Abstract
Glucose isomerase (GIase) catalyzes the isomerization of d-glucose to d-fructose. The GIase from Thermobifida fusca WSH03-11 was expressed in Escherichia coli BL21(DE3), and the purified enzyme took the form of a tetramer in solution and displayed a pI value of 5.05. The temperature optimum of GIase was 80 °C and its half life was about 2 h at 80 °C or 15 h at 70 °C. The pH optimum of GIase was 10 and the enzyme retained 95 % activity over the pH range of 5–10 after incubating at 4 °C for 24 h. Kinetic studies showed that the K m and K cat values of the enzyme are 197 mM and 1,688 min−1, respectively. The maximum conversion yield of glucose (45 %, w/v) to fructose of the enzyme was 53 % at pH 7.5 and 70 °C. The present study provides the basis for the industrial application of recombinant T. fusca GIase in the production of high fructose syrup.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
d-Xylose isomerase (EC 5.3.1.5) is an intracellular enzyme whose physiological function is to isomerize d-xylose to d-xylulose. In addition, this enzyme also catalyzes the isomerization of d-glucose to d-fructose. Hence, it is often referred to as glucose isomerase (GIase). GIase is one of the key enzymes in the industrial production of high fructose syrup (HFS), which is a mixture of glucose and fructose. As it is very sweet and has a low caloric content, HFS is widely used in soft drinks and other food products as a sucrose succedaneum [1]. Usually, HFS is produced from corn starch by three consecutive enzymatic steps: liquefaction, saccharification and isomerization [2]. In the first two steps, corn starch is converted into dextrin and glucose. In the last step, glucose is converted into fructose by GIase [3, 4].
GIases have been found to exist in both prokaryotes and eukaryotes [5]. Based on primary amino acid sequence homology, GIases are classified as classes I or II. The class II enzymes contain an additional insert of about 50 amino acid residues at N-terminus, which is not present in the class I enzymes [6]. Although the homology of the primary sequences of the two classes is low (25–30 %), their three-dimensional structures are similar [7]. Most forms of GIase are homo-tetramers composed of two tightly bound dimers, which are associated with non-covalent bonds [5]. Each subunit contains two domains: a N-terminal major domain folded as an eight-stranded α/β catalytic pocket and a C-terminal minor domain folded as a large loop that embraces the adjacent subunit [8].
Currently, most GIases used in HFS production are from microorganisms, such as Streptomyces sp. [9, 10], Streptomyces olivaceoviridis [11], Streptomyces olivochromogenes [12–14], Arthrobacter sp. [15] and Actinoplanes missouriensis [16–18]. All of these strains belong to Actinomycetales and the GIases produced by them belong to class I. These GIases perform well under the production conditions of the HFS industry, in which the temperature is usually 55–65 °C and the pH is 7.0–8.5.
In the present study, a GIase from a thermophilic actinomycete, Thermobifida fusca, was cloned and expressed in Escherichia coli. The recombinant protein was purified and its biochemical properties were investigated in detail.
Materials and methods
Strains and reagents
The T. fusca WSH03-11 strain [19] was isolated from soil samples and stored in our laboratory. E. coli BL21(DE3) and the plasmid pET24a (+) were purchased from Novagen (Madison, WI, USA). The EZ-10 Spin Column Plasmid Mini-Preps kit and agarose gel DNA purification kit were purchased from Tiangen Biotech Co. Ltd. (Beijing, China). Prime STAR®HS DNA polymerase, restriction enzymes, CIAP, and T4 DNA ligase were obtained from TakaRa (Dalian, China). DNA primer syntheses and DNA sequencing were performed by Shanghai Sangon Biological Engineering Technology & Services Co. Ltd. (Shanghai, China). Tryptone and yeast extract were purchased from Oxoid (Hampshire, UK). Isopropyl β-D-1-thiogalactopyranoside (IPTG) was purchased from Sigma-Aldrich (Milwaukee, WI, USA). Middle molecular weight markers for SDS-PAGE were purchased from Generay (Shanghai, China). Molecular weight markers for gel filtration were purchased from Sigma. pI markers and broad pI kit 3–10 were purchased from GE Healthcare. Other chemicals were obtained from Sinopharm Chemical Reagent Co. Ltd. (Shanghai, China).
Gene cloning
The gene encoding GIase was amplified by standard polymerase chain reaction methods using the T. fusca WSH03-11 genomic DNA as template. The cloning primer sequences were designed according to gene Tfu-1603 (GeneID: CP000088.1) as follows: CATATGAGCAACTACCAGCCCACACCCGAG (forward primer) and AAGCTTTTAGCGCACGCCCAGGAGGTAGT (reverse primer). The NdeI and HindIII restriction sites (underlined) were designed into the primers. The amplification was carried out under the following conditions: the first step was at 95 °C for 4 min; followed by 30 cycles of 98 °C for 10 s, 55 °C for 5 s, and 72 °C for 2 min; and the final extension was carried out at 72 °C for 8 min. The PCR product was gel purified and then ligated into pMD18-T. The recombinant plasmid pMD18-T/xylA was transformed into E. coli JM109 and was identified by restriction analysis and sequencing. pMD18-T/xylA was digested with NdeI and HindIII. The target gene was gel purified and then ligated into plasmid pET24a (+), which was subjected to a similar treatment. The recombinant plasmid pET24a (+)/xylA was transformed into E. coli JM109 and was identified by restriction analysis. The resulting expression vector was transformed into E. coli BL21(DE3) for expression.
Expression of recombinant GIase
To express the recombinant enzymes, recombinant E. coli BL21(DE3) was inoculated into 100 mL of Luria–Bertani (LB) medium containing 30 μg/mL kanamycin (Kana) and grown overnight at 37 °C in an orbital shaker at 200 rpm. The overnight culture was inoculated into fresh TB medium and shaken (200 rpm) at 37 °C until the OD600 reached 1.5. The recombinant enzyme was induced by the addition of IPTG to a final concentration of 0.05 mM. Incubation was continued for another 25 h at 25 °C, and the cells were harvested by centrifugation at 8,000 rpm for 10 min at 4 °C.
Purification of recombinant GIase
Cells were separated from the fermentation medium by centrifugation at 10,000 rpm for 5 min and suspended in the same volume of 50 mM Na2HPO4–KH2PO4 buffer (pH 7.5). The suspension was sonicated on ice for 10 min at 38 % output power and centrifuged (13,000 rpm, 30 min, 4 °C) to remove insoluble denatured proteins. The supernatant was used as the crude enzyme preparation. The activity of the crude enzyme preparation was determined as the yield of GIase in the fermentation broth.
The crude enzyme preparation was heated for 10 min at 70 °C, and the soluble fractions were recovered after centrifugation at 13,000 rpm at 4 °C for 30 min. The enzyme preparation was then filtered (0.22 μm membrane) and loaded on to a DEAE-Sepharose Fast Flow column that had been pre-equilibrated with 30 mM Na2HPO4–KH2PO4 buffer (pH 7.5). The column was eluted using a linear gradient of 0–1 M NaCl in the same buffer using an ÄKTA™ protein purification system. The active-peak fractions were pooled and loaded onto a Superose G-75 gel filtration column and eluted with Na2HPO4–KH2PO4 buffer, and the active-peak fractions were pooled, concentrated, and stored at −80 °C.
Enzyme activity assay
The activity of GIase was measured according to the amount of d-fructose isomerized from d-glucose. The reaction mixture (1 mL) contained 50 mM Na2HPO4–KH2PO4 buffer (pH 7.5), 800 mM d-glucose, 5 mM MgSO4, and 0.1 mL of enzyme solution diluted appropriately. It was then incubated at 70 °C for 10 min and terminated by adding 1 mL of 0.5 M HClO4. The mixture was further diluted with double-distilled water. d-fructose formed in the reaction mixture was measured by the cysteine–carbazole–sulfuric acid method using a spectrophotometer [20, 21]. One unit (U) of enzyme activity was defined as the amount that produced 1 μM d-fructose per min under the above conditions.
Biomass determination
After incubation, the cell mass was obtained by centrifuging the culture broth at 12,000 rpm for 20 min and washing twice with distilled water in centrifuge tube of predetermined weights. The tube was dried in a hot air oven at 100 °C to a constant weight. The dry cell weight (DCW) of culture broth was then calculated.
Molecular weight determination
The molecular weight (M r) of the subunit of GIase was determined by SDS-PAGE. The M r of GIase in its native state was determined by gel filtration utilizing a Superdex 200 10/300GL gel filtration column, and β-amylase (200,000), alcohol dehydrogenase (M r 150,000), albumin bovine serum (M r 66,000), carbonic anhydrase (M r 29,000), and cytochrome C (M r 12,400) were used as M r standards. The elution volume was determined in triplicate for all samples and standards.
Isoelectric point determination
The isoelectric point of GIase was determined by 7.5 % polyacrylamide gel isoelectric focusing. Standard proteins and samples were mixed with gel, cathode solution (1 M NaOH), and anode solution (1 M H3PO4). Standard proteins were amyloglucosidase (pI 3.50), trypsin inhibitor (pI 4.55), b-lactoglobulin A (pI 5.20), carbonic anhydrase B (bovine) (pH 5.85), myoglobin acidic band (pI 6.85). The relative distance was determined in triplicate for all samples and standards.
Temperature optimum and thermostability
The optimal temperature of the GIase was measured in the temperatures range of 40–95 °C. The thermostability was determined by incubating the enzyme in 30 mM Na2HPO4–KH2PO4 buffer (pH 7.5) containing 5 mM MgSO4 at 70 and 80 °C. At different intervals, samples were taken and assayed at 70 °C for residual isomerizing activity. The measurements were carried out in three independent experiments.
pH optimum and stability
The pH optimum of the GIase was measured over a pH range of 4.0–11.0 using 30 mM NaAc–HAc buffer (pH 4.0–5.0), 30 mM Na2HPO4–KH2PO4 buffer (pH 5.0–9.0), and 30 mM Gly–NaOH buffer (pH 9.0–11.0). To determine the pH stability, the enzyme was preincubated in the various buffers described above at 4 °C for 48 h, and then assayed for residual isomerizing activity at pH 7.5. The measurements were carried out in three independent experiments.
Determination of kinetic parameters
The catalytic properties of the enzyme were determined in Na2HPO4–KH2PO4 buffer (pH 7.5) at 70 °C using glucose as the substrate. Substrate concentrations were in the range of 30–900 mM. The Michaelis–Menten parameters, V max, K m, and K cat, were calculated from double reciprocal plots of the reaction curve [22]. The measurements were carried out in three independent experiments.
Isomerization of glucose to fructose
Fructose was produced by incubation of 45 % (w/v) glucose in 30 mM Na2HPO4–KH2PO4 buffer (pH 7.5) containing 5 mM MgSO4 and GIase (at a final activity of 20 U/g glucose) at 70 °C. Samples were withdrawn and analyzed by HPLC.
HPLC analysis
At different intervals, 100 μL of reaction mixture was taken and incubated at 100 °C for 10 min to inactivate the enzyme. The mixture was diluted to 1/100, centrifuged at 12,000 rpm for 10 min, and filtered (0.22 μm). Treated samples of 5 μL were loaded on to an Aminex HPX-87H ion Exclusion Column (7.8 mm × 300 mm) pre-equilibrated with the buffer (0.001 M H2SO4 solution). The column was eluted with the same buffer using an Agilent separation module (model 1200), the temperature of the column was set at 50 °C, and a refractive index detector (model 1200) was used. The measurements were carried out in three independent experiments.
Fed-batch fermentation
The recombinant E. coli BL21(DE3) was cultured in LB medium (containing 30 μg/mL Kana). A 10 % (v/v) inoculum concentration was inoculated into the fermentation medium for fed-batch cultivation in a 3-L fermentor (BioFlo 110, New Brunswick Scientific Co, Edison, NJ, USA). Fed-batch cultivation consisted of three phases: the first phase was batch cultivation with an initial glycerol concentration of 8 g/L at 37 °C. After inoculation, the pH and glycerol content decreased gradually. The end of glycerol consumption was detected by a sudden increase in both dissolved oxygen (DO) and pH value, and then the second phase (the pre-induction phase of fed cultivation) started. When a DCW of 25 g/L was reached, the inducer was fed at 0.2 g/L/h and the temperature was lowered to 30 °C for GIase production, and then the third phase (the induction phase of fed cultivation) began. In the present study, a two-stage feeding strategy was applied. During the pre-induction phase, the glycerol feeding rate was increased exponentially according to the exponential feeding method [23], and cell growth was controlled at a specific growth rate of 0.22. When the induction phase began, the feeding rate was shifted to a gradient-decreasing method. During the entire process, the pH was maintained at 7.2 by the addition of 100 % (v/v) ammonia solution. The DO level was kept at 30 % of air saturation by adjusting the cascading impeller speed and supplementing with oxygen. The DO concentration, pH, temperature, and impeller speed were recorded using advanced fermentation software (AFS) from New Brunswick Scientific Co. Inc.
Results and discussion
Selection of the encoding gene of GIase
T. fusca is a moderately thermophilic soil actinomycete that grows in an environment with a temperature range of 40–70 °C and pH range of 4–10 [24]. A search for the T. fusca YX genome in the NCBI database yielded two possible GIases: one is Tfu_1603, which is 385 amino acids long; the other is Tfu_2709, which is 305 amino acids long. The structure modeling showed that Tfu_1603 exhibits a typical GIase structure with an N-terminal major domain and a C-terminal minor domain (Fig. 1a), whereas Tfu_2709 only contains the N-terminal domain of a GIase and lacks the C-terminal domain (Fig. 1b).
Cloning, expression, and purification of GIase
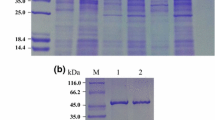
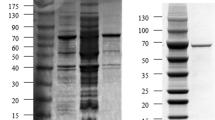
The GIase gene was amplified from the genomic DNA of T. fusca WSH03-11 using primers designed according to the Tfu_1603 gene. For expression, the gene was ligated into the expression vector of pET24a(+), and the resulting plasmid was transformed into E. coli BL21(DE3). After 25 h of induction, the activity of GIase reached 21 U/mL. The recombinant GIase was isolated and purified from the cultured cells by sonication, thermo-denaturation, ion exchange chromatography, and gel filtration. The final specific activity of the purified GIase was 33.8 U/mg.
Physical properties of the recombinant GIase
The M r of GIase was found to be 165 kDa, as determined by Superdex 200 gel filtration chromatography, whereas the M r of the subunit of GIase, as determined by SDS-PAGE, was found to be 42 kDa. Thus, it seemed that GIase was composed of homo-tetrameric subunits, which is consistent with a previous study that reported GIase is usually a tetramer or dimer [5]. The pI value of T. fusca GIase is 5.05, as determined by polyacrylamide gel isoelectric focusing.
Effect of metal ions on recombinant GIase
It was reported that GIase is usually activated by Co2+, Mg2+, or Mn2+ ions and inhibited by Ca2+ ions [5]. To determine the effect of various metal ions on recombinant T. fusca GIase, the purified GIase was incubated with 1 mM EDTA at 4 °C for 30 min and the remaining activity was measured. The results showed that the residual enzyme activity was 6 %. When various metal ions (2 mM) were added to the treated sample, different activation effects on the GIase were found, in which Co2+ > Mg2+ > Mn2+ (Table 1). The role of these metal ions is to form a bridge between the enzyme and the sugar in the enzyme-substrate complex. From the various 3D structures of the enzyme that were determined in the presence of metal ions and substrate, product, or their analogs, it has been concluded that there are two metal-ion-binding sites. Metal site 1 is four-coordinated and tetrahedral in the absence of substrate and is six-coordinated and octahedral in its presence. Metal site 2 is octahedral in all cases and is involved in the binding of oxygen atoms O1 and O2 of the substrate [8]. It has been reported that metal cations that have ionic radii ≤0.8 Å, such as Mg2+, Mn2+, Co2+, and Fe2+, can activate GIase, while larger alkali earth metal cations Ca2+, Ba2+, Sr2+, and transition metal cations Hg2+, Pb2+, Ni2+, Cu2+, Cd2+ inhibit GIase [25, 26]. The current experimental results verified these theories, the Mg2+ or Co2+ enhance the catalytic activity of GIases, while Ca2+ inhibits the activity of GIases.
Optimal temperature and thermostability of recombinant GIase
The influence of temperature on the enzyme activity was investigated in the range 40–95 °C in 30 mM Na2HPO4–KH2PO4 buffer (pH 7.5) containing 5 mM MgSO4. The results showed that the optimal temperature of the purified GIase is 80 °C, and the activity decreased rapidly with further increases in temperature, at 70 °C, was found to be about 75 % of its maximal activity (Fig. 2a). The thermostability experiments showed that the enzyme retains 50 % of its activity after 2.0 h at 80 °C or 15 h at 70 °C (Fig. 2b).
Optimal pH and stability of recombinant GIase
The pH optimum and stability of GIase were measured over a pH range of 4.0–11.0. The optimum pH of the GIase was 10.0, and the activity of the enzyme at pH 7.5 was 78 % of its maximal activity (Fig. 3a). The GIase retained more than 95 % of its initial activity after incubation in the pH range 5.0–10.0 at 4 °C for 24 h, but the residual activity decreased rapidly at pH values lower than 5 or higher than 10 (Fig. 3b).
Effects of pH on the activity and stability of GIase. a pH optimum. The activity of the GIase at pH 10.0 was defined as 100 %. b pH stability. The activity of the GIase without treatment was defined as 100 %. NaAc–HAc buffer (unfilled square), Na2HPO4–KH2PO4 buffer (unfilled triangle) and Gly–NaOH buffer (unfilled circle)
Kinetic analysis of recombinant GIase
The kinetic properties of the purified GIase were analyzed using glucose as the substrate through the double reciprocal curve of the initial conversion rate and substrate concentration. At pH 7.5 and 70 °C, the K m, K cat, and K cat /K m values of the GIase were 197 mM, 1,680 min−1, and 8.56 min−1 mM−1, respectively (Table 2). To our knowledge, this is the highest catalytic efficiency reported for class I GIases.
To explain why T. fusca GIase is more stable and have a high catalytic efficiency, the crystal simulation structure models of T. fusca GIase and other GIases were studied. These GIases come from the strains: Streptomyces olivaceoviridis [10], Streptomyces olivochromogenes [9, 12–14, 27], Streptomyces rubiginosus [28–30], Arthrobacter sp. [15], and Actinoplanes missouriensis [12, 15, 16, 31–36], which all have good performance under application conditions of HFCS industrial production. T. fusca GIase structures model (Fig. 4) was simulated by Swiss Modle according to the known GIs crystal structures. It was found that nine amino acid residues (88P, 89M, 134T, 180A, 214G, 243F, 244H, 284P, 286H) intensively located at or near to the bottom of “active center pocket” in T. fusca were conservative in corresponding sites of above GIases, but the nine conserved amino acids were not all exist in the GIases with poor performance under application conditions. It was interesting that some amino acid residues containing annular structure were introduced to T. fusca GIase sites 88P, 243F, 244H, 284P, 286H, which might increases steric constraints for substrate glucose. However, these amino acid might used their ring structures to make glucose molecule form a more favorable posture while enter active center, hereby to speed up catalytic action. In addition, these rings may increase the stability of substrate domain. What the other structural features of T. fusca GIase make it more stable and more high catalytic efficient, here need more experiments to test.
Isomerization of glucose to fructose by recombinant GIase
The proportion of d-glucose isomerized to d-fructose by GIase is critical to its industrial application to HFS production. To investigate the conversion ability of the T. fusca GIase, the recombinant GIase was utilized to isomerize glucose to fructose at pH 7.5 and 70 °C, with 45 % (w/v) glucose as substrate. The results showed that the concentration of fructose in the reaction mixture increased rapidly during the initial reaction phase and the isomerization reached equilibrium within 4 h. The yield of fructose reached 53 % (Fig. 5), which was comparable to the yields reported in the literature at similar reaction temperatures. Previously, although the yields of fructose using some thermostable GIases from hyperthermophiles were reported to reach 55 % [37–41], the reaction temperature used by these authors was not less than 90 °C. Such high temperatures would increase the by-products and energy consumption. Moreover, the activity of these thermostable GIases declined sharply as the reaction temperature increased.
High cell-density fermentation
An excessive production of enzyme via a high-density fermentation strategy is an effective way to reduce production costs. High-density fermentation of engineered E. coli was performed in a 3-L fermentor. The fed-batch cultivation consisted of three phases: a batch cultivation phase, a pre-induction phase of fed cultivation, and the induction phase of fed cultivation. The culture temperature during the batch phase was 37 °C. When the carbon source of the medium was consumed, the pre-induction phase of fed cultivation started, in which the glycerol feeding rate was increased exponentially. When a DCW of 25 g/L was reached, the induction phase of fed cultivation was initiated. The feeding rate was shifted to a gradient-decreasing method, and the temperature was decreased to 30 °C for GIase production. After cultivation for 33 h, the yield of GIase reached 60.6 U/mL (Fig. 6). To our knowledge, the production of GIase in the present study is the highest level.
Conclusions
In summary, the GIase of T. fusca was cloned, expressed, and its biochemical properties were investigated in detail. The GIase displayed a high catalytic rate (8.56 min−1 mM−1) and good thermal stability at pH 7.5 and 70 °C. The glucose conversion rate by the recombinant GIase reached 53 % (w/w) with 45 % glucose as substrate. In addition, the activity of the GIase reached 60.6 U/mL in a 3-L fermentor, which represents the highest yield reported so far. All of these properties of the T. fusa GIase indicate that it may be utilized in HFS production.
References
Seyhan Tükel S, Alagöz D (2008) Catalytic efficiency of immobilized glucose isomerase in isomerization of glucose to fructose. Food Chem 111:658–662
Tosa T, Shibatani T (1995) Industrial application of immobilized biocatalysts in Japan. Ann N Y Acad Sci 750:364–375
Chandrakant P, Bisaria VS (1998) Simultaneous bioconversion of cellulose and hemicellulose to ethanol. Crit Rev Biotechnol 18:295–331
Vieille C, Zeikus GJ (2001) Hyperthermophilic enzymes: sources, uses, and molecular mechanisms for thermostability. Microbiol Mol Biol Rev 65:1–43
Bhosale SH, Rao MB, Deshpande VV (1996) Molecular and industrial aspects of glucose isomerase. Microbiol Rev 60:280–300
Vangrysperre W, Van Damme J, Vandekerckhove J, De Bruyne CK, Cornelis R, Kersters-Hilderson H (1990) Localization of the essential histidine and carboxylate group in d-xylose isomerases. Biochem J 265:699–705
Hartley BS, Hanlon N, Jackson RJ, Rangarajan M (2000) Glucose isomerase: insights into protein engineering for increased thermostability. Biochimica et biophysica acta 1543:294–335
Jenkins J, Janin J, Rey F, Chiadmi M, van Tilbeurgh H, Lasters I, De Maeyer M, Van Belle D, Wodak SJ, Lauwereys M et al (1992) Protein engineering of xylose (glucose) isomerase from Actinoplanes missouriensis. 1. Crystallography and site-directed mutagenesis of metal binding sites. Biochemistry 31:5449–5458
Borgi MA, Srih-Belguith K, Ben Ali M, Mezghani M, Tranier S, Haser R, Bejar S (2004) Glucose isomerase of the Streptomyces sp. SK strain: purification, sequence analysis and implication of alanine 103 residue in the enzyme thermostability and acidotolerance. Biochimie 86:561–568
Kaneko T, Takahashi S, Saito K (2000) Characterization of acid-stable glucose isomerase from Streptomyces sp., and development of single-step processes for high-fructose corn sweetener (HFCS) production. Biosci Biotechnol Biochem 64:940–947
Kaneko T, Saito K, Kawamura Y, Takahashi S (2001) Molecular cloning of acid-stable glucose isomerase gene from Streptomyces olivaceoviridis E-86 by a simple two-step PCR method, and its expression in Escherichia coli. Biosci Biotechnol Biochem 65:1054–1062
Hashemiravan M, Barikani AS (2005) Biotechnological production of glucose isomerase enzyme with Streptomyces olivochromogenes for production of fructose syrup from hydrol. J Biotechnol 118:S144
Sicard PJ, Leleu JB, Duflot P, Drocourt D, Martin F, Tiraby G, Petsko G, Glasfeld A (1990) Site-directed mutagenesis applied to glucose isomerase from Streptomyces violaceusniger and Streptomyces olivochromogenes. Ann N Y Acad Sci 613:371–375
Simkhada JR, Lee HJ, Jang SY, Cho SS, Park EJ, Sohng JK, Yoo JC (2009) A novel alkalo- and thermostable phospholipase D from Streptomyces olivochromogenes. Biotechnol Lett 31:429–435
Sapunova LI, Lobanok AG, Parakhnia EV, Kazakevich IO (2003) Xylose(glucose) isomerase reactivity of immobilized Arthrobacter sp. Mikrobiologiia 72:395–399
van Bastelaere PB, Kersters-Hilderson HL, Lambeir AM (1995) Wild-type and mutant d-xylose isomerase from Actinoplanes missouriensis: metal-ion dissociation constants, kinetic parameters of deuterated and non-deuterated substrates and solvent-isotope effects. Biochem J 307(Pt 1):135–142
Mrabet NT (1992) One-step purification of Actinoplanes missouriensis d-xylose isomerase by high-performance immobilized copper-affinity chromatography: functional analysis of surface histidine residues by site-directed mutagenesis. Biochemistry 31:2690–2702
Lambeir AM, Lauwereys M, Stanssens P, Mrabet NT, Snauwaert J, van Tilbeurgh H, Matthyssens G, Lasters I, De Maeyer M, Wodak SJ et al (1992) Protein engineering of xylose (glucose) isomerase from Actinoplanes missouriensis. 2. Site-directed mutagenesis of the xylose binding site. Biochemistry 31:5459–5466
Chen S, Tong X, Woodard RW, Du G, Wu J, Chen J (2008) Identification and characterization of bacterial cutinase. J Biol Chem 283:25854–25862
Michal G, Moellering H, Gruber W (1968) Enzymatic determination of fructose-1-phosphate. Enzymologia biologica et clinica 9:154–159
Dische Z, Borenfreund E (1951) A new spectrophotometric method for the detection and determination of keto sugars and trioses. J Biol Chem 192:583–587
Tseng SJ, Hsu JP (1990) A comparison of the parameter estimating procedures for the Michaelis-Menten model. J Theor Biol 145:457–464
Ramalingam S, Gautam P, Mukherjee KJ, Jayaraman G (2007) Effects of post-induction feed strategies on secretory production of recombinant streptokinase in Escherichia coli. Biochem Eng J 33:34–41
Lykidis A, Mavromatis K, Ivanova N, Anderson I, Land M, DiBartolo G, Martinez M, Lapidus A, Lucas S, Copeland A, Richardson P, Wilson DB, Kyrpides N (2007) Genome sequence and analysis of the soil cellulolytic actinomycete Thermobifida fusca YX. J Bacteriol 189:2477–2486
Chen W-P (1980) Glucose isomerase (a review). Process Biochem 15:30–35
Danno GI (1970) Studies on d-Glucose-isomerizing Enzyme from Bacillus coagulans, strain HN-68. Agric Biol Chem 34:1795–1804
Joo GJ, Shin JH, Heo GY, Kim YM, Rhee IK (2005) Molecular cloning and expression of a thermostable xylose (glucose) isomerase gene, xylA, from Streptomyces chibaensis J-59. J Microbiol 43:34–37
Carrell HL, Rubin BH, Hurley TJ, Glusker JP (1984) X-ray crystal structure of d-xylose isomerase at 4-A resolution. J Biol Chem 259:3230–3236
Kaneko T, Saito K, Kawamura Y, Takahashi S (2001) Molecular cloning of acid-stable glucose isomerase gene from Streptomyces olivaceoviridis E-86 by a simple two-step PCR method, and its expression in Escherichia coli. Biosci Biotech Biochem 65:1054–1062
Karimaki J, Parkkinen T, Santa H, Pastinen O, Leisola M, Rouvinen J, Turunen O (2004) Engineering the substrate specificity of xylose isomerase. Protein Eng Des Sel 17:861–869
Amore R, Hollenberg CP (1989) Xylose isomerase from Actinoplanes missouriensis: primary structure of the gene and the protein. Nucleic Acids Res 17:7515
Smith CA, Rangarajan M, Hartley BS (1991) d-Xylose (d-glucose) isomerase from Arthrobacter strain N.R.R.L. B3728. Purification and properties. Biochem J 277(Pt 1):255–261
Kluskens LD, Zeilstra J, Geerling ACM, de Vos WM, van der Oost J (2010) Molecular characterization of the glucose isomerase from the thermophilic bacterium Fervidobacterium gondwanense. Environ Technol 31:1083–1090
Rudolph B, Hansen T, Schonheit P (2004) Glucose-6-phosphate isomerase from the hyperthermophilic archaeon Methanococcus jannaschii: characterization of the first archaeal member of the phosphoglucose isomerase superfamily. Arch Microbiol 181:82–87
Lama L, Nicolaus B, Calandrelli V, Romano I, Basile R, Gambacorta A (2001) Purification and characterization of thermostable xylose(glucose) isomerase from Bacillus thermoantarcticus. J Ind Microbiol Biot 27:234–240
Raykovska V, Dolashka-Angelova P, Paskaleva D, Stoeva S, Abashev J, Kirkov L, Voelter W (2001) Isolation and characterization of a xylose-glucose isomerase from a new strain Streptomyces thermovulgaris 127, var. 7-86. Biochem Cell Biol 79:195–205
Meng M, Lee C, Bagdasarian M, Zeikus JG (1991) Switching substrate preference of thermophilic xylose isomerase from d-xylose to d-glucose by redesigning the substrate binding pocket. Proc Natl Acad Sci USA 88:4015–4019
Meng M, Bagdasarian M, Zeikus JG (1993) The role of active-site aromatic and polar residues in catalysis and substrate discrimination by xylose isomerase. Proc Natl Acad Sci USA 90:8459–8463
Park BC, Koh S, Chang C, Suh SW, Lee DS, Byun SM (1997) Cloning and expression of the gene for xylose isomerase from Thermus flavus AT62 in Escherichia coli. Appl Biochem Biotechnol 62:15–27
Dhungel B, Subedi M, Tiwari KB, Shrestha UT, Pokhrel S, Agrawal VP (2009) Thermostable glucose isomerase from psychrotolerant Streptomyces sp. Int J Life Sci 1:6–10
Sriprapundh D, Vieille C, Zeikus JG (2000) Molecular determinants of xylose isomerase thermal stability and activity: analysis of thermozymes by site-directed mutagenesis. Protein Eng 13:259–265
Lee CY, Zeikus JG (1991) Purification and characterization of thermostable glucose isomerase from Clostridium thermosulfurogenes and Thermoanaerobacter strain B6A. Biochem J 273(Pt 3):565–571
Brown SH, Sjoholm C, Kelly RM (1993) Purification and characterization of a highly thermostable glucose isomerase produced by the extremely thermophilic eubacterium, Thermotoga maritima. Biotechnol Bioeng 41(9):878–886
Vieille C, Hess JM, Kelly RM, Zeikus JG (1995) xylA cloning and sequencing and biochemical characterization of xylose isomerase from Thermotoga neapolitana. Appl Environ Microbiol 61(5):1867–1875
Acknowledgments
This work was supported financially by the National High-tech Research and Development Program of China (863 program No. 2012AA021500).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Deng, H., Chen, S., Wu, D. et al. Heterologous expression and biochemical characterization of glucose isomerase from Thermobifida fusca . Bioprocess Biosyst Eng 37, 1211–1219 (2014). https://doi.org/10.1007/s00449-013-1093-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-013-1093-1