Abstract
Microbial lipids are sustainable feedstock for the production of oleochemicals and biodiesel. Oleaginous yeasts have recently been proposed as alternative lipid producers to plants and animals to promote sustainability in the chemical and fuel industries. The oleaginous yeast Lipomyces starkeyi has great industrial potential as an excellent lipid producer. However, improvement of its lipid productivity is essential for the cost-effective production of oleochemicals and fuels. Genetic and metabolic engineering of L. starkeyi via gene manipulation techniques may result in improvements in lipid production and our understanding of the mechanisms behind lipid biosynthesis pathways. We previously described an integrative transformation system using a drug-resistant marker for L. starkeyi. However, gene-targeting frequencies were very low because non-homologous recombination is probably predominant in L. starkeyi. Genetic engineering tools for L. starkeyi have not been sufficiently developed. In this study, we describe a new genetic tool and its application in L. starkeyi. To develop a highly efficient gene-targeting system for L. starkeyi, we constructed a series of mutants by disrupting genes for LsKu70p, LsKu80p, and/or LsLig4p, which share homology with other yeasts Ku70p, Ku80p, and Lig4p, respectively, being involved in non-homologous end-joining pathway. Deletion of the LsLIG4 gene dramatically improved the homologous recombination efficiency (80.0%) at the LsURA3 locus compared with that in the wild-type strain (1.4%), when 2000-bp homologous flanking regions were used. The homologous recombination efficiencies of the double mutant ∆l sku70∆lslig4 and the triple mutant ∆lsku70∆lsku80∆lslig4 were also markedly enhanced. Therefore, the L. starkeyi ∆lslig4 background strains have promise as efficient recipient strains for genetic and metabolic engineering approaches in this yeast.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The increasing greenhouse gas emissions and the impending shortage of fossil fuels are due to the excessive consumption of fossil fuel resources, so it is becoming increasingly important to identify renewable substitutes. Biodiesel could be one such alternative, but the use of plant oils or animal fats as feedstock for producing biodiesel is often controversial because of the large land area required for their production, their potential competition with food production, and their high cost (Liang and Jiang 2013). However, microbial oils have many advantages over plant oils or animal fats, such as the avoidance of competition with food production, independence of climate and season, ease of scaling up, and high growth rates on various carbon sources, and simplicity of metabolic regulation of lipid-producing microbes by genetic engineering tools compared with that in plants and animals (Liang and Jiang 2013). Therefore, oleaginous microorganisms with lipids in excess of 20% of their biomass dry weight might become excellent oil feedstocks (Ratledge and Wynn 2002). Some yeasts, such as Lipomyces starkeyi, Rhodosporidium toruloides, Rhodotorula glutinis, Cryptococcus curvatus, Zygolipomyces lactosus, and Trichosporon cutaneum, can store intracellular lipids within cells to up to 60% of their cell dry weight (Ageitos et al. 2011). In these oleaginous yeasts, triacylglycerols (TAGs), which have a fatty acid composition similar to that of plant oils used as food and energy sources, mainly accumulate as storage lipids (Kosa and Ragauskas 2011; Papanikolaou and Aggelis 2011). L. starkeyi is one of the most widely known oleaginous microorganisms and is able to accumulate TAGs to up to 75.2% of its dry cell weight (Angerbauer et al. 2008). Thus, L. starkeyi is a unique yeast strain of great industrial potential as a lipid producer. The complete genome sequence of L. starkeyi is now available, and genetic tools for the transformation, multicopy integration, and expression of heterologous genes have recently been reported for this yeast (Calvey et al. 2014; Oguro et al. 2015). One of the disadvantages of L. starkeyi is the low efficiency of gene targeting mediated by homologous recombination, unlike that in Saccharomyces cerevisiae (Rothstein 1991). Gene targeting, which can be used for gene deletion, gene replacement, or the integration of DNA fragments encoding for epitope tags or fluorescent proteins into the genome, is one of the most important approaches for analyzing gene function or blocking metabolic pathways by genetic engineering. It is also a cornerstone skill in many industrially important microorganisms, such as the oleaginous yeast L. starkeyi.
The targeted integration of a DNA fragment is mainly dependent on one of the mechanisms by which double-strand breaks (DSBs) are repaired. In eukaryotes, there are at least two pathways that can repair DSBs: homologous recombination (HR) and non-homologous end joining (NHEJ). In S. cerevisiae, DSB repair primarily occurs by HR, but most fungi seem to prefer NHEJ. S. cerevisiae additionally exhibits efficient gene targeting, which requires only about a 40-bp homologous DNA flanking sequence (Baudin et al. 1993; Wach et al. 1994). This efficient gene targeting leads to the construction of a whole-genome knockout collection, which improves our understanding of the function of genes (Giaever et al. 2002). Most fungi are known to have low gene-targeting efficiency, owing to the predominance of NHEJ over HR. In S. cerevisiae, NHEJ is essential for maintaining genome stability during G1 phase (Moore and Haber 1996; Takata et al. 1998). Repair of DSBs by NHEJ is accomplished via three steps: (1) recognition and seizure of broken DNA ends; (2) formation of a bridge to bring broken DNA ends together; and (3) ligation and repair of DSBs (Chiruvella et al. 2013). Three protein complexes mediate the yeast NHEJ pathway: Ku70/Ku80, Mre11/Rad50/Xrs2 (MRX complex), and Lig4/Lif1/Nej1 (DNA ligase IV). DSBs are recognized by the Ku70/Ku80 heterodimer (Milne et al. 1996), which sequence-independently bind to the broken DNA ends protecting the free ends from extensive degradation (Feldmann et al. 2000). Subsequently, the MRX complex mediates DNA end-bridging via the zinc-hook in Rad50 (Lobachev et al. 2004), triggering DNA ligase IV to target the DNA ends in combination with the ku70/ku80 heterodimers (Chen et al. 2001). The final step is the end-joining of broken DNA strands, which requires for DNA ligase IV for ligation occur. DNA ligase IV is composed of DNA ligase Lig4p (or Dnl4p) (Teo and Jackson 1997; Wilson et al. 1997), Lif1p (yeast XRCC4 homologue) (Herrmann et al. 1998), and Nej1p (Valencia et al. 2001). Lig4p is an ATP-dependent DNA ligase that is strongly associated with a coiled-coil region of Lif1p (Dore et al. 2006).
Thus, the deletion of genes closely related to the NHEJ pathway, such as KU70, KU80, or LIG4, is assumed to increase the frequency of HR caused by reduction of the random integration of DNA fragments. Recently, the enhanced gene-targeting efficiency of mutants with deletion in KU70, KU80, or LIG4 was observed in fungi capable of contributing to the biotechnology industry, including R. toruloides (Koh et al. 2014), Y. lipolytica (Kretzschmar et al. 2013; Verbeke et al. 2013), Candida glabrata (Ueno et al. 2007) (Cen et al. 2015), Candida guilliermondii (Foureau et al. 2013), Cryptococcus neoformans (Goins et al. 2006), Kluyveromyces marxianus (Choo et al. 2014), Pichia pastoris (Naatsaari et al. 2012), P. stipites (Maassen et al. 2008), P. ciferrii (Schorsch et al. 2009), Mortierella alpina (Kikukawa et al. 2015), and Lecanicillium sp. (Ishidoh et al. 2014).
Here, we describe the development of efficient gene targeting leading to the functional analysis of individual genes and metabolic engineering in the oleaginous yeast L. starkeyi. The predicted genes LsKU70, LsKU80, and LsLIG4 were obtained from the L. starkeyi genome. Then, L. starkeyi mutants with the deletion of LsKU70 (∆lsku70), LsKU80 (∆lsku80), LsLIG4 (∆lslig4), both LsKU70 and LsLIG4 (∆lsku70∆lslig4), or all of LsKU70, LsKU80, and LsLIG4 genes (∆lsku70∆lsku80∆lslig4) were constructed, and the gene-targeting efficiency of these mutants was analyzed by deleting the LsURA3 gene. The results indicated that LsLig4p is crucial for gene-targeting efficiency. Therefore, the LsLIG4-deleted strain is a useful host for metabolic engineering and comprehensive investigations of the functional genome of L. starkeyi.
Materials and methods
Strains and media
The bacterial and yeast strains used in this study are listed in Table 1. L-broth [1% Bacto™ Tryptone (BD Biosciences, Franklin Lakes, NJ, USA), 0.5% Bacto™ Yeast Extract (BD Biosciences), and 1% NaCl] was used to grow the Escherichia coli strain. YPD [1% Yeast Extract (Kyokutou, Tokyo, Japan), Polypeptone (Nihonseiyaku, Tokyo, Japan), and 2% glucose] and SD [0.17% Difco™ yeast nitrogen base without amino acids and ammonium sulfate (BD Biosciences), which was supplemented with 0.5% ammonium sulfate, and 2% glucose] media were used to grow the yeast strains. Solid media contained 2% agar (Wako Pure Chemical, Osaka, Japan). Selective YPD media contained 100 µg/ml hygromycin B (Wako Pure Chemical), 50 µg/ml zeocin (Invitrogen, Carlsbad, CA, USA), 30 µg/ml nourseothricin (Cosmo Bio, Tokyo, Japan), and/or 100 µg/ml geneticin (Invitrogen). For the determination of the uracil auxotroph, strains were grown on SD agar supplemented with 20 mM uracil and 5-fluoroorotic acid (5-FOA; Wako Pure Chemical). For comparison of the growth and lipid productivity between the wild-type and mutant strains were grown in YD [1% Yeast Extract (Kyokutou) and 10% glucose].
General molecular biology techniques
Genomic DNA from L. starkeyi strains was prepared following the method described for S. cerevisiae (Hereford et al. 1979), except that zymolyase for digestion of the yeast cell wall was replaced with westase (Takara Bio, Kyoto, Japan). Plasmid DNA from E. coli was prepared by the alkaline extraction method (Bimboim and Doly 1979). Restriction enzymes, ligase, and DNA-modifying enzymes were purchased from Takara Bio. KOD-Plus DNA polymerase (Toyobo, Osaka, Japan) was used for PCR amplification, in accordance with the manufacturer’s instructions. Amplified DNA fragments were recovered from agarose gels with the FastGene Gel/PCR extraction kit (Nippon Genetics, Tokyo, Japan) for purification.
Construction of expression cassettes of drug resistance markers
The open reading frames of drug resistance markers were amplified from plasmids and fused to L. starkeyi TDH3 promoter and terminator for expression, as described previously (Oguro et al. 2015). The primers used in this study are listed in Table 2. The geneticin resistance marker (kanR) was amplified from the plasmid pPIC9K (Invitrogen) using the primer set KanR ORF Fw and KanR ORF Rv. The hygromycin B resistance marker (hph) was amplified from the plasmid pCB1004 (Carroll et al. 1994) using the primer set hph ORF Fw and hph ORF Rv. The zeocin resistance marker (Sh ble) was amplified from the plasmid pGAPZαA (Invitrogen) using the primer set Sh ble ORF Fw and Sh ble ORF Rv. The nourseothricin resistance marker (sNAT1) was amplified from the plasmid pTH421 (a gift from Dr. Marie Nishimura) using the primer set sNAT1 ORF Fw and sNAT1 ORF Rv. The PCR products of L. starkeyi-derived TDH3 promoter and terminator regions were amplified with the primer sets PTDH3Fw/PTDH3Rv and TTDH3Fw/TTDH3Rv, respectively, using genomic DNA of L. starkeyi CBS1807 as a template.
PCR-amplified drug marker DNA fragments (kanR, hph, Sh ble, sNAT1) were phosphorylated by T4 polynucleotide kinase. The LsTDH3 promoter/kanR/LsTDH3 terminator DNA fragment was amplified with the primer set PTDH3Fw and TTDH3Rv using the ligated DNA fragments (LsTDH3 promoter region, phosphorylated kanR, LsTDH3 terminator region) as a template. For construction of the LsTDH3 promoter/kanR/LsTDH3 terminator DNA fragment for expression in L. starkeyi, the other DNA fragments (LsTDH3 promoter/hph/LsTDH3 terminator, LsTDH3 promoter/Sh ble/LsTDH3 terminator, LsTDH3 promoter/sNAT1/LsTDH3 terminator) were amplified with the primer set PTDH3Fw and TTDH3Rv using the ligated DNA fragments (LsTDH3 promoter region, phosphorylated drug markers, LsTDH3 terminator region). The orientation of drug marker ORFs with respect to the LsTDH3 promoter and terminator regions was confirmed by sequencing analysis.
The plasmid vector pBluescript KS (+) was used for the cloning of expression cassettes for the drug resistance markers. To construct the drug marker gene expression vector (pKS-hph, pKS-Sh ble, pKS-sNAT1, and pKS-kanR), pBluescript KS (+) was digested with EcoRV. The above LsTDH3 promoter and terminator fused-drug marker expression DNA fragments were ligated to the obtained EcoRV-digested 3-kbp pBluescript KS (+) fragment.
Construction of the disruption cassette plasmid pKS-kanR-LsKU70, pKS-hph-LsKU80, and pKS-Sh ble-LsLIG4
The non-coding regions of the LsKU70, LsKU80, and LsLIG4 genes were amplified with the primers shown in Table 2 using L. starkeyi genomic DNA as a template. The PCR fragments of 5′- and 3′-non-coding regions of the LsKU70 gene were amplified using the 5′-phosphorylated primer sets of 5′nonLsKU70Fw and 5′nonLsKU70Rv, and 3′nonLsKU70Fw and 3′nonLsKU70Rv, respectively. The DNA fragment containing the 5′-non-coding region of the LsKU70 gene connected with the 3′-non-coding region was amplified with the primer set 5′nonLsKU70Fw and 3′nonLsKU70Rv using the two obtained ligated DNA fragments as a template and successively cloned into the EcoRV site of the vector pBluescript KS (+) to yield pKS-LsKU70 (non-coding). To obtain the disruption cassette plasmid pKS-kanR-LsKU70, the PCR product amplified with the primer set 5′nonLsKU70Rv and 3′nonLsKU70Fw using the vector pKS-LsKU70 (non-coding) as a template was ligated with the above phosphorylated LsTDH3 promoter and terminator fused-kanR gene expression DNA fragment.
The PCR fragments of 5′- and 3′-non-coding regions of the LsKU80 gene were amplified using the 5′-phosphorylated primer sets 5′nonLsKU80Fw and 5′nonLsKU80Rv, and 3′nonLsKU80Fw and 3′nonLsKU80Rv, respectively. The DNA fragment containing the 5′-non-coding region of the LsKU80 gene connected with the 3′-non-coding region was amplified with the primer set 5′nonLsKU80Fw and 3′nonLsKU80Rv using the two obtained ligated DNA fragments as a template and successively cloned into the EcoRV site of the vector pBluescript KS (+) to yield pKS-LsKU80 (non-coding). To obtain the disruption cassette plasmid pKS-hph-LsKU80, the PCR product amplified with the primer set 5′nonLsKU80Rv and 3′nonLsKU80Fw using the vector pKS-LsKU80 (non-coding) as a template was ligated with the above phosphorylated LsTDH3 promoter and terminator fused-hph gene expression DNA fragment.
Two PCR fragments of 5′- and 3′-non-coding regions of the LsLIG4 gene amplified with the primer sets 5′nonLsLIG4Fw and 5′nonLsLIG4Rv, and 3′nonLsLIG4Fw and 3′nonLsLIG4Rv using genome DNA as a template, were digested with HindIII and XbaI, respectively, and successively cloned into the HindIII site and the XbaI site of the vector pKS-Sh ble to yield the disruption cassette plasmid pKS-Sh ble-LsLIG4. The orientation of drug marker with respect to the 5′- and 3′-non-coding regions of the LsLIG4 gene was confirmed by sequencing analysis.
Yeast transformation
Transformation of L. starkeyi was performed as described previously (Oguro et al. 2015). Briefly, L. starkeyi was cultured in YPD medium to the log-phase (OD600, 1.0). The cells were collected, washed with sterilized distilled water, and re-suspended in 0.4 M Na-tartrate in McIlvaine buffer (pH 6.0) supplemented with 5 mg/ml Westase (Takara Bio) at a density of 3.0 × 106 cells/ml. For the formation of spheroplasts, the cells were incubated at 30 °C for 90 min. The spheroplasts were collected by gentle centrifugation (1000 × g) for 5 min at room temperature. They were then washed twice in STC buffer [1.2 M sorbitol, 50 mM Tris–HCl (pH 8.0), and 50 mM CaCl2·2H2O] and re-suspended in 0.5 ml of STC buffer. Samples of spheroplasts (500 µl) were incubated with 20 µg of DNA fragments for 20 min at room temperature. Then, 1 ml of PEG solution [40% PEG 4000 (Wako Pure Chemical), 1.2 M sorbitol, and 50 mM CaCl2–2H2O in 50 mM Tris–HCl, pH 8.0] was added and incubated for 20 min at room temperature. The regeneration of spheroplasts was carried out in 10 ml of TB3 solution [0.3% yeast extract (Kyokuto), 0.3% vitamin assay casamino acid (DIFCO), 20% sucrose] and incubated at 30 °C for 16 h. After the regeneration, the cells were inoculated on YPD agar plates containing 100 µg/ml hygromycin B, 50 µg/ml zeocin, 30 µg/ml nourseothricin, and/or 100 µg/ml geneticin and incubated at 30 °C for 3–4 days.
Construction of LsKU70, LsKU80, and/or LsLIG4 deletion mutants in L. starkeyi
Initially, PCR was performed using the disruption cassette plasmid pKS-kanR-LsKU70 as a template and the primer set 5′nonLsKU70Fw and 3′nonLsKU70Rv. The PCR product (LsKU70 disruption cassette) contained an LsKU70 flanking sequence (approximately 1 kbp) on either side of the kanR expression cassette. The transformant was selected on YPD agar containing 100 µg/ml geneticin. PCR was performed using the obtained transformant genomic DNA as a template and the primer set 5′KU70Fw and 3′KU70Rv to confirm the correct insertion (Fig. 1a, d). We also confirmed the target disruption of LsLU70 by Southern blot analysis of genomic DNA (Supplementary Fig. S1a, b).
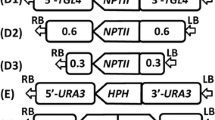
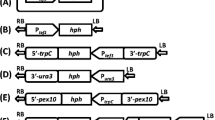
Construction of LsKU70, LsKU80, and/or LsLIG4 deletion mutants. a–c Strategies for homologous recombination of L. starkeyi for LsKU70, LsKU80, and/or LsLIG4 gene disruptions using the kanR, hph, or Sh ble gene as a selectable marker. Black arrows 5′KU70Fw, 3′KU70Rw, 5′KU80Fw, 3′KU80Rw, 5′LIG4Fw, and 3′LIG4Rw indicate the position of oligonucleotide primers used for PCR. d Confirmation of the replacement of the LsKU70 ORF region by the LsTDH3 promoter/kanR ORF region/LsTDH3 terminator DNA fragment by PCR. PCR amplification was performed with 5′KU70Fw and 3′KU70Rv primers and showed distinct bands representing different sizes (3.9 and 5.0 kb). The replacement event results in a 5.0-kb PCR product. e Confirmation of the replacement of the LsKU80 ORF region by the LsTDH3 promoter/hph ORF region/LsTDH3 terminator DNA fragment by PCR. PCR amplification was performed with 5′KU80Fw and 3′KU80Rv primers and showed distinct bands representing different sizes (5.0 and 5.7 kb). The replacement event results in a 5.0-kb PCR product. f Confirmation of the replacement of the LsLIG4 ORF region by the LsTDH3 promoter/Sb ble ORF region/LsTDH3 terminator DNA fragment by PCR. PCR amplification was performed with the 5′LIG4Fw and 3′LIG4Rv primers and showed distinct bands representing different sizes (5.4 and 7.4 kb). The replacement event results in a 5.4-kb PCR product
PCR was performed using the disruption cassette plasmid pKS-hph-LsKU80 as a template and the primer set 5′nonLsKU80Fw and 3′nonLsKU80Rv. The PCR product (LsKU80 disruption cassette) contained an LsKU80 flanking sequence (approximately 1 kbp) on either side of the hph expression cassette. The transformant was selected on YPD agar containing 100 µg/ml hygromycin B. PCR was performed using the obtained transformant genomic DNA as a template and the primer set 5′KU80Fw and 3′KU80Rv to confirm the correct insertion (Fig. 1b, e). We also confirmed the target disruption of LsKU80 by Southern blotting analysis of genomic DNA (Supplementary Fig. S1c, d).
PCR was performed using the disruption cassette plasmid pKS-sh ble-LsLIG4 as a template and the primer set 5′nonLsLIG4Fw and 3′nonLsLIG4Rv. The PCR product (LsLIG4 disruption cassette) contained an LsLIG4 flanking sequence (approximately 1 kbp) on either side of the Sh ble expression cassette. The transformant was selected on YPD agar containing 50 µg/ml zeocin. PCR was performed using the obtained transformant genomic DNA as a template and the primer set 5′LIG4Fw and 3′LIG4Rv to confirm the correct insertion (Fig. 1c, f). We also confirmed the target disruption of LsLIG4 by Southern blot analysis of genomic DNA (Supplementary Fig. S1g, h).
Similarly, an LsKU80 disruption cassette was inserted into the LsKU80 locus of LsKU70-deficient mutant (∆lsku70), to obtain ∆lsku70∆lsku80 mutant strain. An LsLIG4 disruption cassette was inserted into the LsLIG4 locus of LsKU70-deficient mutant (∆lsku70) or LsKU80-deficient mutant (∆lsku80), to obtain ∆lsku70∆lslig4 or ∆lsku80∆lslig4 mutant strain. We also confirmed the correct insertion by PCR and Southern blot analysis (Fig. 1a–f, Supplementary Fig. S1a–e, g, h).
Finally, an LsKU80 disruption cassette was inserted into the LsKU80 locus of LsKU70- and LsLIG4-deficient mutant (∆lsku70∆lslig4) to obtain the ∆lsku70∆lsku80∆lslig4 triple mutant. We confirmed the correct insertion by PCR and/or Southern blotting analysis (Fig. 1a–f, Supplementary Fig. S1a–c, f, g, h).
Construction of the disruption cassette plasmid pKS-sNAT1-LsURA3
Two PCR fragments of 5′- and 3′-non-coding regions of the LsURA3 gene amplified using the primer sets 5′nonLsURA3Fw and 5′nonLsURA3Rv, and 3′nonLsURA3Fw and 3′nonLsURA3Rv, respectively, were digested with HindIII or XbaI, and successively cloned into the HindIII site or XbaI site of the vector pKS-sNAT1 to yield disruption cassette plasmid pKS-sNAT1-LsURA3.
Homologous recombination (HR) frequency experiment
Primer sets of 5′nonUra3Fw-1000 bp and 3′nonUra3Rv-1000 bp, 5′nonUra3Fw-1500 bp and 3′nonUra3Rv-1500 bp, 5′nonUra3Fw-2000 bp and 3′nonUra3Rv-2000 bp, 5′nonUra3Fw-2500 bp and 3′nonUra3Rv-2500 bp, and 5′nonUra3Fw-3000 bp and 3′nonUra3Rv-3000 bp were used to amplify DNA fragments (LsURA3 disruption cassettes) for disrupting the LsURA3 gene using pKS-sNAT1-LsURA3 as a template. These PCR products contained the sNAT1 expression cassette with the 5′- and 3′-LsURA3 flanking sequences of 1000, 1500, 2000, 2500, and 3000 bp, respectively. To evaluate the HR frequency, these PCR products were used to transform the wild-type and the mutant strains (∆lslig4, ∆lsku70∆lslig4, and ∆lsku70∆lsku80∆lslig4). To determine whether the integrated LsURA3 gene had been introduced by HR or NHEJ, the primer sets of 5′Ura3Fw-1000 bp and 3′Ura3Rv-1000 bp, 5′Ura3Fw-1500 bp and 3′Ura3Rv-1500 bp, 5′Ura3Fw-2000 bp and 3′Ura3Rv-2000 bp, 5′Ura3Fw-2500 bp and 3′Ura3Rv-2500 bp, and 5′Ura3Fw-3000 bp and 3′Ura3Rv-3000 bp were used to detect the LsURA3 gene locus with 5′- and 3′-flanking sequences of 1000, 1500, 2000, 2500, and 3000 bp, respectively.
Lipid extraction and quantification
To measure the amount of intracellular lipid (mainly triglycerides), a 1 ml liquid culture was harvested, washed in distilled water three times, and re-suspended at <10 OD660 units/ml in distilled water. Then, 2.5 g of glass beads (1.0 mm in diameter) were added to 1 ml of cell suspension. The sample was then vortexed in a mixer (ASCM-1; Asone, Osaka, Japan) at 1800 rpm for 75 min at room temperature. Next, the mixture was separated by centrifugation at 15,000 rpm for 1 min and the supernatant was subjected to analysis using the TG E-test (Wako Pure Chemical) and the F-kit glycerol (Roche Diagnostics, Tokyo, Japan), in accordance with the manufacturer’s instructions. The intracellular lipid amount was determined as the difference in measured values obtained using the TG E-test and the F-kit glycerol.
Results and discussion
Identification of the target genes LsKU70, LsKU80, and LsLIG4
To enhance the efficiency of HR of L. starkeyi CBS1807 by impairing the function of Ku70p/Ku80p heterodimers and/or ligase IV of the competitive NHEJ pathway, we performed a BLAST search using Yarrowia lipolytica Ylku70p (UniProt Q6CCK2), Ylku80p (UniProt Q6C7B9), and Yllig4p (UniProt Q6C8A3) in the L. starkeyi genome database (http://genome.jgi.doe.gov/Lipst1_1/Lipst1_1.home.html). We found one each of those homologues and designated them as Lsku70p (Protein ID: 106743), Lsku80p (Protein ID: 71617), and Lslig4p (Protein ID: 2300) exhibited 25.9, 28.0, and 37.3% identity to Ylku70p, Ylku80p, and Yllig4p, respectively. Then, we designed their coding genes as LsKU70 (Transcript ID: 106743), LsKU80 (Transcript ID: 71617), and LsLIG4 (Transcript ID: 2300), respectively.
Alignment analyses of the predicted amino acid sequences for Lsku70p, Lsku80p, and Lslig4p, along with those of other yeast homologues from P. pastoris, Y. lipolytica, S. cerevisiae, and C. neoformans are depicted in Supplementary Fig. S2–S4. Structural domains in the target protein were annotated by PFAM domain assignment (Finn et al. 2016). The Lsku70p did not have a Ku70/80 C-terminal alpha/beta domain and SAP domain toward the C terminus, which is generally found in other ku70p homologues. When the amino acid sequence of Lsku70p was compared to other yeast ku70p homologues, similarities in conserved regions were found, although overall similarities were minor (Supplementary Fig. S2). A comparison of Lsku80p to other yeast ku80p homologues indicates high sequence similarity within two well-conserved domains, the Ku70/80 N-terminal alpha/beta domain and the Ku70/80 beta-barrel domain (Supplementary Fig. S3). The alignment of Lslig4p with other yeast Lig4p homologues reveals regions of identity or high similarity throughout the molecules (i.e. DNA ligase N-terminal domain, ATP-dependent DNA ligase domain, ATP-dependent DNA ligase C-terminal region, and BRCT domain), including many residues dispersed throughout the proteins, which are conserved in other yeast Lig4p homologues (Supplementary Fig. S4).
Construction and characterization of LsKU70-, LsKU80-, and/or LsLIG4-disrupted mutants
To develop an efficient gene-targeting method in L. starkeyi, we generated LsKU70-, LsKU80-, and LsLIG4-disrupted mutants (∆lsku70, ∆lsku80, and ∆lslig4) by HR using the LsKU70-, LsKU80-, and LsLIG4-disruption cassettes, respectively, constructed as described in “Materials and methods”. We also generated mutant L. starkeyi strains with deletions of both LsKU70 and LsKU80 genes (∆lsku70∆lsku80), both LsKU70 and LsLIG4 genes (∆lsku70∆lslig4), both LsKU80 and LsLIG4 genes (∆lsku80∆lslig4), and all three of these genes (∆lsku70∆lsku80∆lslig4).
When the wild-type and mutant strains (∆lsku70, ∆lsku80, ∆lslig4, ∆lsku70∆lsku80, ∆lsku70∆lslig4, ∆lsku80∆lslig4, and ∆lsku70∆lsku80∆lslig4) were grown at 30 °C on solid YPD medium, all mutant strains grew as well as the wild type (Supplementary Fig. S5, YPD), which suggested that the LsKU70, LsKU80, and LsLIG4 genes are not required for normal growth. Moreover, we compared the lipid productivity and the growth of the mutant strains (∆lsku70, ∆lsku80, ∆lslig4) to those of the wild type. These mutant strains, ∆lsku70, ∆lsku80, and ∆lslig4, showed similar lipid productivity and growth to the wild type (Supplementary Fig. S6a, b). Thus, we speculate that the deletions of Lsku70, Lsku80, and Lslig4 did not affect the lipid productivity.
As the Ku70p, Ku80p, or Lig4p homologue is involved in the repair of DNA damages in several yeasts, we investigated the sensitivity to the DNA alkylating agents methyl methanesulfonate (MMS), ethyl methanesulfonate (EMS), the topoisomerase I inhibitor camptothecin (CPT), and ultraviolet (UV) light in the wild type and the above-mentioned mutant strains. There were no significant differences in sensitivity among them (Supplementary Fig. S5). It has been reported that the different yeast species showed differential effects of KU70, KU80, and/or LIG4 mutations on the sensitivity to DNA damage stresses, such as MMS, EMS, CPT and UV. The sensitivities to MMS and UV are increased in KU70-deficient mutant of the oleaginous yeast Rhodosporidium toruloides (Koh et al. 2014). The Pichia pastoris ku70-deletion strain has also been shown to be hypersensitive to UV light (Naatsaari et al. 2012). In addition, the Ylku70- and/or Ylku80-deleted strains showed reduced cell viability in comparison with the wild type after UV irradiation in Y. lipolytica. Meanwhile, no significant difference of cell viability to the DNA damaging agent EMS was observed between the wild type and the Ylku70- and/or Ylku80-deleted strains (Kretzschmar et al. 2013). The mutants of MUS-52 (Ku80 homologue) and MUS-53 (Lig4 homologue) in Neurospora crassa are sensitive to MMS, but are not sensitive to CPT and UV (Ishibashi et al. 2006). The KU80- and LIG4-deletion mutants of Candida glabrata showed the same sensitivity to MMS, EMS, and UV as the wild type (Cen et al. 2015; Ueno et al. 2007). These differences in DNA damage stresses are considered to be responsible for differences in their detoxifying potency. Furthermore, these differences may imply that HR is important for the repair of DNA damage. In L. starkeyi, as none of the mutants produced by disrupting Lsku70, Lsku80, and/or Lslig4 showed altered sensitivities to MMS, EMS, CPT and UV relative to the wild-type control, the NHEJ pathway might not be a major pathway for the repair of MMS-, EMS-, CPT- or UV-induced damage. The relationship between those proteins (LsKu70p, LsKu80p, and LsLig4p) and the NHEJ pathway in the oleaginous yeast L. starkeyi remains to be determined in the future studies.
Homologous recombination frequency is significantly increased in ∆lslig4 background strains
NHEJ-deficient mutants have been reported to increase the frequency of HR in non-conventional yeasts (Cen et al. 2015; Choo et al. 2014; Foureau et al. 2013; Goins et al. 2006; Maassen et al. 2008; Naatsaari et al. 2012; Schorsch et al. 2009; Ueno et al. 2007). Therefore, we examined the gene-targeting efficiency of each strain (wild type, ∆lsku70, ∆lsku80, ∆lslig4) after transformation using the LsURA3 disruption cassette as described in “Materials and methods”. LsUra3p (Protein ID: 3918) encoded by LsURA3 (Transcript ID: 3918) is an orthologue of S. cerevisiae Ura3p (57.0% identity). LsURA3 was found to be capable of complementing a ura3 mutant in S. cerevisiae (unpublished data). We tested whether the HR frequency is also increased when the NHEJ pathway is disturbed in L. starkeyi. No transformants with a homologous integrated LsURA3 disruption cassette were obtained in the wild type. However, the HR frequency in ∆lsku70, ∆lsku80, or ∆lslig4 was increased (Table 3). The HR frequency of 17.9% (10 disruptants/56 transformants) in ∆lsku70 was about ninefold higher than that of 2.1% (1 disruptant/48 transformants) in ∆lsku80. This was similar to Y. lipolytica in which the HR of the ku70-disruptant was higher than that of the ku80-disruptant (Verbeke et al. 2013). Those differed from the rates of HR frequency in the ku70-deletion mutants which had a similar effect to ku80-deletion mutants in C. neoformans, A. sojae, and A. oryzae (Goins et al. 2006; Ishibashi et al. 2006). Thus, Lsku80p may have slightly more effect on the L. starkeyi NHEJ pathway than Lsku70p. Furthermore, it may also serve a different role in the L. starkeyi NHEJ pathway.
Moreover, a drastic increase in the frequency of HR (72.2%, 52 disruptants/72 transformants) was observed in ∆lslig4. Furthermore, we investigated the HR frequency in each of the ∆lslig4 background strains, ∆lsku70∆lslig4 and ∆lsku70∆lsku80∆lslig4. The HR frequency (80.6%, 58 disruptants/72 transformants) in ∆lsku70∆lslig4 was modestly increased when compared with that (72.2%, 52 disruptants/72 transformants) in ∆lslig4 (Table 3). There was no significant difference in the HR frequency between ∆lsku70∆lslig4 and ∆lsku70∆lsku80∆lslig4. These findings are supported by the critical role of Lig4p in the nonhomologous recombination pathway in other fungal species (Alshahni et al. 2011; Ishibashi et al. 2006; Schorsch et al. 2009). Meanwhile, the transformation efficiency in each of the ∆lslig4 background strains (∆lslig4, ∆lsku70∆lslig4, ∆lsku70∆lsku80∆lslig4) was nearly half or one-third that of the wild type (Fig. 2). Since the transformation efficiency of ∆lslig4 was the highest in the ∆lslig4 background strains, these results indicated that ∆lslig4 is excellent for genetic engineering in L. starkeyi as a recipient strain.
Efficiency of the transformation in Lipomyces starkeyi wild type, ∆lsku70, ∆lsku80, ∆lslig4, ∆lsku70∆lsku80, ∆lsku70∆lslig4, ∆lsku80∆lslig4, or ∆lsku70∆lsku80∆lslig4 strain. Twenty micrograms of each DNA fragment was added to 3.0 × 106 spheroplasts for transformation by the spheroplast-PEG method. Data are the means ± standard error of mean of three independent experiments in the bar graph
Effect of homologous flanking sequence length on HR frequency
To investigate the correlation between the HR frequency and the length of the flanking sequence, we constructed disruption cassettes with 1000, 1500-, 2000-, 2500-, and 3000-bp regions homologous to 5′- and 3′-flanking DNA of the LsURA3 gene. While no transformants with LsURA3 deletion were obtained using disruption cassettes with 1000 and 1500-bp homologous regions on each side, we obtained transformants with homologous integrated LsURA3 disruption cassettes with homology to 5′- and 3′-flanking sequences of more than 2000 bp in length (Table 4). These results indicated that the minimum homology length of 5′ and 3′ flanking DNA regions for HR in the wild-type strain was 2000 bp. The extension of homology length on each side of the disruption cassette increased the HR frequency at the LsURA3 locus. When the disruption cassette with 3000-bp homology regions was used for LsURA3 deletion, the rates of transformants generated by HR in the wild type, ∆lig4, ∆lsku70∆lslig4, and ∆lsku70∆lsku80∆lslig4 increased up to 11.1, 94.8, 95.2, and 95.8%, respectively (Table 4). In several yeasts, the lengths of 5′ and 3′ flanking sequences required for highly efficient gene targeting have been studied. The HR frequency for the P. pastoris ku70-deletion strain with a flanking sequence length of 650 bp was 100% in the HIS4 locus (Liang and Jiang 2013). Maximal efficiencies (70, 85.4, 95.7, and 91.7%) of HR in K. marxianus, Y. lipolytica, C. neoformans, and R. toruloides NHEJ-deficient strains were achieved with 5′ and 3′ flanking sequences with a 1000-bp homology length for HR (Koh et al. 2014; Kretzschmar et al. 2013; Liang and Jiang 2013). Similarly, HR with high frequencies (70.8–83.3%) was achieved with flanking regions with a 1000-bp homology length in the L. starkeyi ∆lig4 background strains.
The possibility of 5-fluoroorotic acid (5-FOA)-resistant LsURA3 deletion mutant as a useful host the genetic engineering of L. starkeyi
As indicated above, LsURA3-deficient mutants were generated using disruption cassettes with 1000 bp of homology length of the 5′- and 3′-flanking DNA sequences of the LsURA3. The structure of the LsURA3 locus in one of the LsUra3p-deficient mutants, ∆lslig4∆lsura3, was verified by Southern blot analysis (Supplementary Fig. S7), which confirmed that integration had occurred correctly. The wild-type and ∆lslig4∆lsura3 cells were plated onto SD, SD supplemented with uracil, and SD supplemented with uracil and 5-FOA. The ∆lslig4∆lsura3 strain did not grow on SD, but did on SD containing uracil (Fig. 3b). Thus, the ∆lslig4∆lsura3 was shown to be a uracil auxotroph. In most yeast species, because an orotidine 5′-phosphate decarboxylase (Ura3p) converts 5-FOA to one or more toxic intermediates, 5-FOA is toxic to wild-type cells (Boeke et al. 1984). The growth of L. starkeyi wild type was inhibited at a concentration of 1 mg/ml 5-FOA. However, the ∆lslig4∆lsura3 strain could grow in the presence of 1 mg/ml 5-FOA. It is thus suggested that LsUra3p has orotidine 5′-phosphate decarboxylase activity. Furthermore, ∆lslig4∆lsura3 has potential as a host using the URA3-blaster system, which allows both positive and negative selection, based on it being an auxotroph for uracil and resistant to 5-FOA (Alani et al. 1987).
Uracil auxotroph and 5-FOA resistance of L. starkeyi wild type, ∆lsku70, and ∆lsku70∆lsura3. The wild-type, ∆lsku70, and ∆lsku70∆lsura3 cells were pre-cultured for 2 days at 30 °C in SD or SD supplemented with 20 mM uracil. The wild-type and mutant cells were inoculated with a concentration OD600 = 0.1 in SD or SD supplemented with 20 mM uracil liquid medium and cultured for 2 days at 30 °C. Ten-fold serially diluted cultured cells were spotted onto agar plates with SD, SD supplemented with 20 mM uracil, or selective 5-FOA (SD containing 1.5 mg/ml 5-FOA and 20 mM uracil), and then grown for 5 days at 30 °C
Conclusion
We have successfully improved the gene-targeting efficiency in the oleaginous yeast L. starkeyi using ∆lslig4 background strains. The growth and lipid productivity did not differ between the wild type and ∆lig4 under normal conditions. ∆lslig4 also showed the highest transformation efficiency among the ∆lslig4 background strains. These features are adequate for a host strain for use in genetic engineering. Furthermore, the gene-targeting system in this study should contribute to revealing the mechanisms of lipid biosynthesis, degradation, and accumulation and to improve the lipid productivity via metabolic engineering in the oleaginous yeast L. starkeyi.
Abbreviations
- YPD:
-
Yeast extract/peptone/dextrose
- PCR:
-
Polymerase chain reaction
- OD:
-
Optical density
References
Ageitos JM, Vallejo JA, Veiga-Crespo P, Villa TG (2011) Oily yeasts as oleaginous cell factories. Appl Microbiol Biotechnol 90:1219–1227
Alani E, Cao L, Kleckner N (1987) A method for gene disruption that allows repeated use of URA3 selection in the construction of multiply disrupted yeast strains. Genetics 116:541–545
Alshahni MM, Yamada T, Takatori K, Sawada T, Makimura K (2011) Insights into a nonhomologous integration pathway in the dermatophyte Trichophyton mentagrophytes: efficient targeted gene disruption by use of mutants lacking ligase IV. Microbiol Immunol 55:34–43
Angerbauer C, Siebenhofer M, Mittelbach M, Guebitz GM (2008) Conversion of sewage sludge into lipids by Lipomyces starkeyi for biodiesel production. Bioresour Technol 99:3051–3056
Baudin A, Ozier-Kalogeropoulos O, Denouel A, Lacroute F, Cullin C (1993) A simple and efficient method for direct gene deletion in Saccharomyces cerevisiae. Nucleic Acids Res 21:3329–3330
Bimboim H, Doly J (1979) A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res 7:1513–1523
Boeke JD, LaCroute F, Fink GR (1984) A positive selection for mutants lacking orotidine-5′-phosphate decarboxylase activity in yeast: 5-fluoro-orotic acid resistance. Mol Gen Genet 197:345–346
Calvey CH, Willis LB, Jeffries TW (2014) An optimized transformation protocol for Lipomyces starkeyi. Curr Genet 60:223–230
Carroll A, Sweigard J, Valent B (1994) Improved vectors for selecting resistance to hygromycin. Fungal Genet Newsl 41: 22
Cen Y, Fiori A, Van Dijck P (2015) Deletion of the DNA Ligase IV Gene in Candida glabrata Significantly Increases Gene-Targeting Efficiency. Eukaryot Cell 14:783–791
Chen L, Trujillo K, Ramos W, Sung P, Tomkinson AE (2001) Promotion of Dnl4-catalyzed DNA end-joining by the Rad50/Mre11/Xrs2 and Hdf1/Hdf2 complexes. Mol Cell 8:1105–1115
Chiruvella KK, Liang Z, Wilson TE (2013) Repair of double-strand breaks by end joining. Cold Spring Harbor Perspect Biol 5: a012757
Choo JH, Han C, Kim JY, Kang HA (2014) Deletion of a KU80 homolog enhances homologous recombination in the thermotolerant yeast Kluyveromyces marxianus. Biotechnol Lett 36:2059–2067
Feldmann E, Schmiemann V, Goedecke W, Reichenberger S, Pfeiffer P (2000) DNA double-strand break repair in cell-free extracts from Ku80-deficient cells: implications for Ku serving as an alignment factor in non-homologous DNA end joining. Nucleic Acids Res 28:2585–2596
Finn RD, Coggill P, Eberhardt RY, Eddy SR, Mistry J, Mitchell AL, Potter SC, Punta M, Qureshi M, Sangrador-Vegas A (2016) The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res 44:D279–D285
Foureau E, Courdavault V, Rojas LF, Dutilleul C, Simkin AJ, Creche J, Atehortua L, Giglioli-Guivarc’h N, Clastre M, Papon N (2013) Efficient gene targeting in a Candida guilliermondii non-homologous end-joining pathway-deficient strain. Biotechnol Lett 35:1035–1043
Giaever G, Chu AM, Ni L, Connelly C, Riles L, Veronneau S, Dow S, Lucau-Danila A, Anderson K, Andre B, Arkin AP, Astromoff A, El-Bakkoury M, Bangham R, Benito R, Brachat S, Campanaro S, Curtiss M, Davis K, Deutschbauer A, Entian KD, Flaherty P, Foury F, Garfinkel DJ, Gerstein M, Gotte D, Guldener U, Hegemann JH, Hempel S, Herman Z, Jaramillo DF, Kelly DE, Kelly SL, Kotter P, LaBonte D, Lamb DC, Lan N, Liang H, Liao H, Liu L, Luo C, Lussier M, Mao R, Menard P, Ooi SL, Revuelta JL, Roberts CJ, Rose M, Ross-Macdonald P, Scherens B, Schimmack G, Shafer B, Shoemaker DD, Sookhai-Mahadeo S, Storms RK, Strathern JN, Valle G, Voet M, Volckaert G, Wang CY, Ward TR, Wilhelmy J, Winzeler EA, Yang Y, Yen G, Youngman E, Yu K, Bussey H, Boeke JD, Snyder M, Philippsen P, Davis RW, Johnston M (2002) Functional profiling of the Saccharomyces cerevisiae genome. Nature 418:387–391
Goins CL, Gerik KJ, Lodge JK (2006) Improvements to gene deletion in the fungal pathogen Cryptococcus neoformans: absence of Ku proteins increases homologous recombination, and co-transformation of independent DNA molecules allows rapid complementation of deletion phenotypes. Fungal Genet Biol 43:531–544
Hereford L, Fahrner K, Woolford J, Rosbash M, Kaback DB (1979) Isolation of yeast histone genes H2A and H2B. Cell 18:1261–1271
Herrmann G, Lindahl T, Schar P (1998) Saccharomyces cerevisiae LIF1: a function involved in DNA double-strand break repair related to mammalian XRCC4. EMBO J 17:4188–4198
Ishibashi K, Suzuki K, Ando Y, Takakura C, Inoue H (2006) Nonhomologous chromosomal integration of foreign DNA is completely dependent on MUS-53 (human Lig4 homolog) in Neurospora. Proc Natl Acad Sci 103:14871–14876
Ishidoh K-i, Kinoshita H, Ihara F, Nihira T (2014) Efficient and versatile transformation systems in entomopathogenic fungus Lecanicillium species. Curr Genet 60:99–108
Kikukawa H, Sakuradani E, Nakatani M, Ando A, Okuda T, Sakamoto T, Ochiai M, Shimizu S, Ogawa J (2015) Gene targeting in the oil-producing fungus Mortierella alpina 1S-4 and construction of a strain producing a valuable polyunsaturated fatty acid. Curr Genet 61:579–589
Koh CM, Liu Y, Moehninsi, Du M, Ji L (2014) Molecular characterization of KU70 and KU80 homologues and exploitation of a KU70-deficient mutant for improving gene deletion frequency in Rhodosporidium toruloides. BMC Microbiol 14:50
Kosa M, Ragauskas AJ (2011) Lipids from heterotrophic microbes: advances in metabolism research. Trends Biotechnol 29:53–61
Kretzschmar A, Otto C, Holz M, Werner S, Hubner L, Barth G (2013) Increased homologous integration frequency in Yarrowia lipolytica strains defective in non-homologous end-joining. Curr Genet 59:63–72
Liang MH, Jiang JG (2013) Advancing oleaginous microorganisms to produce lipid via metabolic engineering technology. Prog Lipid Res 52:395–408
Lobachev K, Vitriol E, Stemple J, Resnick MA, Bloom K (2004) Chromosome fragmentation after induction of a double-strand break is an active process prevented by the RMX repair complex. Curr Biol 14:2107–2112
Maassen N, Freese S, Schruff B, Passoth V, Klinner U (2008) Nonhomologous end joining and homologous recombination DNA repair pathways in integration mutagenesis in the xylose-fermenting yeast Pichia stipitis. FEMS Yeast Res 8:735–743
Milne GT, Jin S, Shannon KB, Weaver DT (1996) Mutations in two Ku homologs define a DNA end-joining repair pathway in Saccharomyces cerevisiae. Mol Cell Biol 16:4189–4198
Moore JK, Haber JE (1996) Cell cycle and genetic requirements of two pathways of nonhomologous end-joining repair of double-strand breaks in Saccharomyces cerevisiae. Mol Cell Biol 16:2164–2173
Naatsaari L, Mistlberger B, Ruth C, Hajek T, Hartner FS, Glieder A (2012) Deletion of the Pichia pastoris KU70 homologue facilitates platform strain generation for gene expression and synthetic biology. PLoS One 7:e39720
Oguro Y, Yamazaki H, Shida Y, Ogasawara W, Takagi M, Takaku H (2015) Multicopy integration and expression of heterologous genes in the oleaginous yeast, Lipomyces starkeyi. Biosci Biotechnol Biochem 79:512–515
Papanikolaou S, Aggelis G (2011) Lipids of oleaginous yeasts. Part I: biochemistry of single cell oil production. Eur J Lipid Sci Technol 113:1031–1051
Ratledge C, Wynn JP (2002) The biochemistry and molecular biology of lipid accumulation in oleaginous microorganisms. Adv Appl Microbiol 51:1–51
Rothstein R (1991) Targeting, disruption, replacement, and allele rescue: integrative DNA transformation in yeast. Methods Enzymol 194:281–301
Schorsch C, Kohler T, Boles E (2009) Knockout of the DNA ligase IV homolog gene in the sphingoid base producing yeast Pichia ciferrii significantly increases gene targeting efficiency. Curr Genet 55:381–389
Takata M, Sasaki MS, Sonoda E, Morrison C, Hashimoto M, Utsumi H, Yamaguchi-Iwai Y, Shinohara A, Takeda S (1998) Homologous recombination and non-homologous end-joining pathways of DNA double-strand break repair have overlapping roles in the maintenance of chromosomal integrity in vertebrate cells. EMBO J 17:5497–5508
Teo SH, Jackson SP (1997) Identification of Saccharomyces cerevisiae DNA ligase IV: involvement in DNA double-strand break repair. EMBO J 16:4788–4795
Ueno K, Uno J, Nakayama H, Sasamoto K, Mikami Y, Chibana H (2007) Development of a highly efficient gene targeting system induced by transient repression of YKU80 expression in Candida glabrata. Eukaryot Cell 6:1239–1247. doi:10.1128/EC.00414-06
Valencia M, Bentele M, Vaze MB, Herrmann G, Kraus E, Lee SE, Schar P, Haber JE (2001) NEJ1 controls non-homologous end joining in Saccharomyces cerevisiae. Nature 414:666–669
Verbeke J, Beopoulos A, Nicaud JM (2013) Efficient homologous recombination with short length flanking fragments in Ku70 deficient Yarrowia lipolytica strains. Biotechnol Lett 35:571–576
Wach A, Brachat A, Pohlmann R, Philippsen P (1994) New heterologous modules for classical or PCR-based gene disruptions in Saccharomyces cerevisiae. Yeast 10:1793–1808
Wilson TE, Grawunder U, Lieber MR (1997) Yeast DNA ligase IV mediates non-homologous DNA end joining. Nature 388:495–498
Acknowledgements
We thank Dr. Marie Nishimura for kindly providing the plasmid. This study was supported in part by a grant of the Science and Technology Research Promotion Program for Agriculture, Forestry, Fisheries and Food Industry, a grant of the Development of Production Techniques for Highly Functional Biomaterials Using Smart Cells of Plants and Other Organisms, a grant from JSPS KAKENHI (Grant Number 15K07372), and a grant from Nagase Science Technology Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Oguro, Y., Yamazaki, H., Ara, S. et al. Efficient gene targeting in non-homologous end-joining-deficient Lipomyces starkeyi strains. Curr Genet 63, 751–763 (2017). https://doi.org/10.1007/s00294-017-0679-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-017-0679-6