Abstract
Purpose
The aim of the present study was to determine the profile of mismatch repair (MMR) defects in Iranian colorectal cancer patients by using immunohistochemical staining for products of four MMR genes: MLH1, MSH2, PMS2, and MSH6.
Methods
Tissue samples of 343 patients were immunostained for MLH1, MSH2, PMS2, and MSH6. Clinical and family history and survival data were compared between normal and abnormal staining patterns.
Results
Fourteen percent of the patients had abnormal nuclear staining for MMR proteins. MLH1 was absent in four, MLH1/PMS2 in 15, PMS2 in five, MSH2 in 12, and MSH2/MSH6 in 12 patients. These tumors were more proximal, had a nonsignificant better survival, and were more associated with positive family history. Estimation of this study of prevalence of hereditary nonpolyposis colorectal cancer in Iran was 5.5% of the total colorectal cancers.
Conclusions
Along with the recommendations of the National Institute of Cancer, we recommend immunohistochemistry staining for MLH1, MSH2, PMS2, and MSH6 for determining the eligibility of patients for mutation analysis of MMR genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hereditary nonpolyposis colorectal cancer (HNPCC) is an inherited autosomal dominant disorder [1] that accounts for 1–6% of colorectal cancers [2]. It is characterized by early onset colorectal cancer (CRC) [3], more mucinous histology [3], and accelerated transformation from adenoma to carcinoma [4]. It is also associated with synchronous and metachronous colorectal tumors [5], and extra colonic malignancies of brain [6], genitourinary tract, and gastrointestinal system [6, 7].
HNPCC has been shown to be caused by inherited genetic defects in post replication DNA mismatch repair (MMR) system [8] that lead to microsatellite instability (MSI) which is the hallmark of HNPCC [9, 10]. Specific microsatellite markers and PCR can identify MSI [11]. When the instability exceeds 30% of the examined loci, colorectal tumors are classified as high-frequency MSI or MSI-H and otherwise as MSI-L or low-frequency MSI. Absence of alteration in the length of DNA sequence corresponds to be classified as microsatellite stable (MSS). MSI-H is present in about 90% of HNPCC-associated CRCs and four to 15% of sporadic type [10, 12, 13].
The genetic mechanism by which the MSI-H colorectal adenocarcinomas develop is different from that of MSI-L and MSS tumors [14, 15]. These tumors usually involve the proximal colon and express characteristic histopathologic features including poor differentiation, marked peritumoral and intratumoral lymphoid reaction, and mucinous and medullary histology [16, 17]. Moreover, MSI-H tumors show better clinical outcome irrespective of their stage [7, 17, 18] and are less aggressive than common colorectal tumors [18, 19]. Given these data, the assessment of MSI is exceedingly proposed as a provider of useful prognostic information in CRCs [10, 20]. Moreover, MSI has been shown to be a valuable predictor of double primary malignancies in colorectal patients [21].
In a portion of HNPCC subjects, predisposition to cancer have been associated to germline mutations in four MMR genes: MSH2, MLH1, PMS2 [22, 23], and MSH6 [24]. Inactivation of MSH2 and MLH1 is responsible for defects in MMR system in the large majority of MSI-H tumors [10, 22] and can be detected in 50% to 70% of HNPCC families who fulfill the Amsterdam criteria and 30% of atypical HNPCC families [23]. Germline mutations of MSH6 and PMS2 have been found in atypical and a minority of typical HNPCC families [24–26] who show a dominant trend towards familial risk of CRC with minimal or no MSI [26].
MMR proteins interact in the form of heterodimers of MSH2/MSH6 and MLH1/PMS2 [27]. Mutations that affect MLH1 and MSH2 typically cause concurrent immunohistochemical loss of PMS1 and MSH6 through degeneration of heterodimerizing protein partner, respectively [26, 28]. On the contrary, germline mutations affecting the PMS1 or MSH6 result, in the majority of cases, in isolated loss of affected protein on immunohistochemistry (IHC) staining, and can be missed by MSI analysis [26].
A growing body of evidence advocates the potential of immunohistochemical analysis of MLH1 and MSH2 gene products to identify MSI-H colorectal adenocarcinomas specifically [29–33]. Given the advantage of added value of MSH6 and PMS2 and its availability, applicability and lower cost, IHC can be regarded as an effective prescreening tool for identifying HNPCC [30, 34, 35]. The aim of the present study was to determine and study the pattern of MMR defects in a population of CRC patients by using IHC, and to compare the familial, clinical, and survival data of tumors with normal and abnormal MMR status.
Patients and methods
Patients
This study included all colorectal adenocarcinoma patients who had undergone surgical resection from January 2001 to December 2005, at Taleghani Hospital (Tehran, Iran). Data regarding the gender; age at diagnosis; grading, staging, and location of the tumor; histopathology report; and follow up was retrieved from available hospital records for every patient and was completed by telephone interview with the patients or their family members, if necessary. Tumors were originally staged according to TNM system [36] and were graded according to the criteria of the World Health Organization [37]. Tumors were classified as proximal or distal in reference to the splenic flexure of colon.
Paraffin-embedded tissue blocks and histopathology slides were accessed, in advance, from the histopathology archive. Original hematoxylin and eosin slides were reexamined for congruency with previously issued reports or for confirming the diagnosis in case that the original reports were missed. Patients whose pathology specimens were missed and those with history of familial adenomatous polyposis, presurgical radiation therapy, and inflammatory bowel disease were excluded from the study. IHC was employed on tissue specimens of the remaining 343 patients.
Information on family history of CRC was obtained as much as possible from the hospital records of patients in the first place, and measures were taken to verify and complete the familial data by telephone interview with the patients or their close relatives. However, because there was no national registry for cancer in Iran, the CRC patients with a positive family history could not be accurately entered in HNPCC criteria.
Immunohistochemical staining
One tumor specimen from each patient was used for analysis. Four micron-thick sections were obtained from formalin-fixed paraffin-embedded tissue blocks. The tissue sections were deparaffinized in xylene and were rehydrated in graded concentrations of alcohol. Endogenous peroxidase activity was blocked by treating the sections with a blocking solution. For antigen retrieval, the sections were treated while boiling in citrate buffer (pH6.0) in a microwave oven.
Sections were incubated afterwards with primary antibodies against MLH1 (BD Biosciences Pharmingen, clone:G168-15, dilution of 1:100), MSH2 (Calbiochem, Oncogene sciences, clone FE11, dilution of 1:100), MSH6 (BD Trasduction Laboratory, clone: 44, dilution of 1:1000), and PMS2 (BD Pharmingen, clone:A16-4, dilution of 1:500). After each step, slides were rinsed with TBS buffer for 3 min. Then, slides were treated with Envision (DAKO, REAL Envision) for 60 min. To visualize immunoreactivity, 3,2’-diaminobenzidine was used and samples were counterstained with hematoxylin.
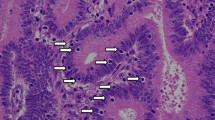
Two different blinded pathologists reviewed all of the slides. Normal epithelial cells, stromal cells, or intramucosal lymphocytes in the same slide were used as internal control for evaluation of IHC staining for MSH2, MLH1, PMS2, and MSH6 (Fig. 1, a1–d1). Slides were evaluated for the presence of nuclear staining, and complete absence of nuclear staining for any of the mentioned MMR gene products (MMRP) was considered abnormal MMRP (Fig. 1, a2–d2). The whole procedure was repeated exactly for certain specimens of which the internal control staining was not satisfactory to ensure preservation of maximum sample size and for one pattern of abnormal MMRP (PMS2 Negative, MLH1 and MSH2 positive).
Normal and abnormal staining patterns of four MMR proteins are presented as eight pictures: a1, 2 (MLH1); b1, 2 (MSH2); c1,2 (MSH6); d1, 2 (PMS2). a1–d1 represent normal immunohistochemical staining for MLH1, MSH2, MSH6, and PMS2, respectively. Normal nuclear staining for MMR proteins is notable both in stromal cells (black arrows) and epithelial tumor cells (white arrows), as brownish accumulation of dye in the nucleus of the mentioned cells. a2–d2 represent abnormal staining for MLH1, MSH2, MSH6, and PMS2, respectively. In these series of pictures, stromal cells (black arrows) still show normal staining for MMR proteins, while there is no dye accumulation in nucleus of epithelial tumor cells (white arrows)
Statistical analysis
Differences of distribution between the categorical variables were examined with chi-square test and Fisher's exact test in case of need. For quantitative variables, Student's t test was employed. Multivariate logistic regression analysis was performed with covariates of age at onset, family history, and tumor site included. Multivariate survival analysis was performed with Cox's proportional hazard model while the covariates of gender, age at diagnosis, tumor site, and tumor stage and grade were included. Reported P values of less than 0.05 were considered to represent the statistical significance.
Results
Clinical data
Of 343 examined colorectal adenocarcinomas, 295 (86%) showed normal nuclear expression of MLH1, MSH6, PMS2, and MSH6, and 48 (14%) showed abnormal staining patterns of MMRP. Tumors with abnormal MMRP staining tended more to be proximal and occur at lower ages; family history of CRC was also higher in these tumors. No significant difference of staging and grading could be found between normal and abnormal MMRP tumors (Table 1). We could find no significant relationship between early onset of CRC (onset before the age of 50) and positive family history.
In order to determine how three variables of age at onset, family history for CRC, and tumor site could have predicted abnormalcy of MMRP patterns, we employed multivariate logistic regression analysis with the mentioned factors included (Table 2). It could be implied from the analysis that there were independent associations between onset age of less than 50 years, positive family history, and proximal tumors with the risk of having abnormal MMRP status.
Staining patterns
Among abnormal MMRP tumors, 19 (39.5%) showed complete loss of MLH1 with normal MSH2 immunoreactivity (MLH1 negative), and 24 (50%) expressed complete loss of MSH2 with normal MLH1 immunohistochemical staining (MSH2 negative). Five tumors (10.4%) showed complete loss of PMS2 while expressing positive immunoreactivity for both MLH1 and MSH2. Of 19 MLH1-negative tumors, 15 had simultaneous loss of PMS2, and 12 tumors out of 24 MSH2-negative tumors had simultaneous loss of MSH6 expression.
According to our findings, 80 CRC patients (23%) had a positive family history of CRC, of whom 47 patients (13.7%) had a cancer onset age of less than 50 years old. Out of these 47 patients, 19 (5.5% of total) showed abnormal IHC staining of MMR proteins. The pattern of abnormal IHC staining in this group of CRC patients was as follows: absence of MLH1 was observed in one, MLH1/PMS2 in five, PMS2 in two, MSH2 in six, and MSH2/MSH6 in five patients.
Survival analysis
Twenty-eight (8.2%) patients died because of distant metastasis or local recurrence, four of whom had abnormal MMRP patterns and 24 did not. Multivariate survival analysis was performed on 342 CRC patients with covariates of gender, tumor site, tumor stage and grade, and age at diagnosis included (Table 3). Five-year follow-up was completed in 24 patients including 20 patients with normal MMRP status and four abnormal MMRP cases. A nonsignificant inclination towards better survival was observed among patients with abnormal MMRP (Fig. 2).
Cancer specific survival of colorectal cancer patients with normal and abnormal immunohistochemistry for mismatch repair proteins according to Cox's proportional hazard model. The cumulative 5-year survival of patients with abnormal MMRP (green line) is better than that of the normal MMRP patients (blue line); however, results are not significant.
Discussion
While a worldwide familial basis of about 15–20% has been reported for CRC [2, 7], recent studies from Iran have unveiled a familial trend of 29.4–35.1% for these type of cancers [38, 39], which would rise to 53% with the family history of other cancers included [39]. In this study, 23.3% of CRC patient had a positive family history of CRC, and 46% had a positive family history of cancer in general. Our results confirmed the findings of previous studies regarding the high-familial inheritance of CRC in Iran.
The present study was unique because for the first time in Iran, this strived to merge the clinical and familial profile of CRC patients to immunohistochemical evidences of ongoing genetic abnormalities. However, like the previous studies that contended with CRC in Iran, this study faced a major limitation: the bias generated by lack of a national cancer registry system [38–41]. This limitation impeded the inclusion of CRC patients into the accepted HNPCC criteria, just based on positive family history or age of cancer onset confidently, because the recall bias could decrease the accuracy of the data.
Of previous studies about CRC in Iran, only one has tried to depict an overview of HNPCC [39]. The mentioned study, reported a prevalence of 4.7% based on the Amsterdam II criteria. If we hypothetically consider the CRC patients with positive family history, early onset age and an abnormal MMRP on IHC as HNPCC, then the estimation of this study of prevalence of HNPCC will be 5.5% of the total CRC patients. This estimation is the highest among those recorded in Iran and is in accordance with the previously reported prevalence; however, studies from other parts of the world have generally reported lower estimates [2, 3, 6, 11, 42].
In our study, the sensitivity and specificity of IHC in contrast to microsatellite analysis could have been determined only by performing both procedures on every specimen, which was not feasible because of limitation of financial resources. However, high rate of isolated absence of PMS2, five (10.4%) of 48 abnormal MMRP, in this study allows us to predict a possible added value of about 10% to IHC over microsatellite analysis. Absence of PMS2 staining might be a sign of germline mutations in MLH1 or PMS2 [26]; this means that by using immunohistochemical staining for PMS2, more carriers of MLH1 mutations can be identified [43]. Previously, staining for PMS2 was recommended only in case of high suspicion for HNPCC in the absence of MSI. However, given an extra benefit of 23% in detecting the MLH1 mutations, a recent study recommended the inclusion of PMS2 staining in the panel of antibodies to identify families eligible for mutation analysis [43].
No isolated absence of MSH6 was found in this study. Four tumors had isolated loss of MLH1, and 15 had loss of both MLH1/PMS2. Twelve tumors had isolated abnormal staining for MSH2, and 12 had absence of MSH2/MSH6 heterodimer. However, isolated absence of MLH1 was observed in just one of the 19 assumed HNPCC patients. Germline mutations are usually responsible for the absence of MSH2 proteins [44]; whereas, the absence of MLH1 can be indicative of a germline mutation or of somatic hypermethylation of its promoter that happens in sporadic CRC [45, 46]. Therefore, a likely explanation for the less frequent loss of MLH1 in this study is the higher prevalence of hereditary cancer in Iran.
To conclude, this study demonstrated the importance and feasibility of running large-scale CRC-screening programs in Iran. The guidelines of the National Institute of Cancer for screening CRC is advocated by the present study; moreover, we recommend IHC as a prescreening tool for determining the eligibility of patients for mutation analysis of MMR genes specially in populations with high-familial incidence of CRC like Iran [47]. According to the present study, in addition to MLH1 and MSH2, PMS2 should be entered into the panel of antibodies used for immunohistochemichal analysis of MMR genes. Although this might exact a substantial financial toll on the prescreening programs, the IHC analysis of PMS2 seems to be inevitable especially in populations with higher incidence of hereditary colorectal cancer. Moreover, measures should be taken in future studies, to confirm the IHC results by MSI analysis, and to determine the concordance of IHC results with the mutation analysis measures.
References
Green SE, Bradburn DM, Varma JS et al (1998) Hereditary non-polyposis colorectal cancer. Int J Colorectal Dis 13:3–12
Aaltonen LA, Salovaara R, Kristo P et al (1998) Incidence of hereditary nonpolyposis colorectal cancer and the feasibility of molecular screening for the disease. N Engl J Med 338:1481–1487
Lynch HT, Lynch JF, Lynch PM (2007) Toward a consensus in molecular diagnosis of hereditary nonpolyposis colorectal cancer (Lynch syndrome). J Natl Cancer Inst 99:261–263
Jass JR, Stewart SM (1992) Evolution of hereditary non-polyposis colorectal cancer. Gut 33:783–786
Jin HY, Liu X, Li VK et al (2008) Detection of mismatch repair gene germline mutation carrier among Chinese population with colorectal cancer. BMC Cancer 7(8):44
Furukawa T, Konishi F, Shitoh K et al (2003) An early stage small bowel adenocarcinoma with microsatellite instability phenotype in a case of hereditary nonpolyposis colorectal cancer. Int J Colorectal Dis 18:267–270
Lynch HT, Smyrk T (1996) Hereditary nonpolyposis colorectal cancer (Lynch syndrome). An updated review. Cancer 78:1149–1167
Jiricny J, Nystron-Lahti M (2000) Mismatch repair defects in cancer. Curr Opin Genet Dev 10:157–161
Chung DC, Rustgi AK (2003) The hereditary nonpolyposis colorectal cancer syndrome: genetics and clinical implications. Ann Intern Med 138:560–570
Müller CI, Schulmann K, Reinacher-Schick A et al (2008) Predictive and prognostic value of microsatellite instability in patients with advanced colorectal cancer treated with a fluoropyrimidine and oxaliplatin containing first-line chemotherapy. A report of the AIO Colorectal Study Group. Int J Colorectal Dis 23:1033–1039
Boland CR, Thibodeau SN, Hamilton SR et al (1998) A National Cancer Institute Workshop on Microsatellite Instability for cancer detection and familial predisposition: development of international criteria for the determination of microsatellite instability in colorectal cancer. Cancer Res 58:5248–5257
Moslein G, Tester DJ, Lindor NM et al (1996) Microsatellite instability and mutation analysis of hMSH2 and hMLH1 in patients with sporadic, familial and hereditary colorectal cancer. Hum Mol Genet 5:1245–1252
Aaltonen LA, Peltomaki P, Mecklin JP et al (1994) Replication errors in benign and malignant tumors from hereditary non-polyposis colorectal cancer patients. Cancer Res 54:1645–1648
Whitehall VL, Walsh MD, Young J (2001) Methylation of O-6-methylguanine DNA methyltransferase characterizes a subset of colorectal cancer with low-level DNA microsatellite instability. Cancer Res 61:827–830
Mori Y, Selaru FM, Sato F et al (2003) The impact of microsatellite instability on the molecular phenotype of colorectal tumors. Cancer Res 63:4577–4582
Jass JR, Do KA, Simms LA et al (1998) Morphology of sporadic colorectal cancer with DNA replication errors. Gut 42:673–679
Lanza G, Gafa R, Santini A et al (2006) ImmunohistochemicalTest for MLH1 and MSH2 expression predicts clinical outcome in stage II and III colorectal cancer patients. J Clin Oncol 24:2359–2367
Gafà R, Maestri I, Matteuzzi M et al (2000) Sporadic colorectal adenocarcinomas with high-frequency microsatellite instability: Pathobiologic features, hMLH1 and hMSH2 expression, and clinical outcome. Cancer 89:2025–2037
Ward R, Meagher A, Tomlinson I et al (2001) Microsatellite instability and the clinicopathological features of sporadic colorectal cancer. Gut 48:821–829
Nehls O, Hass HG, Okech T et al (2009) Prognostic implications of BAX protein expression and microsatellite instability in all non-metastatic stages of primary colon cancer treated by surgery alone. Int J Colorectal Dis 24:655–663
Yun HR, Yi LJ, Cho YK et al (2009) Double primary malignancy in colorectal cancer patients—MSI is the useful marker for predicting double primary tumors. Int J Colorectal Dis 24:369–375
Wheeler JM, Bodmer WF, Mortensen NJ (2000) DNA mismatch repair genes and colorectal cancer. Gut 47:148–153
Wijnen J, Khan PM, Vasen H et al (1997) Hereditary nonpolyposis colorectal cancer families not complying with the Amsterdam criteria show extremely low frequency of mismatch-repair-gene mutations. Am J Hum Genet 61:329–335
Kolodner RD, Tytell JD, Schmeits JL et al (1999) Germline msh6 mutations in colorectal cancer families. Cancer Res 59:5068–5074
Verma L, Kane MF, Brassett C et al (1999) Mononucleotide microsatellite instability and germline MSH6 mutation analysis in early onset colorectal cancer. J Med Genet 36:678–682
Wu Y, Berends MJ, Mensink RG et al (1999) Association of hereditary nonpolyposis colorectal cancer-related tumors displaying low microsatellite instability with MSH6 germline mutations. Am J Hum Genet 65:1291–1298
Kolodner RD, Marsischky GT (1999) Eukaryotic DNA mismatch repair. Curr Opin Genet Dev 9:89–96
Young J, Simms LA, Biden KG et al (2001) Features of colorectal cancers with high-level microsatellite instability occurring in familial and sporadic settings: parallel pathways of tumorigenesis. Am J Pathol 159:2107–2116
Cawkwell L, Gray S, Murgatroyd H et al (1999) Choice of management strategy for colorectal cancer based on a diagnostic immunohistochemical test for defective mismatch repair. Gut 45:409–415
de la Chapelle A (2002) Microsatellite instability phenotype of tumors: genotyping or immunohisto-chemistry? The jury is still out. J Clin Oncol 20:897–899
Lanza G, Gafa R, Maestri I et al (2002) Immunohistochemical pattern of MLH1/MSH2 expression is related to clinical and pathological features in colorectal adenocarcinomas with microsatellite instability. Mod Pathol 15:741–749
Lindor NM, Burgart LJ, Leontovich O et al (2002) Immunohistochemistry versus microsatellite instability testing in phenotyping colorectal tumors. J Clin Oncol 20:1043–1048
Shia J, Ellis NA, Klimstra DS (2004) The utility of immunohistochemical detection of DNA mismatch repair gene proteins. Virchows Arch 445:431–441
Müller W, Burgart LJ, Krause-Paulus R et al (2001) The reliability of immunohistochemistry as a prescreening method for the diagnosis of hereditary nonpolyposis colorectal cancer (HNPCC): results of an international collaborative study. Fam Cancer 1:87–92
Umar A, Boland CR, Terdiman JP et al (2004) Revised bethesda guidelines for hereditary nonpolyposis colorectal cancer (Lynch syndrome) and microsatellite instability. J Nat Cancer Inst 96:261–268
Sobin LH, Wittekind C (eds) (1997) UICC TNM classification of malignant tumours, 5th edn. Wiley, New York
Jass JR, Sobin LH (eds) (1989) World Health Organization international histological classification of tumours. Histological typing of intestinal tumours, 2nd edn. Springer, Berlin
Azadeh S, Moghimi-Dehkordi B et al (2008) Colorectal cancer in Iran: an epidemiological study. Asian Pac J Cancer Prev 9:123–126
Mahdavinia M, Bishehsari F, Ansari R et al (2005) Family history of colorectal cancer in Iran. BMC Cancer 5:112
Pahlavan PS, Kanthan R (2006) The epidemiology and clinical findings of colorectal cancer in Iran. J Gastrointestin Liver Dis 15:15–19
Hosseini SV, Izadpanah A, Yarmohammadi H (2004) Epidemiological changes in colorectal cancer in Shiraz, Iran: 1980–2000. ANZ J Surg 74:547–544
Hampel H, Frankel WL, Martin E et al (2005) Screening for the Lynch syndrome (hereditary nonpolyposis colorectal cancer). N Engl J Med 352:1851–1860
de Jong AE, van Puijenbroek M, Hendriks Y et al (2004) Microsatellite Instability, Immunohistochemistry, and Additional PMS2 staining in suspected hereditary nonpolyposis colorectal cancer. Clin Cancer Res 10:972–980
Mangold E, Pagenstecher C, Friedl W et al (2005) Tumours from MSH2 mutation carriers show loss of MSH2 expression but many tumours from MLH1 mutation carriers exhibit weak positive MLH1 staining. J Pathol 207:385–395
Cunningham JM, Christensen ER, Tester DJ et al (1998) Hypermethylation of the hMLH1 promoter in colon cancer with microsatellite instability. Cancer Res 58:3455–3460
Thibodeau SN, French AJ, Cunningham JM et al (1998) Microsatellite instability in colorectal cancer: Different mutator phenotypes and the principal involvement of hMLH1. Cancer Res 58:1713–1718
Müller A, Beckmann C, Westphal G et al (2006) Prevalence of the mismatch-repair-deficient phenotype in colonic adenomas arising in HNPCC patients: results of a 5-year follow-up study. Int J Colorectal Dis 21:632–641
Funding
This study was funded by the Research Institute for Gastroenterology and Liver Diseases of Shahid Beheshti University M.C.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Molaei, M., Mansoori, B.K., Ghiasi, S. et al. Colorectal cancer in Iran: immunohistochemical profiles of four mismatch repair proteins. Int J Colorectal Dis 25, 63–69 (2010). https://doi.org/10.1007/s00384-009-0784-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00384-009-0784-1