Abstract
A recurrent glycine-to-arginine/valine alteration at codon 34 (G34R/V) within H3F3A, a gene that encodes the replication-independent histone variant H3.3, reportedly occurs exclusively in pediatric glioblastomas. However, the clinicopathological and biological significances of this mutation have not been completely elucidated; especially, no such data exist for tumor samples from Japanese patients. We analyzed 411 consecutive glioma cases representing patients of all ages. Our results demonstrated that 14 patients (3.4%) harbored H3F3A mutations, of which four had G34R mutations and 10 had K27M mutations. G34R-mutant tumors were located in the parietal region in two patients and the basal ganglia in one patient. One patient showed multi-lobular extension similar to the pattern observed in gliomatosis cerebri. Regarding neuroradiological features, intratumoral calcification was evident in two cases and all cases showed no or scarce contrast enhancement on MRI. Histopathologically, the four G34R-mutant cases included three glioblastomas and one astroblastoma. We have also investigated alterations in histone methylation including H3K27me3, H3K9me3, and H3K4me3 in G34R-mutant samples by immunohistochemistry. These results indicate that G34R-mutant tumors are likely to show extensive infiltration and alterations in global histone trimethylation might also play an important role in G34R mutant tumors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
H3F3A is a gene that encodes the replication-independent histone variant H3.3. Two recurrent mutations including a lysine-to-methionine alteration at codon 27 (K27M) and a glycine-to-arginine/valine at codon 34 (G34R/V) within this gene have been identified exclusively in a subset of pediatric glioblastomas (GBM) [1,2,3]. Current integrated molecular classification has demonstrated the clinicopathological differences between K27M and G34R/V-mutant tumors [4,5,6]. Anatomically, K27M mutant tumors are located in the midline region such as the brain stem and thalamus, whereas G34R/V-mutant tumors are typically located in the hemispheric region. In addition, patients with the K27M mutation tend to be younger and have clinically worse prognosis compared to the patients with the G34R/V alteration. Based on the diagnostic and clinical significance of K27M mutations [7], a novel diagnostic entity entitled “diffuse midline glioma, H3 K27M-mutant” has been added to the revised WHO classification of brain tumors. However, the clinicopathological and biological significances of G34R/V have not been completely elucidated [8]. In addition, although the prevalence and clinicopathological features of G34R/V-mutant tumors have been reported in North American and European countries, these clinical characteristics have not been investigated in tumor samples from Japanese patients.
Epigenetic investigation has revealed that post-translational histone modification can activate or repress gene expression [9]. Histone lysine (H3K) methylation is a representative histone modification that activates or represses gene expression, depending on the particular histone residue. For example, methylation of H3K27 and H3K9 is associated with transcriptional silencing, whereas methylation of H3K4 is generally associated with transcriptional activation. Recent studies have reported that H3F3A K27M mutant tumors exclusively exhibit a reduction in H3K27 trimethylation (H3K27me3), suggesting that reduced H3K27me3 is the major driving force behind K27M mutant tumors [9,10,11,12,13,14]. However, alterations in histone methylation including H3K27me3, H3K9me3, and H3K4me3 have rarely been evaluated in G34R/V-mutant tumors.
In this study, we examined the prevalence and clinicopathological features of H3F3A G34R/V-mutant high-grade gliomas by analysis of 411 consecutive glioma cases representing patients of all ages. In addition, we investigated histone methylation to gain insight into the associated epigenetic mechanisms in H3F3A G34R/V-mutant high-grade gliomas.
Methods
Patients
At our institute, all patients with gliomas are registered in the brain tumor database of our department. Tumors were histologically diagnosed by experienced neuropathologists (SOS and TI) and graded according to WHO criteria (2007). From this database, we extracted information on patients diagnosed with glioma and glioneuronal tumors between 2002 and 2016, including 411 patients with the following gliomas (type, number of patients): GBM, 190; gliosarcoma, 5; anaplastic astrocytoma, 33; anaplastic oligodendroglioma, 27; anaplastic oligoastrocytoma, 3; diffuse astrocytoma, 39; oligodendroglioma, 23; oligoastrocytoma, 2; gliomatosis cerebri (GC), 7; chordoid glioma, 1; astroblastoma, 2; pilomyxoid astrocytoma, 3; pilocytic astrocytoma 28; pleomorphic xanthoastrocytoma, 3; ganglioglioma, 13; dysembryoplastic neuroepithelial tumor, 7; rosette-forming glioneuronal tumor, 1; and undefined glioma and glioneuronal tumor, 23. Recurrent tumors were included when surgery for glioma was the first surgery at our institution. However, recurrent tumors in the same patient were excluded except when the recurrent tumor showed histological malignant transformation compared to the previous tumor. Patient age ranged from 1 to 87 years (median 46 years). Regarding sex, 219 patients were males and 188 were females. Sixty-nine patients were younger than 20 years of age. Genetic alterations were determined using snap-frozen tumor tissue samples obtained at the time of surgery.
Samples and DNA preparation
A portion of each sample was saved for histopathological examination and the remainder was snap-frozen in liquid nitrogen and stored at −80 °C. DNA was extracted from tumors using a QIAamp DNA mini kit (Qiagen Science, Germantown, MD, USA) according to the manufacturer’s protocol. This investigation was approved by the Ethics Committee of Kyushu University.
Histopathology
Tissues were fixed in 10% formalin and formalin-fixed, paraffin-embedded glioma tissues were cut into 6-µm-thick sections, which were routinely stained with hematoxylin and eosin. Immunohistochemical analysis was performed using the following antibodies: rabbit anti-GFAP (1:1000; Dako A/S, Glostrup, Denmark), mouse anti-p53 (1:100; ab-2, Calbiochem, Darmstadt, Germany), mouse anti-Ki-67 (1:100; MIB-1, Dako, Santa Clara, USA), rabbit anti-ATRX (1:200; Sigma, Darmstadt, Germany), rabbit anti-H3K27me3 (1:500; Millipore, Darmstadt, Germany), rabbit anti-H3K9me3 (1:400; Abcam, Cambridge, UK), and rabbit anti-H3K4me3 (1:200; Abcam, Cambridge, UK). Immunostaining was performed using either a Bond III autostainer (Leica Microsystems, Wetzlar, Germany) with the Bond Polymer Refine Detection kit (Leica Microsystems) for GFAP, p53, and Ki-67, or was performed manually using a Vectastain Elite ABC Kit (Vector Laboratories, CA, USA) according to the manufacturer’s instructions for ATRX, H3K27me3, H3K9me3, and H3K4me3.
Genetic analysis
To screen for H3F3A mutations, we performed high-resolution melting (HRM) analysis as described previously [15]. Briefly, all HRM reactions were performed using MeltDoctor HRM Master Mix (Applied Biosystems, Tokyo, Japan) according to the manufacturer’s protocol. PCR reactions were performed using a 7500 Fast Real-Time PCR system (Applied Biosystems) with the following primers (forward primer: TAAAGCACCCAGGAAGCAAC, reverse primer: CAAGAGAGACTTTGTCCCATTTT). The HRM v2.0 software (Applied Biosystems) was used for HRM analysis. Subsequently, we used direct sequencing to determine the sequences of the mutations. HRM products were purified using ExoSAP-IT (Affymetrix/USB, Santa Clara, CA, USA), followed by cycle sequencing using BigDye® Terminator v3.1 Cycle Sequencing Kits (Applied Biosystems). Following purification, electrophoresis and analysis were conducted using a PRISM® 310 Genetic Analyzer (Applied Biosystems). For all H3F3A-mutant tumors, HIST1H3B, IDH1, IDH2, and BRAF V600E were assessed by HRM and sequencing, and TERT promoter mutations were analyzed by direct sequencing [16]. Loss of heterozygosity (LOH) on chromosome 1p and 19q was analyzed as described previously [16].
Results
Frequency and patient demographics for H3F3A mutations
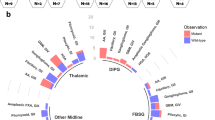
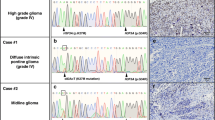
HRM analyses revealed that 14 patients (3.4%) harbored H3F3A mutations among the 411 gliomas examined in this study. Direct Sanger sequencing identified that four cases contained the G34R mutation (Fig. 1) and ten patients harbored the K27M mutation. No G34V mutations were found. We identified two K27R mutant tumors that had not previously been reported: one was anaplastic astrocytoma, discovered in a patient of 37 years of age, and the other was oligodendroglioma, in a patient of 51 years of age. For all H3F3A-mutant tumors, HIST1H3B, IDH1, IDH2, BRAF V600E, and TERT promoter mutations were not identified. None of the H3F3A-mutant tumors had 1p/19q codeletion. Among young patients (under the age of 20), the frequency of G34R and K27M was 4.3 and 14.5%, respectively. Patient demographics are summarized in Table 1. The ages of patients ranged from 8 to 37 years (median, 10.5) for G34R-mutant tumors, whereas they ranged from 5 to 66 years (median, 15) for K27M mutant tumors. Concerning histopathological diagnosis, G34R-mutant tumors included three GBMs and one astroblastoma, whereas K27M mutant tumors were all diagnosed as GBM. When assessing the 12 pediatric GBM patients (under the age of 15), an H3F3A mutation was detected in 10 of 12 cases (G34R:3, K27M:7). Anatomically, G34R-mutant tumors were located in the parietal region in two patients and the basal ganglia in one patient, and were diagnosed as the multi-lobular type in one patient. In K27M-mutant tumors, the tumors were located in the thalamus in six patients, the pons in one patient, the spine in one patient, and the basal ganglia in one patient, and were diagnosed as the multi-lobular type in two patients. Overall survival was analyzed in the patients for whom the results were available. Four G34R-mutant tumors and nine K27M-mutant tumors were analyzed. Results showed that the overall survival was not statistically different between the two groups (Fig. 2).
Case illustrations of G34R-mutant gliomas
Case 1 (Fig. 3)
An 8-year-old boy presented with a headache, right motor weakness, and bilateral oculomotor palsy. MRI demonstrated marked extension of a hyperintensity lesion in a T2 weighted and fluid attenuated inversion recovery (FLAIR) image involving the left temporal lobe, basal ganglia, corona radiata, and right thalamus, suggesting a GC growth pattern. The tumor was not contrast enhanced. Left temporal lobectomy was performed and diagnosis was GBM. Postoperatively, he was treated by concomitant temozolomide (TMZ) and radiotherapy (60 Gy). However, the tumor rapidly progressed, with the identification of a marked contrast-enhanced lesion in the left frontal lobe. Intractable generalized seizures became evident and the patient died 304 days after surgery.
MRI features of case 1. Preoperative MRI demonstrated marked extension of a hyperintensity lesion in the fluid attenuated inversion recovery (FLAIR) image involving the left temporal lobe (a), basal ganglia and corona radiata (b), and right thalamus without contrast enhancement (c). A contrast-enhanced lesion was evident in the left frontal lobe after tumor progression (d)
Case 2 (Fig. 4)
A 10-year-old girl was admitted to a nearby hospital because of a generalized seizure. Upon transfer to our hospital, she suffered from mild left hemiparesis. An MRI FLAIR image demonstrated that the hyperintensity lesion extended from the right parietal lobe to the thalamus, compressing the lateral ventricle. The tumor was slightly contrast-enhanced in the center with restricted signal in the diffusion-weighted image. With CT, intratumoral calcification was evident. The tumor was partially removed by craniotomy and the pathological diagnosis was GBM. Although radiotherapy (52 Gy) and chemotherapy with cisplatin and vincristine were administered, the tumor gradually progressed and extensively invaded the corpus callosum, left temporal lobe, and right frontal lobe. Her general condition got worse and she died 501 days after surgery.
Neuroradiological features of case 2. Preoperative CT imaging showed a slight high-density lesion with intratumoral calcification in the right parietal lobe extending to the thalamus (a). An MRI fluid attenuated inversion recovery (FLAIR) image demonstrated tumor hyperintensity (b). Slight contrast enhancement was observed in the center of the tumor. After progression, the tumor extended along the corpus callosum with marked enhancement (c). The slightly enhanced tumor invaded the right frontal lobe and left temporal lobe (d)
Case 3 (Fig. 5)
An 11-year-old autistic boy presented with generalized seizure. The CT showed a low-density lesion in the left parietal lobe with intratumoral calcification. MRI identified a cystic tumor in the left parietal-to-temporal lobe with indefinite tumor margins, indicating invasive tumor growth. The tumors showed scarce contrast enhancement. Partial tumor resection was performed and it was diagnosed as astroblastoma by pathological examination. Postoperatively, he was treated with concomitant TMZ and radiotherapy (60 Gy). However, he died 439 days after surgery because of tumor progression.
Neuroradiological features of case 3. A preoperative CT showed a low-density lesion in the left parietal lobe with intratumoral calcification (a). MRI fluid attenuated inversion recovery (FLAIR) imaging demonstrated a cystic tumor in the left parietal-to-temporal lobe with indefinite tumor margins (b). The tumor shows scarce contrast enhancement (c). After progression, the tumor invaded into the contralateral side of the FLAIR image (d)
Case 4 (Fig. 6)
A 37-year-old woman presented with mild right motor weakness and sensory disturbance. MRI imaging showed an FLAIR hyperintensity region extending from the left basal ganglia to the paraventricular region involving the frontal lobe with scarce contrast enhancement. Stereotactic biopsy of the tumor in the basal ganglia revealed that the tumor was GBM. Although she was treated by nimustine chloride (ACNU) and radiotherapy post-surgery, the tumor progressed, and she died 526 days after surgery.
Histomorphological characteristics of G34R-mutant gliomas
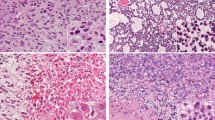
In all four cases, the specimens showed diffuse proliferation of anaplastic glioma cells with nuclear atypia at a high density (Fig. 7). Multinucleated giant cells were recognized in two cases (cases 1 and 2). Well-developed cytoplasmic processes were identified in all cases. However, part of the tumor in case 2 showed densely packed undifferentiated cells of small size with scant cytoplasm and few cytoplasmic processes. The histopathological diagnosis of cases 1, 2, and 4 was GBM with highly anaplastic cells. In case 3, the tumor cells frequently showed astroblastic rosette formation as well as short spindle cytology, and the case was diagnosed as astroblastoma. Immunohistochemically, three of the four tumors showed nuclear expression of p53. Nuclear staining for ATRX was completely lost in all four specimens. H3K27me3 staining was lost in two specimens and was absent and retained in the other two, although the two immunopositive specimens showed heterogeneous expression (Table 2; Fig. 8). Stained endothelial cells or normal cells were used as positive controls. All four specimens demonstrated nuclear immunostaining to H3K4me3, and three specimens showed an immunoreaction to H3K9me3.
Histomorphological features of G34R-mutant specimens. Anaplastic glioma cells show diffuse proliferation with nuclear atypia (a) and positive p53 immunostaining (b) in case 1. The tumor cells show diffuse proliferation (c) with an undifferentiated small cell component with sparse cytoplasm and few cytoplasmic processes in some parts (d) of case 2. The tumor cells frequently show astroblastic rosette formation as well as short spindle cytology (e) with GFAP expression (f) in case 3. Staining for p53 is also detected in this case (g). The tumor cells show diffuse proliferation with cytoplasmic processes in case 4 (h)
Discussion
In this study, we first demonstrated the frequency of G34R mutations in Japanese patients with glioma and found that they occurred less frequently than K27M mutations. When confined to pediatric and young adults (less than 20 years), we detected G34R mutations at a frequency of 4.3% of all glioma cases, whereas K27M mutations were found to have a frequency of 14.5%. In the first report of H3F3A mutations in 48 pediatric GBMs, the frequency of G34R/V mutation and K27M mutation was found to be 13 and 19%, respectively [1]. The G34V mutation is rare and has been detected in only a single patient [1] and cell line [8]. A German group study analyzing 77 GBM patients under the age of 30 showed that G34R/V and K27M mutations were detected at a frequency of 16 and 32%, respectively [17]. These results imply that G34R/V mutations might be less frequent in Japan than in western countries. Sturm et al. demonstrated that the average age of patients harboring G34R/V mutations is slightly higher than that for patients with K27M mutations; the median age for the G34R/V mutation is 18 years (range 9–42 years) and that for K27M is 10.5 years (range 5–23 years) [6]. However, our study showed that patient age was similar (median age 10 vs. 15 years) in both cases. In addition, although most studies have reported that patient prognosis for G34R/V mutations is better than that of K27M mutations [5, 6], our results did not show a significant difference. This discrepancy might be due to the small number of the G34R-mutant tumors assessed in our study.
We have demonstrated some important findings in this study. First, G34R-mutant tumors can be located not only in the hemisphere, but also in the basal ganglia and deep-seated region, away from the thalamus. Importantly, despite the presence of histopathological high-grade tumors, four specimens with G34R mutations showed weak or no contrast enhancement. Furthermore, in one case, the tumor extensively invaded into more than three lobes including deep gray matter structures such as the basal ganglia and thalamus, which is consistent with a GC growth pattern. Broniscer et al. have recently identified four G34R-mutant tumors from 32 GC cases; however, a K27M mutation was not identified in this subset [18]. Our results, in combination with this report, suggest that G34R/V mutations play a role in the pathogenesis of a subset of GC cases. Given that the diagnostic entity of GC was deleted and described as a growth pattern in the recently revised version of the WHO’s classification of central nervous system tumors [19], and the fact that a previous nosological entity of GC is now classified depending on molecular alterations shared with GBM, we hypothesize that a subset of GC cases could be caused by G34R/V mutations.
Second, our results suggest that G34R/V mutations also play a role in the pathogenesis of astroblastoma. Astroblastoma is a rare morphological entity characterized by the existence of GFAP-positive tumor cells radiating towards central blood vessels (astroblastic pseudorosettes) [20]. Reports on the molecular analysis of astroblastoma are scarce and no characteristic genetic alterations have been reported. Fu et al. reported an astroblastoma case that lacked mutations in both IDH1/2 and TP53, suggesting that molecular alterations in astroblastoma are different from those of adult astrocytic tumors [21]. Our group and Lehman et al. recently reported BRAF V600E mutation in astroblastoma [16, 20]. In addition, Strum et al. recently reported frequent MN1 alterations in astroblastoma, suggesting that a subset of this disease is characterized by genetic alterations distinct from conventional diffuse glial tumors [22]. In this study, we first demonstrated that the G34R/V mutation is a genetic alteration that might be associated with the oncogenesis of astroblastoma.
Concerning genotype–phenotype correlations for G34R/V mutations, Gessi et al. reported that G34R/V is detected not only in pediatric GBM but also in pediatric primitive neuroectodermal tumors of the central nervous system (CNS-PNET) [23]. Korshunov et al. extended this observation and demonstrated that G34R/V-mutant CNS tumors comprise a single biological entity with variable morphological appearances, and are either Glioblastoma-like or PNET-like, because G34R/V-mutant GBM and CNS-PNET specimens show similar histopathological and molecular features and are associated with the same prognosis; this was based on an analysis of 81 G34R/V-mutant CNS tumors [24]. However, their subsequent study also demonstrated that histomorphological features such as PNET-like components can occur in different molecular subgroups, based on IDH, K27M, and G34R/V mutations, although a PNET-like morphology is more common in G34R/V-mutant tumors [17]. Given that recent studies (including those mentioned herein) support the notion that CNS-PNET represents a molecular heterogeneous group of tumors [22], the term “CNS-PNET” was deleted from the diagnostic lexicon of the WHO’s classification in 2016 [25]. Therefore, we suggest that G34R/V mutations could be recognized in a subset of glioblastoma with a primitive neuronal component.
Immunohistochemically, all four G34R mutant tumors showed loss of ATRX expression, which is consistent with previous reports [1, 5, 6, 26]. These results suggest that not only K27M mutation but also G34R/V mutation is associated with alternative lengthening of telomeres [1]. Regarding histone methylation, our results differed from those of the original report, concluding that H3K27me3 aberrations are an exclusive feature of K27M-mutant tumors and are not found in G34R-mutant tumors [13, 14]. However, a subsequent study demonstrated that alterations in H3K27me3 could also be detected in a subset of G34R-mutant tumors [9], which is consistent with the results obtained in our study; reduction of H3K9me3 was observed in one G34R-mutant tumor. Given that H3K27me3 and H3K9me3 are generally associated with repression, reduction of H3K27me3 and H3K9me3 can have some effect on gene expression in G34R-mutant tumors. In addition, it has been known that G34R/V mutation affects the methylation of H3K36, which is an activator of gene expression [8, 12]. Therefore, these results suggest that altered histone methylation is a common phenomenon in G34R/V-mutant tumors.
In summary, G34R/V mutations are relatively rare even in pediatric high-grade glioma patients. Although we reported on a limited number of patients with a G34R/V mutation, we demonstrated that the G34R/V mutation can be detected not only in typical GBM but also in astroblastoma and in a subset of anaplastic glioma with a GC growth pattern. This suggests that the relationship between the G34R/V-mutant genotype and phenotype is more complex. In addition, we identified an epigenetic effect, and specifically that altered global histone trimethylation can play an important role not only in K27M-mutant tumors but also in G34R/V-mutant tumors. It is expected that future studies will clarify the genetic and epigenetic effects that result from H3F3A G34R/V mutations, and that target therapy will be developed based on this mutation in the near future.
References
Schwartzentruber J, Korshunov A, Liu XY et al (2012) Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482:226–231
Wu G, Broniscer A, McEachron TA et al (2012) Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat Genet 44:251–253
Wu G, Diaz AK, Paugh BS et al (2014) The genomic landscape of diffuse intrinsic pontine glioma and pediatric non-brainstem high-grade glioma. Nat Genet 46:444–450
Fontebasso AM, Liu XY, Sturm D et al (2013) Chromatin remodeling defects in pediatric and young adult glioblastoma: a tale of a variant histone 3 tail. Brain Pathol 23:210–216
Sturm D, Bender S, Jones DT et al (2014) Paediatric and adult glioblastoma: multiform (epi)genomic culprits emerge. Nat Rev Cancer 14:92–107
Sturm D, Witt H, Hovestadt V et al (2012) Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell 22:425–437
Khuong-Quang DA, Buczkowicz P, Rakopoulos P et al (2012) K27M mutation in histone H3.3 defines clinically and biologically distinct subgroups of pediatric diffuse intrinsic pontine gliomas. Acta Neuropathol 124:439–447
Bjerke L, Mackay A, Nandhabalan M et al (2013) Histone H3.3. mutations drive pediatric glioblastoma through upregulation of MYCN. Cancer Discov 3:512–519
Pathak P, Jha P, Purkait S et al (2015) Altered global histone-trimethylation code and H3F3A-ATRX mutation in pediatric GBM. J Neurooncol 121:489–497
Bender S, Tang Y, Lindroth AM et al (2013) Reduced H3K27me3 and DNA hypomethylation are major drivers of gene expression in K27M mutant pediatric high-grade gliomas. Cancer Cell 24:660–672
Chan KM, Fang D, Gan H et al (2013) The histone H3.3K27M mutation in pediatric glioma reprograms H3K27 methylation and gene expression. Genes Dev 27:985–990
Lewis PW, Muller MM, Koletsky MS et al (2013) Inhibition of PRC2 activity by a gain-of-function H3 mutation found in pediatric glioblastoma. Science 340:857–861
Venneti S, Garimella MT, Sullivan LM et al (2013) Evaluation of histone 3 lysine 27 trimethylation (H3K27me3) and enhancer of Zest 2 (EZH2) in pediatric glial and glioneuronal tumors shows decreased H3K27me3 in H3F3A K27M mutant glioblastomas. Brain Pathol 23:558–564
Venneti S, Santi M, Felicella MM et al (2014) A sensitive and specific histopathologic prognostic marker for H3F3A K27M mutant pediatric glioblastomas. Acta Neuropathol 128:743–753
Hatae R, Hata N, Yoshimoto K et al (2016) Precise detection of IDH1/2 and BRAF hotspot mutations in clinical glioma tissues by a differential calculus analysis of high-resolution melting data. PLoS One 11:e0160489
Hatae R, Hata N, Suzuki SO et al (2016) A comprehensive analysis identifies BRAF hotspot mutations associated with gliomas with peculiar epithelial morphology. Neuropathology
Neumann JE, Dorostkar MM, Korshunov A et al (2016) Distinct histomorphology in molecular subgroups of glioblastomas in young patients. J Neuropathol Exp Neurol 75:408–414
Broniscer A, Chamdine O, Hwang S et al (2016) Gliomatosis cerebri in children shares molecular characteristics with other pediatric gliomas. Acta Neuropathol 131:299–307
Herrlinger U, Jones DT, Glas M et al (2016) Gliomatosis cerebri: no evidence for a separate brain tumor entity. Acta Neuropathol 131:309–319
Lehman NL, Hattab EM, Mobley BC et al (2017) Morphological and molecular features of astroblastoma, including BRAFV600E mutations, suggest an ontological relationship to other cortical-based gliomas of children and young adults. Neuro Oncol 19:31–42
Fu YJ, Taniguchi Y, Takeuchi S et al (2013) Cerebral astroblastoma in an adult: an immunohistochemical, ultrastructural and genetic study. Neuropathology 33:312–319
Sturm D, Orr BA, Toprak UH et al (2016) New brain tumor entities emerge from molecular classification of CNS-PNETs. Cell 164:1060–1072
Gessi M, Gielen GH, Hammes J et al (2013) H3.3 G34R mutations in pediatric primitive neuroectodermal tumors of central nervous system (CNS-PNET) and pediatric glioblastomas: possible diagnostic and therapeutic implications? J Neurooncol 112:67–72
Korshunov A, Capper D, Reuss D et al (2016) Histologically distinct neuroepithelial tumors with histone 3 G34 mutation are molecularly similar and comprise a single nosologic entity. Acta Neuropathol 131:137–146
Louis DN, Perry A, Reifenberger G et al (2016) The 2016 world health organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820
Ebrahimi A, Skardelly M, Bonzheim I et al (2016) ATRX immunostaining predicts IDH and H3F3A status in gliomas. Acta Neuropathol Commun 4:60
Acknowledgements
This study was funded by the Ministry of Education, Culture, Sports, Science and Technology of Japan (Grant No. 26462185).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Rights and permissions
About this article
Cite this article
Yoshimoto, K., Hatae, R., Sangatsuda, Y. et al. Prevalence and clinicopathological features of H3.3 G34-mutant high-grade gliomas: a retrospective study of 411 consecutive glioma cases in a single institution. Brain Tumor Pathol 34, 103–112 (2017). https://doi.org/10.1007/s10014-017-0287-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10014-017-0287-7