Abstract
Dictyotophycidae is a subclass of brown algae containing 395 species that are distributed worldwide. A complete plastid (chloroplast) genome (ptDNA or cpDNA) had not previously been sequenced from this group. In this study, the complete plastid genome of Dictyopteris divaricata (Okamura) Okamura (Dictyotales, Phaeophyceae) was characterized and compared to other brown algal ptDNAs. This plastid genome was 126,099 bp in size with two inverted repeats (IRs) of 6026 bp. The D. divaricata IRs contained rpl21, making its IRs larger than representatives from the orders Fucales and Laminariales, but was smaller than that from Ectocarpales. The G + C content of D. divaricata (31.19%) was the highest of the known ptDNAs of brown algae (28.94–31.05%). Two protein-coding genes, rbcR and rpl32, were present in ptDNAs of Laminariales, Ectocarpales (Ectocarpus siliculosus), and Fucales (LEF) but were absent in D. divaricata. Reduced intergenic space (13.11%) and eight pairs of overlapping genes in D. divaricata ptDNA made it the most compact plastid genome in brown algae so far. The architecture of D. divaricata ptDNA showed higher similarity to that of Laminariales compared with Fucales and Ectocarpales. The difference in general features, gene content, and architecture among the ptDNAs of D. divaricata and LEF clade revealed the diversity and evolutionary trends of plastid genomes in brown algae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Brown algae (Phaeophyceae) are complex photosynthetic organisms and have independently evolved complex multicellularity among the heterokont lineage (Cock and Collén 2015; Terauchi et al. 2017). The brown algae evolved in a distinct lineage from groups containing a primary plastid (i.e., green algae, land plants, rhodophytes, and glaucophytes) (Rodriguez-Ezpeleta et al. 2005; Ševčíková et al. 2015; Dorrell et al. 2017). The chloroplasts of brown algae were derived by secondary endosymbiosis in which red algae were taken up by a non-photosynthetic eukaryote (Keeling 2004, 2010). Phylogenetic studies based on multimarker datasets have provided comprehensive evolutionary trees of the Phaeophyceae (Silberfeld et al. 2010). Several monophyletic early-diverging lineages, such as Discosporangiales and Ishigeales, could be resolved in the phylogeny, but most other brown algae form two super-clades representing two subclasses, Dictyotophycidae and Fucophycidae (Guiry and Guiry 2017). Dictyotophycidae includes four orders Syringodermatales, Sphacelariales, Dictyotales, and Onslowiales (SSDO), and Fucophycidae is a crown group consisting of 13 orders (Charrier et al. 2012; Silberfeld et al. 2014).
There are currently more than 2000 brown algal species that display a great diversity in morphology, physiology, and sexually dimorphic traits and also serve many ecological roles in marine environments (Charrier et al. 2012; Luthringer et al. 2014). Thus far, plastid (chloroplast) genomes (ptDNA or cpDNA) of nine brown algae have been completely sequenced, including Fucus vesiculosus, Sargassum horneri, Sargassum thunbergii, Coccophora langsdorfii (order Fucales); Saccharina japonica, Costaria costata, Undaria pinnatifida (order Laminariales); and Ectocarpus siliculosus, Pleurocladia lacustris (order Ectocarpales) (Le Corguillé et al. 2009; Wang et al. 2013; Zhang et al. 2015a, b; Liu and Pang 2016). This data has allowed for a better understanding of the evolution of plastid genomes and phylogenetic relationships of the Phaeophyceae. However, all nine known plastid genomes of brown algae belong to Fucophycidae and no complete ptDNA of Dictyotophycidae, a group that contains 395 species of brown algae worldwide (Guiry and Guiry 2017).
The sizes of known brown algal plastid genomes are 124.1–125.0 kb in Fucales, 129.9–130.6 kb in Laminariales, and 138.8–140.0 kb in Ectocarpales (Table 1). These ptDNAs are mapped as a canonical quadripartite structure with two large inverted repeats (IRs), which divide the circular molecule into a small single copy region (SSC) and a large single copy region (LSC) (Wang et al. 2013). These ptDNAs contain 6 ribosomal RNA (rRNA) genes, 27–31 transfer RNA (tRNA) genes, 137–139 protein-coding genes, and 2–6 open reading frames (ORFs). The architecture of plastid genomes is highly conserved within the brown algal orders of Fucales (Liu and Pang 2016) and Laminariales (Zhang et al. 2015a, b), while multiple genome rearrangements occurred in the evolution of this eukaryotic lineage (Le Corguillé et al. 2009).
Dictyopteris divaricata (Okamura) Okamura (Dictyotales) is a cosmopolitan brown seaweed that usually inhabits littoral and sublittoral rock zones (Guiry and Guiry 2017). The thallus of this alga is often flat with regular dichotomous branches and bifid tips (Abbas and Shameel 2011) and contains structurally unique sesquiterpenes with important biological functions (Ji et al. 2009). However, species-level taxonomy is difficult and has been troubled for a long time due to morphological plasticity of species in Dictyotales (Tronholm et al. 2010). Fortunately, methods of molecular markers have been used to unveil intraspecific and interspecific relationships, and several new species have been identified (Lozano-Orozco et al. 2015). However, there is still limited genomic information in Dictyotales, which restricts our understanding of the taxonomic status and evolution of this group.
Although the physical map of plastid DNA in Dictyota dichotoma (Hudson) J.V.Lamouroux and size estimation of this genome had been established by Kuhsel and Kowallik (1985) for more than 30 years, no further information has been reported on the plastid genome sequence in the lineage of Dictyotales. To further understand the evolution of plastid genomes in brown algae, the complete plastid genome of D. divaricata was sequenced with next-generation sequencing. This ptDNA sequence represents the first plastid genome from the subclass Dictyotophycidae.
Materials and Methods
Sample Collection and Identification
Mature plants of Dictyopteris divaricata (Okamura) Okamura were initially collected from the rocky shore of No. 3 bathing beach in Qingdao, Shandong Province, China (36° 03′ N, 120° 22′ E) in July 2016 (Supplementary Fig. S1). Samples were transported to the laboratory in coolers (5–8 °C) within 24 h after collection. Frozen tissue from the original algal samples was used for DNA extraction. Algal tissue was ground to fine powder in liquid nitrogen. Total DNA was extracted using a Plant Genomic DNA Kit (Tiangen Biotech, Beijing, China) according to the manufacturer’s instructions. The concentration and the quality of isolated DNA were assessed by electrophoresis on 1.0% agarose gel. Species identification was performed according to morphological features and based on the analyses of plastid-encoded psbA gene. Sequence dataset of the D. divaricata sample (Dd-Qingdao) and other data from GenBank were aligned using a ClustalW with MEGA 7.0 software (Kumar et al. 2016). Maximum likelihood (ML) and neighbor-joining (NJ) analyses were performed with 1000 bootstrap replicates. The ML trees were obtained based on the Kimura two-parameter model (Kimura 1980) and the NJ trees using the maximum composite likelihood method (Tamura et al. 2004) for the psbA dataset in nucleotides. The NJ and ML phylogenetic analyses were performed in MEGA 7.0. This analysis confirmed the identification of D. divaricata (Supplementary Fig. S2).
Plastid DNA Extraction, Sequencing, and Assembly
The plastids of D. divaricata were isolated using the Plant Chloroplast Purification Kit according to the manufacturer’s instructions (Baiaolaibo, Beijing, China). Then, the plastid DNA was extracted using this kit. The Ultra II DNA Library Prep Kit (NEB, USA) was used for library construction for Illumina sequencing. The plastid DNA was fragmented into 350 bp and sequenced using Illumina HiSeq platform. The sequencing run produced ca. 783 Mb raw data with reads length of 150 bp. Poor quality sequences and sequencing adapters were removed using Trim Galore! v0.3.7 (http://www.bioinformatics.babraham.ac.uk/projects/trim_galore/), leaving 730 Mb clean data. De novo assemblies were run using SOAPdenovo v2.04 and GapCloser v1.12 (Luo et al. 2012) with the trimmed sequences. The final plastid assembly contained 156,285 reads with the mean coverage depth of 178. This resulted in one scaffold of 126,099 bp.
Genome Annotation and Comparative Analysis
Protein-coding genes and open reading frames (ORFs) were annotated using Dual Organellar Genome Annotator (DOGMA) (Wyman et al. 2004), NCBI ORF Finder and BLAST similarity searches of the non-redundant databases at NCBI (Altschul et al. 1997). Ribosomal RNA genes were delimited by direct comparison to sequenced brown algal orthologues using MEGA7. Transfer RNA genes were searched for by reconstructing their cloverleaf structures using the tRNAscan-SE 1.21 software with default parameters (Schattner et al. 2005). The physical map of the circular plastid genome was generated with Organellar GenomeDRAW (OGDraw) (Lohse et al. 2013). The genome sequence has been deposited in GenBank with the accession number KY433579. Base composition was determined by the MEGA7.0 software (Kumar et al. 2016). Tandem repeats (TRs) were found with Tandem Repeats Finder using default settings (Benson 1999). Small inverted repeats (SIRs) were identified with Inverted Repeats Finder using the default settings and the additional constraint that repeats had to be > 75% similar (http://tandem.bu.edu/cgi-bin/irdb/irdb.exe). Multiple sequence alignment of the ten brown algal plastid genome sequences (Table 1) was performed using the Mauve Genome Alignment v2.3.1 (Darling et al. 2004) with progressive Mauve algorithm (Darling et al. 2010).
Phylogenetic Analysis
Phylogenetic relationships within the brown algae were analyzed based on ptDNA protein-coding gene (PCG) datasets, which were composed of amino acid (aa) sequences of 18 photosystem II PCGs (psb28, psbA, psbB, psbC, psbD, psbE, psbF, psbH, psbI, psbJ, psbK, psbL, psbN, psbT, psbV, psbX, psbY, and ycf12) that were the most conserved group among PCGs in brown algal ptDNAs (Liu and Pang 2016). The aa sequences of 18 photosystem II PCGs were subjected to concatenated alignments using ClustalX 1.83 with the default settings (Thompson et al. 1997). Vaucheria litorea (Xanthophyceae; Rumpho et al. 2008) was selected as an out-group taxon for analysis of the aa dataset. The evolutionary history was inferred by using the ML method based on the JTT matrix-based model (Jones et al. 1992) and the NJ method (Saitou and Nei 1987) based on the Poisson correction model (Zuckerkandl and Pauling 1965) with 1000 bootstrap replicates, respectively, using MEGA 7.0 software (Kumar et al. 2016). Bayesian Inference (BI) analyses of the aa dataset were performed based on the best scoring alternative model of MtREV + G + I using MrBayes v.3.2 (Huelsenbeck and Ronquist 2001). One million generations were run for tree reconstructions and posterior probabilities using the Markov chain Monte Carlo (MCMC) method. Every 1000th generation was saved and the first 100 generations were discarded as burn-in. Posterior probability values for the majority-rule consensus trees constructed were calculated.
Results and Discussion
Genome Size and Inverted Repeats
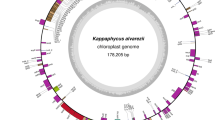
The complete plastid genome of D. divaricata was 126,099 bp in size (Fig. 1), which was smaller than that of Laminariales (129.9–130.6 kb) and Ectocarpales (138.8–140.0 kb), but larger than that of Fucales (124.1–125.0 kb). The size of D. divaricata ptDNA was close to D. dichotoma plastid DNA, which was predicted to be 123 kb by using electron microscopy and gel electrophoresis (Kuhsel and Kowallik 1985). Like most plastid genomes, the D. divaricata ptDNA mapped as a canonical quadripartite structure with two large inverted repeats of 6026 bp dividing single circular genome into regions of a small single copy (SSC 41,399 bp) and a large single copy (LSC 72,648 bp) (Table 1). The nucleotide composition of brown algal ptDNAs was conserved and displayed low G + C content. The G + C content of D. divaricata ptDNAs was 31.19% and slightly higher than that of other brown algae ranging from 28.94% in Fucus vesiculosus to 31.05% in Saccharina japonica.
The plastid genome map of Dictyopteris divaricata. Annotated genes are colored according to the functional categories. Genes on the inside are transcribed in the clockwise direction, whereas genes on the outside are transcribed in the counterclockwise direction. The ring of bar graphs on the inner circle shows the GC content in dark gray
Variation in plastid genome size was mainly due to expansion and contraction of the inverted repeats (IRs), intron number, gene transfer and loss, and size of intergenic spacer regions (Baurain et al. 2010; Tanaka et al. 2011; Brembu et al. 2014; Sabir et al. 2014). The brown algal IRs were comprised by the core rrn5-rnl-trnA-trnI-rns gene cluster and the additional genes that flanked the ribosomal gene operon. The size of IRs in ten sequenced brown algal ptDNAs ranged from 5370 bp in F. vesiculosus to 8616 bp in E. siliculosus. The structure of IRs was conserved at the order level in brown algae (Fig. 2). The D. divaricata IRs was 6026 bp in size and contained the rpl21-rrn5-rnl-trnA-trnI-rns gene cluster. The D. divaricata IR was larger than that of Fucales and Laminariales due to the presence of rpl21 in the IR of D. divaricata but was smaller than that of Ectocarpales. Some genes (e.g., psbA, rpl32, trnL, and trnE) and orfs (e.g., orf53 and orf258) were only present in the IRs of Ectocarpales ptDNAs, which contributed to longer IRs of 8616 bp in E. siliculosus and 8084 bp in P. lacustris.
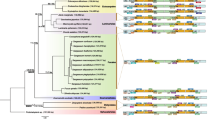
Comparison of inverted repeat (IRs) boundaries in the ten brown algal ptDNAs. a IRa. b IRb. The “p” in the bracket represents the partial of the related genes of Fucus vesiculosus (Fv), Sargassum horneri (Sh), Sargassum thunbergii (St), Coccophora langsdorfii (Cl), Saccharina japonica (Sj), Costaria costata (Cc), Undaria pinnatifida (Up), Ectocarpus siliculosus (Es), Pleurocladia lacustris (Pl), and Dictyopteris divaricata (Dd) ptDNAs. Annotated genes are colored according to the functional categories
Gene Content
The D. divaricata ptDNA contained 174 genes, including six ribosomal RNA genes (rRNA), 28 transfer RNA genes (tRNA), 138 protein-coding genes (two rpl21 genes in IRs), and two conserved open reading frames (orfs). Only one intron was present in the D. divaricata trnL2 gene encoding tRNA-Leu. This intron was also present in the homologous genes of the sequenced Fucales and Laminariales ptDNAs but was absent in two members of the Ectocarpales (Le Corguillé et al. 2009). All protein-coding genes encoded by D. divaricata ptDNA started with the ATG codon with the exception of psbF with GTG and ycf66 with TTG. A total of 113 protein-coding genes were terminated by a TAA stop codon, 20 with TAG, and seven with TGA.
The ten brown algal plastid genomes shared a core set of 133 protein-coding genes (duplicated genes were only counted once), while the other protein-coding genes only occurred in certain lineages (Table 2). Two protein-coding genes, rbcR and rpl32, were found in Laminariales, Ectocarpales (E. siliculosus), and Fucales (LEF) but were absent in D. divaricata. The rpl32 gene was also lost in the P. lacustris ptDNA. The absence of this gene might be due to gene transfer to the nucleus or gene loss. Two genes, syfB and ycf17, were absent in Fucales but present in D. divaricata as well as Laminariales and Ectocarpales. Another two genes, petL and ycf54, were only absent in Laminariales. Besides two conserved orfs shared by all brown algal ptDNAs, some specific orfs with unknown function were identified only in the Ectocarpales ptDNAs (Table 2).
Intergenic Spacer and Overlapping Regions
The intergenic spacer region in D. divaricata ptDNA was a total of 16,528 bp in size and constituted 13.11% of the whole genome, which was slightly less than that in Fucales (13.68–14.16%), Laminariales (16.48–16.74%), and Ectocarpales (19.13–19.60%). Five conserved overlapping regions had been noted in the previously sequenced brown algal plastid genomes (Wang et al. 2013; Zhang et al. 2015a, b), while eight pairs of overlapping genes were found in the D. divaricata plastid genome, including atpD-atpF (1 bp), ycf12-ftrB (6 bp), rpl4-rpl23 (8 bp), rpl29-rps17 (4 bp), rpl1-rpl11 (4 bp), rps1-ycf40 (23 bp), ycf24(sufB)-ycf16(sufC) (4 bp), and psbD-psbC (53 bp) (Table 3). Reduced content of intergenic spacer regions and the increase in overlapping regions made the D. divaricata ptDNA the most compact plastid genome in brown algae so far.
Three overlapping regions, i.e., rpl4-rpl23 (8 bp), ycf24(sufB)-ycf16(sufC) (4 bp), and psbD-psbC (53 bp), were present in all sequenced ptDNAs of brown algae with the same overlapping size, indicating the conservative characteristics of brown algal genome structure. The ycf12-ftrB (6 bp) overlapping region was present in Fucales, Laminariales and D. divaricata, but absent in Ectocarpales. The rps1-thiS (4 bp) overlapping region was only found in the Fucales. Four new overlapping regions identified in D. divaricata, i.e., atpD-atpF (1 bp), rpl29-rps17 (4 bp), rpl1-rpl11(4 bp), and rps1-ycf40 (23 bp), were not detected in plastid genomes of LEF clade, indicating that they appeared in ptDNA of D. divaricata after its divergence from LEF clade and highlighting the diversity of evolutionary trends in brown algal plastid genomes. The psbD-psbC overlapping region was the most conserved in terms of size and sequence even among the ochrophytes which harbored Phaeophyceae, Xanthophyceae, Raphidophyceae, Eusgmatophyceae, Chrysophyceae, Pelagophyceae, and Bacillariophyceae (Ševčíková et al. 2015).
Tandem and Small Inverted Repeats
Numerous tandem (TRs) and small inverted repeats (SIRs) were previously found in plastid genomes (Cattolico et al. 2008; Liu and Pang 2016). In D. divaricata ptDNA, a total of 5 tandem repeats (TRs) and 15 small inverted repeats (SIRs) were identified. Two of TRs were located in the atpA gene, one in ycf46, one in psbA-psbK spacer, and one in rpl11-trnW(cca) spacer. The average size of D. divaricata TR elements was 43.8 ± 11.6 bp, which was similar to that of other brown algae (Liu and Pang 2016). The TRs had a period of 9–24 bp with a copy number of 2–3.5 with low GC content ranging from 5.13 to 25.49%. The D. divaricata SIRs were all localized within intergenic spacer regions and some overlapped partially with adjacent genes. The stem length of SIRs ranged from 21 to 68 bp and the small loop domain from 0 to 18 bp.
Eight SIRs of D. divaricata were located at the termini of two genes transcribed on opposite coding strands, including rbcS/ccsA, orf531/petA, psaI/psbJ, ftsH/psbH, psbN/psbT, psbB/petF, petF/rpl12, and psbC/ycf41. Exploring the biological significance of SIRs and their hairpin structure is a fascinating question. The IRs of stramenopile ptDNAs are usually located adjacent to genes that are related to photosynthesis or energy production (Cattolico et al. 2008) and are likely to play an important role for regulation of transcription, translation, and other biological functions (Lillo et al. 2002). The placement conservation was considered to be likely associated with the functional constraint (Ong et al. 2010; Hovde et al. 2014).
Genome Organization
The quadripartite plastid genomes in brown algae exhibited several rearrangements among species, especially at the order level, which was similar to other observed red algal-derived ptDNAs (Ruck et al. 2014; Starkenburg et al. 2014; Yurchenko et al. 2016). By comparing the genome organization of 10 brown algal species from four orders, 22 conserved gene blocks (CGBs) were identified (Fig. 3). Two large CGBs over 30 kb were noted. The ribosomal gene block (30.5 kb) contained 48 genes from rpl9 to rns. The highly conserved ribosomal gene block was also observed in other red algal-derived plastid genomes (Stoebe and Kowallik 1999; Oudot-Le Secq et al. 2007; Tajima et al. 2016). Another CGB was 33.3 kb that contained 36 genes (rpl27-atpA). Most of the genes in this CGB were mostly transcribed on the same strand.
Genome organization of brown algal ptDNAs was highly conserved at the order level, as was shown by the nine ptDNAs sampled from Laminariales, Ectocarpales, and Fucales. Ectocarpales had displayed a higher number of rearrangements than other lineages. The architecture of D. divaricata ptDNA was more similar to that of Laminariales than Fucales and Ectocarpales. The plastid genomes of D. divaricata and Laminariales showed a high degree of similarity in gene arrangement, indicating the key clue to understand the ancestral gene order of brown algae. Relative to the gene order of Laminariales, in D. divaricata, two blocks of six genes (trnE-psaA-psaB-rps14-petG-psbK) and seven genes (rpl19-trnM-ycf47-petM-petN-acsF-ycf42) had been translocated and inverted.
Phylogenetic Analyses
The phylogenetic analyses of 18 photosystem II protein dataset based on ML, NJ, and BI methods generated identical topologies with similar support values (Fig. 4). The limited taxon sampling of ten brown algal species formed four clades representing four orders: Laminariales, Ectocarpales, Fucales, and Dictyotales. D. divaricata (Dictyotales) diverged before a strongly supported clade comprising Laminariales, Ectocarpales, and Fucales (LEF), which was similar to prior phylogenomic comparisons of ten taxa and 35 mitochondrial protein-coding genes (Liu and Pang 2015). The Dictyotophycidae and Fucophycidae had a sister relationship, which has been previously noted (Silberfeld et al. 2014). However, so far, we still have limited plastid genome sequences and especially lack representative plastid genomes of some brown algal lineages (orders) including the basal taxa Discosporangiales and Ishigeales. Thus, a greater taxon sampling will hopefully provide a more comprehensive and general picture of the diversity of plastid genomes in brown algae.
Phylogenetic tree constructed from analyses of amino acid (aa) sequences of 18 photosystem II PCGs. The tree was rooted with Vaucheria litorea (Xanthophyceae). The numbers at internal nodes (ML/NJ/BI) indicated maximum likelihood (ML) and neighbor-joining (NJ) bootstrap values, as well as Bayesian Inference (BI) posterior probability values, respectively. Branch lengths are proportional to the amount of amino acid substitutions per site, which are indicated by the scale bar below the tree
Conclusion
Dictyopteris divaricata was the first species from the subclass Dictyotophycidae to have its plastid genome completely sequenced and annotated. The most important findings in this study were that the ptDNA of D. divaricata was the most compact plastid genome in brown algae so far and its architecture was more similar to that of Laminariales than Fucales and Ectocarpales. Detailed comparative analyses of the conservation and variation of genome characteristics in a larger scale further incited into our understanding of the evolutionary history of brown algal ptDNAs. The difference in general features, gene content, and architecture among D. divaricata and LEF ptDNAs revealed the diversity and evolutionary trends of plastid genomes in brown algae.
References
Abbas A, Shameel M (2011) Anatomy of Dictyopteris divaricata (Phaeophycota) from the coast of Karachi. Pak J Bot 43(4):2207–2210
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402
Baurain D, Brinkmann H, Petersen J, Rodríguez-Ezpeleta N (2010) Phylogenomic evidence for separate acquisition of plastids in cryptophytes, haptophytes, and stramenopiles. Mol Biol Evol 27(7):1698–1709
Benson G (1999) Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res 27(2):573–580
Brembu T, Winge P, Tooming-Klunderud A, Nederbragt AJ, Jakobsen KS, Bones AM (2014) The chloroplast genome of the diatom Seminavis robusta: new features introduced through multiple mechanisms of horizontal gene transfer. Mar Genomics 16:17–27
Cattolico RA, Jacobs MA, Zhou Y, Chang J, Duplessis M, Lybrand T, McKay J, Ong HC, Sims E, Rocap G (2008) Chloroplast genome sequencing analysis of Heterosigma akashiwo CCMP452 (West Atlantic) and NIES293 (West Pacific) strains. BMC Genomics 9(1):211
Charrier B, Bail AL, de Reviers B (2012) Plant Proteus: brown algal morphological plasticity and underlying developmental mechanisms. Trends Plant Sci 17(8):468–477
Cock JM, Collén J (2015) Independent emergence of complex multicellularity in the brown and red algae. In: Ruiz-Trillo I, Nedelcu AM (eds) Evolutionary transitions to multicellular life: principles and mechanisms. Springer Netherlands, Dordrecht, pp 335–361
Darling AE, Mau B, Blatter FR, Perna NT (2004) Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome Res 14(7):1394–1403
Darling AE, Mau B, Perna NT (2010) progressiveMauve: multiple genome alignment with gene gain, loss, and rearrangement. PLoS One 5(6):e11147
Dorrell RG, Gile G, McCallum G, Méheust R, Bapteste EP, Klinger CM, Brillet-Guéguen L, Freeman KD, Richter DJ, Bowler C (2017) Chimeric origins of ochrophytes and haptophytes revealed through an ancient plastid proteome. elife 6:e23717
Guiry MD, Guiry GM (2017) AlgaeBase. World-wide electronic publication. National University of Ireland, Galway. http://www.algaebase.org. Accessed 20 Jan 2017
Hovde BT, Starkenburg SR, Hunsperger HM, Mercer LD, Deodato CR, Jha RK, Chertkov O, Monnat RJ Jr, Cattolico RA (2014) The mitochondrial and chloroplast genomes of the haptophyte Chrysochromulina tobin contain unique repeat structures and gene profiles. BMC Genomics 15(1):604
Huelsenbeck JP, Ronquist F (2001) MrBAYES: Bayesian inference of phylogeny. Bioinformatics 17(8):754–755
Ji NY, Wen W, Li XM, Xue QZ, Xiao HL, Wang BG (2009) Brominated selinane sesquiterpenes from the marine brown alga Dictyopteris divaricata. Mar Drugs 7(3):355–360
Jones DT, Taylor WR, Thornton JM (1992) The rapid generation of mutation data matrices from protein sequences. Comput Appl Biosci 8(3):275–282
Keeling PJ (2004) Diversity and evolutionary history of plastids and their hosts. Am J Bot 91(10):1481–1493
Keeling PJ (2010) The endosymbiotic origin, diversification and fate of plastids. Philos Trans R Soc B 365(1541):729–748
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16(2):111–120
Kuhsel M, Kowallik KV (1985) The plastome of a brown alga, Dictyota dichotoma I. Physical properties and the Barn HI/Sal I/Bgl H cleavage site map. Plant Mol Biol 4(6):365–376
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874
Le Corguillé G, Pearson G, Valente M, Viegas C, Gschloessl B, Corre E, Bailly X, Peters AF, Jubin C, Vaccherie B, Cock JM, Leblanc C (2009) Plastid genomes of two brown algae, Ectocarpus siliculosus and Fucus vesiculosus: further insights on the evolution of red-algal derived plastids. BMC Evol Biol 9(1):253
Lillo F, Basile S, Mantegna RN (2002) Comparative genomics study of inverted repeats in bacteria. Bioinformatics 18(7):971–979
Liu F, Pang SJ (2015) Mitochondrial phylogenomics reveals a close relationship between Petalonia fascia (Scytosiphonaceae, Phaeophyceae) and Ectocarpus siliculosus. J Appl Phycol 27(2):1021–1028
Liu F, Pang SJ (2016) Chloroplast genome of Sargassum horneri (Sargassaceae, Phaeophyceae): comparative chloroplast genomics of brown algae. J Appl Phycol 28(2):1419–1426
Lohse M, Drechsel O, Kahlau S, Bock R (2013) OrganellarGenomeDRAW—a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucleic Acids Res 41:575–581
Lozano-Orozco JG, Sentíes A, Clerck OD, Dreckmann KM, Díaz-Larrea J (2015) Two new species of the genus Dictyota (Phaeophyceae: Dictyotales) from the Mexican Caribbean. Am J Plant Sci 6(15):2492–2501
Luo RB, Liu BH, Xie YL, Li ZY, Huang WH, Yuan JY, He GZ, Chen YX, Pan Q, Liu YJ, Tang JB, Wu GX, Zhang H, Shi YJ, Liu Y, Yu C, Wang B, Lu Y, Han CL, Cheung DW, Yiu SM, Peng SL, Zhu XQ, Liu GM, Liao XK, Li YR, Yang HM, Wang J, Lam TW, Wang J (2012) SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. GigaScience 1(1):18
Luthringer R, Cormier A, Ahmed S, Peters AF, Cock JM, Coelho SM (2014) Sexual dimorphism in the brown algae. Perspect Phycol 1(1):11–25
Ong HC, Wilhelm SW, Gobler CJ, Bullerjahn G, Jacobs MA, Mckay J, Sims EH, Gillett WG, Zhou Y, Haugen E, Rocap G, Cattolico RA (2010) Analyses of the complete chloroplast genome sequences of two members of the Pelagophyceae: Aureococcus anophagefferens CCMP1984 and Aureoumbra lagunensis CCMP1507. J Phycol 46(3):602–615
Oudot-Le Secq MP, Grimwood J, Shapiro H, Armbrust EV, Bowler C, Green BR (2007) Chloroplast genomes of the diatoms Phaeodactylum tricornutum and Thalassiosira pseudonana: comparison with other plastid genomes of the red lineage. Mol Gen Genomics 277(4):427–439
Rodriguez-Ezpeleta N, Brinkmann H, Burey SC, Roure B, Burger G, Loffelhardt W, Bohnert HJ, Philippe H, Lang BF (2005) Monophyly of primary photosynthetic eukaryotes: green plants red algae and glaucophytes. Curr Biol 15(14):1325–1330
Ruck EC, Nakov T, Jansen RK, Theriot EC, Alverson AJ (2014) Serial gene losses and foreign DNA underlie size and sequence variation in the plastid genomes of diatoms. Genome Biol Evol 6(3):644–654
Rumpho ME, Worful JM, Lee J, Kannan K, Tyler MS, Bhattacharya D, Moustafa A, Manhart JR (2008) Horizontal gene transfer of the algal nuclear gene psbO to the photosynthetic sea slug Elysia chlorotica. PNAS 105(46):17867–17871
Sabir JSM, Yu M, Ashworth MP, Baeshen NA, Baeshen MN, Bahieldin A, Theriot EC, Jansen RK (2014) Conserved gene order and expanded inverted repeats characterize plastid genomes of Thalassiosirales. PLoS One 9(9):e107854
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4):406–425
Schattner P, Brooks AN, Lowe TM (2005) The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res 33:686–689
Ševčíková T, Horák A, Klimeš V, Zbránková V, Demir-Hilton E, Sudek S, Jenkins J, Schmutz J, Přibyl P, Fousek J, Vlček ČB, Lang BF, Oborník M, Worden AZ, Eliáš M (2015) Updating algal evolutionary relationships through plastid genome sequencing: did alveolate plastids emerge through endosymbiosis of an ochrophyte? Sci Rep 5(1):10134
Silberfeld T, Leigh JW, Verbruggen H, Cruaud C, de Reviers B, Rousseau F (2010) A multi-locus time-calibrated phylogeny of the brown algae (Heterokonta, Ochrophyta, Phaeophyceae): investigating the evolutionary nature of the “brown algal crown radiation”. Mol Phylogenet Evol 56(2):659–674
Silberfeld T, Rousseau F, de Reviers B (2014) An updated classification of brown algae (Ochrophyta, Phaeophyceae). Cryptogam Algol 35(2):117–156
Starkenburg SR, Kwon KJ, Jha RK, McKay C, Jacobs M, Chertkov O, Twary S, Rocap G, Cattolico RA (2014) A pangenomic analysis of the Nannochloropsis organellar genomes reveals novel genetic variations in key metabolic genes. BMC Genomics 15(1):212
Stoebe B, Kowallik KV (1999) Gene-cluster analysis in chloroplast genomics. Trends Genet 15(9):344–347
Tajima N, Saitoh K, Sato S, Maruyama F, Ichinomiya M, Yoshikawa S, Kurokawa K, Ohta H, Tabata S, Kuwata A, Sato N (2016) Sequencing and analysis of the complete organellar genomes of Parmales, a closely related group to Bacillariophyta (diatoms). Curr Genet 62(4):887–896
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci U S A 101(30):11030–11035
Tanaka T, Fukuda Y, Yoshino T, Maeda Y, Muto M, Matsumoto M, Mayama S, Matsunaga T (2011) High-throughput pyrosequencing of the chloroplast genome of a highly neutral-lipid-producing marine pennate diatom, Fistulifera sp. strain JPCC DA0580. Photosynth Res 109(1-3):223–229
Terauchi M, Yamagishi T, Hanyuda T, Kawai H (2017) Genome-wide computational analysis of the secretome of brown algae (Phaeophyceae). Mar Genomics 32:49–59
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25(24):4876–4882
Tronholm A, Steen F, Tyberghein L, Leliaert F, Verbruggen H, Ribera Siguan MA, Clerck OD (2010) Species delimitation, taxonomy, and biogeography of Dictyota in Europe (Dictyotales, Phaeophyceae). J Phycol 46(6):1301–1321
Wang X, Shao Z, Fu W, Yao J, Hu Q, Duan D (2013) Chloroplast genome of one brown seaweed, Saccharina japonica (Laminariales, Phaeophyta): its structural features and phylogenetic analyses with other photosynthetic plastids. Mar Genomics 10:1–9
Wyman S, Jansen R, Boore J (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20(17):3252–3255
Yurchenko T, Ševčíková T, Strnad H, Butenko A, Eliáš M (2016) The plastid genome of some eustigmatophyte algae harbours a bacteria-derived six-gene cluster for biosynthesis of a novel secondary metabolite. Open Bio 6(11):160249
Zhang L, Wang XM, Liu T, Wang GL, Chi S, Liu C, Wang HY (2015a) Complete plastid genome sequence of the brown alga Undaria pinnatifida. PLoS One 10(10):e0139366
Zhang L, Wang XM, Liu T, Wang HY, Wang GL, Chi S, Liu C (2015b) Complete plastid genome of the brown alga Costaria costata (Laminariales, Phaeophyceae). PLoS One 10(10):e0140144
Zuckerkandl E, Pauling L (1965) Evolutionary divergence and convergence in proteins. In: Bryson V, Vogel HJ (eds) Evolving genes and proteins. Academic Press, New York, pp 97–166
Acknowledgements
The authors wish to thank Wei Luan for his assistance in algal collection and data analysis.
Funding
This work was supported by the Key Research Program of Frontier Sciences, Chinese Academy of Sciences (No. QYZDB-SSW-DQC023), the Scientific and Technological Innovation Project Financially Supported by Qingdao National Laboratory for Marine Science and Technology (No. 2016ASKJ02), the Key Research and Development Project of Shandong Province, China (No. 2016GSF115041), the Strategic Priority Research Program, Chinese Academy of Sciences (No. XDA11020304), the Youth Innovation Promotion Association, Chinese Academy of Sciences (No. 2015164), the Foundation for Huiquan Young Scholar of Institute of Oceanology, Chinese Academy of Sciences (No. 2015), the Open Research Fund of Key Laboratory of Integrated Marine Monitoring and Applied Technologies for Harmful Algal Blooms, S.O.A. (No. MATHAB201701), and the Earmarked Fund for Modern Agro-industry Technology Research System in Shandong Province of China (No. SDAIT-26-09).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. This article does not contain any studies with human participants or animals performed by any of the authors. Informed consent was obtained from all individual participants included in the study.
Conflict of Interest
The authors declare that they have no conflict of interest.
Electronic Supplementary Material
Fig. S1
The whole plant of Dictyopteris divaricata collected at Qingdao coast. (PPTX 757 kb)
Fig. S2
Phylogenetic tree constructed from analyses of plastid-encoded psbA gene. The tree was rooted with Dictyota dichotoma. The numbers at internal nodes were bootstrap values greater than 70% for 1000 replicates in Maximum Likelihood (ML, above) and Neighbor-Joining (NJ, below) analysis. Branch lengths were proportional to the amount of sequence change, which were indicated by the scale bar below the tree. (PPTX 471 kb)
Rights and permissions
About this article
Cite this article
Liu, F., Jin, Z., Wang, Y. et al. Plastid Genome of Dictyopteris divaricata (Dictyotales, Phaeophyceae): Understanding the Evolution of Plastid Genomes in Brown Algae. Mar Biotechnol 19, 627–637 (2017). https://doi.org/10.1007/s10126-017-9781-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-017-9781-5