Abstract
In the nuclei of human lymphocytes, chromosome territories are distributed according to the average gene density of each chromosome. However, chromosomes are very heterogeneous in size and base composition, and can contain both very gene-dense and very gene-poor regions. Thus, a precise analysis of chromosome organisation in the nuclei should consider also the distribution of DNA belonging to the chromosomal bands in each chromosome. To improve our understanding of the chromatin organisation, we localised chromosome 7 DNA regions, endowed with different gene densities, in the nuclei of human lymphocytes. Our results showed that this chromosome in cell nuclei is arranged radially with the gene-dense/GC-richest regions exposed towards the nuclear interior and the gene-poorest/GC-poorest ones located at the nuclear periphery. Moreover, we found that chromatin fibres from the 7p22.3 and the 7q22.1 bands are not confined to the territory of the bulk of this chromosome, protruding towards the inner part of the nucleus. Overall, our work demonstrates the radial arrangement of the territory of chromosome 7 in the lymphocyte nucleus and confirms that human genes occupy specific radial positions, presumably to enhance intra- and inter-chromosomal interaction among loci displaying a similar expression pattern, and/or similar replication timing.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chromosomal DNA in the cell nucleus is distributed according to its nucleotide composition (guanine + cytosine content or GC level), the GC-poorest DNA being located more peripherally with respect to the GC-richest DNA (Sadoni et al. 1999; Cremer and Cremer 2001; Saccone et al. 2002). This type of DNA distribution defines two nuclear compartments having different GC levels as well as different structural and functional properties (see Bernardi 2004 for a review). In fact, the chromosomal DNA showing the highest GC level is endowed with the highest gene densities, early replication during the S phase of the cell cycle and a more open chromatin structure, in contrast to the GC-poorest DNA that has the opposite features (Ferreira et al. 1997; Sadoni et al. 1999; Saccone et al., 2002; Tanabe et al. 2002a; Boutanaev et al. 2005; Foster and Bridger 2005; Petrova et al. 2006). The compartmentalisation of the cell nucleus is also associated with gene expression activity, with transcripts and splicing small nuclear ribonucleoproteins preferentially located at the nuclear interior, whereas the nuclear periphery, largely occupied by heterochromatin, is generally transcriptionally inactive (Strouboulis and Wolffe 1996; Brown et al. 1997; Andrulis et al. 1998; Cockell and Gasser 1999; Lukasova et al. 2002; Foster and Bridger 2005). A striking example, in this regard, concerns the inactive X chromosome in mammalian female cells, which is typically located towards the nuclear envelope forming the very compact structure of the Barr body but with a large number of genes located away from the bulk of the chromosome territory (Clemson et al. 2006). Similarly, studies of the immunoglobulin heavy-chain locus in B-lineage cells showed a clear correlation for nuclear location, replication timing and transcriptional activity (Zhou et al. 2002).
A number of studies demonstrated that, in spherically shaped nuclei (such as those of lymphocytes), the radial positioning of human chromosome territories is generally in accordance with their gene density (Cremer and Cremer 2001; Boyle et al. 2001). This type of nuclear chromosome distribution was first shown for the human chromosomes 18 and 19, which are located in different radial positions, namely at the periphery and in the interior of the nucleus, respectively (Croft et al. 1999). Indeed, in spite of their similar size, these two chromosomes are endowed with very different gene densities, the former being very gene-poor and the latter very gene-rich (Saccone et al. 1992; Federico et al. 2000; IHGSC 2004). These data are confirmed also in other primates where the nuclear location of the syntenic chromosomes shows a similar distribution in the spherical nuclei of lymphoblastoid cells, indicating that there is a highly conserved evolutionary trend in this specific cell type (Tanabe et al. 2002b; Neusser et al. 2007). In the case of cell types with flat-ellipsoidal nuclei (such as fibroblasts), the radial chromosome territory positioning is less clear. Data showing arrangements correlating with gene density and chromosome size have been reported (Bridger et al. 2000; Bolzer et al. 2005).
Despite the validity of such results, it should be stressed that the typical or average nuclear location of entire chromosomes is an approximation of the real situation because the human chromosomes, especially the large ones, are not homogeneous in their base composition or their gene density. In fact, the majority of human chromosomes contain two compositionally very different sets of bands, the GC-richest and the GC-poorest bands, called H3+ and L1+ bands, respectively, on the basis of the isochores they contain. The remaining chromosomal bands (the H3− and the L1− bands) are largely composed of DNA from the other isochore families (i.e. L2, H1 and H2; Federico et al. 2000; see Bernardi 2004 for a review). In general, the H3+ and L1+ chromosomal bands are not contiguous along the metaphase chromosome and are typically separated by the compositionally intermediate H3− and L1− bands (Saccone et al. 2001; IHGSC 2004). This type of chromosome organisation is highly conserved evolutionarily and has been described not only in mammalian and avian species (Saccone et al. 1997; Andreozzi et al. 2001; Federico et al. 2004; 2005) but also in reptiles and amphibians (Federico et al. 2006). The non-contiguity in a chromosome of the GC-richest H3+ and the GC-poorest L1+ bands emphasises the very different location in the nuclei of these two types of band. Different locations have been demonstrated, for example, for the two telomeric regions of the human chromosome 9, which are compositionally different. The GC-rich/gene-rich telomeric end of the long arm is located in the nuclear interior with an open chromatin conformation, whereas the GC-poor/gene-poor telomeric end of the short arm is located at the periphery of the cell nuclei with a very compact conformation (Saccone et al. 2002).
To improve our understanding of the chromatin organisation in the nucleus, we focused our attention on human chromosome 7, a compositionally very heterogeneous chromosome containing approximately 159 Mb of DNA (Scherer et al. 2003) and consisting, at the chromosomal resolution of 850 bands per haploid genome, of 42 euchromatic bands (Francke 1994). Although the chromosome 7 territory was previously shown to be one of the most peripheral in the human cell nucleus (Boyle et al. 2001), it contains gene-dense/GC-rich regions that we expected to be located in the nuclear interior (Saccone et al. 2002). In the present study, we used DNA probes specific for different regions along the human chromosome 7 to define the radial nuclear locations of the compositionally different types of band. The radial location of each locus was then correlated with gene density and GC level of the 2-Mb DNA region surrounding each probe, and with the chromosomal band containing the probe. Finally, we also investigated the relation between the nuclear location of each locus and the replication timing of its chromosomal band.
Materials and methods
Definition of chromosomal band sizes and their GC level/gene density
According to cytogenetic methods of chromosome staining, different band patterns can be obtained. Following the compositional method previously described (Saccone et al. 2001; Costantini et al. 2007), we defined as H3+ bands the R bands (or G-pale bands) with the highest GC level/gene density and L1+ bands the G bands (G-dark bands) with the lowest GC level/gene density. Among the R and G bands, we also included H3− and L1− bands, which are characterised by an intermediate GC level/gene density. To define localisation and size of chromosome 7 bands, we referred to the DNA sequence as reported in the Human Genome Browser (HGB) Gateway (http://genome.ucsc.edu/ accessed on 24 August 2007).
Preparation of chromosomes and nuclei
Metaphase chromosomes and nuclei were prepared from phytohaemagglutinin (PHA)-stimulated peripheral blood lymphocytes according to standard cytogenetic procedures and fixed with methanol/acetic acid, as described previously (Saccone et al. 2002; D’Antoni et al. 2004). A neuroblastoma cell line (SK–N–BE cells) and two lymphoblastoid cell lines were also used as controls. These cell types were prepared employing standard cytogenetic procedures used for the lymphocytes. Moreover, the SK–N–BE cells were also prepared under conditions to preserve the 3D large-scale chromatin structure in the nuclei (Solovei et al. 2002). This latter procedure involves the use of freshly prepared 4% buffered paraformaldehyde to fix the cells.
DNA probes, in situ hybridisation, and detection
Probes used for the fluorescence in situ hybridisations were mainly BAC clones. These were kindly provided by M. Rocchi, University of Bari, Italy (probes 1–11, 15–17), and by S. Scherer, Hospital for Sick Children, Toronto, Canada (probes 12–14, 18, 19; for a complete list of probes refer to Table 1). Probes were chosen on the basis of their location along the chromosome to obtain a representative spectrum for all chromosome 7 bands.
DNA probes were extracted using a commercial kit (Qiagen, Milan, Italy), biotin-labelled by nick translation (Roche, Mannheim, Germany) and hybridised as previously described (Saccone et al. 2002; D’Antoni et al. 2004). Chromosome painting was performed using a commercial probe for chromosome 7 (Appligene Oncor, UK), directly labelled with fluorescein. Hybridisation detection of biotin-labelled probes was performed using fluorescein- or rhodamine-conjugated avidin. Images were captured using a Photometrics cooled charge-coupled device camera and MacProbe v4.2 software (Applied Imaging, Newcastle, UK).
DNA probes were first tested on metaphase chromosomes to verify (1) the location on the correct chromosomal band and (2) the absence of additional signals on other sites. Cell nuclei were then analysed for the evaluation of radial location of each probe.
Localisation of fluorescent in situ hybridization signals within the cell nuclei and statistical analysis
We determined the radial nuclear location of each locus using a two-dimensional (2D) analysis, following previously described criteria (Croft et al. 1999; Bridger et al. 2000; Roix et al. 2003; D’Antoni et al. 2004; Masny et al. 2004; Ono et al. 2007). More precisely, a high number of hybridised nuclei were captured with MacProbe v4.2 software, and the hybridisation signals were localised in each cell nucleus using a dedicated computer software developed in our lab at the University of Catania (freely available on request). To avoid possible misassignments of the hybridisation signals as due to DNA replication, only nuclei with the unreplicated loci were analysed. That is, only nuclei showing the hybridisation signals as singlets were used for our nuclear location studies (Fig. 1).
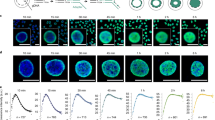
Scheme of the radial nuclear location using 2D analysis. a Three representative nuclei hybridised with BAC probes, showing the signals (indicated by arrows) from the two alleles. Each allele could be unreplicated or replicated, as determined by the presence of singlet (white arrows) or doublet (red arrows) signals. The position of each signal relative to the nuclear centre was defined as a ratio of the nuclear radius, and a high number of hybridised nuclei were analysed for each probe. Only nuclei with unreplicated DNA (singlet–singlet signals) were used to define the radial location. Hybridisations of the biotin-labelled probes were detected by fluorescein-conjugated avidin (green signals), and the nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI). Scale bar, 2 μm. b Computational visualisation of all the signals localised for the indicated probes. CF14 and RP11-213E22 probes (probes 14 and 11) are representative of probes located in the peripheral and internal part of the nucleus, respectively. RP11-673O11 (probe 13) is representative of probes with an intermediate position between these two extremes. c Histograms showing the distribution of the hybridisation signals presented in b. d Percentile graph of the data obtained for the three representative loci. e Median values (and the relative confidence interval, CI) of the data for each of the above three probes
To assign a locus to a defined radial nuclear position, we statistically analysed the hybridisation results (see Fig. 1). A precise position was first assigned to each hybridisation signal as a ratio of the nuclear radius (0 and 1 indicate the centre and the peripheral rim of the nucleus, respectively). For each probe, we evaluated the median value ± confidence interval (CI) of at least 300 hybridised nuclei. The results were also visualised as a radial graph and as a frequency distribution histogram. Percentile analysis was also performed. The statistical analysis and the corresponding graphs were carried out using Microsoft Excel and StatView software. Data from a larger number of probes located throughout the human genome, in addition to those presented here, allowed us to establish that median values lower than 0.65 indicate loci located more internally in the lymphocyte nuclei.
Assessment of the replication timing
The replication timing of chromosome 7 bands was previously assessed after using a BrdU treatment (Dutrillaux et al. 1976). In this study, we confirmed the replication timing of some representative loci using a flourescent in situ hybridization-based method as previously described (Boogs and Chinoult 1997; D’Antoni et al. 2004). Briefly, for each probe, at least 300 nuclei were chosen randomly. In these, we verified the presence of singlet (DNA not yet replicated) or closely spaced doublet signals (replicated DNA) on each allele (see Fig. 1). With this method, a high ratio of singlet/singlet (SS) nuclei with respect to the doublet/doublet (DD) nuclei indicates a late replicated locus. On the other hand, a high ratio of DD nuclei with respect to the SS nuclei indicates an early replicated locus (Boogs and Chinoult 1997).
Results
Definition of R and G bands and their GC level/gene density
On the basis of GC level, we localised the transition points (borders) from G to R bands. This enabled us to define the GC-richest/gene-richest H3+ and the GC-poorest/gene-poorest L1+ bands in human chromosome 7, which cytogenetically correspond to R and G bands, respectively. A detailed map of chromosome 7 bands, based on the isochores they contain, is given in Fig. 2. We defined a total of 42 bands, of which six were H3+ (R bands) and 11 were L1+ (G bands). The remaining bands had intermediate GC level and gene density, consisting largely of DNA from the L2 and H1 isochore families. The H3+ bands were p22.3, p22.1, p13, q11.23, q22.1 and q36.3, and the L1+ bands were p21.3, p21.1, p12.3, p12.1, q21.11, q21.13, q21.3, q31.1, q31.31, q31.33 and q35.
Chromosome 7 features and location of probes. From bottom to top: (1) ideogram of the chromosome 7, indicating the H3+ (red) and the L1+ (blue) bands (modified from Federico et al. 2000). (2) GC level profile of the chromosome 7 sequence and correspondence with the cytogenetic bands. The five isochore families are indicated by different colours: blue, pale blue, yellow, orange and red correspond to the L1, L2, H1, H2 and H3 isochore families, respectively. The correlation between the chromosomal bands and the DNA sequence was performed according to previously defined criteria (Saccone et al. 2001; Costantini et al. 2007), and the position of each band border (Mb from the telomeric end of the short arm) is specified. (3) Gene density along the chromosome 7 on 1 Mb non-overlapping windows starting from the telomeric end of the short arm. The arrows and relative numbers indicate the position of the corresponding probes (see reference in Table 1). (4) Representative images of chromosome 7 hybridised with a probe from the band indicated. Hybridisations of the biotin-labelled probes were detected by fluorescein-conjugated avidin (green signals), and the chromosomes were stained with DAPI (reproduced in grey scale)
The transitions from G to R bands (see Fig. 2) as defined via the compositional method coincided largely with those obtained using other methods (Furey and Haussler 2003), with only few exceptions. For example, we positioned band 7q22.1 (an H3+ band) on the DNA segment between 97.6 and 102.1 Mb, compared to the 97.9 and 104.4 Mb indicated in the HGB Gateway. In fact, the segment between 97.6 and 102.1 Mb is largely composed of H3 and H2 isochores, and surrounded by very long stretches of L1 and L2 isochores. Therefore, in this case, the transition between the R band 7q22.1 (an H3+ band) and the G-band 7q22.2 (an L1− band) did not coincide with the position indicated by the HGB, with approximately 2.3 Mb difference (Fig. 2).
In accordance with the general features of the human genome, the gene density in chromosome 7 showed a high level of correlation with GC (R = 0.75). This means that the highest gene densities are in the H3+ bands, and the “gene deserts” are in the GC-poorest L1+ bands. The highest peak of gene density observed on chromosome 7 was located in 7q22.1, an H3+ band. Moreover, we evaluated the GC level and gene density of each probe-containing locus used in this study (excluding the centromeric probe pZ7.5, 7 in Table 1). More precisely, we defined GC level and gene density (1) of the 2-Mb DNA around each DNA probe and (2) of the chromosomal band containing the DNA probe. Data are shown in Table 1.
Nuclear localisation of chromosome 7 loci
All probes listed in Table 1 were used to determine the nuclear localisation of the corresponding chromosome 7 loci in human lymphocytes. The data obtained are summarised in Fig. 3. Sixteen out of 19 loci showed a median value higher than 0.65, whereas probes located in the H3+ bands 7p22.3 (probe 1, RP11-6A1) and 7q22.1 (probes 11 and 12, RP11-231E22, and 8b.1, respectively) were localised more internally in the cell nuclei, with a median value lower than 0.60.
Radial nuclear location of chromosome 7 loci. The radial location of each probe in the nucleus was statistically defined by the median value of the nuclear locations determined for all the hybridisation signals identified on more than 300 nuclei. On the x-axis, the numbers indicate the probes listed in Table 1. On the y-axis, the median values (and relative CI) of all the probes are shown. The different colour dots indicate loci belonging to the H3+ bands (red dots), to the L1+ bands (blue dots) and to the compositionally intermediate bands (grey dots). The black dot indicates the centromeric region. Tel-p and tel-q indicate the telomeric end of the short and long arm, respectively. The upper histogram indicates the gene density of the 2-Mb region around each probe
Figure 3 shows that (1) the more peripheral loci are generally in the L1+ bands, with median values higher than 0.70; (2) the centromeric region is located peripherally with median value of about 0.70; (3) the majority of loci from H3+ bands are more internal in the nucleus, with three loci below 0.60 median value and only one of the two 7q36.3 band probes with a median value very close to 0.70; (4) probes 1, 11 and 12 (RP11-6A1, RP11-213E22 and 8b.1 respectively), belonging to H3+ bands are localised more in the nuclear interior than all the others; and (5) with exclusion of the above three loci, the largest internal-peripheral distance between points within the chromosome 7 territory is less than 0.10, as defined by the difference between the median value of probes 14 (CF14, median value of 0.748) and 16 (RP11-148k1, median value of 0.653). Moreover, the relative radial distance between probes 1 (RP11-6A1, median value of 0.599) and 16 (RP11-148k1, median value of 0.653) was approximately 0.05. This radial distance increases when one considers the more internal probe 11 (RP11-213E22, median value of 0.552) being 0.10 in distance from probe 16 (RP11-148k1, median value of 0.653). These measurements suggested that the DNA belonging to probes 1, 11 and 12 (RP11-6A1, RP11-213E22 and 8b.1, respectively) could extend away from the bulk of chromosome 7, perhaps forming loops. This was shown by images obtained by dual colour in situ hybridisation (Fig. 4).
Distribution of chromosome 7 in the nucleus. a–e Dual colour in situ hybridisations of human chromosome 7 DNA (chromosome painting) and some representative probes: probe 1 (RP11-6A1) (a), probe 11 (RP11-213E22) (b), probe 3 (RP11-79O21) (c and d) and probes 11 + 12 (RP11-213E22 + 8b1, respectively) (e). Hybridisation of the biotin-labelled probes was detected with rhodamine-conjugated avidin (red signals), chromosome painting DNAs were directly labelled with fluorescein (green signals) and the nuclei were stained with DAPI (blue). Scale bar, 2 μm. f Cartoon showing a nucleus with the representation of the chromosome 7 territories and a possible zig-zag conformation of the DNA according to the data presented in Fig. 3 and according to the above dual-colour in situ hybridisation results. Tel-p, tel-q and cen indicate the telomeric end of the short arm, the telomeric end of the long arm and the centromere, respectively. The three differently stained compartments (pale blue, yellow and pink) roughly indicate the more peripheral, the intermediate and the more internal part of the nucleus. g Enlargement of the bottom-right sector of the cartoon shown in f with the indication of the more probable position of the loci analysed in the present study (the numbers indicate the loci listed in Table 1, and data are that presented in Fig. 3). The different colours of the dots indicate the loci belonging to the H3+ bands (red dots), to the L1+ bands (blue dots) and to the compositionally intermediate bands (grey dots). The black dot (probe 7, pZ7.5) indicates the centromere region. Loci within about 1–2 Mb of each other were connected by a continuous line, whereas larger distances were drawn by a dotted line, indicating that chromosomal DNA can make additional loops between two very distant loci
Overall, the above data indicate an organisation of chromosome 7 DNA in the nucleus that is largely but not entirely confined to the nuclear periphery, with the territories polarised with the GC-poorest/gene-poorest regions located at the nuclear periphery and the GC-richest/gene-richest ones towards the nuclear interior. In this latter case, two regions extend some distance from the surface of the chromosome territory, protruding to the innner part of the nucleus.
Radial nuclear location on different cell types
One aim of our study was to verify whether the radial position of compositionally different probes was consistent in different cell types. For this reason, we chose five out of the 19 probes used on lymphocytes. These were RP11-6A1, RP11-79O21, RP11-213E22, CF14 and 227e6 (probes 1, 3, 11, 14 and 19, respectively). These probes were hybridised onto 2D nuclei derived from (1) peripheral blood lymphocytes from three different individuals and (2) lymphoblastoid cell lines prepared from two different subjects. The probes RP11-6A1, RP11-79O21 and RP11-213E22 (probes 1, 3 and 11, respectively) were also hybridised in a further cell type, namely on a neuroblastoma cell line (SK–N–BE). All cell types were prepared using the standard cytogenetic techniques. Furthermore, probes RP11-6A1 and RP11-79O21 (probes 1 and 3, respectively) were hybridised using nuclei from the neuroblastoma cell line SK–N–BE that were obtained using a procedure previously shown to preserve large-scale chromatin/nuclear structure (see “Materials and methods”).
It is known that the standard cytogenetic techniques used in the present work destroys the 3D structure of the nucleus. However, information on the spatial arrangement of chromosomal regions in the nucleus can be obtained reliably using this fixing method if a sufficiently large number of nuclei are evaluated. To check whether the above procedure and the following hybridisation, impacted on cellular size and morphology, the size of the nuclei from all cell types was measured before and after the hybridisation procedure. This size was approximately 5–9 μm for the lymphocyte nuclei and was not altered significantly by the hybridisation procedure (data not shown).
Our results indicated that a specific locus shows a similar radial location in the different cell lines (Fig. 5). Only in one of the two lymphoblastoid cell lines were the RP11-6A1 and the CF14 probes (probes 1 and 14) located more peripherally and more internally, respectively, compared to what was observed in the other cell lines. More precisely, the RP11-6A1 and RP11-213E22 probes (probes 1 and 11, respectively) showed an internal location in the nuclei, with median values almost always lower than 0.65 in all cell types, whereas the RP11-79O21, CF14 and 227e6 probes (probes 3, 14 and 19, respectively) showed a peripheral location, as indicated by median values always higher than 0.65 in all the cell types. In these latter cases, the loci detected by the RP11-79O21 and CF14 probes (probes 3 and 14) are located peripherally with relative median values always higher than 0.70 and, in some cases, reaching around 0.80, while the locus detected by the 227e6 probe (probe 19) is located close to a median value of 0.65, i.e. close to the internal region of the nucleus. It should be noted that the locus corresponding to this last probe, located in the telomeric 7q36.3 band, is GC-rich but unexpectedly not gene-dense (see Table 1).
Radial nuclear location on different cell types. Five loci from human chromosome 7, identified by the probes RP11-6A1 (probe 1), RP11-79O21 (probe 3), RP11-213E22 (probe 11), CF14 (probe 14) and 227e6 (probe 19; see Table 1 for their features), were tested on different cell types, as indicated on the x-axis by capital letters. A, B and C are peripheral lymphocytes from three different individuals, D and E are two different lymphoblastoid cell lines, F and G are the neuroblastoma SK–N–BE cell line prepared using the standard method of preparation and the method to preserve large-scale chromatin structure (Solovei et al. 2002), respectively. The radial location of each probe in the different cell types was determined by considering the median value (and the relative confidence interval, CI) of the hybridisation signal distribution. The different colour dots indicate the median value observed for each cell type
Association between nuclear location and GC level/gene density
To establish how nuclear location correlates with GC level/gene density, we performed two different analyses. We first considered the 2-Mb region surrounding each probe, and a second analysis was carried out considering the entire chromosomal band containing the probe. The latter analysis allowed us to study a genomic region of relatively homogeneous GC level/gene density, regardless of its size, whereas the former focused on a DNA region of fixed length, regardless of its compositional homogeneity or heterogeneity. Our data indicated a good correlation between nuclear location and both GC level and gene density, with correlation coefficients ranging from 0.73 to 0.89 (Fig. 6). The highest levels of correlations were observed between the nuclear location of the probes and gene density, with the highest R value observed for the correlation between the nuclear location of the loci and the gene density of the entire chromosomal bands.
Correlations among nuclear location and GC level, gene density and replication timing. The radial nuclear location of each locus is defined by the median value shown in Fig. 3. In these correlations, the centromeric locus was not considered. Left Correlation between the nuclear location of each probe and the GC level of the 2 Mb DNA segment surrounding the probe (2 Mb, R 1) and the GC level of the chromosomal band containing the probe (band, R2). Center Correlation between the nuclear location of each probe and the gene density of the 2-Mb DNA region surrounding the probe (2 Mb, R 3) and the gene density of the chromosomal band containing the probe (band, R 4). Right Correlation between the nuclear location of each probe and the replication timing of the relative band (R 5). The replication timing of each chromosomal band was defined by a value ranging from 1 to 18, representing the 18 classes of replication (1 and 18 comprising the chromosomal bands replicated at the onset and at the end of the S phase, respectively) identified by Dutrillaux et al. (1976). R 1, R 2, R 3, R 4 and R 5 are the correlation coefficients of each of the data sets indicated
Association between nuclear location and replication timing
To establish a relation between nuclear localisation of the investigated loci and the replication timing of the corresponding chromosomal bands, we considered the 18 replication classes previously described (Dutrillaux et al. 1976). According to this classification, the H3+ and the L1+ bands replicate in the very early and very late replication classes, respectively (Federico et al. 1998; 2000). In this study, we assessed the replication timing of five chromosomal regions, representing different types of bands. More precisely, we experimentally verified the replication timing of the chromosomal bands 7p22.3, 7p21.3, 7q22.1, 7q31.31 and 7q36.3 using the probes RP11-6A1, RP11-79O21, RP11-213E22, CF14 and 227e6 (probes 1, 3, 11, 14 and 19; for their features, see Table 1 and Fig. 2).
Replication timing of each locus was defined using the previously described cytogenetic method (Boogs and Chinoult 1997; D’Antoni et al. 2004). This involves hybridisation of a probe on the cell nuclei and subsequent identification of the relative proportion of nuclei with singlet (unreplicated locus) or doublet (replicated locus) hybridisation signals. Our results indicated very early and very late replication timing for the H3+ bands 7p22.3 and 7q22.1, and for the L1+ bands 7p21.3 and 7q31.31, respectively. However, the H3+ band 7q36.3 replicates later with respect to the other two H3+ bands (Fig. 7). Incidentally, band 7q36.3 is very GC-rich but gene-poor. Thus, the replication timing of the above five loci was, as expected, in agreement with the previous findings (Dutrillaux et al. 1976).
Assessment of replication timing. SS (yellow) indicates nuclei where both alleles observed were unreplicated (i.e. two singlet signals were seen), SD (pink) indicates nuclei where one allele was in replicated status and the other one unreplicated (i.e. one singlet and one doublet signals were seen), DD (pale blue) indicates nuclei where both alleles observed were in replicated status (i.e. two doublet signals were seen). On the x-axis, the probe number is indicated in reference to five different loci, RP11-6A1 (probe 1), RP11-79O21 (probe 3), RP11-213E22 (probe 11), CF14 (probe 14) and 227e6 (probe 19; for details see Table 1); on the y-axis, the percentages of the nuclei with SS, SD and DD signals, respectively, are shown in the assigned colour code. Low SS ratio indicates early replication timing during the S phase of the cell cycle of the locus detected by a specific probe. Increasing the ratio of the SS nuclei indicates later replication timing
The radial nuclear location of chromosome 7 loci and, more precisely, the median values as defined in the present work, was compared with the replication class of the respective chromosomal band. The resulting correlation was again high (R = 0.80; Fig. 6), indicating that replication of chromosome 7 DNA starts with the chromosomal regions located more internally in the nucleus and ends with the more peripheral ones.
Discussion
The first study on the distribution of chromosomes in spherically shaped nuclei was performed using the GC-rich chromosome 19 and the GC-poor chromosome 18 (Croft et al. 1999). This and subsequent studies (Boyle et al. 2001) showed that chromosomes are distributed according to their gene densities, with the gene-poor chromosome territories located at the nuclear periphery and the gene-dense ones towards the nuclear interior. In general, the radial sub-nuclear location of the entire chromosomes is an approximation of the chromosome organisation in the nuclei. A precise estimate should consider the distribution of the DNA belonging to the chromosomal bands that compose the chromosome itself. In fact, the large chromosomes are generally composed of sub-regions (the H3+ and the L1+ bands) endowed with very different compositional properties. In the small chromosomes 18 and 19, this issue does not exist because they are both compositionally homogeneous (Federico et al. 2000; Saccone et al. 2001; IHGSC 2004). On the other hand, the nuclear location of chromosomal bands belonging to chromosomes 6, 9 and 12 showed that the H3+ and the L1+ band DNAs are located more internally in the nucleus with a very open chromatin conformation and more peripherally with a very compact structure, respectively (Saccone et al. 2002). This reflects the general chromatin arrangement and has also been shown using different procedures (Sadoni et al. 1999; Gilbert et al. 2004). Moreover, very recent results on the human chromosomes 11, 12, 18 and 19, obtained using 3D analyses, showed a polarised distribution of gene-dense versus gene-poor chromatin within chromosome territories in the cell nucleus (Küpper et al. 2007).
Our present data show that the bulk of chromosome 7 does not reach the inner part of the nucleus; in fact, 16 out of 19 loci showed a median value higher than 0.65, in agreement with previous findings on the very peripheral location of the entire human chromosome 7 territory in the nuclei (Boyle et al. 2001). However, we noted that the 7p22.3 and 7q22.1 bands are unexpectedly located in the nuclear interior and thus extend out of the chromosome 7 territory. A similar phenomenon has been described for other chromosomal regions, such as the 6p21 band that loops out of the human chromosome 6 territory (Volpi et al. 2000). Our conclusion is that chromosome 7 in the PHA-stimulated lymphocyte nucleus is arranged with a zig-zag feature, as previously predicted by Zink (2006). This means that the gene-poorest regions are located together on the portion of chromosome territory that lies close to the nuclear envelope, and the gene-dense chromosomal regions are located together in the part of the chromosome territory distributed towards the nuclear interior, with the 7p22.3 and the 7q22.1 band DNA looping out from the bulk of the chromosome territory (see Fig. 4). This once more confirms the relevance of gene density in the intranuclear positioning of the chromosomal band DNA and could explain the more peripheral location of the H3+ band 7q36.3, with respect to the other H3+ bands, due to its exceptionally low gene density.
This type of chromatin arrangement permits sub-regions endowed with similar compositional properties (such as GC level, gene density, CpG island distribution, replication timing, etc.) but located distantly on the same chromosome to share a close reciprocal position. This property is probably related to the different expression/replication features displayed by the different bands, as suggested by a number of studies reporting that the higher-order chromatin arrangement and the related dynamic interactions with other nuclear components are associated with gene expression (see Cremer and Cremer 2001 for review). This means that DNA sequences endowed with similar properties but located on different chromosomes or distant on the same chromosome could be positioned close to each other in the cell nucleus, allowing the coordination of their replication/transcription. This was also recently suggested by D’Onofrio et al. (2007) who described a significant clustering of human genes according to several functional classes (cellular metabolism, information storage and processing, and cellular processes and signalling) in the H3+ bands.
In the present study, we observed a high level of correlation between the nuclear location of a probe and the replication timing of the chromosomal band containing the probe. This was not surprising because replication timing is one of the functional properties of the human genome largely related to the GC level of the DNA (Federico et al. 1998). In this regard, our present experimental data are in agreement with the literature, with the 7p22.3, 7p21.3, 7q22.1, 7q31.31 and 7q36.3 bands replicating in the fourth, 11th, first, 12th , and sixth classes as defined by Dutrillaux et al. (1976). Thus, DNA replication starts in the GC-richest bands and finishes in the GC-poorest ones. As these types of chromosomal bands are located differently in cell nuclei, the DNA replication follows a precise direction, from the nuclear interior to the nuclear periphery, as previously suggested by other authors (Sadoni et al. 1999; Zink 2006).
A recent piece of work on chromosomes 11, 12, 18 and 19, performed using similar analyses to those we describe here but using a different procedure, showed that a good predictor for the radial positioning of chromatin in the nucleus is the local gene density within a 2-Mb window around the probe rather than the local gene density within a 10-Mb window (Küpper et al. 2007). Our findings confirm a high level of correlation between the radial nuclear location of a locus and the 2-Mb local gene density, as well as between the radial nuclear location and the gene-density of the entire chromosomal band containing the probe. In our study, we preferred to use, as a large segment window, the entire chromosomal band containing the probe independently of the band size, as it allows one to correlate the nuclear location with compositionally homogeneous chromosomal regions. The results suggest that large homogeneous DNA segments, namely the chromosomal bands, should be considered the largest chromosomal unit with specific properties, such as a defined nuclear location. Our results, obtained using a 2D analysis, show a large degree of correspondence with those of Küpper et al. (2007), which were obtained on nuclei prepared under conditions to preserve the 3D large-scale chromatin structure. After having performed our experimental controls on nuclei with preserved large-scale chromatin structure, we can confirm that the analysis and statistical evaluation of a high number of nuclei (prepared with the standard cytogenetic procedure) is a reliable method to identify the radial nuclear positioning of specific loci.
The radial organisation of the DNA in the cell nucleus according to GC level was shown to be highly conserved during the evolution of warm-blooded vertebrates (Saccone et al. 2002; Federico et al. 2004; 2005); therefore, the location of specific chromosomal band DNAs in the nucleus should be relevant from an evolutionary point of view. The radial location of a chromosomal band DNA appears to be largely independent of any other band located in the same chromosome but endowed with different compositional properties. Thus, in the case of large chromosome territories that occupy a large extension within the nuclear space, the GC-rich/gene-rich bands occupy the more internal part of the chromosome territory, and the GC-poor/gene-poor bands occupy that part of the chromosome territory near the nuclear periphery. The tendency for a chromosomal band to localise in a specific nuclear sub-domain seems to depend on its intrinsic properties, such as GC level, gene density or, possibly, local epigenetic modifications of the chromatin such as histone acetylation and/or DNA methylation. Then, large chromosomal rearrangements should have only minor effects on the positioning of affected chromosomal sub-regions. This is supported by experimental data from the study of reciprocal translocations between the peripheral human chromosome 18 and the more internal chromosome 19 that did not show alterations in the radial positioning of the translocated sub-regions (Croft et al. 1999).
Assuming that the nuclear location of specific large chromosomal regions and subregions remains invariant, we expect that the nuclear positioning of specific genes might be affected in cases of chromosome rearrangement. It has been suggested that epigenetic alterations of the chromatin structure, related to the nuclear positioning of specific genes, may play a role in human genetic diseases (Bickmore and van der Maarel 2003; D’Antoni et al. 2004; Boutanaev et al. 2005; Foster and Bridger 2005; Ono et al. 2007). It remains to be investigated whether an alteration of the radial position of a specific locus in the interphase nucleus, as a consequence of a chromosomal rearrangement involving very small chromosomal regions, could lead to genetic diseases via a position effect type mechanism.
References
Andreozzi L, Federico C, Motta S, Saccone S, Sazanova AL, Sazanov AA, Smirnov AF, Galkina SA, Lukina NA, Rodionov AV, Carels N, Bernardi G (2001) Compositional mapping of chicken chromosomes and identification of the gene-richest regions. Chromosom Res 9:521–532
Andrulis ED, Neiman AM, Zappulla DC, Sternglanz R (1998) Perinuclear localization of chromatin facilitates transcriptional silencing. Nature 394:592–595
Bernardi G (2004) Structural and evolutionary genomics. Natural selection in genome evolution. Elsevier, Amsterdam, The Netherlands
Bickmore WA, van der Maarel SM (2003) Perturbations of chromatin structure in human genetic disease: recent advances. Hum Mol Genet 12:207–213
Boggs BA, Chinault AC (1997) Analysis of DNA replication by fluorescence in situ hybridization. Methods 13:259–270
Bolzer A, Kreth G, Solovei I, Koehler D, Saracoglu K, Fauth C, Muller S, Eils R, Cremer C, Speicher MR, Cremer T (2005) Three-dimensional maps of all chromosomes in human male fibroblast nuclei and prometaphase rosettes. PLoS Biol 3(5):e157
Boutanaev AM, Mikhaylova LM, Nurminsky DI (2005) The pattern of chromosome folding in interphase is outlined by the linear gene density profile. Mol Cell Biol 25:8379–8386
Boyle S, Gilchrist S, Bridger JM, Mahy NL, Elis JA, Bickmore WA (2001) The spatial organization of human chromosome within the nuclei of normal and emerin-mutant cells. Hum Mol Genet 10:211–219
Bridger JM, Boyle S, Kill IR, Bickmore WA (2000) Re-modelling of nuclear architecture in quiescent and senescent human fibroblasts. Curr Biol 10:149–152
Brown KE, Guest SS, Smale ST, Hahm K, Merkenschlager M, Fisher AG (1997) Association of transcriptionally silent genes with ikaros complexes at centromeric heterochromatin. Cell 91:845–854
Clemson CM, Hall LL, Byron M, McNeil J, Lawrence B (2006) The X chromosome is organized into a gene-rich outer rim and an internal core containing silenced nongenic sequences. Proc Natl Acad Sci USA 103:7688–7693
Cockell M, Gasser SM (1999) Nuclear compartments and gene regulation. Curr Op Genet Dev 9:199–205
Costantini M, Clay O, Federico C, Saccone S, Auletta F, Bernardi G (2007) Human chromosomal bands: nested structure, high definition map and molecular basis. Chromosoma 116:36–49
Cremer T, Cremer C (2001) Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nature Genet 2:292–301
Croft JA, Bridger JM, Boyle S, Perry P, Teague P, Bickmore WA (1999) Differences in the localization and morphology of chromosomes in the human nucleus. J Cell Biol 145:1119–1131
D’Antoni S, Mattina T, Di Mare P, Federico C, Motta S, Saccone S (2004) Altered replication timing of the HIRA/Tuple1 locus in the DiGeorge and velocardiofacial syndromes. Gene 333:111–119
D’Onofrio G, Ghosh TC, Saccone S (2007) Different functional classes of genes are characterized by different compositional properties. FEBS Lett 581:5819–5824
Dutrillaux B, Couturier J, Richer C-L, Viegas-Pequinot E (1976) Sequence of DNA replication in 277 R- and Q-bands of human chromosomes using a BrdU treatment. Chromosoma 58:51–61
Federico C, Saccone S, Bernardi G (1998) The gene-richest bands of human chromosomes replicate at the onset of the S-phase. Cytogenet Cell Genet 80:83–88
Federico C, Andreozzi L, Saccone S, Bernardi G (2000) Gene density in the Giemsa bands of human chromosomes. Chromosom Res 8:737–746
Federico C, Saccone S, Andreozzi L, Motta S, Russo V, Carels N, Bernardi G (2004) The pig genome: compositional analysis and indentification of the gene-richest regions in chromosomes and nuclei. Gene 343:245–251
Federico C, Cantarella CD, Scavo C, Saccone S, Bed’Hom B, Bernardi G (2005) Avian genomes: different karyotypes but a similar distribution of the GC-richest chromosome regions at interphase. Chromosom Res 13:785–793
Federico C, Scavo C, Cantarella CD, Motta S, Saccone S, Bernardi G (2006) Gene-rich and gene-poor chromosomal regions have different locations in the interphase nuclei of cold-blooded vertebrates. Chromosoma 115:123–128
Ferreira J, Paolella G, Ramos C, Lamond AI (1997) Spatial organization of large-scale chromatin domains in the nucleus: a magnified view of single chromosome territories. Cell Biol 139:1597–1610
Foster HA, Bridger JM (2005) The genome and the nucleus: a marriage made by evolution. Genome organization and nuclear architecture. Chromosoma 114:212–229
Francke U (1994) Digitized and differentially shaded human chromosome ideograms for genomic applications. Cytogenet Cell Genet 6:206–219
Furey TS, Haussler D (2003) Integration of the cytogenetic map with the draft human genome sequence. Hum Mol Genet 12:1037–1044
Gilbert N, Boyle S, Fiegler H, Woodfine K, Carter NP, Bickmore WA (2004) Chromatin architecture of the human genome: gene-rich domains are enriched in open chromatin fibers. Cell 118:555–566
IHGSC (International Human Genome Sequencing Consortium) (2004) Finishing the euchromatic sequence of the human genome. Nature 431:931–945
Kupper K, Kolbl A, Biener D, Dittrich S, von Hase J, Thormeyer T, Fiegler H, Carter NP, Speicher MR, Cremer T, Cremer M (2007) Radial chromatin positioning is shaped by local gene density, not by gene expression. Chromosoma 116:285–306
Lukasova E, Kozubek S, Kozubek M, Falk M, Amrichova J (2002) The 3D structure of human chromosomes in cell nuclei. Chromosom Res 10:535–48
Masny PS, Bengtsson U, Chung S-A, Martin JH, van Engelen B, van der Maarel SM, Winokur ST (2004) Localization of 4q35.2 to the nuclear periphery: is FSHD a nuclear envelope disease? Hum Mol Genet 13:1857–1871
Neusser M, Schubel V, Koch A, Cremer T, Müller S (2007) Evolutionary conserved, cell type and species-specific higher order chromatin arrangements in interphase nuclei of primates. Chromosoma 116:307–320
Ono A, Kono K, Ikebe D, Muto A, Sun J, Kobayashi M, Ueda K, Melo JV, Igarashi K, Tashiro S (2007) Nuclear positioning of the BACH2 gene in BCR-ABL positive leukemic cells. Genes Chromosomes Cancer 46:67–74
Petrova NV, Yarovaya OV, Razin SV (2006) Specific spatial organization of chromosomes in nuclei of primary human fibroblasts is maintained by nuclear matrix. Dokl Biochem Biophys 406:4–6
Roix JJ, McQueen PG, Munson P, Parada LA, Misteli T (2003) Spatial proximity of traslocation-prone gene loci in human lymphomas. Nat. Genet 34:287–291
Saccone S, De Sario A, Della Valle G, Bernardi G (1992) The highest gene concentrations in the human genome are in telomeric bands of metaphase chromosomes. Proc Natl Acad Sci USA 89:4913–4917
Saccone S, Cacciò S, Perani P, Andreozzi L, Rapisarda A, Motta S, Bernardi G (1997) Compositional mapping of mouse chromosomes and identification of the gene-rich regions. Chromosom Res 5:293–300
Saccone S, Pavlicek A, Federico C, Paces J, Bernardi G (2001) Genes, isochores and bands in human chromosomes 21 and 22. Chromosom Res 9:533–539
Saccone S, Federico C, Bernardi G (2002) Localization of the gene-richest and the gene-poorest isochores in the interphase nuclei of mammals and birds. Gene 300:169–178
Sadoni N, Langer S, Fauth C, Bernardi G, Cremer T, Turner BM, Zink D (1999) Nuclear organization of mammalian genomes: polar chromosome territories build up functionally distinct higher order compartments. J Cell Biol 146:1211–1226
Scherer SW, Cheung J, MacDonald JR, Osborne LR, Nakabayashi K et al (2003) Human chromosome 7: DNA sequence and biology. Science 300:767–772
Solovei I, Cavallo A, Schermelleh L, Jaunin F, Scasselati C, Cmarko D, Cremer C, Fakan S, Cremer T (2002) Spatial preservation of nuclear chromatin architecture during three-dimensional fluorescence in situ hybridization (3D-FISH). Exp Cell Res 276:10–23
Strouboulis J, Wolffe AP (1996) Functional compartmentalization of the nucleus. J Cell Sci 109:1991–2000
Tanabe H, Habermann FA, Solovei I, Cremer M, Cremer T (2002a) Non-random radial arrangements of interphase chromosome territories: evolutionary considerations and functional implications. Mutat Res 504:37–45
Tanabe H, Muller S, Neusser M, von Hase J, Calcagno E, Cremer M, Solovei I, Cremet C, Cremer T (2002b) Evolutionary conservation of chromosome territory arrangements in cell nuclei from higher primates. Proc Natl Acad Sci USA 99:4424–4429
Volpi EV, Chevret E, Jones T, Vatcheva R, Williamson J, Beck S, Campbell RD, Goldsworthy M, Powis SH, Ragoussis J, Trowsdale J, Sheer D (2000) Large-scale chromatin organization of the major histocompatibility complex and other regions of human chromosome 6 and its response to interferon in interphase nuclei. J Cell Sci 113:1565–1576
Zhou J, Ermakova OV, Riblet R, Birshtein BK, Schildkraut CL (2002) Replication and subnuclear location dynamics of the immunoglobulin heavy-chain locus in B-lineage cells. Mol Cell Biol 22:4876–4889
Zink D (2006) The temporal program of DNA replication: new insight into old question. Chromosoma 115:273–287
Acknowledgement
We thank Giorgio Bernardi for very useful comments, Joanna Bridger and Oliver Clay for critical reading of the manuscript, and the anonymous referees for constructive criticisms.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Allshire
Rights and permissions
About this article
Cite this article
Federico, C., Cantarella, C.D., Di Mare, P. et al. The radial arrangement of the human chromosome 7 in the lymphocyte cell nucleus is associated with chromosomal band gene density. Chromosoma 117, 399–410 (2008). https://doi.org/10.1007/s00412-008-0160-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-008-0160-x