Abstract
Studies on the interplay between morphological diversity and genomic duplication are crucial, since novel traits may have profound evolutionary and ecological consequences. Within the Lythraceae, Cuphea stands out as the most karyotypically diverse genus, exhibiting a broad distribution across contrasting ecosystems in the Neotropics such as wetlands, forests, savannas, and prairies, ranging from coastal regions to the central continent, and from sea level to mountainous areas, with a remarkable array of morphological variation in vegetative and reproductive structures. To elucidate the potential relationships between karyotypic diversity and morphological variability in Cuphea, we compiled a morphological database, chromosome numbers and a molecular phylogenetic tree to infer the phylogenetic signal of morphological traits and karyotype evolution in this genus. Our findings confirm the pivotal role of polyploidy in the extensive karyotypic diversification observed in Cuphea. Ancestral state reconstruction revealed that the ancestral chromosomal number for the genus is x = 11 and there is a high incidence of polyploidy in the genus, with at least 11 events of whole-genome duplication, in addition to 20 events of dysploidy (10 ascending and 10 descending). We found evidence to support a positive correlation between chromosome numbers and overall size of selected morphological traits with polyploid species showing longer floral tubes. Our investigations also revealed a potential association between polyploidy and the adaptive radiation of Cuphea during its distribution expansion to the North America. These results reinforce the importance of whole-genome duplication events in producing trait diversity and, consequently, speciation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chromosomes represent the final organizational stage of the nuclear genome and have relative stability throughout the reproductive process. Thus, changes in chromosome numbers serve as key mechanisms for understanding the evolutionary history of a group (King 1995; Graham and Cavalcanti 2001; Carta et al. 2020). Polyploidy, characterized by the whole-genome duplication (WGD), is a widespread phenomenon in flowering plants (Meyers and Levin 2006; Wood et al. 2009). Polyploids are commonly classified into two major categories based on their genotypic composition: autopolyploids, formed by the duplication of homologous chromosomes, and allopolyploids, resulting from interspecific hybridization followed by chromosome set duplication or fusion of unreduced gametes. In both cases, the nuclear content comprises non-homologous chromosomes (Parisod et al. 2010). Together with WGD, dysploidy events, which encompass variations in chromosome number occasionally induced by chromosomal fissions and/or fusions, represent major karyological change phenomena in plant evolutionary history (Escudero et al. 2014).
Considered intrinsic to plant evolution, WGD events are accompanied by gene loss, differential expression, or neofunctionalization, promoting increased genetic and genomic variation (Blanc and Wolfe 2004; Soltis et al. 2009; Ramsey and Ramsey 2014). Polyploid individuals when compared to their diploid ancestors, are known for their enhanced adaptability to new habitats, with a greater capacity to occupy new ecological niches (Baniaga et al. 2020; Otto and Whitton 2000). Although these phenomena typically yield individuals that initially have lower adaptive value than their progenitors, they are subject to natural selection that may favor a small fraction that can better adapt to a distinct environment (Stebbins 1971). Consequently, even though the phenotypic impact may be subtle, the resulting differences are of utmost importance for the evolutionary success of new polyploid lineages (Otto and Whitton 2000).
One possible consequence of polyploidy in plants is the fixation of mutations that can generate morphological variation. This variation is primarily characterized by an increase in cell size (Otto and Whitton 2000), including changes in stomatal size and density (Balao et al. 2011; Beaulieu et al. 2008; Munzbergová 2009), and also affecting various vegetative and reproductive traits, such as larger flowers and seeds (Chansler et al. 2016; Griesbach 1985). Understanding the relationship between morphological diversity and WGD events is crucial, as the selection of novel morphological characteristics can facilitate the adaptation to new environments, leading to evolutionary and ecological consequences (Garbutt and Bazzaz 1983; Li et al. 2010; Chansler et al. 2016; Soltis et al. 2016; Van de Peer et al. 2017; Gloria 2020; Moraes et al. 2022).
Numerous angiosperm families display extensive karyotypic variation, and several polyploidy events, including Asteraceae, Fabaceae, and Orchidaceae (Goldblatt 1980; Bairiganjan and Patnaik 1989; Weiss-Schneeweiss et al. 2008; Doyle 2012; Hedrén et al. 2018; Senderowicz et al. 2021). Lythraceae, a eudicot family of Myrtales (Chase et al. 2016), encompasses 28 genera (Facco and Cavalcanti 2023; Inglis et al. 2023). Among them, Cuphea is the most diverse, comprising around 260 species (Facco and Cavalcanti 2023). Cuphea species are herbaceous or small perennial shrubs known for their substantial morphological variation in vegetative and reproductive organs (Fig. 1). The native geographical distribution of Cuphea is limited to the Neotropics, with accidental introductions in Africa naturalized and possibly in China, spanning diverse ecosystems such as wetlands, forests, savannas, and prairies, ranging from coastal regions to the central continent, and from sea level to mountainous areas (Facco and Cavalcanti 2023). Cuphea represents the most karyotypically diverse genus within Lythraceae, with haploid chromosome numbers ranging from 6 to 86 (Graham 1992; Graham and Cavalcanti 2001; Pozzobon et al. 2022).
Detailed taxonomic studies on Cuphea have been ongoing since the late twentieth century, involving taxonomic revisions and the description of new species (Brauner and Cavalcanti 2018; Cavalcanti and Graham 2008; Facco 2015; Facco and Cavalcanti 2022, 2023; Graham 1988, 1989a, 2017, 2019a, 2019b; Graham and Cavalcanti 2013). The advancement of Cuphea phylogeny, combined with available taxonomic and cytogenetic knowledge, provides ample opportunities for investigating hypotheses concerning the evolution of chromosome numbers. Previous studies hypothesized a potential relationship between chromosomal diversity and the taxonomic and morphological richness of Cuphea, indicating that the genus diversity may be linked to polyploidy and aneuploidy events (Graham and Cavalcanti 2001; Barber et al. 2010).
Thus, we conducted an investigation using a molecular phylogeny to: 1. Explore the evolution of chromosome numbers in the genus by reconstructing ancestral chromosome numbers; 2. Assess the presence of phylogenetic signals for morphological characters and karyotypes; 3. Determine possible correlations between karyotypes and leaf and floral morphology. Considering that Cuphea represents a widely distributed genus across heterogeneous ecosystems, and exhibits diverse morphology with a broad range of karyotypes, we hypothesize that polyploidy events throughout its evolutionary history are associated with the vegetative and reproductive morphological variation observed in the genus.
Material and methods
Taxon sampling
The chromosome number data available for Cuphea species was gathered using the Chromosome Counts Database—CCDB (2021, continuously updated) and the IAPT/IOPB Chromosome Data (2021, continuously updated). These databases provide information from various sources, including Dollon and Hamel (1967), Graham (1968a, b, 1980, 1982, 1987, 1988, 1989a, 1989b, 1992), Graham and Cavalcanti (2001, 2007), Goldblatt (1981), Guha (1972), Molero et al. (2002), Ornduff (1967), and Pozzobon et al. (2022). A total of 104 species with reliable data were sampled, representing approximately 40% of the genus. In cases where different karyotypic counts were reported for the same species (3%), the highest recorded chromosome number was considered. Species with supernumerary chromosomes were included, although the B chromosomes were not counted (Online Resource 1).
A morphological database was established for the 104 Cuphea species with available chromosomal data. Morphological traits related with species environment adaptations, such as leaf length and width (Knight and Ackerly 2003; Ackerly 2004), along with reproductive biology traits, floral tube length was measured (Vichiato et al. 2014, Caruso et al. 2019). These measurements were obtained from taxonomic revisions of the genus and other published sources (Graham 1968a, b, 1988, 1989a, 1990, 1998, 2017; Cavalcanti and Graham 2008; Facco 2015; Brauner 2018) and, when necessary, supplemented with images from online botanical collections (eg. K, P, MO, NY, RB, R, UB, UFMT) utilizing the software ImageJ for accurate measurements (Rasband 2012).
Out of the 104 characterized species, sequences from 54 species were retrieved from the Genbank database for constructing a phylogenetic tree. This selection, although representing ca. 21% of the genus, encompassed nearly all sections of Cuphea, with the exception of C. sect. Amazoniana for which karyotype data were not available in the literature (Graham 2019b). Geographically, the sampling covered the entire distribution range of the genus (Graham 1968a, b, 1988, 1989a, 1990, 1998, 2017; Cavalcanti and Graham 2008; Facco 2015; Brauner 2018). Molecular markers, including the plastid spacer trnL-trnF, plastid intron rpl16, and nuclear spacer (rDNA) ITS (including 5.8S), were used for the phylogenetic analysis. Most species have at least one nuclear and one plastid marker. Pleurophora anomala (A.St.-Hil.) Koehne was chosen as the outgroup based in Barber et al. (2010).
Phylogenetic analyses
Sequences of each marker were aligned into independent matrices using the MUSCLE (Edgar 2004) on the Geneious platform (Biomatters, Auckland, New Zealand), and alignments were subsequently manually optimized. The best-fitting nucleotide substitution model for each data matrix (ITS, trnL-trnF, and rpl16) was selected using JModelTest v.2.1.5 (Darriba et al. 2012) under the Akaike information criterion (AIC). The GTR + G model was found to be the most appropriate for all three matrices.
The matrices were then concatenated, and Bayesian inference (BI) analyses were performed using Mr. Bayes v3.2 (Ronquist et al. 2012) implemented on CIPRES Science Gateway (Miller et al. 2009) for both each individual data matrix and the concatenated matrix. BI was performed with two simultaneous runs and four chains each. The Markov chain Monte Carlo (MCMC) parameters were set to 20 million generations, sampling every 1000 trees. The initial 25% of trees were discarded as burn-in. Convergence between the two independent runs was checked with Tracer 1.6 (Rambaut et al. 2018) based on the estimated sample size value (> 200). Clades with a posterior probability (PP) greater than or equal to 0.95 were considered to have strong support; while, clades with PP values between 0.94 and 0.90 were considered moderately supported (Cummings et al. 2003; Erixon et al. 2003).
The resulting phylogenetic trees were generated only for the purpose of assessing branch support, identifying well-supported incongruences, and validating the robustness of the inferred phylogenetic relationships. During this process, 10 species were excluded from the analyses due to strongly supported incongruences observed among the independent marker trees (ITS, trnL-trnF, and rpl16), which promoted the occurrence of polytomies in the concatenated data tree. Incongruent phylogenetic patterns can be the result of hybridization, technical issues like misidentification, contamination, uncertainty in phylogenetic reconstruction, inaccurate orthology assessment, analytical artefacts such as branch attraction, horizontal gene transfer, and incomplete lineage sorting (Guo et al. 2018; Davis and Xi 2015; Wendel and Doyle 1998). Thus, to scape these possibilities the final dataset included 44 species (approximately 17% of the genus) for subsequent analyses (Fig. 2).
Consensus phylogram derived from a Bayesian inference analysis using nuclear (ITS) and plastidial (trnL-trnF and rpl16) marker data for Cuphea species. Posterior probability (PP) values are displayed above the branches. The current classification of species sections is positioned next to the scientific name and follows the following acronyms: ARC (C. sect. Archocuphea); BRA (C. sect. Brachyandra); CUP (C. sect. Cuphea); DIP (C. sect. Diploptychia); EUA (C. sect. Euandra); HTD (C. sect. Heterodon); LEP (C. sect. Leptocalyx); MEL (C. sect. Melvilla); and TRI (C. sect. Trispermum). The five major clades are identified by the black bars
Cytogenetics
To reconstruct changes in basic chromosomic numbers, an ultrametric tree including all 44 species was constructed using both ITS and plastid datasets, employing BEAST 1.8.4 (Drummond et al. 2012) on CIPRES Science Gateway (Miller et al. 2009). The BEAUti 1.8 program was used to create the input files, applying the same evolutionary models cited above. A lognormal relaxed molecular clock model was applied for this analysis, following the Yule process prior. The MCMC parameters were set to perform 150,000,000 generations, sampling every 1500 iterations. The resulting tree was evaluated using Tracer v1.6 (Rambaut et al. 2018), and the final tree was constructed by processing the post-burn-in trees using TreeAnnotator v1.10.4 (Bouckaert et al. 2019).
The reconstruction of ancestral states for chromosome number was performed using ChromEvol v.2 software (Mayrose et al. 2009; Glick and Mayrose 2014) implemented in the RASP platform (Yu et al. 2015). To accomplish this, we employed the ultrametric tree generated by our analysis and the available chromosome number data provided in Online Resource 1. All available models of chromosome number evolution in ChromEvol were evaluated to determine the best model to explain the observed variation in chromosome numbers. The optimal model, as determined by the Akaike information criterion (AIC), was identified as CONST_RATE. The analysis encompassed 10,000 simulations; while, the basic number considered for analysis was x = 8.
Phylogenetic signal
In order to examine potential associations between the evolutionary history of the genus and the distribution of morphological characteristics, as well as karyotypes, phylogenetic signal analyses were employed. To accomplish this, we used our ultrametric tree and the morphological traits investigated included the minimum and maximum measurements of leaf length and width, and floral tube length. Subsequently, Pagel’s λ coefficient (Pagel 1999) was estimated for each selected trait using the 'phylosig' function on the 'phytools' package v1.9-16 (Revell 2012). Pagel’s λ represents a measure of phylogenetic signal, indicating the extent to which the evolutionary history accounts for the observed distribution of traits among the terminal taxa. Furthermore, this function conducts a hypothesis test to assess the statistical significance of the obtained lambda values. That means, if closely related species resembles each other more closely than expected by chance alone. The lambda values range from 0 to 1, where 0 indicates that trait evolution is independent of phylogeny, while 1 signifies the maximum degree of phylogenetic signal, e.g., trait evolution follows the exact pattern of the phylogenetic tree (Münkemüller et al. 2012).
These analyses were instrumental in evaluating whether the potential correlations between morphological traits and karyotypes could be attributed to the evolutionary history of the group, rather than implying a causative relationship. If both the morphological and cytogenetic datasets exhibit significant phylogenetic signal, it becomes challenging to ascertain whether the observed correlation arose due to ancestral traits shared among descendants (indirect relationship, mere causality) or whether it is a consequence of recent independent events, such as genomic structural changes (e.g., polyploidy) in terminal nodes, thereby enhancing adaptive processes that favor the fixation of morphological traits. Conversely, if no phylogenetic signal is detected in one of the datasets, any putative correlation cannot be explained by indirect factors such as the conservation of ancestral traits and is more likely to possess a cause-and-effect relationship between morphological traits and karyotypes.
The mapping of morphological characters into the ultrametric tree was executed utilizing the 'contMap' function on the 'phytools' package v1.9-16 (Revell 2012). This function, proposed by Revell (2012), facilitates the visualization of ancestral estimations for continuous characters through maximum likelihood (Schluter et al. 1997), interpolating states along branches utilizing the equation outlined in Felsenstein (1985). In the context of this study, its utility lies in examining whether the characters demonstrating phylogenetic signal in Cuphea exhibit a congruent topological distribution when compared to ancestral genomic alteration events related to chromosome numbers. The analyses for ascertaining the phylogenetic signal and character mapping within morphological datasets were executed using R software (R Core Team 2023).
Statistical analyses
The normality of the variables was verified using the Shapiro–Wilk test. Non-normal distributions were observed across all attribute sets, thus, nonparametric tests were performed. The analyses were undertaken in two distinct manners to investigate potential correlations between ploidy levels and the observed variables. To this end, the species were examined separately within two groups: diploids alone (59 spp.) and diploids combined with polyploids (104 spp.), following a similar approach to that employed by Moraes et al. (2022). In this context, both ascending and descending dysploidies were taken into consideration. The diploid group encompassed species with n = 6 to n = 12; whereas, the polyploid group encompassed species with n = 13 to n = 56. Spearman test was used to proceed with the correlation analyses between the chromosome number and morphological variables. Mann–Whitney test was applied to evaluate the potential differences between two morphological datasets, namely diploids (D) and polyploids (P). The statistical analyses were conducted using the R software (R Core Team 2023).
Results
Phylogenetic analyses
Our final phylogram, based on three markers (ITS, trnL-trnF, and rpl16) and derived from Bayesian Inference (Fig. 2), provides good support for five primary clades within Cuphea (clade 1, PP = 1.0; clade 2, PP = 1.0; clade 3, PP = 0.98; clade 4, PP = 1.0; clade 5, PP = 1.0) that were found in previous phylogenetic studies conducted by Graham et al. (2006), Graham and Cavalcanti (2007). Despite our smaller sample size, our analysis demonstrated a good resolution (Fig. 2).
According to Koehne (1903) classification, the first divergent lineage corresponds to C. subg. Cuphea (clade 1, PP = 1.0), including a species from C. subg. Bracteolatae sect. Brachyandra (C. circaeoides Sm. ex Sims) nested among the other taxa. Consequently, C. subg. Cuphea may be found to be non-monophyletic, as well as its two sections, C. sect. Cupheaand C. sect. Archocuphea, which are also not monophyletic. Together, clades 2, 3 and 5 represent C. subg. Bracteolatae and display moderate support (PP = 0.93). However, the sections C. sect. Brachyandra, C. sect. Diploptychia, C. sect. Heterodon, C. sect. Leptocalix, and C. sect. Melvilla, are also non-monophyletic. Notably, only C. sect. Trispermum is confirmed to be monophyletic as part of clade 3. A significant observation is the presence of a clade composed solely of North American species (clade 5). Here, we contribute to this clade by including C. salicifolia Schltdl. & Cham.
Cytogenetics
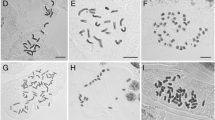
The retrieved chromosomal numbers exhibit a range from n = 6 (C. lanceolata W.T.Aiton) to n = 56 (C. retroscabra S.Watson), being n = 8 and n = 16 the most frequent (18 and 19 species respectively) (Fig. 3). Within Cuphea, the majority of chromosomal numbers are multiples of eight, suggesting a basic chromosomal number of x = 8. However, ancestral state reconstruction of chromosomal numbers revealed that the ancestral chromosomal number for the genus is x = 11 (n = 11–87.09%; n = 8–7.37%; Online Resources 2, 3 and 4). This number is primarily supported by its prevalence as the common karyotype in clades C1 and C2, which represent the earliest diverging lineages in the genus phylogeny (Fig. 4). Conversely, chromosomal numbers multiples of eight are more frequently observed in clades C3, C4, and C5, which contain most species with available molecular and chromosomal data. Chromosome numbers close to n = 8 largely originate from a descending dysploidy event in clade 10 (Fig. 4; n = 8–100%, Online Resources 2, 3 and 4), which represents the most recent common ancestor of C3 + C4 + C5.
Reconstruction of chromosome numbers in the genus Cuphea. The colors filling the circles represent the most probable ancestral chromosome number at each node of the tree. The numbers inside the circles identify the nodes of the tree. The recorded chromosome number changes are represented by squares with P↑ polyploidy; Dip↓ diploidization; D↑ ascending Dysploidy; D↓ descending dysploidy, *Cuphea node. **North American clade node
The examination of ancestral karyotypes in Cuphea reveals a high incidence of polyploidy in the genus, with at least 11 events of whole-genome duplication occurring in clades 23 (C4, Fig. 4; n = 8–35.46%; n = 14–32.82%, Online Resources 2, 3 and 4), 29 (C5, Fig. 4; n = 16–81.49%, n = 8–18.51%; Online Resources 2, 3 and 4), and 37 (C5, Fig. 4; n = 8–54.19%, n = 8–45.81%; Online Resources 2, 3 and 4). Additionally, polyploidy events are observed in terminal taxa such as C. pseudosilene Griseb., C. parsonsia (L.) R.Br. ex Steud., C. thymoides Cham. & Schltdl., C. hyssopifolia Kunth, C. spectabilis S.A.Graham, C. aequipetala Cav., C. appendiculata Benth., C. schumannii Koehne, and C. salicifolia Schltdl. & Cham (Fig. 4). The North American clade (C5) accounted for approximately 59% of the polyploidy events in the genus; while, C4 and C3 accounted for 41% of the events.
Dysploidy, characterized by changes in chromosomal numbers, also occurs recurrently in the group. Specifically, 10 events of ascending dysploidy were identified in clades 14 and 43, as well as in terminal taxa such as C. melvilla Lindl., C. pseudosilene Griseb., C. bustamanta Lex., C. spectabilis S.A.Graham, C. tolucana Peyr., C. appendiculata Benth., C. paucipetala S.A.Graham, and C. salicifolia Schltdl. & Cham. Furthermore, 10 occurrences of descending dysploidy were observed within the genus in clades 10, 20, 23, 24, 34, as well as in terminal taxa such as C. gaumeri Koehne, C. denticulata Kunth, C. avigera B.L.Rob. & Seaton, C. aequipetala Cav., and C. lutea Rose ex Koehne. Lastly, a diploidization event was noted in clade 34 (C5, Fig. 4; n = 7–100%, Online Resources 3 and 4).
Overall, it is apparent that alterations in chromosomal numbers in Cuphea primarily occur closer to the terminal branches of the phylogeny. The tree’s backbone consists of clades with the most recent common ancestor possessing n = 8 or n = 11 chromosomes. Additionally, these events are not clustered to specific clades but are dispersed across all major clades (Fig. 4).
Phylogenetic signal
The analyses demonstrated that, apart from maximum leaf width, all other morphological characteristics sampled in this study exhibited significant phylogenetic signal (λ = 0.73–0.45; p < 0.05) (Table 1). However, the chromosomal data exhibit no phylogenetic signal (λ < 0.05). Therefore, any observed correlation between morphology and karyotypes cannot be attributed to indirect factors such as the retention of ancestral characters. By observing the mapping of morphological traits into the phylogenetic tree topology, it is noted that species displaying the greatest lengths of floral tubes and the largest leaves are situated within the North American clade, as well as clade C2. Contrariwise, the smallest values are predominantly observed in clades C1 and C4 (Fig. 5).
Mapping of all leaf and flower morphological traits that exhibited phylogenetic signal in Cuphea. a Maximum floral tube length; b minimum floral tube length; c maximum leaf length; d minimum floral tube length; e minimum leaf width. The values a 4.5–40, b 3.5–25, c 8–140, d 2–70 and e 0.5–34 represent the range of sizes sampled in millimeters
Statistical analyses
The results obtained from the correlation tests indicate a statistically significant relationship between chromosomal numbers and certain morphological characteristics in Cuphea (ρ = 0.22–0.32). Analysis of the data with all ploidy species revealed a positive correlation between the chromosomal number and the minimum leaf length, as well as the minimum and maximum lengths of the floral tube. However, when considering only diploid species, no correlation was observed between the floral traits and the chromosomal number (Table 2). Furthermore, our investigation revealed significant variations in the length of the floral tube between polyploid and diploid species. Specifically, polyploid species exhibited longer floral tubes when compared to diploid species (Table 3; Fig. 6).
Discussion
Our research confirmed that the chromosomal evolution of the genus Cuphea exhibits multiple independent polyploidy events, and the pivotal role of WGD and dysploidy in the extensive karyotypic variation observed. We found phylogenetic signal for some morphological features but not for chromosome numbers, so the correlations found cannot be explained by conservation of ancestral traits and is more likely to possess a cause-and-effect relationship. There is compelling evidence suggesting that the evolution of floral tube and leaf sizes has been influenced by the occurrence of WGD events, as evidenced by a positive correlation with higher chromosomal numbers. The North American clade within Cuphea represents a noteworthy model for potential future deliberations regarding the contribution of polyploidy to adaptive radiations.
This study aimed to examine the potential relationship between karyotypic evolution and morphological character variation in Cuphea. The presence of different chromosome numbers within the genus can be attributed to events of WGD accompanied by chromosomal rearrangements, including descending and ascending dysploidy and subsequent diploidization (Lysák and Schubert 2013). Our findings demonstrate that polyploidy and dysploidy events played a crucial role in the karyotypic diversification of Cuphea, with 20% of the studied species exhibiting polyploidy events in terminal branches. This prevalence of polyploidy in terminal branches is an uncommon occurrence among angiosperms but has been observed in select genera such as Opuntia (Cactaceae, Majure et al. 2012), Epidendrum (Orchidaceae, Cordeiro et al. 2022), Turnera (Passifloraceae, Solis Nefa et al. 2022), Ulex (Fabaceae, Bellot et al. 2023), and Hieracium (Asteraceae, Krahulcová et al. 2009).
We were able to confirm several dysploidy events in the evolutionary history of Cuphea. Dysploidy has been proposed as one of the primary forces triggering karyotypic variation in plants (Carta et al. 2018), as observed in Asteraceae (Mas de Xaxars et al. 2016) and Marantaceae (Winterfeld et al. 2020). Changes in chromosomal numbers created by dysploidy events persist longer through evolutionary time compared to polyploid changes (Escudero et al. 2014), it may explain the high number of dysploidy events found in our analysis. We found at least nine ascending or descending events in terminal clades, indicating that they could be related to speciation as described to Artemisia L. (Asteraceae, Mas de Xaxars et al. 2016), Reichardia Roth (Asteraceae, Cho et al. 2022) Chamaecrista Moench (Fabaceae, Braz et al. 2024), and Passiflora L. (Passifloraceae, Sader et al. 2019), playing a crucial role in the diversification (De Storme and Mason 2014).
Our phylogenetic analysis exhibited higher resolution compared to that of Barber et al. (2010), who chose not to concatenate the matrices of plastidial and nuclear markers. By excluding taxa that caused significant incongruence between the matrices, we were able to produce a concatenated analysis with robust branch supports. The incongruences observed may have arisen from past hybridization and introgression events, retention of ancestral polymorphism (Pessoa et al. 2022), technical issues as misidentification, contamination, horizontal gene transfer, and incomplete lineage sorting (Wendel and Doyle 1998; Davis and Xi 2015; Guo et al. 2018).
Notably, we confirmed the presence of a clade comprising solely North American endemic species (Clade 5; Fig. 2). These species belong to the second center of diversification within the genus and are predominantly found in the dry subtropical forests of the southern Mexican mountain ranges, known for their remarkable biodiversity and richness of endemic species (Marshall and Liebherr 2001; Morrone 2014; Montano-Arias et al. 2018). This clade had been previously identified by Barber et al. (2010), who differentiated it based on its larger flowers and seeds compared to taxa originating from South America. Our analyses align with these previous observations as this clade encompasses species with larger floral and leaf measurements. Additionally, this clade accounts for 59% of the identified polyploidy events within the genus.
Although autopolyploidy was also previously recorded to Cuphea in natural conditions (Gathman and Ray 1987), one of the main phenomena related to the origin of polyploid species is hybridization (allopolyploidy) (Soltis and Soltis 2009). Hybrids have been described for Cuphea since the only completed taxonomic monograph of the genus (Koehne 1903), furthermore significant efforts have been made to produce hybrids with economic potential (Gathman and Ray 1987). However, most of these hybrids are not fertile due to meiotic irregularities, nevertheless, the uncommon fertile hybrids display greater vigor and can have morphological differences (Ray et al. 1988). Gathman and Ray (1987) indicates that when hybridization happens between closely related species with genomes that are little differentiated, with close chromosome homology, diploidization occurs quite rapidly. The occurrence of fertile hybrids in Cuphea suggests a potential mechanism underlying the remarkable presence of polyploids within the genus (Robertson et al. 2010; Bertier et al. 2013; Alix et al. 2017; Morales-Briones et al. 2018).
Overall, we did not detect significant phylogenetic signal in the variation of chromosome numbers (Table 1). This is consistent with the reconstructed karyotypic evolution (Fig. 3), which indicates recent and rapid diversification of chromosome numbers in Cuphea, with most dysploidy and polyploidy events occurring near the terminal branches. Our results stand against the common pattern observed in angiosperms, where karyotypes distribution exhibit phylogenetic signal (Carta et al. 2018). But, it is important to highlight that clade 5 include most of the WDG events and perhaps new analysis restricted to this clade may provide opposite results. WGD and dysploidy phenomena are known to induce meiotic failures (Bomblies et al. 2015). Meiotic adaptations can lead to high karyotypic diversity among closely related species (Haiduk and Baker 1982) which may provide a non-clear phylogenetic organization, as evidenced in Cuphea.
Conversely, we observed significant phylogenetic signal in the morphological characteristics of the genus (Table 1), indicating that closely related species share similar sizes of vegetative and reproductive organs. This pattern is expected as morphological characters tends to correspond to the phylogeny (Fougère-Danezan et al. 2010). Correlations between WGD events and specific morphological characters have been well-documented in the literature (Balao et al. 2011; Beaulieu et al. 2008; Munzbergová 2009). Previous studies in the genus also suggested a relationship between floral tube size and increased chromosome number in certain sections, such as C. sect. Melvilla (Graham 1980). However, a comprehensive analysis integrating phylogenetic, cytogenetic, and statistical approaches is not available for this taxon.
Our sampling effectively represented the discrepancy in floral morphologies within the genus (vide Graham 1990, 1998, 2017; Cavalcanti and Graham 2008; Facco 2015), as well as the variation in chromosome numbers. Based on our data, we were able to confirm that polyploid Cuphea species possess longer floral tubes (Fig. 6). Furthermore, there is a positive correlation between chromosome numbers and floral tube lengths (Table 2), supporting the hypothesis suggested by Graham (1980). This correlation is consistent with observations made by Carta et al. (2018) for angiosperms in general, where smaller chromosome numbers are associated with smaller flowers.
The production of flowers with longer floral tubes may entail trade-offs, such as increased water and nutrient demands and higher carbon allocation (Teixido and Valladares 2014). Bigger flowers are more expensive because of the high demand of energy needed for their production, development, and to keep then physiologically active (Patiño and Grace 2002; Galen 1999). In Cuphea some species of longer flower tubes are known to be pollinated by hummingbirds (Melazzo and Oliveira 2012; Feinsinger 1978). Ornithophilous flowers are characterized by low sugar concentration and high nectar volume (Reed Haisworth and Wolf 1976). Nevertheless, polyploids are known to exhibit enhanced ecological tolerance and adaptive potential (Balao et al. 2011), often accompanied by apomictic reproductive mechanisms that support further population expansion and establishment. Additionally, the fixation of larger floral characteristics throughout evolutionary processes may be linked to the ability of these flowers to attract more pollinators, suggesting a potential positive selective pressure (Galen 1999). However, environmental conditions can neutralize selective pressure of pollinators (Caruso et al. 2019).
WGD phenomena can also contribute to variations in vegetative organs, with polyploid plants often exhibiting larger leaf sizes and higher stomatal densities (Otto and Whitton 2000; Beaulieu et al. 2008). It is important to note that multiple selective pressures can influence the development of these characters (Givnish 1987). However, our study revealed a clear positive correlation between chromosome numbers and leaf length (represented by minimal leaf length) in the genus, similar to the relationship observed with reproductive characters (Table 2). As individuals can have leaves of different sizes, minimum length recognizes better species with bigger leaves from the ones with smaller leaves. Furthermore, the absence of phylogenetic signal for karyotypes (Table 1) excludes the possibility of these correlations being explained by the retention of ancestral characters.
Considering that WGD phenomena serve as facilitators of speciation and evolution in plants (Alix et al. 2017), the polyploidy events in Cuphea may have enabled and driven the emergence and fixation of longer floral tubes and larger leaves as crucial mechanisms during the adaptive radiation of the genus, when expanding its distribution northward on the American continent, given that Cuphea has likely a South America origin (Inglis et al. 2023). Therefore, the colonization of new North American niches may have been facilitated by WGD events and subsequent chromosomal rearrangements.
Conclusions
We found evidence to support a positive correlation between chromosome numbers and overall size of selected morphological traits with polyploid species showing longer floral tubes. Our investigations also revealed a potential association between polyploidy and the adaptive radiation of Cuphea during its distribution expansion to the North America. These results reinforce the importance of whole-genome duplication events in producing trait diversity and, consequently, speciation. We propose that further studies be undertaken, focusing on this clade, to gain deeper insights into the role of polyploidy in Cuphea diversification. A comprehensive analysis, encompassing genome size data, physiological and anatomical information, and an expanded phylogenetic framework, holds promise for elucidating the mechanisms by which polyploidy may have facilitated the diversification of Cuphea.
References
Ackerly DD (2004) Adaptation, niche conservatism, and convergence: comparative studies of leaf evolution in the California chaparral. Amer Nat 163:654–671. https://doi.org/10.1086/383062
Alix K, Gérard PR, Schwarzacher T, Heslop-Harrison JS (2017) Polyploidy and interspecific hybridization: partners for adaptation, speciation and evolution in plants. Ann Bot (Oxford) 120:183–194. https://doi.org/10.1093/aob/mcx079
Bairiganjan GC, Patnaik SN (1989) Chromosomal evolution in Fabaceae. Cytologia 54:51–64. https://doi.org/10.1508/cytologia.54.51
Balao F, Herrera J, Talavera S (2011) Phenotypic consequences of polyploidy and genome size at the microevolutionary scale: a multivariate morphological approach. New Phytol 192:256–265. https://doi.org/10.1111/j.1469-8137.2011.03787.x
Baniaga AE, Marx HE, Arrigo N, Barker MS (2020) Polyploid plants have faster rates of multivariate niche differentiation than their diploid relatives. Ecol Lett 23:68–78. https://doi.org/10.1111/ele.13402
Barber JC, Ghebretinsae A, Graham SA (2010) An expanded phylogeny of Cuphea (Lythraceae) and a North American monophyly. Pl Syst Evol 289:35–44. https://doi.org/10.1007/s00606-010-0329-7
Beaulieu JM, Leitch IJ, Patel S, Pendharkar A, Knight CA (2008) Genome size is a strong predictor of cell size and stomatal density in angiosperms. New Phytol 179:975–986. https://doi.org/10.1111/j.1469-8137.2008.02528.x
Bellot S, Dias PMB, Affagard M, Aïnouche ML, Misset MT, Aïnouche A (2023) Molecular phylogenetics shed light on polyploid speciation in gorses (Ulex, Fabaceae: Genisteae) and on the origin of the invasive Ulex europaeus. Bot J Linn Soc 202:52–75. https://doi.org/10.1093/botlinnean/boac061
Bertier L, Leus L, D’hondt L, De Cock AWAM, Höfte M (2013) Host adaptation and speciation through hybridization and polyploidy in Phytophthora. PLoS ONE 8:e85385. https://doi.org/10.1371/journal.pone.0085385
Blanc G, Wolfe KH (2004) Functional divergence of duplicated genes formed by polyploidy during Arabidopsis evolution. Pl Cell 16:1679–1691. https://doi.org/10.1105/tpc.021410
Bomblies K, Higgins JD, Yant L (2015) Meiosis evolves: adaptation to external and internal environments. New Phytol 208:306–323. https://doi.org/10.1111/nph.13499
Bouckaert R, Vaughan TG, Barido-Sottani J, Duchêne S, Fourment M, Gavryushkina A, Heled J, Jones G, Kühnert D, De Maio N, Matschiner M, Mendes FK, Müller NF, Ogilvie HA, Du Plessis L, Popinga A, Rambaut A, Rasmussen D, Siveroni I, Suchard MA, Wu CH, Xie D, Zhang C, Stadler T, Drummond AJ (2019) BEAST 2.5: AN advanced software platform for Bayesian evolutionary analysis. PLOS Comput Biol 15:e1006650. https://doi.org/10.1371/journal.pcbi.1006650
Brauner L, Cavalcanti T (2018) A new species, a new synonym and lectotypification in Cuphea (Lythraceae) from Brazil. Phytotaxa 350:155. https://doi.org/10.11646/phytotaxa.350.2.4
Brauner LM (2018) O gênero Cuphea P. Browne na Chapada Diamantina. MSc Thesis, Universidade de Brasilia, Brazil
Braz GT, Van-Lume B, de Resende KFM, Cardoso FP, Oliveira L, De Andrade MJG, Souza G, Torres GA (2024) Cytomolecular trends in Chamaecrista Moench (Caesalpinioideae, Leguminosae) diversification. Genetica 152:51–61. https://doi.org/10.1007/s10709-024-00205-4
Carta A, Bedini G, Peruzzi L (2018) Unscrambling phylogenetic effects and ecological determinants of chromosome number in major angiosperm clades. Sci Rep 8:14258. https://doi.org/10.1038/s41598-018-32515-x
Carta A, Bedini G, Peruzzi L (2020) A deep dive into the ancestral chromosome number and genome size of flowering plants. New Phytol 228:1097–1106. https://doi.org/10.1111/nph.16668
Caruso CM, Eisen KE, Martin RA, Sletvold N (2019) A meta-analysis of the agents of selection on floral traits. Evolution 73:4–14. https://doi.org/10.1111/evo.13639
Chase MW, Christenhusz MJ, Fay MF, Byng JW, Judd WS, Soltis DE, Mabberley DJ, Sennikov AN, Soltis PS, Stevens PF, The Angiosperm Phylogeny Group (2016) An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG IV. Bot J Linn Soc 181:1–20. https://doi.org/10.1111/boj.12385
Cavalcanti TB, Graham SA (2008) New Species, varieties, and combinations in Cuphea (Lythraceae) from Brazil. Novon 18:298–314. https://doi.org/10.3417/2006163
Chansler MT, Ferguson CJ, Fehlberg SD, Prather LA (2016) The role of polyploidy in shaping morphological diversity in natural populations of Phlox amabilis. Amer J Bot 103:1546–1558. https://doi.org/10.3732/ajb.1600183
Cho M-S, Yang J, Mejías JA, Kim S-C (2022) Phylogenomic insight into dysploidy, speciation, and plastome evolution of a small Mediterranean genus Reichardia (Cichorieae; Asteraceae). Sci Rep 12:11030. https://doi.org/10.1038/s41598-022-15235-1
Cordeiro JMP, Chase MW, Hágsater E, Almeida EM, Costa L, Souza G, Nollet F, Felix LP (2022) Chromosome number, heterochromatin, and genome size support recent polyploid origin of the Epidendrum nocturnum group and reveal a new species (Laeliinae, Orchidaceae). Botany 100:409–421. https://doi.org/10.1139/cjb-2021-0113
Cummings MP, Handley SA, Myers DS, Reed DL, Rokas A, Winka K (2003) Comparing bootstrap and posterior probability values in the four-taxon case. Syst Biol 52:477–487. https://doi.org/10.1080/10635150390218213
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Meth 9:772–772. https://doi.org/10.1038/nmeth.2109
Davis CC, Xi Z (2015) Horizontal gene transfer in parasitic plants. Curr Opin Pl Biol 26:14–19. https://doi.org/10.1016/j.pbi.2015.05.008
De Storme N, Mason A (2014) Plant speciation through chromosome instability and ploidy change: cellular mechanisms, molecular factors and evolutionary relevance. Curr Pl Biol 1:10–33. https://doi.org/10.1016/j.cpb.2014.09.002
Dollon T, Hamel J-L (1967) Contribution a l’etude caryotaxonomique des Lythracees et des Punicacees. Bull Mus Hist Nat 39:793–818
Doyle JJ (2012) Polyploidy in legumes. In: Soltis PS, Soltis DE (eds) Polyploidy and genome evolution. Springer, Berlin, pp 147–180
Drummond AJ, Suchard MA, Xie D, Rambaut A (2012) Bayesian phylogenetics with BEAUti and the BEAST 1.7. Molec Biol Evol 29:1969–1973. https://doi.org/10.1093/molbev/mss075
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucl Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Erixon P, Svennblad B, Britton T, Oxelman B (2003) Reliability of Bayesian posterior probabilities and bootstrap frequencies in phylogenetics. Syst Biol 52:665–673. https://doi.org/10.1080/10635150390235485
Escudero M, Martín-Bravo S, Mayrose I, Fernández-Mazuecos M, Fiz-Palacios O, Hipp AL, Pimentel M, Jiménez-Mejías P, Valcárcel V, Vargas P, Luceño M (2014) Karyotypic changes through dysploidy persist longer over evolutionary time than polyploid changes. PLoS ONE 9:e85266. https://doi.org/10.1371/journal.pone.0085266
Facco MG (2015) Estudo taxonômico do gênero Cuphea P. Browne (Lythraceae) no Rio Grande do Sul. MSc Thesis, Universidade Federal do Rio Grande do Sul, Brazil
Facco MG, Cavalcanti TB (2022) Novelties in Cuphea (Lythraceae) from Serra do Espinhaço and Serra da Mantiqueira, and a key to the species from Minas Gerais state, Brazil. Phytotaxa 568:267–276. https://doi.org/10.11646/phytotaxa.568.3.3
Facco MG, Cavalcanti TB (2023) Taxonomic revision of Cuphea sect. Trispermum s.l. (Lythraceae). Phytotaxa 588:1–67. https://doi.org/10.11646/phytotaxa.588.1.1
Feinsinger P (1978) Ecological interactions between plants and hummingbirds in a successional tropical community. Ecol Monogr 48:269–287. https://doi.org/10.2307/2937231
Felsenstein J (1985) Phylogenies and the comparative method. Amer Nat 125:1–15. https://doi.org/10.1086/284325
Fougère-Danezan M, Herendeen PS, Maumont S, Bruneau A (2010) Morphological evolution in the variable resin-producing Detarieae (Fabaceae): do morphological characters retain a phylogenetic signal? Ann Bot (Oxford) 105:311–325. https://doi.org/10.1093/aob/mcp280
Galen C (1999) Why do flowers vary?: the functional ecology of variation in flower size and form within natural plant populations. Bioscience 49:631–640. https://doi.org/10.2307/1313439
Garbutt K, Bazzaz FA (1983) Leaf demography, flower production and biomass of diploid and tetraploid populations of Phlox Drummondii Hook. on a soil moisture gradient. New Phytol 93:129–141. https://doi.org/10.1111/j.1469-8137.1983.tb02698.x
Gathman AC, Ray DT (1987) Meiotic analysis of 14 Cuphea species and two interspecific hybrids. J Heredity 78:315–318. https://doi.org/10.1093/oxfordjournals.jhered.a110394
Glick L, Mayrose I (2014) ChromEvol: Assessing the pattern of chromosome number evolution and the inference of polyploidy along a phylogeny. Mol Biol Evol 31:1914–1922. https://doi.org/10.1093/molbev/msu122
Givnish TJ (1987) Comparative studies of leaf form: assessing the relative roles of selective pressures and phylogenetic constraints. New Phytol 106:131–160. https://doi.org/10.1111/j.1469-8137.1987.tb04687.x
Gloria TPR (2020) How can chromosome number variation indicate evolutionary relationships between the Caatinga, the Cerrado and the Atlantic forest? MSc Thesis, Universidade Estadual de Campinas, Brazil
Goldblatt P (1980) Polyploidy in angiosperms: monocotyledons. In: Lewis WH (ed) Polyploidy: biological relevance. Springer, Boston, pp 219–239
Goldblatt P (1981) Index to Plant chromosome numbers 1975–1978. Monogr Syst Bot Missouri Bot Gard 5:553
Graham SA (1968a) New species of Cuphea (Lythraceae) from Mexico. Brittonia 20:1–10. https://doi.org/10.2307/2805456
Graham SA (1968b) IOPB chromosome number reports XVI. Taxon 17:199–204
Graham SA (1980) Cuphea inflata (Lythraceae), a New species from Mexico. Syst Bot 5:322–325. https://doi.org/10.2307/2418376
Graham SA (1982) In: IOPB chromosome number reports LXXIV. Taxon 31:119
Graham SA (1987) Chromosome Number Reports 94. Taxon 36:282–283
Graham SA (1988) Revision of Cuphea section Heterodon (Lythraceae). Syst Bot Monogr 20:1–168. https://doi.org/10.2307/25027697
Graham SA (1989a) Revision of Cuphea sect. Leptocalyx (Lythraceae). Syst Bot 14:43–76. https://doi.org/10.2307/2419051
Graham SA (1989b) Chromosome numbers in Cuphea (Lythraceae): new counts and a summary. Amer J Bot 76:1530–1540. https://doi.org/10.1002/j.1537-2197.1989.tb15135.x
Graham SA (1990) New species of Cuphea section Melvilla (Lythraceae) and an annotated key to the section. Brittonia 42:12–32. https://doi.org/10.2307/2807021
Graham SA (1992) New chromosome counts in Lythraceae—systematic and evolutionary implications. Acta Bot Mex 17:45–51. https://doi.org/10.21829/abm17.1992.635
Graham SA (1998) Revision of Cuphea section Diploptychia (Lythraceae). Syst Bot Monogr 53:1–96. https://doi.org/10.2307/25096642
Graham SA (2017) A Revision of Cuphea section Brachyandra s. s. (Lythraceae). Syst Bot 42:859–919. https://doi.org/10.1600/036364417X696528
Graham SA (2019a) A revision of Cuphea section Melvilla (Lythraceae). Syst Bot Monogr 105:1–146
Graham SA (2019b) A Revision of Cuphea section Amazoniana s. s. (Lythraceae). Syst Bot 44:146–183. https://doi.org/10.1600/036364419X697994
Graham SA, Cavalcanti TB (2001) New chromosome counts in the Lythraceae and a review of chromosome numbers in the family. Syst Bot 26:445–458. https://doi.org/10.1043/0363-6445-26.3.445
Graham SA, Cavalcanti TB (2007) Lythraceae in IAPT/IOPB chromosome data 3. Taxon 56:209
Graham SA, Cavalcanti TB (2013) Taxonomic revision of Cuphea sect. Euandra subsect. Oidemation (Lythraceae). Phytotaxa 113:1–86. https://doi.org/10.11646/phytotaxa.113.1.1
Graham SA, Freudenstein JV, Luker M (2006) A phylogenetic study of Cuphea (Lythraceae) based on morphology and nuclear rDNA ITS sequences. Syst Bot 31:764–778. https://doi.org/10.1600/036364406779696004
Griesbach RJ (1985) Polypioidy in Phalaenopsis orchid improvement. J Heredity 76:74–75. https://doi.org/10.1093/oxfordjournals.jhered.a110029
Guha S (1972) Cytotaxonomical studies on the family Lythraceae. Proc Indian Sci Congr Assoc 59:344–345
Guo X, Thomas DC, Saunders RMK (2018) Gene tree discordance and coalescent methods support ancient intergeneric hybridisation between Dasymaschalon and Friesodielsia (Annonaceae). Molec Phylogen Evol 127:14–29. https://doi.org/10.1016/j.ympev.2018.04.009
Haiduk MW, Baker RJ (1982) Cladistical analysis of G-Banded chromosomes of nectar feeding bats (Glossophaginae: Phyllostomidae). Syst Biol 31:252–265. https://doi.org/10.1093/sysbio/31.3.252
Hedrén M, Lorenz R, Teppner H, Dolinar B, Giotta C, Griebl N, Hansson S, Heidtke U, Klein E, Perazza G, Ståhlberg D, Surina B (2018) Evolution and systematics of polyploid Nigritella (Orchidaceae). Nordic J Bot 36:njb-01539. https://doi.org/10.1111/njb.01539
Inglis PW, Cavalcanti TB, Facco MG, Bakker FT, Graham SA (2023) A comprehensive genus-level phylogeny and biogeographical history of the Lythraceae based on whole plastome sequences. Ann Bot (Oxford) 132:293–318. https://doi.org/10.1093/aob/mcad091
King M (1995) Species evolution: the role of chromosome change. Cambridge University Press, New York
Knight CA, Ackerly DD (2003) Evolution and plasticity of photosynthetic thermal tolerance, specific leaf area and leaf size: congeneric species from desert and coastal environments. New Phytol 160:337–347. https://doi.org/10.1046/j.1469-8137.2003.00880.x
Koehne E (1903) Lythraceae. In: Engler A (ed) Das Pflanzenreich, vol. 4. Wilhelm Engelmann, Leipzig, pp 1–326
Krahulcová A, Rotreklová O, Krahulec F, Rosenbaumová R, Plačková I (2009) Enriching ploidy level diversity: the role of apomictic and sexual biotypes of Hieracium subgen. Pilosella (Asteraceae) that coexist in polyploid populations. Folia Geobot 44:281–306. https://doi.org/10.1007/s12224-009-9041-1
Li D, Liu Y, Zhong C, Huang H (2010) Morphological and cytotype variation of wild kiwifruit (Actinidia chinensis complex) along an altitudinal and longitudinal gradient in central-west China. Bot J Linn Soc 164:72–83. https://doi.org/10.1111/j.1095-8339.2010.01073.x
Lysák MA, Schubert I (2013) Mechanisms of chromosome rearrangements. In: Greilhuber J, Dolezel J, Wendel JF (eds) Plant genome diversity, vol. 2: physical structure, behaviour and evolution of plant genomes. Springer, Vienna, pp 137–147
Majure LC, Judd WS, Soltis PS, Soltis DE (2012) Cytogeography of the Humifusa clade of Opuntia s.s. Mill. 1754 (Cactaceae, Opuntioideae, Opuntieae): correlations with pleistocene refugia and morphological traits in a polyploid complex. Comp Cytogenet 6:53–77. https://doi.org/10.3897/CompCytogen.v6i1.2523
Marshall CJ, Liebherr JK (2001) Cladistic biogeography of the Mexican transition zone. J Biogeogr 27:203–216. https://doi.org/10.1046/j.1365-2699.2000.00388.x
Mas de Xaxars G, Garnatje T, Pellicer J, Siljak-Yakovlev S, Vallès J, Garcia S (2016) Impact of dysploidy and polyploidy on the diversification of high mountain Artemisia (Asteraceae) and allies. Alpine Bot 126:35–48. https://doi.org/10.1007/s00035-015-0159-x
Mayrose I, Barker MS, Otto SP (2009) Probabilistic models of chromosome number evolution and the inference of polyploidy. Syst Biol 59:132–144. https://doi.org/10.1093/sysbio/syp083
Melazzo AFO, Oliveira PE (2012) Cuphea melvilla Lindlay (Lythraceae): uma espécie do Cerrado polinizada por beija-flores. Acta Bot Brasil 26:281–289. https://doi.org/10.1590/S0102-33062012000200005
Meyers LA, Levin DA (2006) On the abundance of polyploids in flowering plants. Evolution 60:1198–1206. https://doi.org/10.1111/j.0014-3820.2006.tb01198.x
Miller MA, Holder MT, Vos R, Midford PE, Liebowitz T, Chan L, Hoover P, Warnow T (2009) The CIPRES Portals. CIPRES. Available at: http://www.phylo.org/subsections/portal
Molero J, Rovira A, Simón J, Duré R, Franco D (2002) IOPB chromosome data 18. Newslett Int Organ Pl Biosyst 34:22–24
Montaño-Arias G, Luna-Vega I, Morrone JJ, Espinosa D (2018) Biogeographical identity of the Mesoamerican dominion with emphasis on seasonally dry tropical forests. Phytotaxa 376:277–290. https://doi.org/10.11646/phytotaxa.376.6.3
Moraes AP, Engel TBJ, Forni-Martins ER, Barros F, Felix LP, Cabral JS (2022) Are chromosome number and genome size associated with habit and environmental niche variables? Insights from the Neotropical orchids. Ann Bot (Oxford) 130:11–25. https://doi.org/10.1093/aob/mcac021
Morales-Briones DF, Liston A, Tank DC (2018) Phylogenomic analyses reveal a deep history of hybridization and polyploidy in the Neotropical genus Lachemilla (Rosaceae). New Phytol 218:1668–1684. https://doi.org/10.1111/nph.15099
Morrone JJ (2014) Cladistic biogeography of the Neotropical region: identifying the main events in the diversification of the terrestrial biota. Cladistics 30:202–214. https://doi.org/10.1111/cla.12039
Münkemüller T, Lavergne S, Bzeznik B, Dray S, Jombart T, Schiffers K, Thuiller W (2012) How to measure and test phylogenetic signal. Methods Ecol Evol 3:743–756. https://doi.org/10.1111/j.2041-210X.2012.00196.x
Münzbergová Z (2009) The effect of genome size on detailed species traits within closely related species of the same habitat. Bot J Linn Soc 160:290–298. https://doi.org/10.1111/j.1095-8339.2009.00973.x
Ornduff R (1967) Index to plant chromosome numbers for 1965. Regnum Veg 50:1–128.
Otto SP, Whitton J (2000) Polyploid incidence and evolution. Annual Rev Genet 34:401–437. https://doi.org/10.1146/annurev.genet.34.1.401
Pagel M (1999) Inferring the historical patterns of biological evolution. Nature 401:877–884. https://doi.org/10.1038/44766
Parisod C, Holderegger R, Brochmann C (2010) Evolutionary consequences of autopolyploidy. New Phytol 186:5–17. https://doi.org/10.1111/j.1469-8137.2009.03142.x
Patiño S, Grace J (2002) The cooling of convolvulaceous flowers in a tropical environment. Pl Cell Environm 25:41–51. https://doi.org/10.1046/j.0016-8025.2001.00801.x
Pessoa EM, Nollet F, Magalhães RF, Viruel J, Pinheiro F, Chase MW (2022) Nuclear–plastid discordance indicates past introgression in Epidendrum species (Laeliinae: Orchidaceae) with highly variable chromosome numbers. Bot J Linn Soc 199:357–371. https://doi.org/10.1093/botlinnean/boab080
Pozzobon MT, Facco MG, Cordeiro JMP, Santos AMS, Graham SA, Cavalcanti TB (2022) IAPT chromosome data 33/17, Lythraceae. In: Marhold, K., Kucera, J. (eds.) & al., IAPT chromosome data 33. Taxon 69:1394–1405
R Core Team (2023) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. Available at: http://www.R-project.org
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67:901–904. https://doi.org/10.1093/sysbio/syy032
Ramsey J, Ramsey TS (2014) Ecological studies of polyploidy in the 100 years following its discovery. Philos Trans Roy Soc B 369:20130352. https://doi.org/10.1098/rstb.2013.0352
Rasband WS (2012) ImageJ: image processing and analysis in Java. Astrophysics Source Code Library ascl: 1206.013
Ray DT, Thompson AE, Gathman AC (1988) Interspecific hybridization in Cuphea. HortScience 23:751–753. https://doi.org/10.21273/HORTSCI.23.4.751
Reed Haisworth F, Wolf LL (1976) Nectar Characteristics and food selection by hummingbirds. Oecologia 25:101–113. https://doi.org/10.1007/BF00368847
Revell LJ (2012) phytools: an R package for phylogenetic comparative biology (and other things). Meth Ecol Evol 3:217–223. https://doi.org/10.1111/j.2041-210X.2011.00169.x
Robertson A, Rich TCG, Allen AM, Houston L, Roberts C, Bridle JR, Harris SA, Hiscock SJ (2010) Hybridization and polyploidy as drivers of continuing evolution and speciation in Sorbus. Molec Ecol 19:1675–1690. https://doi.org/10.1111/j.1365-294X.2010.04585.x
Ronquist F, Teslenko M, Van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Sader MA, Amorim BS, Costa L, Souza G, Pedrosa-Harand A (2019) The role of chromosome changes in the diversification of Passiflora L. (Passifloraceae). Syst Biodivers 17:7–21. https://doi.org/10.1080/14772000.2018.1546777
Schluter D, Price T, Mooers AØ, Ludwig D (1997) Likelihood of ancestor states in adaptive radiation. Evolution 51:1699–1711. https://doi.org/10.1111/j.1558-5646.1997.tb05095.x
Senderowicz M, Nowak T, Rojek-Jelonek M, Bisaga M, Papp L, Weiss-Schneeweiss H, Kolano B (2021) Descending dysploidy and bidirectional changes in genome size accompanied Crepis (Asteraceae) evolution. Genes 12:1436. https://doi.org/10.3390/genes12091436
Solís Neffa VG, Moreno EMS, Silva GC, Kovalsky IE, Via Do Pico GM, Almirón ENA, Luque JMR, Fernández SA, Paredes EN, Chalup LMI, Dobladez GAR, Seijo JG (2022) How important was polyploidy in the diversification of herbs in the Chaquean Domain? The case of the Turnera sidoides autopolyploid complex (Passifloraceae: Turneroideae). Bot J Linn Soc 199:286–311. https://doi.org/10.1093/botlinnean/boab085
Soltis DE, Albert VA, Leebens-Mack J, Bell CD, Paterson AH, Zheng C, Sankoff D, Pamphilis CW, Wall PK, Soltis PS (2009) Polyploidy and angiosperm diversification. Amer J Bot 96:336–348. https://doi.org/10.3732/ajb.0800079
Soltis DE, Visger CJ, Marchant DB, Soltis PS (2016) Polyploidy: pitfalls and paths to a paradigm. Amer J Bot 103:1146–1166. https://doi.org/10.3732/ajb.1500501
Soltis PS, Soltis DE (2009) The role of hybridization in plant speciation. Annual Rev Pl Biol 60:561–588. https://doi.org/10.1146/annurev.arplant.043008.092039
Stebbins GL (1971) Processes of organic evolution. Prentice-Hall, New Jersey
Teixido AL, Valladares F (2014) Disproportionate carbon and water maintenance costs of large corollas in hot Mediterranean ecosystems. Perspect Pl Ecol Evol Syst 16:83–92. https://doi.org/10.1016/j.ppees.2014.02.002
Van de Peer Y, Mizrachi E, Marchal K (2017) The evolutionary significance of polyploidy. Nat Rev Genet 18:411–424. https://doi.org/10.1038/nrg.2017.26
Vichiato MRM, Vichiato M, Pasqual M, Rodrigues FA, Melo De Castro D (2014) Morphological effects of induced polyploidy in Dendrobium nobile Lindl. (Orchidaceae). Crop Breed Appl Biotechnol 14:154–159. https://doi.org/10.1590/1984-70332014v14n3a23
Weiss-Schneeweiss H, Tremetsberger K, Schneeweiss GM, Parker JS, Stuessy TF (2008) Karyotype diversification and evolution in diploid and polyploid South American Hypochaeris (Asteraceae) inferred from rDNA localization and genetic fingerprint data. Ann Bot (Oxford) 101:909–918. https://doi.org/10.1093/aob/mcn023
Wendel JF, Doyle JJ (1998) Phylogenetic incongruence: window into genome history and molecular evolution. In: Soltis DE, Soltis OS, Doyle JJ (eds) Molecular systematics of plants II: DNA sequencing. Springer, Boston, pp 265–296
Winterfeld G, Ley A, Hoffmann MH, Paule J, Röser M (2020) Dysploidy and polyploidy trigger strong variation of chromosome numbers in the prayer-plant family (Marantaceae). Pl Syst Evol 306:36. https://doi.org/10.1007/s00606-020-01663-x
Wood TE, Takebayashi N, Barker MS, Mayrose I, Greenspoon PB, Rieseberg LH (2009) The frequency of polyploid speciation in vascular plants. Proc Natl Acad Sci USA 106:13875–13879. https://doi.org/10.1073/pnas.0811575106
Yu Y, Harris AJ, Blair C, He X (2015) RASP (Reconstruct Ancestral State in Phylogenies): a tool for historical biogeography. Molec Phylogen Evol 87:46–49. https://doi.org/10.1016/j.ympev.2015.03.008
Acknowledgements
The authors acknowledge the Federal University of Mato Grosso, and those who founded our research, including Conselho Nacional de Desenvolvimento Cientifico e Tecnológico-CNPq and Cordenação de Aperfeçoamento Pessoal de Nivel Superior-CAPES. EMP also acknowledges the productivity grant from CNPq (303556/2022-6).
Author information
Authors and Affiliations
Contributions
LHJF: Conceptualization (equal); Data curation (major); Investigation (major); Methodology (equal); Writing—original draft (major); Writing—review & editing (equal). EMP: Conceptualization (major); Data curation (equal); Investigation (equal); Methodology (equal); Writing—original draft (equal); Writing—review & editing (major); Validation (equal). TBZ: Conceptualization (equal); Data curation (equal); Investigation (equal); Methodology (equal); Writing—original draft (equal); Writing—review & editing (equal); Validation (equal). TC: Conceptualization (equal); Data curation (equal); Investigation (equal); Methodology (equal); Writing—original draft (equal); Writing—review & editing (equal); Validation (equal). SG: Conceptualization (equal); Data curation (equal); Investigation (equal); Methodology (equal); Writing—original draft (equal); Writing—review & editing (equal); Validation (equal).
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Handling Editor: Julius Jeiter.
Supplementary Information
Below is the link to the electronic supplementary material.
Information on Electronic Supplementary Material
Information on Electronic Supplementary Material
Online Resource 1. List of species cataloged by the study and their respective data.
Online Resource 2. GenBank accession numbers for the sequences used in the phylogenetic analysis.
Online Resource 3. Reconstruction of ancestral states regarding chromosomal numbers in Cuphea with the reproduction of probabilities in ChromEvol 2.0.
Online Resource 4. Table indicating the distribution of ancestral chromosomal number probabilities for each node in the reconstruction produced by CromEvol 2.0.
Online Appendix 1. Alignment used to produce the phylogeny of nuclear + plastid dataset.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Feitoza, L.H.J., Zanata, T.B., Cavalcanti, T. et al. Evolution of chromosome numbers in Cuphea (Lythraceae) reveals associations between morphological variation and polyploidy. Plant Syst Evol 310, 19 (2024). https://doi.org/10.1007/s00606-024-01900-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00606-024-01900-7