Abstract
Lung cancer is the leading cause of cancer death in the world. Recently, targeted therapy and anti-programmed cell death receptor 1 (PD-1) and anti-programmed cell death ligand 1 (PD-L1) immunotherapy have made great progress in treatment of lung cancer. However, responses to these therapies are variable, influenced by genetic alterations, high microsatellite instability and mismatch repair deficiency. Liquid biopsy of extracellular vesicles and circulating tumor DNA (ctDNA) emerges as a new promising non-invasive means that enables not only biomarker determination, but also continuous monitoring of cancer treatment. Notably, tumor extracellular vesicles play important roles in tumor formation and progression, and also serve as natural carriers for anti-tumor drugs and short-interfering RNA. In this review, we summarize the latest progress in understanding the relationships of extracellular vesicles and ctDNA in cancer biology, diagnosis and drug delivery. In particular, the application of extracellular vesicles and ctDNA in anti-PD-1/PD-L1 immunotherapy is discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer is one of the most prevalent cancers and the leading cause of cancer-related death worldwide [1, 2]. More than half of lung cancer patients have an advanced form of the disease at the time of diagnosis [2]. Delayed detection and a lack of reliable biomarkers and new drugs are major causes of the high mortality of lung cancer [3]. Despite the high death rate, a large number of patients with non-small cell lung cancer (NSCLC) can benefit from targeted therapies for gene alterations such as EGFR, ALK and ROS1 [4]. Recently, anti-programmed cell death receptor 1 (PD-1) and anti-programmed cell death ligand 1 (PD-L1) immunotherapy, which may be more efficient for cancer patients who have high PD-L1 expressions, high microsatellite instability (MSI-H) or mismatch repair deficiency (dMMR) [5, 6], has made great progress as a form of NSCLC treatment. The detection of genetic alterations, PD-L1 expression, MSI and dMMR status in patients with lung cancer, especially for NSCLC, is critical for subsequent therapy. However, it is always difficult and sometimes impossible to obtain tumor tissues from patients. Further, for patients suffering from a relapse after receiving chemotherapy or targeted therapy, the original somatic mutation may change, but a repeated biopsy is very difficult to obtain [7].
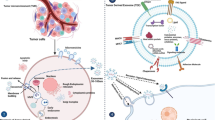
Extracellular vesicles (also called exosomes) were described as microvesicles with 5′-nucleotidase activity, released from neoplastic cells and other cells [8]. These vesicles usually display diameters of 40–120 nm, and are produced from multivesicular bodies (MVBs). Several mechanisms have been suggested to mediate the uptake of exosomes, including an exosome fusion with the cellular membrane of the recipient cell, juxtacrine signaling through receptor–ligand interactions and endocytosis by phagocytosis [9] (Fig. 1). They usually contain the cytosolic milieu, including proteins, lipids, RNA, and DNA, appearing as ‘mini-mes’ of the parent cell [10]. The composition of extracellular vesicles, including many kinds of important biomacromolecules as well as their relative stability makes them ideal candidates for clinical diagnosis [11].
Biogenesis and genetic information foundation of tumor extracellular vesicles and ctDNA. Exosomes are produced from endosomes. The membrane of the endosomes bulges inward to form exosomes. During this process, proteins, nucleic acids and lipids are packed into exosomes. The causes for ctDNA release include apoptosis, necrosis from dying tumor cells, and active release from viable tumor cells through secretion. Occurrence of gene mutations and the statuses of microsatellite instability (MSI), mismatch repair deficiency (dMMR) and methylation can be detected in order to monitor progress of cancer as well as provide guidance for targeted therapy or immunotherapy
Circulating tumor DNA (ctDNA) is tumor-derived fragmented DNA found in the bloodstreams of cancer patients. The mechanisms of ctDNA release include apoptosis and necrosis from dying cells, as well as cell lysis of viable tumor cells [12, 13]. Cell-free DNA (cfDNA), is a broader category, which describes any DNA that is freely circulating in the bloodstream. It is commonly accepted that ctDNA is a subcategory of cfDNA, distinguished by that fact that it is released from tumor cells [13]. It has been demonstrated that analysis of ctDNA enables non-invasive monitoring of tumor evolution over time in clinical practice [14].
In this review, the latest progress in exosome and ctDNA cancer biology, diagnosis and drug delivery is summarized. The application of exosome and ctDNA analysis for choice of anti-PD-1/PD-L1 immunotherapy is also discussed.
Extracellular vesicles in lung cancer
Role of extracellular vesicles in lung cancer
Cancer cells usually secrete factors into their microenvironment to promote their own growth and protect against attacks by the immune system [15]. Exosomes, released from tumor cells, represent certain important characteristics of tumors, and also are reported to play roles in tumor promotion (Fig. 2) [16,17,18]. In the initial phase of cancer, exosomes from malignant cells have the potential to induce transformation of normal cells, playing a role in tumorigenesis. For example, Abd et al. found [19] that exosomes from prostate cancer (PC) cells subvert adipose-derived stem cells to undergo neoplastic transformation. The underlying mechanisms include the downregulation of LATS2 and PDCD4, and association with trafficking by PC cell-derived exosomes of oncogenic factors, including H-ras and K-ras transcripts, onco-miRNAs, as well as the Ras superfamily of GTPases. Melo et al. [20] reported that exosomes derived from cells and sera of patients with breast cancer, which contained pre-miRNAs, RISC complex and Dicer, could cause tumorigenesis in epithelial cells. In this way, cancer extracellular vesicles mediate efficient and rapid silencing of mRNAs to reprogram the target cell transcriptome.
Roles of tumor extracellular vesicles in cancer. Extracellular vesicles play important roles in tumor cell proliferation, mediating drug resistance, evading immune destruction, promoting inflammation, activating invasion and metastasis, inducing angiogenesis, resisting death and deregulating cellular energetics. Most hallmarks of cancer, with the exception of genome instability, can be promoted by tumor extracellular vesicles [16,17,18]
The exosomes also play important roles in proliferation of lung cancer cells. Using a mast cell line HMC-1 constitutively expressing the active form of the KIT receptor, and a NSCLC cell line A549, Xiao et al. [21] found that mast cell exosomes promoted the proliferation of lung adenocarcinoma cells. The exosomes accomplished this by transferring KIT to tumor cells, which initiated receptor–ligand interactions in recipient cells.
Studies show that extracellular vesicles participate in angiogenesis, which is a key requirement for tumor growth. Inhibition of angiogenesis using monoclonal antibodies blocking VEGF–VEGFR binding or small molecule tyrosine kinase inhibitors (TKIs) that inhibit the downstream VEGFR-mediated signaling is an important targeted therapy in NSCLC. Zhuang et al. [22] showed that tumor-secreted exosomes participated in intercellular communication and promoted angiogenesis as well as migration in A549 cells. Mechanistic studies revealed that the exosomal miR-9 secreted by tumor cells activated the JAK-STAT pathway. Moreover, hypoxic multiple myeloma cells produce exosomes containing miR-135b, which directly inhibited HIF-1 and thus enhances tumor angiogenesis [23].
Cai et al. [24] reported that activated T cell extracellular vesicles promoted tumor invasion by increasing the expression of MMP9 in lung cancer and melanoma, revealing a new mechanism of tumor immune escape that is mediated by exosomes. It has been found [25] that exosomal integrins are key tumor factors that regulate organotropic metastasis. Tumor-derived exosomes taken up by organ-specific cells prepare the pre-metastatic niche, and treatment with exosomes from lung-tropic models redirects the metastasis of bone-tropic tumor cells. Rahman et al. [26] observed that exosomes derived from highly metastatic lung cancer cells and serum of patients with distant metastasis in lung cancer induced vimentin expression and epithelial-to-mesenchymal transition (EMT) in human bronchial epithelial cells (HBECs).
Fabbri et al. [27] showed that tumor-secreted exosomal miR-21 and miR-29a bound as ligands to receptors of the toll-like receptor (TLR) family, murine TLR7 and human TLR8, in immune cells, triggering a TLR-mediated prometastatic inflammatory response and ultimately leading to tumor growth and metastasis. Zhou et al. [28] demonstrated that miR-105, which is characteristically expressed and secreted by metastatic breast cancer cells through exosomes, is a potent regulator of migration through targeting the tight junction protein ZO-1. Exosome-mediated transfer of cancer-secreted miR-105 efficiently destroyed the integrity of tight junctions against metastasis. Moreover, miR-105 could be detected in the circulation at the premetastatic stage, and its levels in blood and tumor tissue were associated with ZO-1 expression and metastatic progression in early-stage breast cancer. As paracrine agonists of TLRs, secreted miRNAs were found to be key regulators of the tumor microenvironment. This mechanism of action by exosomal miRNAs is implicated in tumor–immune system interactions, which are important for tumor growth and spread.
Exosomes are also important mediators of drug resistance in lung cancer. Feng and his colleagues found that lung cancer cells A549 increased their exosome secretion after cisplatin treatment, and these exosomes could reduce sensitivity of receptor cells to cisplatin [29]. Further study showed that cisplatin resistance develops in an exosomal miR-100-5p-dependent manner with mTOR as its potential target both in vitro and in vivo [30]. Incorporation of paclitaxel into exosomes increases antineoplastic activity by more than 50 times in multiple drug-resistant tumor cells, including lung cancer cells [31]. An increasing number of reports reveal that long non-coding RNAs (lncRNAs) are critical regulators of diverse biological processes [32, 33]. A recent study conducted by Le et al. [34] revealed that lncARSR can be incorporated into exosomes and transmitted to sensitive cells, thus disseminating sunitinib resistance. After being transferred into sensitive cells, lncARSR promoted sunitinib resistance via competitively binding miR-34/miR-449 to facilitate AXL and c-MET expression in renal cancer cells.
Extracellular vesicles as biomarkers in lung cancer
As previously mentioned, extracellular vesicles contain proteins, nucleic acids and lipids from tumor cells and provide an indication of many characteristics of tumors. They are ideal biomarkers, facilitating early diagnosis of lung cancer. The components of exosomes that could serve as biomarkers have been identified and characterized in several recent studies (Table 1).
Wang et al. [35] investigated differentially expressed exosomal miRNAs in pleural effusions of lung adenocarcinoma, tuberculous, and other benign lesions by using deep sequencing and quantitative polymerase chain reaction. As a result, they identified nine miRNAs which were preferentially represented in exosomes derived from the pleural effusions of lung adenocarcinoma in this study. Jakobsen et al. [36] profiled the exosome proteins from the plasma isolated from 109 NSCLC patients with advanced stage (IIIA–IV) disease and 110 matched control subjects. Using the random forests method and extracellular vesicle arrays containing 37 antibodies targeting lung cancer-related proteins, the authors established a combined 30-marker model which classifies patients with a 75.3% rate of accuracy. A similar strategy was adopted by Sandfeld-Paulsen et al. [37]. By analyzing proteins attached to the exosomal membrane, they found that nine exosomal proteins show potential as prognostic markers in NSCLC. Among them, NY-ESO-1 was a strong prognostic biomarker in NSCLC [hazard rate (HR) 1.78 95% (1.78–2.44); p = 0.0001] after Bonferroni correction.
Besides the exosomal proteins, the exosomal RNA such as miRNA has been extensively investigated in recent years [38, 39]. miRNAs are ~ 22 nt noncoding RNAs that play important roles in regulating gene expression mainly at the posttranscriptional level [40, 41]. Because of their low molecular weights, miRNAs are much more stable than other RNA such us mRNA and lncRNAs in the circulating system, which makes them ideal biomarkers for cancer diagnosis [42, 43].
Rabinowits et al. [44] conducted a study that enrolled 27 lung adenocarcinoma patients and nine controls to study the expression of circulating exosome miRNAs. They found the mean miRNA concentrations from lung adenocarcinoma were much higher than those of the control group (158.6 vs. 68.1 ng/mL, p < 0.001). This significant difference in total exosomal miRNA levels between lung cancer patients and controls indicated that circulating exosomal miRNA might be useful as a screening test for lung adenocarcinoma. However, a correlation between exosomal miRNA levels and disease stage was not observed, which may be due to limited sample size.
Using 30 plasma samples for wide-range miRNAs analysis, Cazzoli et al. [45] identified four miRNAs as biomarkers to distinguish lung adenocarcinoma and carcinoma patients from healthy former smokers. They also identified six miRNAs that segregated lung adenocarcinoma and lung granuloma patients. Furthermore, exosomal miRNA profiling in culture media of normal lung cells Beas-2b and lung cancer H1299 and in subcutaneous primary and recurrent lung cancer xenografts in nude mouse models [46] showed that a total of 77 miRNAs were observed to be significantly modulated in the H1299 cells, and two miRNAs, miR-21 and miR-155, were significantly upregulated in recurrent tumors compared to primary tumors.
Extracellular vesicles as tumor therapeutic cargos
As natural membrane-based vesicles, the lipid bilayer membrane of exosomes forms a natural protective shelter and a sustained release capsule for various anti-cancer drugs [47]. Unlike liposomes and other synthetic drug nanoparticle carriers, exosomes contain membrane-anchored proteins that may enhance endocytosis, and thus have higher efficiency for delivering their internal contents [48, 49]. Moreover, as the natural products of cells, exosomes are biocompatible and biodegradable, and thus have low toxicity and immunogenicity [50]. These characteristics of exosomes make them ideal tumor therapeutic cargos.
Using bovine milk as a scalable source of exosomes, Munagala et al. [51] reported that withaferin A shows enhanced anti-cancer and anti-inflammatory effects by exosomal delivery compared to free withaferin A in lung cancer cell lines and lung tumor xenografts in vivo. In another study [52], celastrol, a plant-derived triterpenoid inhibitor of Hsp90 and NF-κB pathways, displayed therapeutic importance in various cancers. A stronger anti-tumor effect of exosome-loaded celastrol compared to free drug was observed in human lung cancer cell xenograft model. Paclitaxel is one of most wildly used chemotherapy drugs in various cancers including lung cancer. Batrakova and her colleagues demonstrated [31] that incorporation of paclitaxel into exosomes increases antineoplastic activity by more than 50 times in multiple drug resistance cells. Similar results were obtained in a model of murine Lewis lung carcinoma pulmonary metastases.
siRNAs and miRNAs are powerful tools that can downregulate interesting oncogenes and inhibit tumor growth and metastasis. In lung adenocarcinoma cancer mice model, Xue et al. [53] reported that the small RNA combination of miR-34a and K-ras siRNA improves therapeutic responses over those observed with either small RNA alone. Furthermore, small RNA combination plus cisplatin-based chemotherapy prolongs survival in this model compared with chemotherapy alone. However, delivery of siRNA and miRNAs remain significant challenges and thus hamper their applications. Poor stability and short half-life in biological systems are big problems in miRNAs and siRNA delivery. As natural carriers of DNA and RNA, exosomes can protect nucleic acids from various DNases and RNases as well as carry them to target cells. Studies have shown that exosomes do not result in unwanted accumulation of therapeutic cargo in the liver, which is a negative side effect usually observed with nanoparticle therapeutic delivery systems. At the same time, exosomes can completely bypass the liver [54]. Surface proteins present on exosomes provide them with the intrinsic ability to enter into target tissues. In contrast, artificial carriers lack surface proteins, and because of this they are unable to fuse with target cells [55]. For these reasons, exosomes are ideal carriers for delivering siRNA and miRNA to tumors for cancer therapy. An increasing number of studies are being conducted to test the usefulness of exosomes as therapeutic vehicles to successfully deliver miRNAs and siRNAs.
Alvarez-Erviti et al. [56] conducted a study of delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Shtam et al. [57] reported that using exosomes to deliver siRNA targeting RAD51 significantly suppresses the proliferation of cells and causes cell death in vitro. Very recently, it was found [58] that exosomes derived from normal fibroblast-like mesenchymal cells carrying siRNA targeting mutated K-rasG12D, a common mutation in NSCLC and pancreatic cancer, could facilitate therapeutic targeting of oncogenic K-RAS in pancreatic cancer. Compared to liposomes, the siRNA-loaded exosomes target oncogenic K-RAS with an enhanced efficacy that is dependent on CD47, and is facilitated by micropinocytosis. Treatment with siRNA-loaded exosomes suppresses cancer in multiple mouse models of pancreatic cancer, and significantly increases overall survival. This study not only suggests that exosomes exhibit a superior ability to deliver RNAi and suppress tumor growth compared to liposomes, but also offers insight into the therapeutic potential of exosomes in specific targeting of mutated oncogenes, which may minimize the toxicity of siRNAs.
Let-7 is a well-known tumor-suppressing miRNA that can inhibit expression of several oncogenic targets including K-ras [59]. SNPs in the let-7 complementary sites in the 3′UTR of K-ras results in increased levels of K-ras and poor prognosis for patients with lung and breast cancer [60, 61]. Shimbo et al. reported that EGFR-specific binding peptide GE11 could guide Let-7a-containing exosomes to EGFR-positive cancer cells, which dramatically inhibits EGFR-positive human breast cancer cell growth in a xenograft mouse model [55]. The mutation of K-ras is an important cause for resistance to EGFR-TKI [62], a preferred choice of EGFR-mutant NSCLC patients. The let-7-loaded exosomes may provide a novel therapy that may specifically benefit these patients. Moreover, as mentioned above, exosomes that contain mutated Kras-specific siRNA may help reduce TKI-resistance caused by the mutation of K-ras.
Brain metastasis is an important cause of poor prognosis in lung cancer. Approximately 7.4% of NSCLC patients have brain metastases at presentation, and 30–50% develop brain metastases during the course of their disease [63]. For these patients, it is challenging to find a means of delivering drug across the blood–brain barrier (BBB). Because most antineoplastic agents have difficulty crossing the BBB and reaching sufficient concentrations, these drugs usually fail to benefit these patients [64]. Many nanoparticles-based approaches have been introduced to boost intracerebral drug concentration. However, other problems, such as nano-toxicity and rapid drug clearance by the mononuclear phagocyte system (MPS), have been reported [65]. Because exosomes are intracellular membrane-based vesicles from the body’s own cells, they can be tailored to cross the BBB, thus improving intracerebral drug concentrations by decreasing MPS drug clearance [50]. Using zebrafish as an animal mode, Yang et al. [66] delivered anticancer drugs by exosomes derived from brain endothelial cells, showing the potential of exosomes for brain delivery across the BBB and explaining their transport mechanisms.
Extracellular vesicles as tumor therapeutic agents
Besides functioning as tumor therapeutic cargo, due to preserving the molecular composition that bestows them with potent immunostimulatory properties, exosomes from immune cells exhibit anti-tumor activity on their own. For example, dendritic cell (DC)-derived exosomes (Dex) maintain the key functions of DCs in their ability to present tumor-associated antigens (TAAs) and to activate TAA-specific immune responses. The outer membrane of these exosomes contains various antigen presentation (MHC class I, class II, CD1), adhesion (ICAMs), costimulatory (CD86, CD40), and docking (integrin) molecules, which facilitate the in vivo functionality of DC-derived exosomes [67, 68]. Hence, over the last decade, Dex have been developed as clinical cell-free cancer vaccines [69, 70]. Dex-based phase I and II clinical trials have been conducted in advanced malignancies, showing the feasibility and safety of the approach, as well as the propensity of these nanovesicles to mediate T and NK cell-based immune responses in patients [71].
In 2005, a phase I clinical trial using TAA-loaded DC-derived exosomes was conducted in NSCLC patients [72]. Recently, the phase II clinical trial using DC-derived exosomes for maintenance immunotherapy after first-line chemotherapy in NSCLC was reported by Besse et al. [73] (NCT01159288). An important innovation in phase II clinical trial is the use of exosomes derived from TLR4L- or IFN-γ-maturated DCs, following discoveries that such Dex induce greater T cell stimulation compared to DC-derived exosomes from immature DCs. Seven patients (32%) experienced stabilization of their disease for over 4 months. One patient had a grade-3 hepatotoxicity. The phase II trial confirmed the capacity of Dex to boost the NK cell arm of antitumor immunity in patients with advanced NSCLC.
Taken together, these results show that as natural products of cells, exosomal carriers provide advantages of both cell-based drug delivery and nanotechnology for efficient drug transport capable of overcoming various biological barriers [74]. However, several limitations need to be further addressed. One of the major difficulties is the efficient loading of extracellular vesicles with a therapeutic agent without significant changes in the structure and content of exosomal membranes. Another challenge is identifying and removing those tumor-supporting components from extracellular vesicles, which is critical for exosome-mediated cancer therapy [47].
ctDNA as biomarkers in lung cancer
As mentioned above, detection of ctDNA in plasma or serum as biomarker can prevent the need for tumor tissue biopsies, which is advantageous for numerous diagnostic applications. Such a liquid biopsy facilitates repeated blood sampling and thus allows for monitoring of the changes during the natural course of the disease or during cancer treatment [75]. There are several problems that remain to be solved before its application. The major challenge is the sensitivity and specificity of ctDNA detection, exacerbated by its low abundance in blood compared to cfDNA from normal cells [76].
Many attempts have been made to address these issues. Aaron et al. [77] developed a method called CAPP-Seq for NSCLC with a design covering multiple classes of somatic alterations that identified mutations in > 95% of tumors. They detected ctDNA in 100% of patients with stage II–IV NSCLC and in 50% of patients with stage I, with 96% specificity for mutant allele fractions down to ~ 0.02%. Very recently, Abbosh et al. [78] used a tumor-specific phylogenetic approach to profile the ctDNA of the first 100 TRACERx [Tracking NSCLC Evolution Through Therapy (Rx)] study participants. They identified independent predictors of ctDNA release and analyzed the tumor-volume detection limit. The results show that phylogenetic ctDNA profiling tracks the subclonal nature of lung cancer relapse and metastasis, providing a new approach for ctDNA-driven therapeutic studies. More importantly, it was found that ctDNA detection could indicate NSCLC relapse at an earlier stage than could be indicated by CT imaging, with a median interval of 70 days (range 10–346 days). In another TRACERx study, Jamal-Hanjani et al. [79] reported that intratumor heterogeneity, which is mediated through chromosome instability is associated with an increased risk of recurrence or death. This supports the potential value of chromosome instability as a prognostic predictor.
A large portion of NSCLC patients have drug-responsive gene alterations such as EGFR, ALK and ROS1, and as a result, can benefit from targeted therapies. Histological or cytological samples are routinely analyzed for EGFR mutation. However, tumor samples are not always obtainable or of satisfactory quality in advanced NSCLC patients [75]. Further, most NSCLC patients with drug-responsive EGFR mutations develop resistance to first generation EGFR-TKI (e.g., gefitinib, erlotinib, afatinib) after about 10 months of EGFR-TKI therapy. It is preferable to obtain a biopsy in order to detect new mutations and characterize the mechanism of resistance. While mutation analysis can allow for targeting of drug-resistant NSCLC, obtaining sufficient tissue for mutation analysis in patients with advanced disease is challenging, as invasive interventions may be ineffective and unsafe. Furthermore, mutations from the biopsy of a single tumor lesion may not reflect the patient’s complete disease burden, especially in heterogeneous cancers [80, 81]. Still, the benefit of genetic analysis is evident in the fact that EGFR T790M mutation is present in nearly 60% of patients whose disease progresses after initial response to sensitizing EGFR-TKI therapy [82, 83], and osimertinib (AZD9291) is a highly selective, irreversible TKI, that inhibits both EGFR-sensitizing mutations and T790M [84]. Despite the challenges of genetic analysis, in clinical trials, all the selected patients were tested for T790M mutation [85].
ctDNA offers new opportunities to perform mutation analysis in patients for whom a tumor biopsy is unavailable. Several studies have demonstrated that mutations, including the EGFR T790M mutation, detected in plasma ctDNA is highly concordant with those detected in tumor tissue in patients [86,87,88]. This indicates that ctDNA analysis as a liquid biopsy is a feasible and minimally invasive alternative to tissue biopsy. In addition, ctDNA has been used in molecular assessment for diagnosis, serial (real-time) monitoring of resistance mutations [89], and clinical management of patients [90].
The immunotherapeutic significance of extracellular vesicles and ctDNA
Early immunotherapy methods which employed interleukin-2, vaccines, and interferons for NSCLC treatment were not successful [91]. Because smoking- and pollution-associated lung cancers express the highest density of missense mutated genes of any cancer type, suppression of the antigen-presenting machinery may be the cause of immune resistance in these patients [92].
More recently, rapid progress has been made in the use of immune checkpoint inhibitors as a novel immunotherapy approach [93]. Immune checkpoints are proteins on the surface of immune cells, mostly found on the cytotoxic T cells. When bound to a specific ligand from tumor cells, they can transmit inhibitory signals to suppress the cellular adaptive immune response [92]. Many studies suggest that the predominant mechanism by which NSCLC evades detection and elimination by the immune system is exploitation of one such inhibitory pathway through the expression of PD-L1 (B7-H1) [94]. PD-L1 binds to its receptor, PD-1, on surveilling lymphocytes and initiates a signaling cascade which leads to lymphocyte exhaustion, a state of impaired function [95].
Currently, two anti-PD-1 agents, nivolumab and pembrolizumab and one anti-PD-L1 agent, atezolizumab are in advanced stages of development for treatment of advanced or metastatic NSCLC (Table 2). In the CheckMate-057 trial, 582 patients were randomly assigned either to nivolumab group or to docetaxel group, nivolumab treatment demonstrated a survival benefit vs. docetaxel in refractory squamous NSCLC, reporting 41% reduction in risk of death, and better safety profile than standard-of-care chemotherapy [96]. In a study examining patients with previously treated advanced NSCLC and having a minimum of 1% of tumor cells with PD-L1 expression, a total of 1,034 patients received either pembrolizumab at 2 or 10 mg/kg, or docetaxel at 75 mg/m2 every 3 weeks. With pembrolizumab treatment, median overall survival (OS) for patients was 10.4 months at the dosage of 2 mg/kg and 12.7 months at the dosage of 10 mg/kg vs. 8.5 months with docetaxel intervention. OS was improved with both pembrolizumab doses, compared with docetaxel. In patients with at least 50% of tumor cells expressing PD-L1, OS was 14.9 months with pembrolizumab at 2 mg/kg and 17.3 months with pembrolizumab at 10 mg/kg vs. 8.2 months with docetaxel [4]. Based on these results, the FDA approved both nivolumab and pembrolizumab as single agents for the second-line therapy of patients with advanced NSCLC. Nivolumab treatment does not require testing for PD-L1 expression, while PD-L1 overexpression is necessary for pembrolizumab treatment [97].
A deficiency in MMR causes genomes to be unstable and produces high numbers of somatic mutations in cancer cells. As proof-of-concept study, several tests have shown that cancers with MMR deficiency are sensitive to an immune checkpoint blockade with anti-PD-1 antibodies [6, 98, 99]. Recently, Le et al. [6] conducted a study in which 86 patients with MMR deficiency were enrolled across 12 different tumor types to evaluate efficacy of the PD-1 blockade. Objective radiographic responses were observed in 53% of patients and complete responses (CR) were achieved in 21% of patients. Functional analysis in responding patients demonstrated rapid in vivo expansion of neoantigen-specific T cell clones that were reactive to mutant neopeptides found in tumors. These studies revealed that a large proportion of mutant neoantigens in MMR-deficient cancers made them sensitive to an immune checkpoint blockade, regardless of the cancers’ tissue of origin. Recently, the FDA approved the first cancer treatment, pembrolizumab, for any solid tumor with MSI-H or dMMR.
Recently, Rizvi et al. [100] reported that a higher nonsynonymous mutation burden in tumors is associated with improved objective response, durable clinical benefit, and progression-free survival after treatment with pembrolizumab. Because smoking- and pollution-associated lung cancers possess the highest density of missense mutations in expressed genes of any cancer type (roughly 12 mutations per megabase of expressed exonic sequence [92]), lung cancer patients are expected to benefit most from PD-1 inhibitors.
A large portion of patients still cannot benefit from these immunotherapies, and will suffer from disease progression. Notably, both pembrolizumab and nivolumab are very expensive, so the cost-effectiveness should be considered when choosing treatment strategies. The molecular determinants that define this subset of tumors is still unclear, although several markers, including PD-L1 expression, RNA expression signatures, mutational burden and lymphocytic infiltrates, have been evaluated in specific tumor types [94, 100, 101]. The use of pembrolizumab requires the detection of PD-L1 expression or MSI/dMMR status, which is usually determined through protein analysis and tumor tissue DNA analysis, respectively. This is challenging, due to the difficulty of obtaining biopsy tissues. Biopsies may cause significant discomfort to patients, and can sometimes be difficult or impossible to obtain due to location of the tumor.
Liquid biopsy of exosomes or ctDNA may serve as new ways of detecting the PD-L1 expression or MSI/dMMR status. Recently, microsatellite frameshift mutations in exosomal and cellular DNA and exosomal protein profiles were examined by PCR-based DNA fragment analysis and mass spectrometry, respectively [102]. It was found that the coding MSI phenotype of DNA mismatch repair-deficient CRC cells was maintained in their exosomal DNA. This groundbreaking study showed that exosomes and other liquid biopsies may serve as important biomarkers for immunotherapy. Several studies are ongoing, building upon these findings. For example, researchers detect the expression of PD-L1 in circulating tumor cells and white blood cells from patients with advanced NSCLC [103]. Taken together, these results indicate that monitoring the mutation burden is very important for evaluating the response to PD-1-antibody-based immunotherapy. For those patients whose tumor tissues are unavailable, the exosome or ctDNA analysis can provide valuable information about mutation burden.
Conclusions and perspectives
Both ctDNA and tumor extracellular vesicles are natural products that bear many characteristics of the tumors from which they originate. They preserve many valuable pieces of molecular information and serve as promising biomarkers. However, there are still several problems remaining to be solved before their further application. The first challenge is improvement of sensitivity and specificity. There are three major methods to detect mutation via high-throughput sequencing, quantitative PCR (qPCR) and digital PCR. The latter two can only detect one site per reaction. Due to the low abundance of ctDNA and tumor extracellular vesicles in blood, the mutation ratio is sometimes even lower than the sequencing error rate. Some novel sequencing methods such as Duplex Sequencing [104] and o2n-seq [105] have been developed to improve sequencing sensitivity.
Another challenge is the inconsistent results obtained when using ctDNA or exosomal DNA as biomarkers. The inconsistency can probably be explained by low sensitivity, lack of uniformity in technical approaches, variations in sample type preference (serum vs. plasma), differing storage conditions, detection of candidate mutations and insufficiently sensitive detection techniques. A possible solution is to establish standardization of all experimental steps, including storage, purification, detection and analysis. Cross-validation and external validation should also be conducted during the experiments.
The tumor-specific mutations are widely used as markers to distinguish normal cell DNA from ctDNA. However, the information is limited. Recently, Sun et al. [106] and Lehmann-Werman et al. [107] reported that the methylation information of circulating DNA could be used to identify tissue-specific DNA.
With the development of analytical technologies and clinical trials of new targeted drugs, more and more patients will benefit from the use of ctDNA and extracellular vesicles as biomarkers.
References
Torre LA, Siegel RL, Jemal ALung (2016) Cancer statistics. Adv Exp Med Biol 893:1–19. https://doi.org/10.1007/978-3-319-24223-1_1
Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J (2016) Cancer statistics in China, 2015. CA Cancer J Clin 66(2):115–132. https://doi.org/10.3322/caac.21338
Siegel RL, Miller KD, Jemal A (2016) Cancer statistics, 2016. CA Cancer J Clin 66(1):7–30. https://doi.org/10.3322/caac.21332
Hanna N, Johnson D, Temin S, Baker Jr S, Brahmer J, Ellis PM, Giaccone G, Hesketh PJ, Jaiyesimi I, Leighl NB, Riely GJ, Schiller JH, Schneider BJ, Smith TJ, Tashbar J, Biermann WA, Masters G (2017) Systemic therapy for stage IV non-small-cell lung cancer: American Society of Clinical Oncology clinical practice guideline update. J Clin Oncol 35(30):3484–3515. https://doi.org/10.1200/JCO.2017.74..6065
Herbst RS, Baas P, Kim DW, Felip E, Pérez-Gracia JL, Han JY, Molina J, Kim JH, Arvis CD, Ahn MJ, Majem M, Fidler MJ, de Castro Jr G, Garrido M, Lubiniecki GM, Shentu Y, Im E, Dolled-Filhart M, Garon EB (2016) Pembrolizumab versus docetaxel for previously treated, PD-L1-positive, advanced non-small-cell lung cancer (KEYNOTE-010): a randomised controlled trial. Lancet 387(10027):1540–1550. https://doi.org/10.1016/S0140-6736(15)01281-7
Le DT, Durham JN, Smith KN, Wang H, Bartlett BR, Aulakh LK, Lu S, Kemberlin H, Wilt C, Luber BS, Wong F, Azad NS, Rucki AA, Laheru D, Donehower R, Zaheer A, Fisher GA, Crocenzi TS, Lee JJ, Greten TF, Duffy AG, Ciombor KK, Eyring AD, Lam BH, Joe A, Kang SP, Holdhoff M, Danilova L, Cope LC, Zhou S, Goldberg RM, Armstrong DK, Bever KM, Fader AN, Taube J, Housseau F, Spetzler D, Xiao N, Pardoll DM, Papadopoulos N, Kinzler KW, Eshleman JR, Vogelstein B, Anders RA, Diaz LA Jr (2017) Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357(6349):409–413. https://doi.org/10.1126/science.aan6733
Ilié M, Hofman P (2016) Pros: can tissue biopsy be replaced by liquid biopsy? Transl Lung Cancer Res 5(4):420–423. https://doi.org/10.21037/tlcr.2016.08.06
Logozzi M, Angelini DF, Iessi E, Mizzoni D, Di Raimo R, Federici C, Lugini L, Borsellino G, Gentilucci A, Pierella F, Marzio V, Sciarra A, Battistini L, Fais S (2017) Increased PSA expression on prostate cancer exosomes in in vitro condition and in cancer patients. Cancer Lett 403:318–329. https://doi.org/10.1016/j.canlet.2017.06.036
Zhang X, Yuan X, Shi H, Wu L, Qian H, Xu W (2015) Exosomes in cancer: small particle, big player. J Hematol Oncol 8:83. https://doi.org/10.1186/s13045-015-0181-x
Sheridan C (2016) Exosome cancer diagnostic reaches market. Nat Biotechnol 34(4):359–360. https://doi.org/10.1038/nbt0416-359
Zhou L, Lv T, Zhang Q, Zhu Q, Zhan P, Zhu S, Zhang J, Song Y (2017) The biology, function and clinical implications of exosomes in lung cancer. Cancer Lett 407:84–92. https://doi.org/10.1016/j.canlet.2017.08.003
Thierry AR, El Messaoudi S, Gahan PB, Anker P, Stroun M (2016) Origins, structures, and functions of circulating DNA in oncology. Cancer Metastasis Rev 35(3):347–376. https://doi.org/10.1007/s10555-016-9629-x
Schwarzenbach H, Hoon DS, Pantel K (2011) Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer 11(6):426–437. https://doi.org/10.1038/nrc3066
Siravegna G, Marsoni S, Siena S, Bardelli A (2017) Integrating liquid biopsies into the management of cancer. Nat. Rev Clin Oncol 14(9):531–548. https://doi.org/10.1038/nrclinonc.2017.14
Eichmüller SB, Osen W, Mandelboim O, Seliger B Immune, Modulatory (2017) microRNAs involved in tumor attack and tumor immune escape. J Natl Cancer Inst 109(10)
Marinho R, Alcântara PSM, Ottoch JP, Seelaender M (2018) Role of exosomal microRNAs and myomiRs in the development of cancer cachexia-associated muscle wasting. Front Nutr 4:69. https://doi.org/10.3389/fnut.2017.00069 (eCollection 2017)
Meehan K, Vella LJ (2016) The contribution of tumour-derived exosomes to the hallmarks of cancer. Crit Rev. Clin Lab Sci 53(2):121–131. https://doi.org/10.3109/10408363.2015.1092496
Moore C, Kosgodage U, Lange S, Inal JM (2017) The emerging role of exosome and microvesicle- (EMV-) based cancer therapeutics and immunotherapy. Int J Cancer 141(3):428–436. https://doi.org/10.1002/ijc.30672
Abd Elmageed ZY, Yang Y, Thomas R, Ranjan M, Mondal D, Moroz K, Fang Z, Rezk BM, Moparty K, Sikka SC, Sartor O, Abdel-Mageed AB. (2014) Neoplastic reprogramming of patient-derived adipose stem cells by prostate cancer cell-associated. Cells 32(4):983–997. https://doi.org/10.1002/stem.1619
Melo SA, Sugimoto H, O’Connell JT, Kato N, Villanueva A, Vidal A, Qiu L, Vitkin E, Perelman LT, Melo CA, Lucci A, Ivan C, Calin GA, Kalluri R (2014) Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell 26(5):707–721. https://doi.org/10.1016/j.ccell.2014.09.005
Xiao H, Lässer C, Shelke GV, Wang J, Rådinger M, Lunavat TR, Malmhäll C, Lin LH, Li J, Li L, Lötvall J (2014) Mast cell exosomes promote lung adenocarcinoma cell proliferation-role of KIT-stem cell factor signaling. Cell Commun Signal. https://doi.org/10.1186/s12964-014-0064-8
Zhuang G, Wu X, Jiang Z, Kasman I, Yao J, Guan Y, Oeh J, Modrusan Z, Bais C, Sampath D, Ferrara N (2012) Tumour-secreted miR-9 promotes endothelial cell migration and angiogenesis by activating the JAK–STAT pathway. EMBO J 31(17):3513–3523. https://doi.org/10.1038/emboj.2012.183
Umezu T, Tadokoro H, Azuma K, Yoshizawa S, Ohyashiki K, Ohyashiki JH (2014) Exosomal miR-135b shed from hypoxic multiple myeloma cells enhances angiogenesis by targeting factor-inhibiting HIF-1. Blood 124(25):3748–3757. https://doi.org/10.1182/blood-2014-05-576116
Cai Z, Yang F, Yu L, Yu Z, Jiang L, Wang Q, Yang Y, Wang L, Cao X, Wang J (2012) Activated T cell exosomes promote tumor invasion via Fas signaling pathway. J Immunol 188(12):5954–5961. https://doi.org/10.4049/jimmunol.1103466
Hoshino A, Costa-Silva B, Shen TL, Rodrigues G, Hashimoto A, Tesic Mark M, Molina H, Kohsaka S, Di Giannatale A, Ceder S, Singh S, Williams C, Soplop N, Uryu K, Pharmer L, King T, Bojmar L, Davies AE, Ararso Y, Zhang T, Zhang H, Hernandez J, Weiss JM, Dumont-Cole VD, Kramer K, Wexler LH, Narendran A, Schwartz GK, Healey JH, Sandstrom P, Labori KJ, Kure EH, Grandgenett PM, Hollingsworth MA, de Sousa M, Kaur S, Jain M, Mallya K, Batra SK, Jarnagin WR, Brady MS, Fodstad O, Muller V, Pantel K, Minn AJ, Bissell MJ, Garcia Garcia BA, Kang Y, Rajasekhar VK, Ghajar CM, Matei I, Peinado H, Bromberg J, Lyden D (2015) Tumour exosome integrins determine organotropic metastasis. Nature 527(7578):329–335. https://doi.org/10.1038/nature15756
Rahman MA, Barger JF, Lovat F, Gao M, Otterson GA, Nana-Sinkam P (2016) Lung cancer exosomes as drivers of epithelial mesenchymal transition. Oncotarget 7(34):54852–54866. https://doi.org/10.18632/oncotarget.10243
Fabbri M, Paone A, Calore F, Galli R, Gaudio E, Santhanam R, Lovat F, Fadda P, Mao C, Nuovo GJ, Zanesi N, Crawford M, Ozer GH,,Nana-Sinkam P, Perrotti D, Wernicke D, Alder H, Caligiuri MA, Croce CM (2012) MicroRNAs bind to toll-like receptors to induce prometastatic inflammatory response. Proc Natl Acad Sci USA 109(31):E2110–2116. https://doi.org/10.1073/pnas.1209414109
Zhou W, Fong MY, Min Y, Somlo G, Liu L, Palomares MR, Yu Y, Chow A, O’Connor ST, Chin AR, Yen Y, Wang Y, Marcusson EG, Chu P, Wu J, Wu X, Li AX, Li Z, Gao H, Ren X, Boldin MP, Lin PC, Wang SE (2014) Cancer-secreted miR-105 destroys vascular endothelial barriers to promote metastasis. Cancer Cell 25(4):501–515. https://doi.org/10.1016/j.ccr.2014.03.007
Xiao X, Yu S, Li S, Wu J, Ma R, Cao H, Zhu Y, Feng J (2014) Exosomes: decreased sensitivity of lung cancer A549 cells o cisplatin. PLoS One 9(2):e89534. https://doi.org/10.1371/journal.pone.0089534
Qin X, Yu S, Zhou L, Shi M, Hu Y, Xu X, Shen B, Liu S, Yan D, Feng J (2017) Cisplatin-resistant lung cancer cell-derived exosomes increase cisplatin resistance of recipient cells in exosomal miR-100-5p-dependent manner. Int J Nanomed 12:3721–3733. https://doi.org/10.2147/IJN.S131516
Kim MS, Haney MJ, Zhao Y, Mahajan V, Deygen I, Klyachko NL, Inskoe E, Piroyan A, Sokolsky M, Okolie O, Hingtgen SD, Kabanov AV, Batrakova EV (2016) Development of exosome-encapsulated paclitaxel to overcome MDR cancer cells. Nanomedicine 12(3):655–664. https://doi.org/10.1016/j.nano.2015.10.012
Quinn JJ, Chang HY (2016) Unique features of long non-coding RNA biogenesis and function. Nat Rev Genet 17(1):47–62. https://doi.org/10.1038/nrg.2015.10
Kondo Y, Shinjo K, Katsushima K (2017) Long non-coding RNAs as an epigenetic regulator in human cancers. Cancer Sci 108(10):1927–1933. https://doi.org/10.1111/cas.13342
Qu L, Ding J, Chen C, Wu ZJ, Liu B, Gao Y, Chen W, Liu F, Sun W, Li XF, Wang X, Wang Y, Xu ZY, Gao L, Yang Q, Xu B, Li YM, Fang ZY, Xu ZP, Bao Y, Wu DS, Miao X, Sun HY, Sun YH7, Wang HY, Wang LH (2016) Exosome-transmitted lncARSR promotes sunitinib resistance in renal cancer by acting as a competing endogenous RNA. Cancer Cell 29(5):653–668. https://doi.org/10.1016/j.ccell.2016.03.004
Wang Y, Xu YM, Zou YQ, Lin J, Huang B, Liu J, Li J, Zhang J, Yang WM, Min QH, Li SQ, Gao QF, Sun F, Chen QG, Zhang L, Jiang YH, Deng LB, Wang XZ (2017) Identification of differential expressed PE exosomal miRNA in lung adenocarcinoma, tuberculosis, and other benign lesions. Medicine 96(44):e8361. https://doi.org/10.1097/MD.0000000000008361
Jakobsen KR, Paulsen BS, Baek R, Varming K, Sorensen BS, Jrgensen MM (2015) Exosomal proteins as potential diagnostic markers in advanced non-small cell lung carcinoma. J Extracell Vesicles 4:26659. https://doi.org/10.3402//jev.v4.26659
Sandfeld-Paulsen B, Aggerholm-Pedersen N, Baek R, Jakobsen KR, Meldgaard P, Folkersen BH, Rasmussen TR, Varming K, Jrgensen MM, Sorensen BS (2016) Exosomal proteins as prognostic biomarkers in non-small cell lung cancer. Mol Oncol 10(10):1595–1602. https://doi.org/10.1016/j.molonc.2016.10.003
Salehi M, Sharifi M (2018) Exosomal miRNAs as novel cancer biomarkers: challenges and opportunities. J Cell Physiol. https://doi.org/10.1002/jcp.26481 (epub ahead of print)
Vanni I, Alama A, Grossi F, Dal Bello MG, Coco S (2017) Exosomes: a new horizon in lung cancer. Drug Discov Today 22(6):927–936. https://doi.org/10.1016/j.drudis.2017.03.004
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136(2):215–233. https://doi.org/10.1016/j.cell.2009.01.002
Ambros V(2004) The functions of animal microRNAs. Nature 431(7006):350–355. https://doi.org/10.1038/nature02871
Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, Guo J, Zhang Y, Chen J, Guo X, Li Li Q, Li X, Wang W, Zhang Y, Wang J, Jiang X, Xiang Y, Xu C, Zheng P, Zhang J, Li R, Zhang H, Shang X, Gong T, Ning G, Wang J, Zen K, Zhang J, Zhang CY (2008) Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 18(10):997–1006. https://doi.org/10.1038/cr.2008.282
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, Peterson A, Noteboom J, O’Briant KC, Allen A, Lin DW, Urban N, Drescher CW, Knudsen BS, Stirewalt DL, Gentleman R, Vessella RL, Nelson PS, Martin DB, Tewari M (2008) Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA 105(30):10513–10518. https://doi.org/10.1073/pnas.0804549105
Rabinowits G, Gerçel-Taylor C, Day JM, Taylor DD, Kloecker GH (2009) Exosomal microRNA: a diagnostic marker for lung cancer. Clin Lung Cancer 10(1):42–46. https://doi.org/10.3816/CLC.2009.n.006
Cazzoli R, Buttitta F, Di Nicola M, Malatesta S, Marchetti A, Rom WN, Pass HI (2013) microRNAs derived from circulating exosomes as noninvasive biomarkers for screening and diagnosing lung cancer. J Thorac Oncol 8(9):1156–1162. https://doi.org/10.1097/JTO.0b013e318299ac32
Munagala R, Aqil F, Gupta RC (2016) Exosomal miRNAs as biomarkers of recurrent lung cancer. Tumour Biol 37(8):10703–10714. https://doi.org/10.1007/s13277-016-4939-8
Wang J, Zheng Y, Zhao MExosome (2017) Based cancer therapy: implication for targeting cancer stem cells. Front Pharmacol 7:533. https://doi.org/10.3389/fphar.2016.00533
Johnsen KB, Gudbergsson JM, Skov MN, Pilgaard L, Moos T, Duroux M (2014) A comprehensive overview of exosomes as drug delivery vehicles-endogenous nanocarriers for targeted cancer therapy. Biochim Biophys Acta 1846(1):75–87. https://doi.org/10.1016/j.bbcan.2014.04.005
Vader P, Mol EA, Pasterkamp G, Schiffelers RM (2016) Extracellular vesicles for drug delivery. Adv Drug Deliv Rev 106(Pt A):148–156. https://doi.org/10.1016/j.addr.2016.02.006
Ha D, Yang N, Nadithe V (2016) Exosomes as therapeutic drug carriers and delivery vehicles across biological membranes: current perspectives and future challenges. Acta Pharm Sin B 6(4):287–296. https://doi.org/10.1016/j.apsb.2016.02.001
Munagala R, Aqil F, Jeyabalan J, Gupta RC (2016) Bovine milk-derived exosomes for drug delivery. Cancer Lett 371(1):48–61. https://doi.org/10.1016/j.canlet.2015.10.020
Aqil F, Kausar H, Agrawal AK, Jeyabalan J, Kyakulaga AH, Munagala R, Gupta R (2016) Exosomal formulation enhances therapeutic response of celastrol against lung cancer. Exp Mol Pathol 101(1):12–21. https://doi.org/10.1016/j.yexmp.2016.05.013
Xue W, Dahlman JE, Tammela T, Khan OF, Sood S, Dave A, Cai W, Chirino LM, Yang GR, Bronson R, Crowley DG, Sahay G, Schroeder A, Langer R, Anderson DG, Jacks T (2014) Small RNA combination therapy for lung cancer. Proc Natl Acad Sci USA 111(34):E3553–3561. https://doi.org/10.1073/pnas.1412686111
Zhuang X, Xiang X, Grizzle W, Sun D, Zhang S, Axtell RC, Ju S, Mu J, Zhang L, Steinman L, Miller D, Zhang HG (2011) Treatment of brain inflammatory diseases by delivering exosome encapsulated anti-inflammatory drugs from the nasal region to the brain. Mol Ther 19(10):1769–1779. https://doi.org/10.1038/mt.2011.164
Subra C, Laulagnier K, Perret B, Record M (2007) Exosome lipidomics unravels lipid sorting at the level of multivesicular bodies. Biochimie 89(2):205–212.https://doi.org/10.1016/j.biochi.2006.10.014
Alvarez-Erviti L, Seow Y, Yin H, Betts C, Lakhal S, Wood MJ (2011) Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat Biotechnol 29(4):341–345. https://doi.org/10.1038/nbt.1807
Shtam TA, Kovalev RA, Varfolomeeva EY, Makarov EM, Kil YV, Filato MV (2013) Exosomes are natural carriers of exogenous siRNA to human cells in vitro. Cell Commun Signal 11:88. https://doi.org/10.1186/1478-811X-11-88
Kamerkar S, LeBleu VS, Sugimoto H, Yang S, Ruivo CF, Melo SA, Lee JJ, Kalluri R (2017) Exosomes facilitate therapeutic targeting of oncogenic KRAS in pancreatic cancer. Nature 546(7659):498–503. https://doi.org/10.1038/nature22341
Wang X, Cao L, Wang Y, Wang X, Liu N, You Y (2012) Regulation of let-7 and its target oncogenes (review). Oncol Lett 3(5):955–960. https://doi.org/10.3892/ol.2012.609
Chin LJ, Ratner E, Leng S, Zhai R, Nallur S, Babar I, Muller RU, Straka E, Su L, Burki EA, Crowell RE, Patel R, Kulkarni T, Homer R, Zelterman D, Kidd KK, Zhu Y, Christiani DC, Belinsky SA, Slack FJ, Weidhaas JB (2008) A SNP in a let-7 microRNA complementary site in the KRAS 3′ untranslated region increases non-small cell lung cancer risk. Cancer Res 68(20):8535–8540. https://doi.org/10.1158/0008-5472.CAN-08-2129
Nelson HH, Christensen BC, Plaza SL, Wiencke JK, Marsit CJ, Kelsey KT (2010) KRAS mutation, KRAS-LCS6 polymorphism, and non-small cell lung cancer. Lung Cancer 69(1):51–53. https://doi.org/10.1016/j.lungcan.2009.09.008
Zhou B, Tang C, Li J (2017) k-RAS mutation and resistance to epidermal growth factor receptor-tyrosine kinase inhibitor treatment in patients with nonsmall cell lung cancer. J Cancer Res Ther 13(4):699–701. https://doi.org/10.4103/jcrt.JCRT_468_17
Simoff MJ, Lally B, Slade MG, Goldberg WG, Lee P, Michaud GC, Wahidi MM, Chawla M (2013) Symptom management in patients with lung cancer: diagnosis and management of lung cancer, 3rd edn: American College of Chest Physicians evidence-based clinical practice guidelines. Chest 143(5 Suppl):e455S–e497S. https://doi.org/10.1378/chest.12-2366
Pardridge WM (2012) Drug transport across the blood–brain barrier. J Cereb Blood Flow Metab 32(11):1959–1972. https://doi.org/10.1038/jcbfm.2012.126
Peng Q, Zhang S, Yang Q, Zhang T, Wei XQ, Jiang L, Zhang CL, Chen QM, Zhang ZR, Lin YF (2013) Preformed albumin corona, a protective coating for nanoparticles based drug delivery system. Biomaterials 34(33):8521–8530. https://doi.org/10.1016/j.biomaterials.2013.07.102
Yang T, Martin P, Fogarty B, Brown A, Schurman K, Phipps R, Yin VP, Lockman P, Bai S (2015) Exosome delivered anticancer drugs across the blood–brain barrier for brain cancer therapy in Danio rerio. Pharm Res 32(6):2003–2014. https://doi.org/10.1007/s11095-014-1593-y
Pitt JM, Charrier M, Viaud S, André F, Besse B, Chaput N, Zitvogel L (2014) Dendritic cell-derived exosomes as immunotherapies in the fight against cancer. J Immunol 193(3):1006–1011. https://doi.org/10.4049/jimmunol.1400703
Robbins PD, Morelli AE (2014) Regulation of immune responses by extracellular vesicles. Nat Rev Immunol 14(3):195–208. https://doi.org/10.1038/nri3622
Lamparski HG, Metha-Damani A, Yao JY, Patel S, Hsu DH, Ruegg C, Le Pecq JB (2002) Production and characterization of clinical grade exosomes derived from dendritic cells. J Immunol Methods 270(2):211–226
Viaud S, Théry C, Ploix S, Tursz T, Lapierre V, Lantz O, Zitvogel L, Chaput N (2010) Dendritic cell-derived exosomes for cancer immunotherapy: what’s next? Cancer Res 70(4):1281–1285. https://doi.org/10.1158/0008-5472.CAN-09-3276
Pitt JM, André F, Amigorena S, Soria JC, Eggermont A, Kroemer G, Zitvogel L (2016) Dendritic cell-derived exosomes for cancer therapy. J Clin Invest 126(4):1224–1232. https://doi.org/10.1172/JCI81137
Morse MA, Garst J, Osada T, Khan S, Hobeika A, Clay TM, Valente N, Shreeniwas R, Sutton MA, Delcayre A, Hsu DH, Le Pecq JB, Lyerly HK (2005) A phase I study of dexosome immunotherapy in patients with advanced non-small cell lung cancer. J Transl Med 3(1):9. https://doi.org/10.1186/1479-5876-3-9
Besse B, Charrier M, Lapierre V, Dansin E, Lantz O, Planchard D, Le Chevalie T, Livartoski A, Barlesi F, Laplanche A, Ploix S, Vimond N, Peguillet I, Théry C, Lacroix L, Zoernig I, Dhodapkar K, Dhodapkar M, Viaud S, Soria JC, Reiners KS, Pogge von Strandmann E, Vély F, Rusakiewicz S, Eggermont A, Pitt JM, Zitvogel L, Chaput N (2015) Dendritic cell-derived exosomes as maintenance immunotherapy after first line chemotherapy in NSCLC. Oncoimmunology 5(4):e1071008. https://doi.org/10.1080/2162402X.2015.1071008
Kumar L, Verma S, Vaidya B, Gupta VExosomes (2015) Natural carriers for siRNA delivery. Curr Pharm Des 21(31):4556–4565
Vendrell JA, Mau-Them FT, Béganton B, Godreuil S, Coopman P, Solassol J (2017) Circulating cell free tumor DNA detection as a routine tool for lung cancer patient management. Int J Mol Sci. https://doi.org/10.3390/ijms18020264
Ma M, Zhu H, Zhang C, Sun X, Gao X, Chen G (2015) “Liquid biopsy"-ctDNA detection with great potential and challenges. Ann Transl Med 3(16):235. https://doi.org/10.3978/j.issn.2305-5839.2015.09.29
Newman AM, Bratman SV, To J, Wynne JF, Eclov NC, Modlin LA, Liu CL, Neal JW, Wakelee HA, Merritt RE, Shrager JB, Loo BW Jr, Alizadeh AA, Diehn M (2014) An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat Med 20(5):548–554. https://doi.org/10.1038/nm.3519
Abbosh C, Birkbak NJ, Wilson GA, Jamal-Hanjani M, Constantin T, Salari R, Le Quesne J, Moore DA, Veeriah S, Rosenthal R, Marafioti T, Kirkizlar E, Watkins TBK, McGranahan N, Ward S, Martinson L, Riley J, Fraioli F, Al Bakir M, Grönroos E, Zambrana F, Endozo R, Bi WL, Fennessy FM, Sponer N, Johnson D, Laycock J, Shafi S, Czyzewska-Khan J, Rowan A, Chambers T, Matthews N, Turajlic S, Hiley C, Lee SM, Forster MD, Ahmad T, Falzon M, Borg E, Lawrence D, Hayward M, Kolvekar S, Panagiotopoulos N, Janes SM, Thakrar R, Ahmed A, Blackhall F, Summers Y, Hafez D, Naik A, Ganguly A, Kareht S, Shah R, Joseph L, Marie Quinn A, Crosbie PA, Naidu B, Middleton G, Langman G, Trotter S, Nicolson M, Remmen H, Kerr K, Chetty M, Gomersall L, Fennell DA, Nakas A, Rathinam S, Anand G, Khan S, Russell P, Ezhil V, Ismail B, Irvin-Sellers M, Prakash V, Lester JF, Kornaszewska M, Attanoos R, Adams H, Davies H, Oukrif D, Akarca AU, Hartley JA, Lowe HL, Lock S, Iles N, Bell H, Ngai Y, Elgar G, Szallasi Z, Schwarz RF, Herrero J, Stewart A, Quezada SA, Peggs KS, Van Loo P, Dive C, Lin CJ, Rabinowitz M, Aerts HJWL., Hackshaw A, Shaw JA, Zimmermann BG; TRACERx Consortium; PEACE Consortium, Swanton C (2017) Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution. Nature 545(7655):446–451. https://doi.org/10.1038/nature22364
Jamal-Hanjani M, Wilson GA, McGranahan N, Birkbak NJ, Watkins TBK, Veeriah S, Shafi S, Johnson DH, Mitter R, Rosenthal R, Salm M, Horswell S, Escudero M, Matthews N, Rowan A, Chambers T, Moore DA, Turajlic S, Xu H, Lee SM, Forster MD, Ahmad T, Hiley CT, Abbosh C, Falzon M, Borg E, Marafioti T, Lawrence D, Hayward M, Kolvekar S, Panagiotopoulos N, Janes SM, Thakrar R, Ahmed A, Blackhall F, Summers Y, Shah R, Joseph L, Quinn AM, Crosbie PA, Naidu B, Middleton G, Langman G, Trotter S, Nicolson M, Remmen H, Kerr K, Chetty M, Gomersall L, Fennell DA, Nakas A, Rathinam S, Anand G, Khan S, Russell P, Ezhil V, Ismail B, Irvin-Sellers M, Prakash V, Lester JF, Kornaszewska M, Attanoos R, Adams H, Davies H, Dentro S, Taniere P, O’Sullivan B, Lowe HL, Hartley JA, Iles N, Bell H, Ngai Y, Shaw JA, Herrero J, Szallasi Z, Schwarz RF, Stewart A, Quezada SA, Le Quesne J, Van Loo P, Dive C, Hackshaw A, Swanton C; TRACERx Consortium (2017) Tracking the evolution of non-small-cell lung cancer. N Engl J Med 376(22):2109–2121. https://doi.org/10.1056/NEJMoa1616288
Fisher R, Pusztai L, Swanton C (2013) Cancer heterogeneity: implications for targeted therapeutics. Br J Cancer 108(3):479–485. https://doi.org/10.1038/bjc.2012.581
Weber B, Meldgaard P, Hager H, Wu L, Wei W, Tsai J, Khalil A, Nexo E, Sorensen BS (2014) Detection of EGFR mutations in plasma and biopsies from non-small cell lung cancer patients by allele-specific PCR assays. BMC Cancer. https://doi.org/10.1186/1471-2407-14-294
Yu PP, Vose JM, Hayes DF (2015) Genetic cancer susceptibility testing: increased technology, increased complexity. J Clin Oncol 33(31):3533–3534. https://doi.org/10.1200/JCO.2015.63.3628
Riely GJ, Yu Haegfr (2015) The paradigm of an oncogene-driven lung cancer. Clin Cancer Res 21(10):2221–2226. https://doi.org/10.1158/1078-0432.CCR-14-3154
Mok TS, Wu Y-L, Ahn M-J, Garassino MC, Kim HR, Ramalingam SS, Shepherd FA, He Y, Akamatsu H, Theelen WS, Lee CK, Sebastian M, Templeton A, Mann H, Marotti M, Ghiorghiu S, Papadimitrakopoulou VA; AURA3 Investigators (2017) Osimertinib or platinum-pemetrexed in EGFR T790M-positive lung cancer. N Engl J Med 376(7):629–640. https://doi.org/10.1056/NEJMoa1612674
Jänne PA, Yang JC, Kim DW, Planchard D, Ohe Y, Ramalingam SS, Ahn MJ, Kim SW, Su WC, Horn L, Haggstrom D, Felip E, Kim JH, Frewer P, Cantarini M, Brown KH, Dickinson PA, Ghiorghiu S, Ranson M (2015) AZD9291 in EGFR inhibitor-resistant non-small-cell lung cancer. N Engl J Med 372(18):1689–1699. https://doi.org/10.1056/NEJMoa1411817
Castellanos-Rizaldos E, Grimm DG, Tadigotla V, Hurley J, Healy J, Neal PL, Sher M, Venkatesan R, Karlovich C, Raponi M, Krug AK, Noerholm M, Tannous J, Tannous BA, Raez LE, Skog J (2018) Exosome-based detection of EGFR T790M in plasma from non-small cell lung cancer patients. Clin Cancer Res. https://doi.org/10.1158/1078-0432.CCR-17-3369 (epub ahead of print)
Yu Q, Huang F, Zhang M, Ji H, Wu S, Zhao Y, Zhang C, Wu J, Wang B, Pan B, Zhang X, Guo W (2017) Multiplex picoliter-droplet digital PCR for quantitative assessment of EGFR mutations in circulating cell-free DNA derived from advanced non-small cell lung cancer patients. Mol Med Rep 16(2):1157–1166. https://doi.org/10.3892/mmr.2017.6712
Karlovich C, Goldman JW, Sun JM, Mann E, Sequist LV, Konopa K, Wen W, Angenendt P, Horn L, Spigel D, Soria JC, Solomon B, Camidge DR, Gadgeel S, Paweletz C, Wu L, Chien S, O’Donnell P, Matheny S, Despain D, Rolfe L, Raponi M, Allen AR, Park K, Wakelee H (2016) Assessment of EGFR mutation status in matched plasma and tumor tissue of NSCLC patients from a phase i study of rociletinib (CO-1686). Clin Cancer Res. https://doi.org/10.1158/1078-0432.CCR-15-1260
Luo J, Shen L, Zheng D (2014) Diagnostic value of circulating free DNA for the detection of EGFR mutation status in NSCLC: a systematic review and meta-analysis. Sci Rep. https://doi.org/10.1038/srep06269
Vallée A, Marcq M, Bizieux A, Kouri CE, Lacroix H, Bennouna J, Douillard JY, Denis MG (2013) Plasma is a better source of tumor-derived circulating cell-free DNA than serum for the detection of EGFR alterations in lung tumor patients. Lung Cancer 82(2):373–374. https://doi.org/10.1016/j.lungcan.2013.08.014
Al-Moundhri M, O’Brien M, Souberbielle BE (1998) Immunotherapy in lung cancer. Br J Cancer 78(3):282–288
Brahmer JR, Pardoll DM (2013) Immune checkpoint inhibitors: making immunotherapy a reality for the treatment of lung cancer. Cancer Immunol Res 1(2):85–91. https://doi.org/10.1158/2326-6066.CIR-13-0078
Sgambato A, Casaluce F, Sacco PC, Palazzolo G, Maione P, Rossi A, Ciardiello F, Gridelli C (2016) Anti PD-1 and PDL-1 immunotherapy in the treatment of advanced non-small cell lung cancer (NSCLC): a review on toxicity profile and its management. Curr Drug Saf 11(1):62–68
Taube JM, Klein A, Brahmer JR, Xu H, Pan X, Kim JH, Chen L, Pardoll DM, Topalian SL, Anders RA (2014) Association of PD-1, PD-1 ligands, and other features of the tumor immune microenvironment with response to anti-PD-1 therapy. Clin Cancer Res 20(19):5064–5074. https://doi.org/10.1158/1078-0432.CCR-13-3271
Jiang Y, Li Y, Zhu B (2015) T-cell exhaustion in the tumor micro environment. Cell Death Dis 6:e1792. https://doi.org/10.1038/cddis.2015.162
Borghaei H, Paz-Ares L, Horn L, Spigel DR, Steins M, Ready NE, Chow LQ, Vokes EE, Felip E, Holgado E, Barlesi F, Kohlhäufl M, Arrieta O, Burgio MA, Fayette J, Lena H, Poddubskaya E, Gerber DE, Gettinger SN, Rudin CM, Rizvi N, Crinò L, Blumenschein GR Jr, Antonia SJ, Dorange C, Harbison CT, Graf Finckenstein F, Brahmer JR (2015) Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N Engl J Med 373(17):1627–1639. https://doi.org/10.1056/NEJMoa1507643
Pai-Scherf L, Blumenthal GM, Li H, Subramaniam S, Mishra-Kalyani PS, He K, Zhao H, Yu J, Paciga M, Goldberg KB, McKee AE, Keegan P, Pazdur R (2017) FDA approval summary: pembrolizumab for treatment of metastatic non-small cell lung cancer: first-line therapy and beyond. Oncologist 22(11):1392–1399. https://doi.org/10.1634/theoncologist.2017-0078
Le DT, Uram JN, Wang H, Bartlett BR, Kemberling H, Eyring AD, Skora AD, Luber BS, Azad NS, Laheru D, Biedrzycki B, Donehower RC, Zaheer A, Fisher GA, Crocenzi TS, Lee JJ, Duffy SM, Goldberg RM, de la Chapelle A, Koshiji M, Bhaijee F, Huebner T, Hruban RH, Wood LD, Cuka N, Pardoll DM, Papadopoulos N, Kinzler KW, Zhou S, Cornish TC, Taube JM, Anders RA, Eshleman JR, Vogelstein B, Diaz LA Jr (2015) PD-1 blockade in tumors with mismatch-repair deficiency. N Engl J Med 372(26):2509–2520. https://doi.org/10.1056/NEJMoa1500596
Overman MJ, McDermott R, Leach JL, Lonardi S, Lenz HJ, Morse MA, Desa J, Hill A, Axelson M, Moss RA, Goldberg MV, Cao ZA, Ledeine JM, Maglinte GA, Kopetz S, André T (2017) Nivolumab in patients with metastatic DNA mismatch repair-deficient or microsatellite instability-high colorectal cancer (CheckMate 142) :an open-label, multicentre, phase 2 study. Lancet Oncol 18(9):1182–1191. https://doi.org/10.1016/S1470-2045(17)30422-9
Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, Lee W, Yuan J, Wong P, Ho TS, Miller ML, Rekhtman N, Moreira AL, Ibrahim F, Bruggeman C, Gasmi B, Zappasodi R, Maeda Y, Sander C, Garon EB, Merghoub T, Wolchok JD, Schumacher TN, Chan TA (2015) Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348(6230):124–128. https://doi.org/10.1126/science.aaa1348
Herbst RS, Soria JC, Kowanetz M, Fine GD, Hamid O, Gordon MS, Sosman JA, McDermott DF, Powderly JD, Gettinger SN, Kohrt HE, Horn L, Lawrence DP, Rost S, Leabman M, Xiao Y, Mokatrin A, Koeppen H, Hegde PS, Mellman I, Chen DS, Hodi FS. (2014) Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature 515(7528):563–567. https://doi.org/10.11038/nature14011
Fricke F, Lee J, Michalak M, Warnken U, Hausser I, Suarez-Carmona M, Halama N, Schnölzer M, Kopitz J, Gebert J (2017) TGFBR2-dependent alterations of exosomal cargo and functions in DNA mismatch repair-deficient HCT116 colorectal cancer cells. Cell Commun Signal 15(1):14. https://doi.org/10.1186/s12964-017-0169-y
Ilié M, Szafer-Glusman E, Hofman V, Chamorey E, Lalvée S, Selva E, Leroy S, Marquette CH, Kowanetz M, Hedge P, Punnoose E, Hofman P (2018) Detection of PD-L1 in circulating tumor cells and white blood cells from patients with advanced non-small-cell lung cancer. Ann Oncol 29(1):193–199. https://doi.org/10.1093/annonc/mdx636
Kennedy SR, Schmitt MW, Fox EJ, Kohrn BF, Salk JJ, Ahn EH, Prindle MJ, Kuong KJ, Shen JC, Risques RA, Loeb LA (2014) Detecting ultralow-frequency mutations by duplex sequencing. Nat Protoc 9(11):2586–2606. https://doi.org/10.1038/nprot.2014.170
Wang K, Lai S, Yang X, Zhu T, Lu X, Wu CI, Ruan J (2017) Ultrasensitive and high-efficiency screen of de novo low-frequency mutations by o2n-seq. Nat Commun 8:15335. https://doi.org/10.1038/ncomms15335
Sun K, Jiang P, Chan KC, Wong J, Cheng YK, Liang RH, Chan WK, Ma ES, Chan Chan SL, Cheng SH, Chan RW, Tong YK, Ng SS, Wong RS, Hui DS, Leung TN, Leung TY, Lai PB, Chiu RW, Lo YM (2015) Plasma DNA tissue mapping by genome-wide methylation sequencing for noninvasive prenatal, cancer, and transplantation assessments. Proc Natl Acad Sci USA 112(40):E5503–E5512. https://doi.org/10.1073/pnas.1508736112
Lehmann-Werman R, Neiman D, Zemmour H, Moss J, Magenheim J, Vaknin-Dembinsky A, Rubertsson S, Nellgård B, Blennow K, Zetterberg H, Spalding K, Haller MJ, Wasserfall CH, Schatz DA, Greenbaum CJ, Dorrell C, Grompe M, Zick A, Hubert A, Maoz M, Fendrich V, Bartsch DK, Golan T, Ben Sasson SA, Zamir G, Razin A, Cedar H, Shapiro AM, Glaser B, Shemer R, Dor Y (2016) Identification of tissue-specific cell death using methylation patterns of circulating DNA. Proc Natl Acad Sci USA 113(13):E1826–E1834. https://doi.org/10.1073/pnas.1519286113
Funding
Funding information is not applicable.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Chengliang Huang declares that he has no conflict of interest. Sitong Liu declares that she has no conflict of interest. Xiang Tong declares that he has no conflict of interest. Hong Fan declares that she has no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Huang, C., Liu, S., Tong, X. et al. Extracellular vesicles and ctDNA in lung cancer: biomarker sources and therapeutic applications. Cancer Chemother Pharmacol 82, 171–183 (2018). https://doi.org/10.1007/s00280-018-3586-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00280-018-3586-8