drive phenotypic changes in cellular phenotype that influences migratory behavior (Velez et al. 2017; Morris et al. 2016). In the future, we hope to couple the model presented here with intra- and extracellular signaling-driven temporal changes in cellular genotype, phenotype and consequently mechanotype to examine how migrating cellular collectives may adapt to different extracellular environments.
5 Conclusions
We present a model that can simulate collective cell migration long term in 3D with both cell–cell and cell–ECM interactions. Although validation of the model is limited due to the difficulty of performing 3D cluster migration assays, we believe that this physics-based approach works well as an accurate predictive tool for experimental research. Furthermore, the model allows for perturbations to be easily introduced for several parameters affecting cell and ECM properties to study how cellular clusters might optimize the leader–follower dynamics to better adapt for movement through their given environment. Leader–follower dynamics play an important role in any form of collective migration and further research to interrogate which cell types and genetic regulators give rise to the different leading scenarios could present new targets for inhibiting cancer metastasis. Our results highlight some of the migratory phenotypes that should be looked into for in vitro and clinical settings and present possible prognostic phenotypes that could be identified prior to aggressive cancer treatments.
References
Abraham VC, Krishnamurthi V, Taylor DL, Lanni F (1999) The actin-based nanomachine at the leading edge of migrating cells. Biophys J 77(3):1721–1732
Ahmadzadeh H, Webster MR, Behera R, Jimenez Valencia AM, Wirtz D, Weeraratna AT, Shenoy VB (2017) Modeling the two-way feedback between contractility and matrix realignment reveals a nonlinear mode of cancer cell invasion. Proc Natl Acad Sci U S A 114(9):E1617–E1626. https://doi.org/10.1073/pnas.1617037114
Alexander S, Koehl GE, Hirschberg M, Geissler EK, Friedl P (2008) Dynamic imaging of cancer growth and invasion: a modified skin-fold chamber model. Histochem Cell Biol 130(6):1147–54. https://doi.org/10.1007/s00418-008-0529-1
An J, Enomoto A, Weng L, Kato T, Iwakoshi A, Ushida K, Maeda K, Ishida-Takagishi M, Ishii G, Ming S, Sun T, Takahashi M (2013) Significance of cancer-associated fibroblasts in the regulation of gene expression in the leading cells of invasive lung cancer. J Cancer Res Clin Oncol 139(3):379–388. https://doi.org/10.1007/s00432-012-1328-6
Ananthakrishnan R, Ehrlicher A (2007) The forces behind cell movement. Int J Biol Sci 3(5):303
Arima S, Nishiyama K, Ko T, Arima Y, Hakozaki Y, Sugihara K, Koseki H, Uchijima Y, Kurihara Y, Kurihara H (2011) Angiogenic morphogenesis driven by dynamic and heterogeneous collective endothelial cell movement. Development 138(21):4763–76. https://doi.org/10.1242/dev.068023
Arrieumerlou C, Meyer T (2005) A local coupling model and compass parameter for eukaryotic chemotaxis. Dev Cell 8(2):215–227. https://doi.org/10.1016/j.devcel.2004.12.007
Bianco A, Poukkula M, Cliffe A, Mathieu J, Luque CM, Fulga TA, Rørth P (2007) Two distinct modes of guidance signalling during collective migration of border cells. Nature 448(7151):362–365. https://doi.org/10.1038/nature05965
Bosgraaf L, Van Haastert PJM (2009) The ordered extension of pseudopodia by amoeboid cells in the absence of external cues. PLoS ONE 4(4):e5253. https://doi.org/10.1371/journal.pone.0005253
Bronsert P, Enderle-Ammour K, Bader M, Timme S, Kuehs M, Csanadi A, Kayser G, Kohler I, Bausch D, Hoeppner J, Hopt U, Keck T, Stickeler E, Passlick B, Schilling O, Reiss C, Vashist Y, Brabletz T, Berger J, Lotz J, Olesch J, Werner M, Wellner U (2014) Cancer cell invasion and EMT marker expression: a three-dimensional study of the human cancer-host interface: 3d cancer-host interface. J Pathol 234(3):410–422. https://doi.org/10.1002/path.4416
Bruinsma R (2005) Theory of force regulation by nascent adhesion sites. Biophys J 89(1):87–94
Buhl J, Sumpter DJT, Couzin ID, Hale JJ, Despland E, Miller ER, Simpson SJ (2006) From disorder to order in marching locusts. Science 312(5778):1402–6. https://doi.org/10.1126/science.1125142
Burgess BT, Myles JL, Dickinson RB (2000) Quantitative analysis of adhesion-mediated cell migration in three-dimensional gels of RGD-grafted collagen. Ann Biomed Eng 28(1):110–118
Cai D, Chen SC, Prasad M, He L, Wang X, Choesmel-Cadamuro V, Sawyer J, Danuser G, Montell D (2014) Mechanical feedback through E-cadherin promotes direction sensing during collective cell migration. Cell 157(5):1146–1159. https://doi.org/10.1016/j.cell.2014.03.045
Cai D, Dai W, Prasad M, Luo J, Gov NS, Montell DJ (2016) Modeling and analysis of collective cell migration in an in vivo three-dimensional environment. Proc Natl Acad Sci USA 113(15):E2134. https://doi.org/10.1073/pnas.1522656113
Campbell JJ, Husmann A, Hume RD, Watson CJ, Cameron RE (2017) Development of three-dimensional collagen scaffolds with controlled architecture for cell migration studies using breast cancer cell lines. Biomaterials 114:34–43. https://doi.org/10.1016/j.biomaterials.2016.10.048
Carey SP, Kraning-Rush CM, Williams RM, Reinhart-King CA (2012) Biophysical control of invasive tumor cell behavior by extracellular matrix microarchitecture. Biomaterials 33(16):4157–65. https://doi.org/10.1016/j.biomaterials.2012.02.029
Carmona-Fontaine C, Matthews HK, Kuriyama S, Moreno M, Dunn GA, Parsons M, Stern CD, Mayor R (2008) Contact inhibition of locomotion in vivo controls neural crest directional migration. Nature 456(7224):957–961. https://doi.org/10.1038/nature07441
Cavagna A, Cimarelli A, Giardina I, Parisi G, Santagati R, Stefanini F, Viale M (2010) Scale-free correlations in starling flocks. Proc Natl Acad Sci USA 107(26):11865–11870. https://doi.org/10.1073/pnas.1005766107
Chen Z, Zou Y (2017) A multiscale model for heterogeneous tumor spheroid in vitro. Math Biosci Eng 15(2):361–392. https://doi.org/10.3934/mbe.2018016
Chu YS, Thomas WA, Eder O, Pincet F, Perez E, Thiery JP, Dufour S (2004) Force measurements in E-cadherin-mediated cell doublets reveal rapid adhesion strengthened by actin cytoskeleton remodeling through Rac and Cdc42. J Cell Biol 167(6):1183–1194. https://doi.org/10.1083/jcb.200403043
Cooper G (2007) The eukaryotic cell cycle. In: Cooper GM, Hausman RE (eds) The cell: a molecular approach. ASM Press, Washington, DC
Couzin ID, Krause J, Franks NR, Levin SA (2005) Effective leadership and decision-making in animal groups on the move. Nature 433(7025):513–516. https://doi.org/10.1038/nature03236
Cross SE, Jin YS, Tondre J, Wong R, Rao J, Gimzewski JK (2008) AFM-based analysis of human metastatic cancer cells. Nanotechnology 19(38):384003. https://doi.org/10.1088/0957-4484/19/38/384003
Dickinson RB, Tranquillo RT (1993) Optimal estimation of cell movement indices from the statistical analysis of cell tracking data. AIChE J 39(12):1995–2010. https://doi.org/10.1002/aic.690391210
DiMilla P, Barbee K, Lauffenburger D (1991) Mathematical model for the effects of adhesion and mechanics on cell migration speed. Biophys J 60(1):15–37. https://doi.org/10.1016/S0006-3495(91)82027-6
Drasdo D, Hoehme S (2012) Modeling the impact of granular embedding media, and pulling versus pushing cells on growing cell clones. New J Phys 14(5):055025. https://doi.org/10.1088/1367-2630/14/5/055025
Du Roure O, Saez A, Buguin A, Austin RH, Chavrier P, Siberzan P, Ladoux B (2005) Force mapping in epithelial cell migration. Proc Natl Acad Sci 102(7):2390–2395
DuChez BJ, Doyle AD, Dimitriadis EK, Yamada KM (2019) Durotaxis by human cancer cells. Biophys J 116(4):670–683. https://doi.org/10.1016/j.bpj.2019.01.009
Erdmann T, Schwarz US (2006) Bistability of cell-matrix adhesions resulting from nonlinear receptor-ligand dynamics. Biophys J 91(6):L60–L62
Ewald AJ, Brenot A, Duong M, Chan BS, Werb Z (2008) Collective epithelial migration and cell rearrangements drive mammary branching morphogenesis. Dev Cell 14(4):570–581. https://doi.org/10.1016/j.devcel.2008.03.003
Fraley SI, Wu PH, He L, Feng Y, Krisnamurthy R, Longmore GD, Wirtz D (2015) Three-dimensional matrix fiber alignment modulates cell migration and MT1-MMP utility by spatially and temporally directing protrusions. Sci Rep 5(January):14580. https://doi.org/10.1038/srep14580
Frascoli F, Hughes BD, Zaman MH, Landman KA (2013) A computational model for collective cellular motion in three dimensions: general framework and case study for cell pair dynamics. PloS ONE 8(3):e59249. https://doi.org/10.1371/journal.pone.0059249
Friedl P, Mayor R (2017) Tuning collective cell migration by cell–cell junction regulation. Cold Spring Harb Perspect Biol 9(4):1146–1159. https://doi.org/10.1101/cshperspect.a029199
Friedl P, Noble PB, Walton PA, Laird DW, Chauvin PJ, Tabah RJ, Black M, Zanker KS (1995) Migration of coordinated cell clusters in mesenchymal and epithelial cancer explants in vitro. Cancer Res 55(20):4557–4560
Gaggioli C, Hooper S, Hidalgo-Carcedo C, Grosse R, Marshall JF, Harrington K, Sahai E (2007) Fibroblast-led collective invasion of carcinoma cells with differing roles for RhoGTPases in leading and following cells. Nat Cell Biol 9(12):1392–400. https://doi.org/10.1038/ncb1658
Gallant ND, Michael KE, García AJ (2005) Cell adhesion strengthening: contributions of adhesive area, integrin binding, and focal adhesion assembly. Mol Biol Cell 16(9):4329–40. https://doi.org/10.1091/mbc.e05-02-0170
Garcia S, Hannezo E, Elgeti J, Joanny JF, Silberzan P, Gov NS (2015) Physics of active jamming during collective cellular motion in a monolayer. Proc Natl Acad Sci USA 112(50):15314–15319. https://doi.org/10.1073/pnas.1510973112
Gaudet C, Marganski WA, Kim S, Brown CT, Gunderia V, Dembo M, Wong JY (2003) Influence of type I collagen surface density on fibroblast spreading, motility, and contractility. Biophys J 85(5):3329
Gillitzer R, Goebeler M (2001) Chemokines in cutaneous wound healing. J Leukoc Biol 69(4):513–21
Goodenough AE, Little N, Carpenter WS, Hart AG (2017) Birds of a feather flock together: insights into starling murmuration behaviour revealed using citizen science. PLoS ONE 12(6):1–18. https://doi.org/10.1371/journal.pone.0179277
Harjanto D, Zaman MH (2013) Modeling extracellular matrix reorganization in 3d environments. PloS ONE 8(1):e52509
Ilina O, Bakker GJ, Vasaturo A, Hofmann RM, Friedl P (2011) Two-photon laser-generated microtracks in 3d collagen lattices: principles of MMP-dependent and -independent collective cancer cell invasion. Phys Biol 8(1):015010. https://doi.org/10.1088/1478-3975/8/1/015010
Jakobsson L, Franco CA, Bentley K, Collins RT, Ponsioen B, Aspalter IM, Rosewell I, Busse M, Thurston G, Medvinsky A, Schulte-Merker S, Gerhardt H (2010) Endothelial cells dynamically compete for the tip cell position during angiogenic sprouting. Nat Cell Biol 12(10):943–953. https://doi.org/10.1038/ncb2103
Kato T, Enomoto A, Watanabe T, Haga H, Ishida S, Kondo Y, Furukawa K, Urano T, Mii S, Weng L, Ishida-Takagishi M, Asai M, Asai N, Kaibuchi K, Murakumo Y, Takahashi M (2014) TRIM27/MRTF-B-dependent integrin \(\beta\)1 expression defines leading cells in cancer cell collectives. Cell Rep 7(4):1156–1167. https://doi.org/10.1016/j.celrep.2014.03.068
Kim MC, Whisler J, Silberberg YR, Kamm RD, Asada HH (2015) Cell invasion dynamics into a three dimensional extracellular matrix fibre network. PLoS Comput Biol. https://doi.org/10.1371/journal.pcbi.1004535
Knutsdottir H, Condeelis JS, Palsson E (2016) 3-D individual cell based computational modeling of tumor cell-macrophage paracrine signaling mediated by EGF and CSF-1 gradients. Integr Biol Quant Biosci Nano Macro 8(1):104–19. https://doi.org/10.1039/c5ib00201j
Lambert AW, Pattabiraman DR, Weinberg RA (2017) Emerging biological principles of metastasis. Cell 168(4):670–691. https://doi.org/10.1016/j.cell.2016.11.037
Lange JR, Fabry B (2013) Cell and tissue mechanics in cell migration. Exp Cell Res 319(16):2418–23. https://doi.org/10.1016/j.yexcr.2013.04.023
Levental KR, Yu H, Kass L, Lakins JN, Egeblad M, Erler JT, Fong SFT, Csiszar K, Giaccia A, Weninger W, Yamauchi M, Gasser DL, Weaver VM (2009) Matrix crosslinking forces tumor progression by enhancing integrin signaling. Cell 139(5):891–906. https://doi.org/10.1016/j.cell.2009.10.027
Li F, Redick SD, Erickson HP, Moy VT (2003) Force measurements of the \(\alpha\)5\(\beta\)1 integrin–fibronectin interaction. Biophy J 84(2):1252–1262
Lo CM, Wang HB, Dembo M, Yl Wang (2000) Cell movement is guided by the rigidity of the substrate. Biophys J 79(1):144–152. https://doi.org/10.1016/S0006-3495(00)76279-5
Lusche DF, Wessels D, Soll DR (2009) The effects of extracellular calcium on motility, pseudopod and uropod formation, chemotaxis, and the cortical localization of myosin II in Dictyostelium discoideum. Cell Motil Cytoskelet 66(8):567–587. https://doi.org/10.1002/cm.20367
Mallon PSFNE (2001) Individual and collective decision-making during nest site selection by the ant Leptothorax albipennis. Behav Ecol Sociobiol 50(4):352–359. https://doi.org/10.1007/s002650100377
Mansury Y, Kimura M, Lobo J, Deisboeck TS (2002) Emerging patterns in tumor systems: simulating the dynamics of multicellular clusters with an agent-based spatial agglomeration model. J Theor Biol 219(3):343–370. https://doi.org/10.1006/jtbi.2002.3131
Morris BA, Burkel B, Ponik SM, Fan J, Condeelis JS, Aguirre-Ghiso JA, Castracane J, Denu JM, Keely PJ (2016) Collagen matrix density drives the metabolic shift in breast cancer cells. EBioMedicine 13:146–156. https://doi.org/10.1016/j.ebiom.2016.10.012
Mousavi SJ, Doweidar MH, Doblaré M (2014) Computational modelling and analysis of mechanical conditions on cell locomotion and cell–cell interaction. Comput Methods Biomech Biomed Eng 17(6):678–93. https://doi.org/10.1080/10255842.2012.710841
Munjal A, Lecuit T (2014) Actomyosin networks and tissue morphogenesis. Development 141(9):1789–93. https://doi.org/10.1242/dev.091645
Nasrollahi S, Pathak A (2016) Topographic confinement of epithelial clusters induces epithelial-to-mesenchymal transition in compliant matrices. Sci Rep 6(January):18831. https://doi.org/10.1038/srep18831
Palamidessi A, Malinverno C, Frittoli E, Corallino S, Barbieri E, Sigismund S, Beznoussenko GV, Martini E, Garre M, Ferrara I, Tripodo C, Ascione F, Cavalcanti-Adam EA, Li Q, Di Fiore PP, Parazzoli D, Giavazzi F, Cerbino R, Scita G (2019) Unjamming overcomes kinetic and proliferation arrest in terminally differentiated cells and promotes collective motility of carcinoma. Nat Mater 18(11):1252–1263. https://doi.org/10.1038/s41563-019-0425-1
Palecek SP, Loftus JC, Ginsberg MH, Lauffenburger DA, Horwitz AF (1997) Integrin-ligand binding properties govern cell migration speed through cell-substratum adhesiveness. Nature 385(6616):537–540. https://doi.org/10.1038/385537a0
Palsson E (2001) A three-dimensional model of cell movement in multicellular systems. Future Gener Comput Syst 17(7):835–852. https://doi.org/10.1016/S0167-739X(00)00062-5
Palsson E (2008) A 3-D model used to explore how cell adhesion and stiffness affect cell sorting and movement in multicellular systems. J Theor Biol 254(1):1–13. https://doi.org/10.1016/j.jtbi.2008.05.004
Parker KK, Brock AL, Brangwynne C, Mannix RJ, Wang N, Ostuni E, Geisse NA, Adams JC, Whitesides GM, Ingber DE (2002) Directional control of lamellipodia extension by constraining cell shape and orienting cell tractional forces. FASEB J 16(10):1195–1204. https://doi.org/10.1096/fj.02-0038com
Petitjean L, Reffay M, Grasland-Mongrain E, Poujade M, Ladoux B, Buguin A, Silberzan P (2010) Velocity fields in a collectively migrating epithelium. Biophys J 98(9):1790–1800. https://doi.org/10.1016/j.bpj.2010.01.030
Plotnikov SV, Pasapera AM, Sabass B, Waterman CM (2012) Force fluctuations within focal adhesions mediate ECM-rigidity sensing to guide directed cell migration. Cell 151(7):1513–27. https://doi.org/10.1016/j.cell.2012.11.034
Reebs (2000) Can a minority of informed leaders determine the foraging movements of a fish shoal? Anim Behav 59(2):403–409. https://doi.org/10.1006/anbe.1999.1314
Robertson-Tessi M, Gillies RJ, Gatenby RA, Anderson ARA (2015) Impact of metabolic heterogeneity on tumor growth, invasion, and treatment outcomes. Cancer Res 75(8):1567–1579. https://doi.org/10.1158/0008-5472.CAN-14-1428
Saez A, Buguin A, Silberzan P, Ladoux B (2005) Is the mechanical activity of epithelial cells controlled by deformations or forces? Biophys J 89(6):L52–L54. https://doi.org/10.1529/biophysj.105.071217
Sepúlveda N, Petitjean L, Cochet O, Grasland-Mongrain E, Silberzan P, Hakim V (2013) Collective cell motion in an epithelial sheet can be quantitatively described by a stochastic interacting particle model. PLoS Computat Biol 9(3):e1002944. https://doi.org/10.1371/journal.pcbi.1002944
Sumpter DJT (2006) The principles of collective animal behaviour. Philos Trans R Soc Lond B Biol Sci 361(1465):5–22. https://doi.org/10.1098/rstb.2005.1733
Sun M, Bloom AB, Zaman MH (2015) Rapid quantification of 3d collagen fiber alignment and fiber intersection correlations with high sensitivity. PLoS ONE 10(7):1–17
Taubenberger A, Cisneros DA, Friedrichs J, Puech PH, Muller DJ, Franz CM (2007) Revealing early steps of \(\alpha\)2\(\beta\)1 integrin-mediated adhesion to collagen type I by using single-cell force spectroscopy. Mol Biol Cell 18(5):1634–1644
VanderVorst K, Dreyer CA, Konopelski SE, Lee H, Ho HYH, Carraway KL (2019) Wnt/PCP signaling contribution to carcinoma collective cell migration and metastasis. Cancer Res 79(8):1719–1729. https://doi.org/10.1158/0008-5472.CAN-18-2757
Vedula SRK, Leong MC, Lai TL, Hersen P, Kabla AJ, Lim CT, Ladoux B (2012) Emerging modes of collective cell migration induced by geometrical constraints. Proc Natl Acad f Sci 109(32):12974–12979. https://doi.org/10.1073/pnas.1119313109
Velez DO, Tsui B, Goshia T, Chute CL, Han A, Carter H, Fraley SI (2017) 3d collagen architecture induces a conserved migratory and transcriptional response linked to vasculogenic mimicry. Nat Commun 8(1):1651. https://doi.org/10.1038/s41467-017-01556-7
Wang X, Enomoto A, Asai N, Kato T, Takahashi M (2016) Collective invasion of cancer: perspectives from pathology and development. Pathol Int 66(4):183–192. https://doi.org/10.1111/pin.12391
Wolf K, Wu YI, Liu Y, Geiger J, Tam E, Overall C, Stack MS, Friedl P (2007) Multi-step pericellular proteolysis controls the transition from individual to collective cancer cell invasion. Nat Cell Biol. https://doi.org/10.1038/ncb1616
Wu PH, Giri A, Sun SX, Wirtz D (2014) Three-dimensional cell migration does not follow a random walk. Proc Natl Acad Sci USA 111(11):3949–54. https://doi.org/10.1073/pnas.1318967111
Yeoman BM, Katira P (2018) A stochastic algorithm for accurately predicting path persistence of cells migrating in 3d matrix environments. PloS ONE 13(11):e0207216
Zaman MH, Kamm RD, Matsudaira P, Lauffenburger Da (2005) Computational model for cell migration in three-dimensional matrices. Biophys J 89(2):1389–1397. https://doi.org/10.1529/biophysj.105.060723
Zaman MH, Trapani LM, Sieminski AL, MacKellar D, Gong H, Kamm RD, Wells A, Lauffenburger DA, Matsudaira P (2006) Migration of tumor cells in 3d matrices is governed by matrix stiffness along with cell-matrix adhesion and proteolysis. Proc Natl Acad Sci 103(29):10889–10894
Acknowledgements
This work was supported by Grants from the National Science Foundation (BMMB - 1905390, BMMB - 1763132 to P. K.) and the Army Research Office (W911NF-17-1-0413 to P. K.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There are no conflicts of interest for any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10237_2020_1290_MOESM1_ESM.png
Figure S1. Mean Squared Displacement (MSD) Plots for Example Cells at 1.25x10-3 fibers/\(\mu m^3\) . A) MSD plot on a log–log scale for low alignment and low cell–cell adhesion. Fit equations are shown in the same color as their plot, with goodness of fit given by R. Solid blue line is defined leader and solid red line is undefined leader. Colored dashed lines are the fits for their matching color. Dashed black line is \(\langle R^2\rangle = \tau\). B) MSD plot on a log–log scale for high alignment and low cell–cell adhesion. C) MSD plot on a log–log scale for low alignment and high cell–cell adhesion. D) MSD plot on a log–log scale for high alignment and high cell–cell adhesion (png 294 KB)
10237_2020_1290_MOESM2_ESM.png
Figure S2. Mean Squared Displacement (MSD) Plots for Example Cells at 1.63x10-3 fibers/\(\mu m^3\) . A) MSD plot on a log–log scale for low alignment and low cell–cell adhesion. Fit equations are shown in the same color as their plot, with goodness of fit given by R. Solid blue line is defined leader and solid red line is undefined leader. Colored dashed lines are the fits for their matching color. Dashed black line is \(\langle R^2\rangle = \tau\). B) MSD plot on a log–log scale for high alignment and low cell–cell adhesion. C) MSD plot on a log–log scale for low alignment and high cell–cell adhesion. D) MSD plot on a log–log scale for high alignment and high cell–cell adhesion (png 293 KB)
10237_2020_1290_MOESM3_ESM.png
Figure S3. This figure shows MSD results for 5-cell cluster over increasing fiber density. Figure S1 (a), (c), and (e) shows MSD with no fiber alignment, and Figure S1 (b), (d), and (f) shows MSD for clusters with high fiber alignment. Figure S1 (a) and (b) shows MSD with clusters at low adhesion (25 nN), Figure S1 (c) and (d) shows MSD at medium adhesion (50 nN), and Figure S1 (e) and (f) shows MSD at high adhesion (100 nN) (png 761 KB)
10237_2020_1290_MOESM4_ESM.png
Figure S4. This figure shows MSD results for 10-cell cluster over increasing fiber density. Figure S2 (a), (c), and (e) shows MSD with no fiber alignment, and Figure S2 (b), (d), and (f) shows MSD for clusters with high fiber alignment. Figure S2 (a) and (b) shows MSD with clusters at low adhesion (25 nN), Figure S2 (c) and (d) shows MSD at medium adhesion (50 nN), and Figure S2 (e) and (f) shows MSD at high adhesion (100 nN) (png 732 KB)
10237_2020_1290_MOESM5_ESM.png
Figure S5. Cluster Size vs. Lifetime for Defined and Undefined Leading Clusters. A) Defined leader cluster lifetime as a function of cluster size. B) Undefined leader cluster as a function of cluster size. Clusters were simulated with low adhesion (25 nN) in unaligned matrices. Error bars are mean squared error (png 128 KB)
Video S1 shows the migration of a 10-cell cluster with a single defined leader over 18 hours. This sample was run with high adhesion, no fiber alignment, and an intermediate fiber density. The cell color at each time step corresponds to the phases accordingly: contraction (green), outgrowth (yellow), and retraction (red) (mp4 5948 KB)
Video S2 shows the migration of a 10-cell cluster with an undefined leader over 38 hours. This sample was run with high adhesion, no fiber alignment, and an intermediate fiber density. The color at each time step corresponds to the phases accordingly: contraction (green), outgrowth (yellow), and retraction (red) (mp4 1590 KB)
Rights and permissions
About this article
Cite this article
Collins, T.A., Yeoman, B.M. & Katira, P. To lead or to herd: optimal strategies for 3D collective migration of cell clusters. Biomech Model Mechanobiol 19, 1551–1564 (2020). https://doi.org/10.1007/s10237-020-01290-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10237-020-01290-y