Abstract
Changes in the nuclear positioning of specific genes, depending on their expression status, have been observed in a large diversity of physiological processes. However, gene position is poorly documented for immune cells which have been subjected to activation following bacterial infection. Using a pig model, we focused our study on monocyte-derived macrophages and neutrophils, as they are the first lines of defence against pathogens. We examined whether changes in gene expression due to LPS activation imply that genes have repositioned in the nuclear space. We first performed a transcriptomic analysis to identify the differentially expressed genes and then analysed the networks involved during lypopolysaccharide/interferon gamma activation in monocyte-derived macrophages. This allowed us to select four up-regulated (IL1β, IL8, CXCL10 and TNFα) and four down-regulated (VIM, LGALS3, TUBA3 and IGF2) genes. Their expression statuses were verified by quantitative real-time RT-PCR before studying their behaviour in the nuclear space during macrophage activation by means of 3D fluorescence in situ hybridization. No global correlation was found between gene activity and radial positioning. Only TNFα belonging to the highly transcribed MHC region on chromosome 7 became more peripherally localized in relation to the less decondensed state of its chromosome territory (CT) in activated macrophages. The analysis of gene positioning towards their CT revealed that IL8 increases significantly its tendency to be outside its CT during the activation process. In addition, the gene to CT edge distances increase for the three up-regulated genes (IL8, CXCL10 and TNFα) among the four analysed. Contrarily, the four down-regulated genes did not change their position. The analysis of gene behaviour towards their CT was extended to include neutrophils for three (TNFα, IL8 and IL1β) up- and two (IGF2 and TUBA3) down-regulated genes, and similar results were obtained. The analysis was completed by studying the four up-regulated genes in fibroblasts, not involved in immune response. Our data suggest that relocation in the nuclear space of genes that are differentially expressed in activated immune cells is gene and cell type specific but also closely linked to the entire up-regulation status of their chromosomal regions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The development of an immune response is a complex mechanism involving an extensive regulation of cells that are central to the inflammatory process. The first actors of inflammation are monocyte-derived macrophages (reviewed in Geissmann et al. 2010) and neutrophils (Nathan 2006) which are essential for building and modulating the innate response. These phagocytic cells are able to destroy infectious agents and their properties are coupled with remarkably diverse patterns of gene expression. Neutrophils are the first cells to intervene at infection sites but they have a short lifespan, and after phagocytosis they undergo apoptosis and must be eliminated from the inflammatory sites to avoid tissue damage. The second actors to arrive at infection sites are monocytes. These differentiate into macrophages and phagocyte both pathogens and apoptotic neutrophils. To do this, they develop their cytoplasm to make pseudopodia which enables them to internalize foreign particles. This highly dynamic aspect concerns the cytoplasm but the nucleus also needs a certain adaptive plasticity for movement. So in response to a pathogen aggression, these immune cells undergo a series of distinct functional changes in state, resulting in cytoskeletal rearrangements, production of reactive oxygen and release of toxic peptides, all processes which imply variations in gene expression. These variations can be estimated by transcriptomic analyses using microarrays that allow the screening of a large number of changes in mRNA expression. Microarrays have been widely used in immune cells to study the host response at the gene transcription level in several situations such as bacterial infections (Malcolm et al. 2003; Ge et al. 2010), stimulation by various components (Kawano et al. 2010), atherosclerosis (Smith et al. 2006) or ischemic stroke (Tang et al. 2006).
Numerous recent studies have supported the concept of 3D remodelling of chromatin as a determinant for genomic regulation, pinpointing the major role of 3D nuclear organization in gene regulation (for a review, see Cremer et al. 2006; Lanctôt et al. 2007; Misteli 2007; Pai and Engelke 2010). In the higher eukaryotes studied so far, chromosomes have been shown to be located in specific regions in the interphase nucleus called chromosome territories (CTs) and more-detailed mapping studies have confirmed that genome regions (early- or late-replicating DNA) are also non-randomly arranged within the nucleus (reviewed in Cremer and Cremer 2001; Parada and Misteli 2002; Cremer et al. 2006; Misteli 2007). The spatial configuration of chromosomes results in interchromosomal and intrachromosomal interactions that bring distant genomic elements in close proximity. As with chromosomes, the position of single genes is non-random and has been linked to their functional status. Originally supported by the fact that the nuclear envelope is a silent compartment (Kosak et al. 2002), further studies have shown that the repositioning of loci from nuclear periphery to nuclear interior is correlated with gene activation (Zink et al. 2004; Ragoczy et al. 2006; Finlan et al. 2008). However, recent data on Saccharomyces cerevisae indicate that the nuclear periphery does not function exclusively as a repressive environment. In yeast, a large number of genes are repositioned to the periphery when they become activated, by interacting with nuclear-pore components (Casolari et al. 2004; Akhtar and Gasser 2007). Evidence for gene activation at the nuclear periphery has also been observed in mammalian cells (reviewed in Brown and Silver 2007; Lanctôt et al. 2007), and a study using a transgene tethered to the nuclear lamina showed that this transgene maintains transcriptional activity (Kumaran and Spector 2008). Two other studies have shown that attachment to the nuclear periphery does not generally preclude transcriptional activity, but the transcription of a fraction of genes near the tethered locus has shown reduced expression (Finlan et al. 2008; Reddy et al. 2008). Apart from these observations, loci organization within a CT has also been found to be an important factor in gene regulation. While some authors have predicted that active genes can be found outside chromosome territories (Dietzel et al. 1999), others have demonstrated that active or inactive genes can both be found at the periphery of chromosome territories (Kurz et al. 1996; Scheuermann et al. 2004). In addition, upon transcriptional activation, several cases have been documented of genes repositioning relative to their chromosome territories by means of chromatin loop formation (Volpi et al. 2000; Williams et al. 2002; Chambeyron and Bickmore 2004; Szczerbal et al. 2009). However, genes looping out from chromosome territories may not always be linked to up-regulation of gene expression as demonstrated by Morey and collaborators (2009) with their example of the ability of a transposed Hoxb1 to induce some looping out, in the absence of transcription.
Thus over the past 15 years, numerous studies have highlighted the major role of nuclear architecture in the regulation of cellular functions. Among several examples, this role has been studied during cellular differentiation (Bártová et al. 2002; Williams et al. 2002; Stadler et al. 2004; Ragoczy et al. 2006; Brown et al. 2008; Szczerbal et al. 2009), embryonic development (Chambeyron and Bickmore 2004; Chambeyron et al. 2005; Morey et al. 2009) and tumour progression (Meaburn and Misteli 2008). Immune response is another important biological process. However, data available on the spatial organization of the genome concern principally the lymphocytes while little information is available for phagocytes, which also play a key role in this process. Some studies have analysed the 3D nuclear organization of chromosome territories in murine macrophages (Mayer et al. 2005; Hepperger et al. 2008) and in human (Hepperger et al. 2009) and porcine (Yerle-Bouissou et al. 2009) neutrophils. The relation between nuclear position and gene activity status has been investigated in human neutrophils (Bártová et al. 2002), and chicken (Stadler et al. 2004) and murine (Hepperger et al. 2008) macrophages but mainly in the context of cell differentiation.
To address this question, we chose porcine monocyte-derived macrophages as our model and lypopolysaccharide (LPS)/interferon gamma (IFNγ) activation. Using a transcriptomic approach, we first determined which genes are differentially expressed when the macrophages are activated. We then selected four up- and five down-regulated genes and one non-expressed gene to investigate the relationship between nuclear positions and expression levels. The spatial arrangements of differentially expressed genes were then compared in control and activated macrophages using 3D fluorescence in situ hybridization (3D-FISH), confocal microscopy and 3D image analyses. In order to estimate whether the type of repositioning observed is gene specific and/or cell specific, similar analyses were first performed on neutrophils subjected to LPS stimulation as another type of immune cell and finally on fibroblasts not related to immune response.
Materials and methods
Cell isolation and stimulation
Monocytes/macrophages
Macrophages were isolated from Jugular venous blood of 3- to 5-month-old female pigs (synthetic line PIC410), collected aseptically (50 ml) in heparinised tubes. The blood was diluted in Hank’s solution (v/v). Peripheral blood mononuclear cells (PBMCs) were isolated by density gradient centrifugation (1.077 g/l, Lymphoprep, Eurobio, les Ulis, France) for 30 min at 800 g at room temperature. PBMCs were collected and submitted to a lysis solution (NH4Cl 0.15 M–KHCO3 0.01 M–EDTA Na2 1 μM) at 4°C for 5 min to eliminate red cells. The cells were washed twice with PBS and resuspended in RPMI-1640 media (Eurobio). Briefly, PBMCs were adjusted to a concentration of 108 cells/ml with RPMI-1640, and 2 ml of the cell suspension was distributed to each compartment of a 6-well polystyrene plate (CellBIND®, Corning, Bagneaux Sur Loing, France). Four hours later, three PBS washings were undertaken to remove non-adherent cells. For the generation of monocyte-derived macrophages, the cells were cultured in RPMI-1640 supplemented with 10% foetal bovine serum (Eurobio) completed with 1% l-Glutamin (Invitrogen, Cergy Pontoise, France), 1% non-essential amino acids (Sigma-Aldrich, Saint Quentin Fallavier, France), 100 IU/ml penicillin, 100 μg/ml streptomycin (Invitrogen) and CSF-1 growth factor. This factor was released in their growing medium by LADMAC cells (Sklar et al. 1985). Ten percent of the LADMAC-cell medium was added to a normal culture medium. The culture was kept at 39°C with 5% CO2 for 4 days. The cells were then submitted to labelling with anti human CD14 antibody (MACS CD14 microbeads, Miltenyi Biotec, Paris, France). After washing off the excess beads, the purified CD14+ cells were passed through an AutoMACS separation column (Miltenyi Biotec), according to the manufacturer’s instructions.
Purity and viability of the cell population were estimated by flow cytometry using anti-porcine CD163 monoclonal antibody (clone 74-22-15, IgG1, VMRD, Lissieu, France) and anti-porcine SWC3 monoclonal antibody (IgG1, AbD Serotec, Colmar, France). Macrophages accounted for 85% of the counted cells. Cell viability was greater than 85%.
Neutrophils
Neutrophils were isolated as described previously (Yerle-Bouissou et al. 2009) from the same blood samples. Briefly, after the density gradient the pellet containing the neutrophils was put in a lysis solution (NH4Cl 0.15 M–KHCO3 0.01 M–EDTA Na2 1 μM) at 4°C for 10 min. The cell suspension was then centrifuged at 300×g for 20 min, and then the cell pellet was resuspended in 1 ml of RPMI 1640.
Purity of the cell population was estimated at 90% by microscopic examination of May-Grünwald Giemsa (Sigma-Aldrich) stained blood smears. Cell viability was found to be greater than 95% by a Trypan blue dye exclusion test.
Fibroblasts
Primary porcine skin fibroblast cells were grown as monolayers on glass slides in RPMI-1640 (3.7 × 105 cells/slide) supplemented with 15% foetal calf serum (Eurobio) at 37°C in a 5% CO2 atmosphere.
Cell activation
The macrophage sample (107 cells) was divided into two parts, one of which was incubated in a culture medium (control batch), the other in a culture medium supplemented with LPS (10 μg/ml) and IFNγ (1 ng/ml; activated batch), both for 3 h at 37°C. Culture supernatants from these batches (control and activated) were frozen to −20°C for tumour necrosis factor α (TNFα) and IL1β cytokines quantification by ELISA tests using commercially available ELISA kits (DuoSet, R&D Systems, Lille, France), according to the manufacturer’s instructions.
Similarly, two batches (control and LPS activated) were prepared for neutrophils as described previously (Yerle-Bouissou et al. 2009).
RNA isolation (from macrophages and neutrophils) and quality control
Five different samples from five young female pigs were used for RNA isolation. The samples were prepared and activated independently. Cell suspensions were briefly centrifuged, and the cell pellets were put in a 2-ml solution of Extract All (Eurobio) per 20 × 106 cells for macrophages and in 3 ml solution of Extract All (Eurobio) per 100 × 106 cells for neutrophils. All RNAs were extracted simultaneously. Total RNAs were extracted as specified in the manufacturer’s protocol using the RNeasy Mini column (Qiagen, Courtaboeuf, France) for macrophages and in a NucleospinRNAII column (Macherey-Nagel, Hoerdt, France) for neutrophils and fibroblasts. RNA concentrations were determined by Nanodrop quantification (Thermo Fisher Scientific, Illkirch, France). RNA quality was checked on an Agilent 2100 Bioanalyzer (Agilent Technologies, Massy, France). These RNA samples obtained from the five different animals were used both for microarray and quantitative real-time RT-PCR (qRT-PCR) experiments. Only RNAs with a RIN score between 8 and 10 were used. All RNAs were stored at −80°C.
RNA labelling, microarray hybridization and signal quantification
A common reference hybridization scheme was used to compare gene expression of control and LPS/IFNγ activated macrophages. Each pig/condition RNA was labelled with Cy3 and the common reference, created by mixing all samples, was labelled with Cy5. Total RNA (5 μg) was reverse-transcribed and directly labelled using the ChipShot™ Direct Labeling System (Promega, Charbonniere, France). The CyDye-labelled cDNAs were purified using the ChipShot™ Membrane Clean-Up System (Promega). The absorbance at 260, 550 and 650 nm of CyDye-labelled cDNAs was measured by Nanodrop (Thermo Fisher Scientific). Frequency of incorporation and labelling efficiency were checked by referring to standards provided by Labeled cDNA Calculator (http://www.promega.com/applications/ivt/calculator/#ResultsView). The CyDye-labelled cDNAs were resuspended at a final concentration of 2.5 pmol/μl in DNA/long-oligonucleotide hybridization buffer. A total of ten SLA-RI/NRSP8-13K chips (Gao et al. 2010) were used in our study. Chip hybridization was performed using the Corning hybridization system. Prior to hybridization, the slides were treated with the background-reducing Pronto Pre-Soak System (Corning) and then prehybridized using the Pronto Universal Hybridization Solutions and Kits (Corning). Hybridizations were carried out for 16 h at 42°C with the Ventana Discovery XT System. The slides were washed according to the recommended protocol (Corning) and dried by centrifugation at 1,000 rpm for 1 min. The slides were scanned using a GenePix™ 4000B array scanner (Molecular Devices, Saint-Grégoire, France), and the array images were then processed with the GenePix™ Pro software V6.0 (Molecular Devices) to align spots, integrate ID data files and export reports of spot intensity data.
Microarray data statistical analyses
To identify significant differential expressions, the microarray data were analysed using Limma (Linear Models for Microarray Data; Smyth 2004) from the Bioconductor open-source project running under R (Gentleman et al. 2004). After data pre-processing using within-array global loess normalization, the empirical eBayes method in Limma, which computes moderated t statistics and log-odds of differential expression, was applied to identify the significance of differential expression for each culture condition. The adjustment for multiple testing was carried out using the false discovery rate (FDR) method (Reiner et al. 2003) in Limma. Raw and processed microarray data along with details on sample processing and microarray data analysis are available in the Gene Expression Omnibus (www.ncbi.nlm.nih.gov/geo) under the accession number: GSE25491.
Gene network analyses
The differentially expressed genes were analysed using Ingenuity Pathway Analysis (IPA; Ingenuity Systems, USA) and Gene Set Enrichment Analysis (GSEA).
IPA software provides targeted information on genes, proteins, chemicals and drugs, and builds interactive models. We used it to map genes with known human locus IDs with corresponding differential expression values to the corresponding gene object in the Knowledge Base. Gene networks were algorithmically generated based on their connectivity and a significance score was assigned for each network, with an associated p value displayed as −log10.
The GSEA method uses a variation of a Kolmogorov–Smirnov statistic to provide an enrichment score for a gene set (Subramanian et al. 2005).
Quantitative real-time RT-PCR
Two microgrammes of total RNAs was reverse-transcribed using Superscript II enzyme with Oligo (dT) primers. The cDNAs were quantified using a Nanodrop (Thermo Fisher Scientific) and diluted to a working concentration (20 ng/μl). Triplicate reactions were performed in a final volume of 20 μl with 50 ng cDNA, 300 nM primers and SYBR Green PCR Master Mix (Applied Biosystem, Courtaboeuf, France), using a LightCycler® 480 System (Roche, Meylan, France). Primers were chosen with the Primer Express Software (Supplementary Table 1). β-2-Microglobulin was chosen as the internal reference gene, and the Dart-PCR method was used to calculate the fold change in gene expression as had previously been described (Devriendt et al. 2009). In order to make the comparison of the two technologies more reliable, qRT-PCRs were carried out using the same RNA samples that were used for the microarray experiments.
3D-FISH experiment
Preparation of slides
The slides were prepared with cells isolated from one animal. For macrophages, as for neutrophils, a 5 × 106 cell/ml suspension in RPMI 1640 was applied to polylysine® slides (CML, Nemours, France) for 10 min to allow cell adhesion. 3D fixation was performed similarly for macrophages, neutrophils and fibroblasts at room temperature for 10 min with a 4% formaldehyde solution in 1× phosphate-buffered saline (PBS) freshly made from paraformaldehyde followed by a short wash in 1× PBS. Before storage at −20°C, the slides were immersed in 1× PBS containing 20% of glycerol for 30 min.
Preparation of complex DNA probes (chromosome painting and BAC clones)
Porcine chromosome paint probes from flow-sorted chromosomes (Yerle et al. 1993) were individually directly labelled by random priming with Alexa 568 (Invitrogen, Cergy Pontoise, France). BAC clones containing the selected genes were isolated from a porcine BAC library (CRB-GADIE, INRA, Jouy-en-Josas, France) using specific primers (Supplementary Table 1). The BAC numbers are listed in Table 1. These BACs were random priming labelled by incorporation of dUTP Alexa 488 using the Bioprime DNA labeling kit (Invitrogen). Products from the reaction were precipitated with porcine Cot-1 DNA (Applied Genetics Laboratories, Melbourne, USA) and salmon sperm DNA (Eurobio). Each probe was dropped onto slides at a final concentration of 100 ng/μl in a hybridization buffer. The specificity of all probes was tested by 2D-FISH on porcine metaphase spreads prepared from lymphocytes according to protocols which had previously been described (Yerle et al. 1994).
3D-FISH experiment
3D-FISH experiments were carried out according to (Yerle-Bouissou et al. 2009) with some modifications for macrophages and fibroblasts: (a) For macrophages, stored cells (−20°C) were rinsed for 10 min in 1× PBS followed by a 2-min wash in 1× PBS. Cells were then permeabilized with 0.5% Triton X-100 and 0.5% saponin in 1× PBS for 15 min at room temperature. After a brief wash in 1× PBS (2 min), the slides were immersed in Tris–HCl 0.1 M pH 7.2 for 15 min. After a bath in 20% glycerol in 1× PBS for 30 min at room temperature, cells were freeze–thawed (six times) in liquid nitrogen, washed in 1× PBS for 5 min, treated with 200 μg/ml RNase A for 30 min at 37°C. Cells were then washed in 2× saline sodium citrate (SSC) for 5 min and treated with 0.1 N HCl for 2 × 15 min at room temperature. They were washed in 2× SSC for 5 min and incubated in 50% formamide and 2× SSC for at least 3 days at 4°C. (b) For fibroblasts, softer treatments were applied: post-fixation was omitted, only one incubation in 0.1 N HCl for 10 min was performed and the freeze–thaw operations with liquid nitrogen were performed five times. Briefly, cells and probes were simultaneously heat-denatured at 75°C for 8 min and then incubated for 72 h for macrophages and neutrophils or 48 h for fibroblasts at 37°C. The entire procedure was carried out in a DAKO hybridizer. Post-hybridization washes were performed first in 2× SSC at room temperature (twice) for 15 min, then three times for 15 min in 2× SSC 50% formamide pH 7.0 at 45°C and finally, three times for 15 min in 0.1× SSC at 50°C. Nuclei were counterstained with 4′,6′diamidino-2-phenylindole in Vectashield medium (Vector Laboratories, Burlingame, USA).
Confocal microscopy and image analysis
Confocal microscopy was carried out using a Leica TCS SP2 confocal microscope (Leica Instruments, Heidelberg, Germany) equipped with an oil immersion objective (plan achromatic 63× N.A. = 1.4). The Z-stacks were acquired at 1,024 × 1,024 pixels per frame using an 8-bit pixel depth for each channel at a constant voxel size of 0.077 × 0.077 × 0.244 μm. Typically, a stack of 45 confocal planes was acquired. Segmentations and 3D measurements between objects (nucleus, genes and CT) were done using NEMO (Iannuccelli et al. 2010), developed from Smart 3D-FISH software (Gué et al. 2005). .Briefly, NEMO offers a function that automatically detects cells in fields by taking into account circularity and minimum area parameters for individual nuclei. Then processing and segmentation quality estimation parameters are entered. Processing parameters define xyz resolutions and filters to run on raw images: 3D median and 3D mathematical morphology filters were applied to all objects and a TopHat filter was applied to the gene channel. All objects are detected automatically by the intensity of pixels above a globally set threshold. After processing, users visually inspect image segmentation and distance values using the NEMO interface. If necessary, the segmentation threshold can be adjusted manually to improve object detection.
3D-FISH statistical analyses
Macrophage nuclei have a spherical shape, so the radial position of each gene was calculated using the distance between the gene centre and the nucleus centre normalized by the local radius (corresponding to the nucleus radius measured through the gene centre). The cumulative distributions of radial positions for each gene under the two conditions (control and activated) were compared using a two-sided Kolmogorov–Smirnov test (R software, R Development Core Team 2005).
To determine the position of the genes relative to their CT, we defined three categories (inside, edge, outside) taking into account the gene centre to CT edge distances and the percentage of gene co-localization in CT. The edge category comprises genes located at less than one voxel from the edge of the segmented CT. A χ 2 test was used to compare the gene distribution in these categories. p values <0.05 were considered as significant.
Results
Genes differentially expressed in porcine macrophages activated by LPS/IFNγ
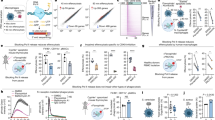
Monocyte-derived macrophages, obtained as described in the “Materials and methods” section, were activated by LPS and IFNγ stimulation. The efficacy of the activation protocol was assessed by measuring the production of pro-inflammatory cytokines, TNFα and IL1β. Macrophages naturally produce TNFα and IL1β but this production increases under activation. In the present experiment, the production of TNFα (from 476 pg/μl to 5930 pg/μl) and IL1β (from 362 pg/μl to 3668 pg/μl) was increased 12 and 10 times, respectively, in LPS/IFNγ treated cells.
The analysis of gene expression profiles of control and activated macrophages was carried out using a dedicated microarray, the porcine SLA-RI/NRSP8-13K chip which includes a dedicated SLA-RI oligonucleotide set (3,773 probes targeting Swine Leukocyte Antigens and Immune Response), the Qiagen-NRSP8 microarray oligonucleotide set and a series of positive and negative controls, totalling 19,200 probes (Gao et al. 2010). The experimental model was designed to compare samples from control macrophages with LPS/IFNγ-activated macrophages using a common reference.
The comparison of control and activated macrophages revealed 68 probes (Fig. 1 and Supplementary Table 2) targeting 60 differentially expressed genes with a 5% FDR. Of these probes, 45 originated from the pangenomic set (66%) and 23 from the SLA-RI set (34%). Seven probes were down-regulated (10%) and 61 were up-regulated (90%) on LPS/IFNγ activation. A low amplitude of log ratio was observed for these genes. Interestingly, in this selection, eight up-regulated genes were targeted with probes from the two sets and concordant expression levels were obtained (Supplementary Table 2). The 60 differentially expressed genes are well distributed on the porcine genome as all chromosomes except SSC16 (Sus scrofa domestica 16), a relatively small chromosome, carry at least one of these genes (http://www.ensembl.org/index.html; Supplementary Fig. 1).
LPS/IFNγ-related gene networks
Using Ingenuity Pathway Analysis (IPA), a web-based software application that enables to analyse, integrate and understand data derived from gene expression (http://www.ingenuity.com/index.html), we found that the 60 differentially expressed genes are associated with three biological function catalogues: “Diseases and Disorders”, “Molecular and Cellular Functions” and “Physiological System Development and Function”. More precisely, in each catalogue, five main categories are defined as shown in Fig. 2. The “Genetic Disorder” class (24 genes) is well represented, probably due to the large number of regulatory elements and inducible factors linked to the 60 selected genes. The other major categories “Antigen presentation” (24 genes), “Hematological system development and function” (23 genes) and “Cell-mediated immune response” (21 genes) are clearly related to our cellular model. Overall, eight of the 15 categories found by the IPA analysis are related to immunity.
IPA analysis of the DNA microarray data: numbers of genes implicated in the principal biological function categories related to the catalogues Diseases and Disorders, Molecular and Cellular Function and Physiological System Development and Function. Categories associated with immunity are shown in blue
Applying IPA to the 60 differentially expressed genes allowed us to define three gene networks with scores over 2 (corresponding to 99% confidence of their not being randomly generated; Fig. 3). Network 1 (score of 49) implies 22 up-regulated genes (Fig. 3a) and is associated with cellular movement, haematological system development and function, and immune cell trafficking. This network comprises a large number of cytokines and chemokines such as interferons. P38 MAP kinase is also strongly implicated in terms of its interactions. Network 2 (score of 25) groups 11 genes, among which ten are up-regulated and only one is down-regulated (Fig. 3b). This network is implicated in the development and function of connective tissue, tissue morphology and cancer, and includes interferon gamma and the proto-oncogene MYC. Network 3, with the lowest score (19) groups nine genes, of which six are up- and three are down-regulated (Fig. 3c). This network, which is related to cellular movement, haematological system development and function and immune cell trafficking, is centred on cytokines IL1β and IL8, and includes the NFKBIA and NFκB complexes.
LPS-related gene networks identified by IPA. a Network 1. b Network 2. c Network 3. The up- and down-regulated input proteins are coloured in red and green, respectively. The colour intensity is directly proportional to fold-change values reported in Supplementary Table 2. The pathway components identified by the algorithm and not detected in the analysis are shown in white
A Gene Set Enrichment Analysis (GSEA; Subramanian et al. 2005) was also carried out in order to have a global view of the activation process. Using this approach, we found 20 positively regulated gene sets, seven of which involve regulation of an interferon (Supplementary Table 3). In addition, 18 gene sets were found to be down-regulated (Supplementary Table 3); two of these are involved in the cytoskeleton and two others in the ribosomes. These analyses confirm the hypothesis that the predominant genes affected by LPS/IFNγ stimulation are related to immunity, cell movement and cell interactions.
Gene selection and validation for nuclear analysis by 3D-FISH
To ascertain whether a variation in gene expression (up- or down-regulation) is associated with repositioning in the nuclear space, we selected from the DNA array results, the top 4 down-regulated genes: NEDD4L, VIM, LGALS3 and TUBA3 and the top four up-regulated ones: IL1β, SLAMF7, IL8 and CXCL10 irrespective of their position in a network. TNFα was added to the list of the up-regulated genes according to data obtained from the literature (Bradley 2008) demonstrating its central role in inflammation. Two additional genes were added to this selection: IGF2, which harbours no significant expression level difference between control and activated states, and ZAR1, which is non-expressed in macrophages. Chromosomal positions and cellular functions of these selected genes are given in Table 1.
Differential expressions of these genes were validated by qRT-PCR. Globally concordant results were obtained with the two methods but larger amplitude ratios were observed in qRT-PCR experiments than with the DNA array approach (Fig. 4a). However, differences were observed for two genes: NEDD4L was found to be down-regulated by the microarray analysis while the qRT-PCR did not confirm this down-regulation. On the contrary, a down-regulation of IGF2 was revealed by the qRT-PCR analysis while it was not found to be differentially expressed in the microarray analysis. We checked the sequence homology of the NEDD4L oligonucleotide present on the microarray used with other porcine ESTs and found a high sequence homology (98%) with another EST (AJ679682) targeting the smooth muscle protein 22-alpha. Similarly the IGF2 oligonucleotide, shares 100% sequence homology with the porcine EST AY242112 targeting tyrosine hydroxylase. This could explain the discrepancy, in other words why the expression levels obtained do not correspond to the expected genes. Since the qRT-PCR is a more accurate method, we selected IGF2 and removed NEDD4L from the list of selected genes for the 3D-FISH analysis. The non-expression of ZAR1 in macrophages was confirmed by classical RT-PCR (Supplementary Fig. 2).
Normalized graphs for mRNA expression levels in macrophages and neutrophils. a Expression level in activated macrophages relative to control macrophages analysed by microarray (blue) and qRT-PCR (red). For each gene, the expression level in control cells is considered as a baseline. b Expression level in activated neutrophils relative to control neutrophils analysed by qRT-PCR (*p < 0.05; **p < 0.01; ***p < 0.001)
Gene-specific positioning during macrophage activation by LPS/IFNγ
In order to correlate gene repositioning with change in gene expression during macrophage activation, we investigated the nuclear localization of the selected genes along with their associated chromosomes by 3D-FISH. The chromosomal localizations of the selected genes were controlled by 2D-FISH on metaphases using the BAC clones (Supplementary Fig. 3). SLAMF7 was thus excluded because the probe used does not target the expected chromosome (SSC4). The selected genes map to 7 different chromosomes: SSC1, SSC2, SSC3, SSC5, SSC7, SSC8 and SSC10. Interestingly, three genes among the nine investigated map to the same chromosome (SSC8) and have a different expression status, two being up-regulated (IL8 and CXCL10) and one non-expressed (ZAR1). We used structurally preserved macrophages in combination with confocal microscopy and 3D image analysis to investigate the nuclear positioning of the selected genes relative both to the nuclear radius and to their associated chromosome territories (Fig. 5). In each 3D-FISH experiment, we generated serial optical sections from around 80 to 140 nuclei of macrophages in both control and activated states. Image stacks were processed with NEMO software (Iannuccelli et al. 2010).
Analyses of gene positioning in the cell nuclei by 3D-FISH and confocal microscopy. Series of photos showing structurally preserved nuclei of control or activated macrophages and neutrophils and of fibroblasts after hybridization with painting probes specific to chromosome territories (red) and BACs containing genes (green). Maximal intensity projections of confocal image stacks are shown. DNA in the nuclei is counterstained with 4′,6′diamidino-2-phenylindole (DAPI, blue)
Radial positioning of genes in the nuclei
Distances between the BAC signal centres and the nucleus centres were measured for the 9 selected genes and were normalized individually to the nuclear radius of each cell. These radial distances are shown in Fig. 6. IGF2 is located in the nuclear interior (Fig. 6e), VIM, TUBA3, TNFα and IL1β in an intermediate position (Fig. 6a, d, f, g,) and LGALS3, IL8, CXCL10 and ZAR1 are located closer to the nuclear periphery (Fig. 6b, h, c, i). To ascertain whether any of these genes had undergone repositioning within the nuclear space when the macrophages were activated, we compared their radial distances in both states using a two-sided Kolmogorov–Smirnov test. Only TNFα changes its radial position significantly (p = 0.04, Fig. 6f), moving from an intermediate point to a more external position. In addition, the comparison of the radial positions of the three genes located on SSC8 revealed no significant difference (p values >0.06), while two were found to be up-regulated (IL8 and CXCL10) and one non-expressed (ZAR1). Similarly, the down-regulated gene VIM occupies a radial position that is not different from the one occupied by TNFα and IL1β, both up-regulated (p = 0.287 and p = 0.816).
Radial distribution of genes in control and activated macrophages. a–i Cumulative frequency graphs of the radial positions for each gene in control (grey) and activated (red for up-regulated genes and green for down-regulated) macrophages. The cumulative frequency curve of a random distribution (P(X < d) = d 3 for 0 < d < 1 where X is the random radial position) is shown in black in each graph for comparison. j Pairwise comparisons (p values) of cumulative radial distributions (two-sided Kolmogorov–Smirnov test) are indicated in each graph. The number of loci analysed in each condition is given at the bottom right of the graphs
We continued by analysing the threshold cycle of qRT-PCR for each gene in each state (Supplementary Fig. 4), which gives a good trend of expression level in relation to radial positioning. No correlation was found between gene activity level and radial positioning as assessed by a Pearson test on activated and control macrophages (p = 0.055 and p = 0.283, respectively).
Positioning within the associated chromosome
We next investigated whether changes in gene activity correlate with changes in positioning relative to the chromosome territories. To address this question, we used the gene probe (BAC) combined with the parent chromosome territory probe (whole chromosome painting probe) in 3D-FISH experiments (Fig. 5). For each “gene/chromosome territory” pair we obtained the distance between the gene centre and the chromosome territory edge and the percentage of gene co-localization in its chromosome territory. We assigned negative values to the distances when the co-localization percentages were greater than 50%. In this way, we created three categories: inside (distances < 0), edge (distances = 0), and outside (distances > 0). The results are given in Fig. 7. The first important observation is that the position of the genes relative to their chromosome territories differs from one gene to another. VIM is clearly positioned outside its chromosome territory (Fig. 7a) in 75% and 74% of control and activated cells, respectively. In the same way, more than 60% of IGF2 signals is located outside SSC2 territory under both conditions (Fig. 7e). Conversely, IL1β and ZAR1 are clearly located in their CT (in 50% of the cells whatever the state analysed) (Fig. 7g, i). Other genes locate preferentially at the edge or outside the segmented chromosome territories under both conditions: this is the case of LGALS3 (Fig. 7b), TNFα (Fig. 7f), IL8 (Fig. 7h) and CXCL10 (Fig. 7c). Finally, no trend emerges for TUBA3 (Fig. 7d), which is distributed equally in the three classes (around 30% to 35% of signals in all classes).
Nuclear positioning of gene loci relative to their home chromosome territories in interphase nuclei of porcine macrophages (a–i) and neutrophils (j–o). The number of loci (n) analysed for each gene under each condition is indicated at the top left of the histograms. The distribution of a given gene was ranked in three classes (inside, edge and outside) and compared between control (grey) and LPS/IFNγ-activated macrophages (green, red or black for down- and up-regulated or non-expressed genes, respectively). The three classes are defined by combining the 3D distances: gene centre-to-chromosome territory edge and the percentage of co-localization (gene in its chromosome territory). These data were generated using NEMO (Iannuccelli et al. 2010). Statistical differences were assessed by a χ 2 test and p values are indicated on the graphs (top right)
The second important point is that among the nine genes, the χ 2 test used between groups of positions revealed that only one gene changes its position in its chromosome territory when the macrophages are activated. This concerns IL8, up-regulated in activated macrophages, which clearly moves outside SSC8 (p = 0.003) upon activation: the number of events in the inside category decreases (from 22% to 11%) while it increases in the outside category (from 40% to 50%). However, even though the number of events in the three categories does not vary significantly for TNFα and CXCL10, we observed that the genes tend to be further from the edge of their chromosome territories. Consequently we then investigated whether the distances from gene centre to CT edge vary when the immune cells are activated. We normalized these distances by the corresponding nucleus radius in the two macrophage batches (control and LPS/IFNγ activated). The results of a Mann–Whitney Wilcoxon test revealed that the distances increase significantly in the activated macrophages for TNFα (p = 0.001), IL8 (p = 0.022) and CXCL10 (p = 0.025). However, no significant variation was detected for IL1β (p = 0.28), which tends to remain inside its CT, or for the four down-regulated genes (TUBA3, p = 0.53; LGALS3, p = 0.24; VIM, p = 0.13; IGF2, p = 0.47; Table 2). The greatest values for the average normalized distances are observed for the two down-regulated genes, VIM and IGF2, which have a more pronounced tendency to be outside their chromosome territories (Fig. 7) both in control and activated macrophages.
To ascertain whether the results concerning gene behaviour relative to their CT during the activation of macrophages are gene specific and cell type specific, we analysed what happens in neutrophils as another immune cell type. Although macrophages and neutrophils both belong to the family of phagocyte cells, their mechanism of action against bacterial infection is quite different. Another important difference is observed at the nucleus level as neutrophils have a polylobed nucleus. So we extended the analysis to neutrophils by investigating the three up-regulated genes, TNFα, IL1β and IL8, two down-regulated genes, TUBA3 and IGF2 and the non-expressed gene ZAR1. We first confirmed by qRT-PCR that these genes have the same expression status in LPS-activated neutrophils and macrophages (Fig. 4b). 3D-FISH analyses were then carried out on control and activated neutrophils (Fig. 5) and revealed that globally the gene positions relative to their chromosome territories are similar: the down-regulated TUBA3 gene is distributed equally across the three positions in both cell types; TNFα, IL8 and IGF2 tend to be outside their chromosome territories (Fig. 7l, n, k) while IL1β remains preferentially in its CT (Fig. 7m). Interestingly, only IL8 shows clear repositioning outside its CT both in activated macrophages (p = 0.003) and neutrophils (p = 10−16; Fig. 7h, n).
Finally to determine whether the observed preferential position at the edge or outside the chromosome territory of the three up-regulated cytokines IL8, CXCL10 and TNFα is specific for cells engaged in immune response, we studied their position in fibroblasts as a control cell line which is not related to immune response. We also analysed the fourth up-regulated cytokine IL1β found inside its chromosome territory both in control and activated macrophages and neutrophils. We first analysed their expression status by qRT-PCR. As expected, this analysis revealed that the four cytokines are expressed much more in immune cells than in fibroblasts: IL8 is 470 times more expressed in activated macrophages than in fibroblasts, CXCL10 2,878 times, TNFα 47 times and finally, IL1β 764 times (Supplementary Fig. 5). 3D analyses were then carried on in fibroblasts (Fig. 5) and revealed that while IL8 and CXCL10 tend to be clearly at the edge or outside their chromosome territories in immune cells whatever the state (control or activated), they do not have the same position in fibroblasts where they are equally distributed in the three classes (Supplementary Fig. 6c, d). TNFα has a tendency to be at the border or outside its CT in the three cell types while IL1β remains in its CT (Supplementary Fig. 6a, b).
Discussion
In order to investigate the role of genome nuclear organization in immune response, we used in vitro immune cells (porcine macrophages and neutrophils) and activated them with LPS/IFNγ or LPS to mimic bacterial infection. Indeed, the pig has proven to be a very important model organism for studying not only various diseases (arteriosclerosis, obesity and diabetes) but also for understanding resistance to disease (Litten-Brown et al. 2010). Compared to the mouse as a model organism for humans, it has the great advantage of more similar genome behaviour, particularly in terms of the immune system (Lunney et al. 2009). Consequently, it is important to improve our basic knowledge of immune response in this species. In this context, the first phase of our study consisted in determining which genes are differentially expressed when immune cells undergo activation, the second in selecting genes with a variation of expression, and the third in studying the nuclear positioning of these target genes to ascertain whether a variation of expression implies a repositioning in the nuclear space. Numerous studies have investigated this important question in the context of cell differentiation (Stadler et al. 2004; Ragoczy et al. 2006; Szczerbal et al. 2009) or malignancy development (Meaburn and Misteli 2008), but we found no detailed information in the available literature for mammalian phagocytic cells subjected to an activation process mimicking a pathogen aggression.
Expression profiles of porcine macrophages
Transcriptomic analyses have been conducted to define the expression profiles of human macrophages under various conditions (Detweiler et al. 2001; Nau et al. 2002), but it was not possible to study these mechanisms in swine, due to the lack of appropriate tools, until a few years ago. Some transcriptomic studies are now available for porcine alveolar macrophages infected by viruses (Zhang et al. 2006; Xiao et al. 2010). Today, progress in gene annotations (Archibald et al. 2010) and DNA chips targeting immunity (Gao et al. 2010) makes it possible to carry the transcriptional analysis of porcine macrophages further. A recent analysis was conducted on peripheral blood mononuclear cells (PBMC) (Gao et al. 2010) using the same dedicated microarray, but our study focuses on swine monocyte-derived macrophages under LPS/IFNγ stimulation. Indeed, LPS, which interacts with a large number of proteins including LPS-binding protein, CD14 and TLR4, has been extensively used to study innate immune response in different species (Boldrick et al. 2002; Nau et al. 2002; Wells et al. 2003) and is particularly appropriate for activating monocytes and macrophages that express the cell surface receptor CD14. IFNγ, principally produced by T-lymphocyte helpers, was added to LPS to mimic an inflammatory context and to promote classical type 1 macrophage activation (Ma et al. 2003). According to this study, most of the mRNAs elicited by LPS/IFNγ encode proteins that are important for the innate immune response. Our analysis revealed that 60 genes of the 17,070 genes present on the array show a significant difference in expression levels when the macrophages are activated by LPS/IFNγ (Supplementary Table 2). This number is the result of a strong selection due to the choice of a stringent false discovery rate (5%) applied to obtain the more likely candidates for 3D-FISH analysis. Our results are consistent with the data obtained on human macrophages activated by LPS (Nau et al. 2002) as ten common genes, including IL1β, IL8, SAAL1 and NFKBIA, were found to be up-regulated. A similar analysis on porcine PBMC activated with LPS revealed a greater number of genes differentially expressed (Gao et al. 2010). This is not surprising as we focused on a particular cell type, obtained by differentiation of monocytes, which represents only a small proportion of the PBMC (8% in swine). It is necessary to identify transcriptome modifications occurring in each cell subtype in order to better decipher immune response, and this is essential in our approach, which aims to correlate levels of expression and nuclear organization.
Genes and networks involved during LPS/IFNγ stimulation
Inflammation is a powerful protective mechanism which is coordinated and controlled by cytokines and chemokines and, as expected, we detected an increase in the expression level of IL1β and IL8, and members of the CXCL family (CXCL10, CXCL11 and CXCL17). One of the three networks (Fig. 3c) obtained by IPA analysis demonstrated the implication of several transcription factors, particularly NFKBIA and RNAPolII related to cytokines (IL1β and IL8), suggesting their important induction during macrophage activation as found in previous studies (Nau et al. 2002; Gao et al. 2010). Indeed, these proteins play a key role in immune response, targeting neutrophils at the inflammation site (IL8) or inducing cell proliferation and differentiation (IL1β). TNFα was found to be up-regulated in our study and is a well-known initiator and enhancer of macrophage activation (Zhang et al. 2010). These cytokine and IFNγ are related to transcription factors found in another network (Fig. 3a): TNFα uses TRAFs (TNF Receptor Associated Factor) to activate NFκB and interferons initiate Janus kinase–signal transducer and activator of transcription and tyrosine kinase cascades. The LPS/IFNγ activation used leads to a classical type 1 activation of macrophages (resulting in nitric oxide and cytokine production and a decrease in phagocytosis) as described previously in other species (Ma et al. 2003; Zhang et al. 2010).
In the context of an activation process investigated by transcriptomic analysis, the number of genes found to be down-regulated is often smaller than the number of up-regulated ones (Jenner and Young 2005). Our data obtained for porcine macrophages activated by LPS/IFNγ are concordant with these observations, as seven genes among the 60 differentially expressed were found to be down-regulated. Two of them (VIM and TUBA3) are factors in cytoskeleton dynamics as shown by the GSEA analysis. Locomotion for phagocytosis and regulation of cell shape are important elements of macrophage functions which partly modulate the inflammatory response. In this context, the cytoskeleton is a fundamental point of regulation and needs to be very flexible in macrophages. Curiously, in our analysis, genes implicated in cytoskeleton structure are down-regulated. We postulate that the synthesis of proteins implicated in basic cellular functions such as cytoskeleton maintenance is reduced to enhance the synthesis of cytokines and other proteins implicated in immune functions. This competition for transcription factors between house-keeping genes and induced genes may serve to reduce the expenditure of energy for internal cellular functions in activated macrophages.
Nuclear positioning of specific genes
Differential positioning of active and inactive genes, relative to nuclear domains, had led to the hypothesis that nucleus positioning is functionally important and influences gene activity (Lanctôt et al. 2007; Sexton et al. 2007; Pai and Engelke 2010). Several studies illustrated that the nuclear periphery can act as an inactive compartment and that repositioning towards the nuclear interior is required to allow efficient transcription, as for the immunoglobulin heavy chain in B lymphocytes (Kosak et al. 2002) and for CFTR in different human cell types (Zink et al. 2004). However, recent studies in human cells demonstrated that location adjacent to the nuclear periphery is not incompatible with active transcription (Finlan et al. 2008; Kumaran and Spector 2008). In addition, cases of lack of repositioning events despite changes in gene activation have also been reported (Küpper et al. 2007) which underlies the difficulty in defining unifying rules.
Another important point is that even though the nuclear arrangement of CT according to gene density has been shown to be conserved in vertebrates (Tanabe et al. 2002), differences have been observed between species in terms of links between nuclear positioning and transcriptional activity (Sadoni et al. 2008).
In this context, our aim was to investigate whether changes in gene activity in activated immune cells were related to repositioning events in the nucleus. The genes we selected for our analysis, except for ZAR1, were expressed both in control and in activated macrophages and were subjected to changes in their expression levels. We first analysed their radial positions and demonstrated that when the macrophages are activated, a significant change in radial positioning was only observed for TNFα, up-regulated, that became more peripherally localized. We then compared the radial positions of the 3 genes located on SSC8 and found no significant difference while two are up-regulated (IL8 and CXCL10) and one non-expressed (ZAR1). Similarly, the down-regulated gene VIM occupies a radial position that is not different from the one occupied by TNFα and IL1β, both up-regulated. These results tend to demonstrate that there is no link between radial positioning and variations of expression in these immune cells.
We continued by analysing the threshold cycle of qRT-PCR for each gene in each state (Supplementary Fig. 4), which gives a good trend of expression level in relation to radial positioning. No correlation was found between gene activity level and radial positioning. Our results agree with previous reports on the lack of correlation between activity level and nuclear radial positioning (Scheuermann et al. 2004; Meaburn and Misteli 2008). When we compared the radial positions of the selected genes, the ones we found more externally situated (LGALS3, IL8, CXCL10 and ZAR1) are those located on chromosomes with a low gene density (6 genes/Mb for SSC1 and 4.9 genes/Mb for SSC8) while IGF2 on SSC2 (12.3 genes/Mb), TNFα on SSC7 (12.3 genes/Mb) and TUBA3 on SSC5 (9.8 genes/Mb) are clearly more central. Even though these estimations of gene density (http://www.ensembl.org/Sus_scrofa) should be treated with caution, as the porcine genome is still being sequenced (Archibald et al. 2010), these data suggest that the specific location of each gene is more closely linked to the position of its home chromosome than to its expression level as previously observed (Federico et al. 2008).
Our data demonstrate that when the macrophages react to a bacterial infection, variations in expression level occur for several genes; however, these variations do not imply a global repositioning of genes in the nuclear space, as we only observed one gene moving significantly towards the nucleus periphery. This gene, TNFα, belongs to the major histocompatibility complex and is located on SSC7. Our hypothesis is that the whole of this highly transcribed MHC genomic region rearranges upon activation in immune cells as observed previously (Volpi et al. 2000) and, more globally, that the chromatin of SSC7 is in a less condensed state (Fig. 5). This phenomenon has been observed previously for chromosome 1 in chicken macrophages (Stadler et al. 2004).
The activation process of immune cells by LPS/IFNγ is not associated with major genome reorganization, contrary to what has been observed in several models of cell differentiation. Indeed it has been shown that during adipocyte differentiation, the position of seven adipogenesis genes changed significantly (Szczerbal et al. 2009). A similar phenomenon was observed during normal differentiation of mammary epithelial cells into acini, as seven out of 11 genes tested occupied distinct radial positions (Meaburn and Misteli 2008). However, in both cases, it occurred during a normal differentiation process. The situation has been shown to be different for malignant differentiation as no major difference was detected when malignant mammary differentiation was compared to normal mammary differentiation (Meaburn and Misteli 2008). Another recently published example illustrates the absence of global nuclear reorganization of genes or chromosome territories after hormone addition and transcriptional activation in human breast epithelial and cancer cell lines (Kocanova et al. 2010). The activation of immune cells by LPS/IFNγ induces inflammatory response and so is closer in terms of cellular reactions to processes that occur during tumourigenesis development or cell activation than during normal cell differentiation.
Gene positions relative to their territory
It has been argued that the active genes are located preferentially at the periphery of chromosome territories or even outside (Kurz et al. 1996; Dietzel et al. 1999; Bártová et al. 2002; Chambeyron and Bickmore 2004; Harnicarová et al. 2006), whereas other reports indicate no preference in the position of active genes that may also be positioned in the CT interior (Mahy et al. 2002; Zink et al. 2004; Küpper et al. 2007). Our data are concordant with the latter view as IL1β, which is expressed in macrophages and moreover up-regulated in activated macrophages, remains preferentially in its CT. However, the comparison of the basal expression levels of IL1β and IL8 (which is clearly outside its CT) estimated by the qRT-PCR threshold cycle, revealed that IL1β is less expressed than IL8 (Supplementary Fig. 4). We obtained exactly the same results in control and activated neutrophils (data not shown).
A key goal of our study was to ascertain whether the variation in expression level for some target genes, due to the activation process, is associated with changes in their behaviour towards their CT. A significant change was detected for IL8 as its tendency to be outside its CT was higher in activated macrophages and neutrophils than in control ones.
The division into three classes (inside, edge and outside) gave a global view of the gene location relative to its CT. We went further by comparing the normalized CT edge to gene centre 3D distances (for the inside and outside classes), in control and activated macrophages. This analysis revealed that for the three up-regulated cytokines, IL8, TNFα and CXCL10, all of which also have a tendency to be outside their CT, these distances increase significantly in activated macrophages (Table 2). Altogether, these results illustrate that the transcriptional activation of IL8, TNFα and CXCL10 by LPS/IFNγ leads to further decondensation and increased accessibility of chromatin, manifested as an increase in the frequency with which the region appears on an external loop (IL8) or in an increase in the distance from the CT edge (IL8, TNFα and CXCL10). Several studies focusing on the MHC region have already demonstrated such phenomenon after transcriptional up-regulation by IFNγ (Volpi et al. 2000; Christova et al. 2007; Kumar et al. 2007; Ottaviani et al. 2008). Our data report a similar behaviour for a gene of the MHC complex in porcine phagocytic cells. Our hypothesis is that it can be extended to other genes whose physiological activation by IFNγ is a key step in the cell-mediated immune response.
Taken together, these observations suggest that individual genomic regions have a specific 3D configuration that is mainly gene dependent but also, in our immune cell models, activity dependent. Another point in favour of this hypothesis is that IL8, CXCL10 and ZAR1 both map to SSC8 but behave differently: the position of IL8 and CXCL10 (both up-regulated) relative to SSC8 is modified by the activation process, while the position of ZAR1 (non-expressed) was not. IL8 and CXCL10, separated by only 3 Mb, are in a region where neighbouring genes (clusters of inflammatory chemokines) are globally active, contrary to ZAR1 (Fig. 8). The existence of such regions with increased gene expression has already been described (Caron et al. 2001; Koutná et al. 2007). Their extension outside the CT has been reported as an important element for their interaction with other elements in the nucleus or their grouping in specific transcription factories (Osborne et al. 2004; Soutoglou and Misteli 2007; Szczerbal and Bridger 2010).
Distribution of the expressed genes on chromosomes. Genes annotated in the Ensembl genome browser and expressed in macrophages (data from microarray) are shown in their chromosomal position, by their differential expression levels (green = low, red = high) in active state compared to control. For each chromosome, the ‘plus’ strand is represented in the left-hand column and the ‘minus’ strand in the right-hand column. Genes selected for the 3D-FISH analyses are represented and named in their corresponding strand
However, our observations also confirmed that external loops might not be an absolute requirement for transcription per se (Christova et al. 2007) as IL1β, which is also an up-regulated cytokine in activated immune cells (macrophages and neutrophils), remains in its CT while it is actively transcribed. Interestingly, IL1β is surrounded by other genes that were not found to be differentially expressed upon stimulation. Taking into account the data available, IL1β remains the only up-regulated gene in this region (Fig. 8).
The results of our study demonstrate that LPS/IFNγ stimulation of macrophages leads to variations in gene expression. A systematic relocation of these genes in the nucleus space was not observed. Indeed, three of the four up-regulated genes studied relocated in the nucleus (TNFα) and/or relatively to their CT (TNFα, IL8 and CXCL10) while the fourth (IL1β) did not move. In addition, the 4 down-regulated genes analysed did not show any significant change in their nuclear positioning. We found the same results in another type of immune cell (neutrophils) despite the particular structure of their nucleus suggesting that this gene behaviour is probably specific to cells involved in immune response. The results we obtained on fibroblasts chosen as a control cell line which is not involved in immune response, confirm this hypothesis. Indeed, the two up-regulated cytokines (IL8, CXCL10), highly expressed in macrophages even before activation, are located rather outside or at the edge of their CT in this cell type while this tendency is not found in fibroblasts, in which they are much less expressed, and where they are equally distributed in the three categories (inside, edge and outside). On the contrary, TNFα retains its tendency to be at the edge or outside its chromosome territory in fibroblasts. This gene belongs to the MHC which has been already found most often at the periphery of its CT or on large external loops in human fibroblasts (Volpi et al. 2000) irrespective of its expression status, due probably to its position in one of the most-gene rich regions of the genome. Altogether our data suggest that relocation in the nuclear space of genes differentially expressed in response to LPS activation is gene and cell type specific, but also closely linked to the entire up-regulation status of their chromosomal regions in immune cells.
References
Akhtar A, Gasser SM (2007) The nuclear envelope and transcriptional control. Nat Rev Genet 8:507–517
Archibald AL, Bolund L, Churcher C, Fredholm M, Groenen MAM, Harlizius B, Lee K-T, Milan D, Rogers J, Rothschild MF, Uenishi H, Wang J, Schook LB, Consortium SGS (2010) Pig genome sequence-analysis and publication strategy. BMC Genomics 11:438
Bártová E, Kozubek S, Jirsová P, Kozubek M, Gajová H, Lukásová E, Skalníková M, Ganová A, Koutná I, Hausmann M (2002) Nuclear structure and gene activity in human differentiated cells. J Struct Biol 139:76–89
Boldrick JC, Alizadeh AA, Diehn M, Dudoit S, Liu CL, Belcher CE, Botstein D, Staudt LM, Brown PO, Relman DA (2002) Stereotyped and specific gene expression programs in human innate immune responses to bacteria. Proc Natl Acad Sci USA 99:972–977
Bradley JR (2008) TNF-mediated inflammatory disease. J Pathol 214:149–160
Brown CR, Silver PA (2007) Transcriptional regulation at the nuclear pore complex. Curr Opin Genet Dev 17:100–106
Brown JM, Green J, das Neves RP, Wallace HAC, Smith AJH, Hughes J, Gray N, Taylor S, Wood WG, Higgs DR, Iborra FJ, Buckle VJ (2008) Association between active genes occurs at nuclear speckles and is modulated by chromatin environment. J Cell Biol 182:1083–1097
Caron H, van Schaik B, van der Mee M, Baas F, Riggins G, van Sluis P, Hermus MC, van Asperen R, Boon K, Voûte PA, Heisterkamp S, van Kampen A, Versteeg R (2001) The human transcriptome map: clustering of highly expressed genes in chromosomal domains. Science 291:1289–1292
Casolari JM, Brown CR, Komili S, West J, Hieronymus H, Silver PA (2004) Genome-wide localization of the nuclear transport machinery couples transcriptional status and nuclear organization. Cell 117:427–439
Chambeyron S, Bickmore WA (2004) Chromatin decondensation and nuclear reorganization of the HoxB locus upon induction of transcription. Genes Dev 18:1119–1130
Chambeyron S, Silva NRD, Lawson KA, Bickmore WA (2005) Nuclear re-organisation of the Hoxb complex during mouse embryonic development. Development 132:2215–2223
Christova R, Jones T, Wu P-J, Bolzer A, Costa-Pereira AP, Watling D, Kerr IM, Sheer D (2007) P-STAT1 mediates higher-order chromatin remodelling of the human MHC in response to IFNgamma. J Cell Sci 120:3262–3270
Cremer T, Cremer C (2001) Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat Rev Genet 2:292–301
Cremer T, Cremer M, Dietzel S, Müller S, Solovei I, Fakan S (2006) Chromosome territories—a functional nuclear landscape. Curr Opin Cell Biol 18:307–316
Detweiler CS, Cunanan DB, Falkow S (2001) Host microarray analysis reveals a role for the Salmonella response regulator phoP in human macrophage cell death. Proc Natl Acad Sci USA 98:5850–5855
R Development Core Team (2005) R: A language and environment for statistical computing. In: R Foundation for Statistical Computing, Vienna, Austria. ISBN 3-900051-07-0. http://www.R-project.org
Devriendt B, Gallois M, Verdonck F, Wache Y, Bimczok D, Oswald IP, Goddeeris BM, Cox E (2009) The food contaminant fumonisin B(1) reduces the maturation of porcine CD11R1(+) intestinal antigen presenting cells and antigen-specific immune responses, leading to a prolonged intestinal ETEC infection. Vet Res 40:40
Dietzel S, Schiebel K, Little G, Edelmann P, Rappold GA, Eils R, Cremer C, Cremer T (1999) The 3D positioning of ANT2 and ANT3 genes within female X chromosome territories correlates with gene activity. Exp Cell Res 252:363–375
Federico C, Cantarella CD, Mare PD, Tosi S, Saccone S (2008) The radial arrangement of the human chromosome 7 in the lymphocyte cell nucleus is associated with chromosomal band gene density. Chromosoma 117:399–410
Finlan LE, Sproul D, Thomson I, Boyle S, Kerr E, Perry P, Ylstra B, Chubb JR, Bickmore WA (2008) Recruitment to the nuclear periphery can alter expression of genes in human cells. PLoS Genet 4:e1000039
Gao Y, Flori L, Lecardonnel J, Esquerré D, Hu ZL, Teillaud A, Lemonnier G, Lefèvre F, Oswald IP, Rogel-Gaillard C (2010) Transcriptome analysis of porcine PBMCs after in vitro stimulation by LPS or PMA/ionomycin using an expression array targeting the pig immune response. BMC Genomics 11:292
Ge S, Danino V, He Q, Hinton JCD, Granfors K (2010) Microarray analysis of response of Salmonella during infection of HLA-B27- transfected human macrophage-like U937 cells. BMC Genomics 11:456
Geissmann F, Manz MG, Jung S, Sieweke MH, Merad M, Ley K (2010) Development of monocytes, macrophages, and dendritic cells. Science 327:656–661
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J, Hornik K, Hothorn T, Huber W, Iacus S, Irizarry R, Leisch F, Li C, Maechler M, Rossini AJ, Sawitzki G, Smith C, Smyth G, Tierney L, Yang JYH, Zhang J (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5:R80
Gué M, Messaoudi C, Sun JS, Boudier T (2005) Smart 3D-FISH: automation of distance analysis in nuclei of interphase cells by image processing. Cytometry 67:18–26
Harnicarová A, Kozubek S, Pacherník J, Krejci J, Bártová E (2006) Distinct nuclear arrangement of active and inactive c-myc genes in control and differentiated colon carcinoma cells. Exp Cell Res 312:4019–4035
Hepperger C, Mannes A, Merz J, Peters J, Dietzel S (2008) Three-dimensional positioning of genes in mouse cell nuclei. Chromosoma 117:535–551
Hepperger C, Mayer A, Merz J, Vanderwall DK, Dietzel S (2009) Parental genomes mix in mule and human cell nuclei. Chromosoma 118:335–347
Iannuccelli E, Mompart F, Gellin J, Lahbib-Mansais Y, Yerle M, Boudier T (2010) NEMO: a tool for analyzing gene and chromosome territory distributions from 3D-FISH experiments. Bioinformatics 26:696–697
Jenner RG, Young RA (2005) Insights into host responses against pathogens from transcriptional profiling. Nat Rev Microbiol 3:281–294
Kawano M, Thet MM, Makino T, Kushida T, Sakagami H (2010) DNA microarray analysis of signaling pathway in macrophages stimulated by lignin-carbohydrate complex from lentinus edodes mycelia (LEM) extract. Anticancer Res 30:2567–2576
Kocanova S, Kerr EA, Rafique S, Boyle S, Katz E, Caze-Subra S, Bickmore WA, Bystricky K (2010) Activation of estrogen-responsive genes does not require their nuclear co-localization. PLoS Genet 6:e1000922
Kosak ST, Skok JA, Medina KL, Riblet R, Beau MML, Fisher AG, Singh H (2002) Subnuclear compartmentalization of immunoglobulin loci during lymphocyte development. Science 296:158–162
Koutná I, Krontorád P, Svoboda Z, Bártová E, Kozubek M, Kozubek S (2007) New insights into gene positional clustering and its properties supported by large-scale analysis of various differentiation pathways. Genomics 89:81–88
Kumar PP, Bischof O, Purbey PK, Notani D, Urlaub H, Dejean A, Galande S (2007) Functional interaction between PML and SATB1 regulates chromatin-loop architecture and transcription of the MHC class I locus. Nat Cell Biol 9:45–56
Kumaran RI, Spector DL (2008) A genetic locus targeted to the nuclear periphery in living cells maintains its transcriptional competence. J Cell Biol 180:51–65
Küpper K, Kölbl A, Biener D, Dittrich S, von Hase J, Thormeyer T, Fiegler H, Carter NP, Speicher MR, Cremer T, Cremer M (2007) Radial chromatin positioning is shaped by local gene density, not by gene expression. Chromosoma 116:285–306
Kurz A, Lampel S, Nickolenko JE, Bradl J, Benner A, Zirbel RM, Cremer T, Lichter P (1996) Active and inactive genes localize preferentially in the periphery of chromosome territories. J Cell Biol 135:1195–1205
Lanctôt C, Cheutin T, Cremer M, Cavalli G, Cremer T (2007) Dynamic genome architecture in the nuclear space: regulation of gene expression in three dimensions. Nat Rev Genet 8:104–115
Litten-Brown JC, Corson AM, Clarke L (2010) Porcine models for the metabolic syndrome, digestive and bone disorders: a general overview. Animal 4:899–920
Lunney JK, Ho CS, Wysocki M, Smith DM (2009) Molecular genetics of the swine major histocompatibility complex, the SLA complex. Dev Comp Immunol 33:362–374
Ma J, Chen T, Mandelin J, Ceponis A, Miller NE, Hukkanen M, Ma GF, Konttinen YT (2003) Regulation of macrophage activation. Cell Mol Life Sci 60:2334–2346
Mahy NL, Perry PE, Gilchrist S, Baldock RA, Bickmore WA (2002) Spatial organization of active and inactive genes and noncoding DNA within chromosome territories. J Cell Biol 157:579–589
Malcolm KC, Arndt PG, Manos EJ, Jones DA, Worthen GS (2003) Microarray analysis of lipopolysaccharide-treated human neutrophils. Am J Physiol Lung Cell Mol Physiol 284:L663–L670
Mayer R, Brero A, von Hase J, Schroeder T, Cremer T, Dietzel S (2005) Common themes and cell type specific variations of higher order chromatin arrangements in the mouse. BMC Cell Biol 6:44
Meaburn KJ, Misteli T (2008) Locus-specific and activity-independent gene repositioning during early tumorigenesis. J Cell Biol 180:39–50
Misteli T (2007) Beyond the sequence: cellular organization of genome function. Cell 128:787–800
Morey C, Kress C, Bickmore WA (2009) Lack of bystander activation shows that localization exterior to chromosome territories is not sufficient to up-regulate gene expression. Genome Res 19:1184–1194
Nathan C (2006) Neutrophils and immunity: challenges and opportunities. Nat Rev Immunol 6:173–182
Nau GJ, Richmond JFL, Schlesinger A, Jennings EG, Lander ES, Young RA (2002) Human macrophage activation programs induced by bacterial pathogens. Proc Natl Acad Sci USA 99:1503–1508
Osborne CS, Chakalova L, Brown KE, Carter D, Horton A, Debrand E, Goyenechea B, Mitchell JA, Lopes S, Reik W, Fraser P (2004) Active genes dynamically colocalize to shared sites of ongoing transcription. Nat Genet 36:1065–1071
Ottaviani D, Lever E, Mitter R, Jones T, Forshew T, Christova R, Tomazou EM, Rakyan VK, Krawetz SA, Platts AE, Segarane B, Beck S, Sheer D (2008) Reconfiguration of genomic anchors upon transcriptional activation of the human major histocompatibility complex. Genome Res 18:1778–1786
Pai DA, Engelke DR (2010) Spatial organization of genes as a component of regulated expression. Chromosoma 119:13–25
Parada L, Misteli T (2002) Chromosome positioning in the interphase nucleus. Trends Cell Biol 12:425–432
Ragoczy T, Bender MA, Telling A, Byron R, Groudine M (2006) The locus control region is required for association of the murine beta-globin locus with engaged transcription factories during erythroid maturation. Genes Dev 20:1447–1457
Reddy KL, Zullo JM, Bertolino E, Singh H (2008) Transcriptional repression mediated by repositioning of genes to the nuclear lamina. Nature 452:243–247
Reiner A, Yekutieli D, Benjamini Y (2003) Identifying differentially expressed genes using false discovery rate controlling procedures. Bioinformatics 19:368–375
Sadoni N, Targosz B-S, Englmann A, Fesser S, Koch J, Schindelhauer D, Zink D (2008) Transcription-dependent spatial arrangements of CFTR and conserved adjacent loci are not conserved in human and murine nuclei. Chromosoma 117:381–397
Scheuermann MO, Tajbakhsh J, Kurz A, Saracoglu K, Eils R, Lichter P (2004) Topology of genes and nontranscribed sequences in human interphase nuclei. Exp Cell Res 301:266–279
Sexton T, Schober H, Fraser P, Gasser SM (2007) Gene regulation through nuclear organization. Nat Struct Mol Biol 14:1049–1055
Sklar MD, Tereba A, Chen BD, Walker WS (1985) Transformation of mouse bone marrow cells by transfection with a human oncogene related to c-myc is associated with the endogenous production of macrophage colony stimulating factor 1. J Cell Physiol 125:403–412
Smith JD, Peng DQ, Dansky HM, Settle M, Baglione J, Goff WL, Chakrabarti E, Xu Y, Peng X (2006) Transcriptome profile of macrophages from atherosclerosis-sensitive and atherosclerosis-resistant mice. Mamm Genome 17:220–229
Smyth GK (2004) Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3: Article3
Soutoglou E, Misteli T (2007) Mobility and immobility of chromatin in transcription and genome stability. Curr Opin Genet Dev 17:435–442
Stadler S, Schnapp V, Mayer R, Stein S, Cremer C, Bonifer C, Cremer T, Dietzel S (2004) The architecture of chicken chromosome territories changes during differentiation. BMC Cell Biol 5:44
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550
Szczerbal I, Bridger JM (2010) Association of adipogenic genes with SC-35 domains during porcine adipogenis. Chromosome Res 18:887–895
Szczerbal I, Foster HA, Bridger JM (2009) The spatial repositioning of adipogenesis genes is correlated with their expression status in a porcine mesenchymal stem cell adipogenesis model system. Chromosoma 118:647–663
Tanabe H, Müller S, Neusser M, von Hase J, Calcagno E, Cremer M, Solovei I, Cremer C, Cremer T (2002) Evolutionary conservation of chromosome territory arrangements in cell nuclei from higher primates. Proc Natl Acad Sci USA 99:4424–4429
Tang Y, Xu H, Du X, Lit L, Walker W, Lu A, Ran R, Gregg JP, Reilly M, Pancioli A, Khoury JC, Sauerbeck LR, Carrozzella JA, Spilker J, Clark J, Wagner KR, Jauch EC, Chang DJ, Verro P, Broderick JP, Sharp FR (2006) Gene expression in blood changes rapidly in neutrophils and monocytes after ischemic stroke in humans: a microarray study. J Cereb Blood Flow Metab 26:1089–1102
Volpi EV, Chevret E, Jones T, Vatcheva R, Williamson J, Beck S, Campbell RD, Goldsworthy M, Powis SH, Ragoussis J, Trowsdale J, Sheer D (2000) Large-scale chromatin organization of the major histocompatibility complex and other regions of human chromosome 6 and its response to interferon in interphase nuclei. J Cell Sci 113:1565–1576
Wells CA, Ravasi T, Faulkner GJ, Carninci P, Okazaki Y, Hayashizaki Y, Sweet M, Wainwright BJ, Hume DA (2003) Genetic control of the innate immune response. BMC Immunol 4:5
Williams RRE, Broad S, Sheer D, Ragoussis J (2002) Subchromosomal positioning of the epidermal differentiation complex (EDC) in keratinocyte and lymphoblast interphase nuclei. Exp Cell Res 272:163–175
Xiao S, Jia J, Mo D, Wang Q, Qin L, He Z, Zhao X, Huang Y, Li A, Yu J, Niu Y, Liu X, Chen Y (2010) Understanding PRRSV infection in porcine lung based on genome-wide transcriptome response identified by deep sequencing. PLoS One 5:e11377
Yerle M, Schmitz A, Milan D, Chaput B, Monteagudo L, Vaiman M, Frelat G, Gellin J (1993) Accurate characterization of porcine bivariate flow karyotype by PCR and fluorescence in situ hybridization. Genomics 16:97–103
Yerle M, Goureau A, Gellin J, Tissier PL, Moran C (1994) Rapid mapping of cosmid clones on pig chromosomes by fluorescence in situ hybridization. Mamm Genome 5:34–37
Yerle-Bouissou M, Mompart F, Iannuccelli E, Robelin D, Jauneau A, Lahbib-Mansais Y, Delcros C, Oswald IP, Gellin J (2009) Nuclear architecture of resting and LPS-stimulated porcine neutrophils by 3D FISH. Chromosome Res 17:847–862
Zhang F, Hopwood P, Abrams CC, Downing A, Murray F, Talbot R, Archibald A, Lowden S, Dixon LK (2006) Macrophage transcriptional responses following in vitro infection with a highly virulent African swine fever virus isolate. J Virol 80:10514–10521
Zhang S, Kim CC, Batra S, McKerrow JH, Pn L (2010) Delineation of diverse macrophage activation programs in response to intracellular parasites and cytokines. PLoS Negl Trop Dis 4:e648
Zink D, Amaral MD, Englmann A, Lang S, Clarke LA, Rudolph C, Alt F, Luther K, Braz C, Sadoni N, Rosenecker J, Schindelhauer D (2004) Transcription-dependent spatial arrangements of CFTR and adjacent genes in human cell nuclei. J Cell Biol 166:815–825
Acknowledgements
We would like to thank Agnès Bonnet for the technical advice on neutrophil RNA analyses, for the helpful discussions on transcriptome analysis and for critically reading the manuscript. The authors would like to thank the CRB-GADIE (INRA, UMR GABI, France) for providing the BAC clones. We are grateful to A. Jauneau and C. Pouzet for their help and the use of the IFR40 platform for confocal microscopy facility. This work was supported by a grant to Romain Solinhac from the French Ministère de l'Education Nationale, de la Recherche et de la Technologie and by a complementary grant from INRA (Animal Genetics and Animal Health Departments).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by G. Matera
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
(JPEG 95 kb)
Supplementary Fig. 2
(JPEG 68 kb)
Supplementary Fig. 3
(JPEG 124 kb)
Supplementary Fig. 4
(JPEG 68 kb)
Supplementary Fig. 5
(JPEG 92 kb)
Supplementary Fig. 6
(JPEG 109 kb)
Supplementary Table 1
(DOC 63 kb)
Supplementary Table 2
(DOC 154 kb)
Supplementary Table 3
(DOC 109 kb)
Supplementary Table 4
(DOC 30 kb)
Rights and permissions
About this article
Cite this article
Solinhac, R., Mompart, F., Martin, P. et al. Transcriptomic and nuclear architecture of immune cells after LPS activation. Chromosoma 120, 501–520 (2011). https://doi.org/10.1007/s00412-011-0328-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-011-0328-7