Abstract
This work reports the capacity of 137 strains of starter and non-starter LAB belonging to nine species of the genera Lactobacillus, Lactococcus, Streptococcus and Leuconostoc (all isolated from artisanal cheeses) to produce histamine, tyramine, putrescine and β-phenylethylamine, the biogenic amines (BA) most commonly found in dairy products. Production assays were performed in liquid media supplemented with the appropriate precursor amino acid; culture supernatants were then tested for BA by (U)HPLC. In addition, the presence of key genes involved in the biosynthetic pathways of the target BA, including the production of putrescine via the agmatine deiminase pathway, was assessed by PCR. Twenty strains were shown to have genes involved in the synthesis of BA; these belonged to the species Lactobacillus brevis (4), Lactobacillus curvatus (3), Lactococcus lactis (11) and Streptococcus thermophilus (2). With the exception of the two S. thermophilus strains, all those possessing genes involved in BA production synthesized the corresponding compound. Remarkably, all the putrescine-producing strains used the agmatine deiminase pathway. Four L. brevis and two L. curvatus strains were found able to produce both tyramine and putrescine. There is increasing interest in the use of autochthonous LAB strains in starter and adjunct cultures for producing dairy products with ‘particular geographic indication’ status. Such strains should not produce BA; the present results show that BA production capacity should be checked by (U)HPLC and PCR.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lactic acid bacteria (LAB) play an essential role in the production of fermented dairy products, with Lactococcus lactis and Streptococcus thermophilus being the species most commonly used as primary fermentation starters [1]. Their major function is the rapid production of lactic acid from lactose, resulting in a lowering the pH.
The so-called non-starter lactic acid bacteria (NSLAB) participate in the development of the final organoleptic properties of fermented dairy products [2]. NSLAB may be present in the milk itself, be part of the flora of dairy facilities or be added to fermentations as adjunct cultures [3]. These bacteria are frequently facultative, heterofermentative lactobacilli belonging to the species Lactobacillus casei/paracasei, Lactobacillus plantarum or Lactobacillus curvatus [4, 5]. Leuconostoc may be involved in the development of aroma components [6]. There is increasing interest in the characterization and use of NSLAB from artisanal products for use in tailored cultures to be employed in the manufacture of dairy products with ‘protected geographic indication’ (PGI) status. Their use would help maintain their typical organoleptic characteristics [6–9].
The long and safe history of the use of LAB in dairy products has resulted in the assignment of Qualified Presumption of Safety (QPS) status [awarded by the European Food Safety Authority (EFSA)] to the majority of LAB. However, some properties and enzymatic activities can generate undesirable flavors [10] or even toxic compounds such as biogenic amines (BA) [11], the presence of which should be avoided in dairy products.
BA are low molecular weight nitrogenous compounds formed by the decarboxylation of certain amino acids that may be present in foods. The consumption of foods with high BA concentrations may cause intoxications manifested as headache, nausea or vomiting, alterations in blood pressure, rashes, etc. [12]. Cheese is the fermented food most commonly associated with BA poisoning; indeed, the term cheese reaction was coined to refer to it [13]. Tyramine, putrescine and histamine are the most commonly encountered and abundant BA, in cheese [11, 14, 15]. Certainly, cheese provides an ideal matrix for the production and accumulation of BA since the amino acid substrates required are made easily available by casein proteolysis, and the low pH favors decarboxylase gene transcription and enzyme activity [11]. Further, cheese naturally contains milk-derived Gram positive LAB, generally of the genera Lactobacillus and Enterococcus, which possess decarboxylating activity [11, 16]. BA-producing strains have also been described among the species most commonly used as dairy starters, such as L. lactis, S. thermophilus and Lactobacillus delbrueckii [17–19]. BA producers may also enter dairy products via contamination [20, 21].
The selection of starter strains with no BA-producing capacity would be a good starting point for reducing BA accumulation in dairy products [22]. Different methods have been devised for assessing the capacity of LAB to produce BA, including the use of differential media and pH indicators [23]. Unfortunately, the strong acidification of the medium occasioned by harmless LAB can result in false negatives. Moreover, these methods target the presence of amino acid decarboxylases and do not test the presence of deimination routes involved in the production of some BA such as putrescine [11]. Analytical methods that directly detect BA compounds in culture supernatants after incubation with an amino acid precursor have also been commonly used [24, 25]. However, culture-independent methods based on PCR techniques, aimed to detect the genetic determinants involved in the synthesis of BA, are now regarded as the most suitable for screening collections of isolates [26]. Agreement between the results obtained by analytical and molecular methods strengthens the case for the use of the latter [27, 28].
In the present work, Ultra-High-Performance Liquid Chromatography [(U)HPLC] and PCR methods were used to examine the capacity of 137 LAB strains (four genera, nine species), isolated from artisanal cheeses, and all with potential for use in dairy starter or adjunct cultures designed for the production of artisanal cheeses with PGI status, to produce histamine, tyramine, putrescine and β-phenylethylamine.
Materials and methods
Bacterial strains
One hundred and thirty-seven strains isolated from different artisanal cheeses [29, 30], identified by comparison of partial 16S rRNA gene sequences, and belonging to four different genera—Lactococcus, Streptococcus, Leuconostoc and Lactobacillus—were assessed for their capacity to produce BA (Table 1). L. lactis, S. thermophilus and Leuconostoc mesenteroides strains were grown statically in M17 (Oxoid) supplemented with 0.5 % glucose and 0.5 % lactose (w/v) at either 30 (L. lactis, L. mesenteroides) or 37 °C (S. thermophilus strains). All Lactobacillus strains, which belonged to six species (Table 1), were grown statically in MRS (Oxoid) at 30 °C, except those belonging to L. delbrueckii which were grown at 37 °C.
In vivo BA production capacity
BA production was assessed in triplicate in culture supernatants of the LAB strains grown for 24 h in 10 ml M17 or MRS broth supplemented with 1 mM tyrosine (M17/MRS-T), 1 mM histidine (M17/MRS-H), 1 mM ornithine (M17/MRS-O) or 1 mM agmatine (M17/MRS-A). Both ornithine and agmatine are precursors of putrescine, although via different pathways. Tyramine, histamine and putrescine detection was performed as previously described [31] after the centrifugation of the cultures (2000×g for 15 min) and filtering of the supernatant through a 0.2-μm pore diameter membrane (Pall, USA), followed by derivatization of 100 μl with diethyl ethoxymethylene malonate. Derivatized samples were analyzed by (U)HPLC in a Waters H-Class ACQUITY UPLC apparatus with a UV detector (Waters, USA) controlled by Empower 2.0 software (Waters), under the conditions described by Redruello et al. [32].
Detection of BA-producing genes
The presence of the tyrosine decarboxylase gene tdcA, the histidine decarboxylase gene hdcA, the ornithine decarboxylase gene odc and the aguA and aguD genes from the agmatine deiminase cluster (AGDI) was checked by PCR using the primer pairs P2-for and P1-rev [33], JV16HC and JV17HC [34], ODC3 and ODC16 [35], and Seq1 and Seq2 [17], respectively. The PCR conditions were those described in [17, 33–35], respectively, and were performed in a MyCycler™ thermal cycler (Bio-Rad, Spain) using DreamTaq polymerase (Fermentas, Lithuania). Total DNA from the strains was obtained as previously described [36] and used as a template in PCR. Total DNA from the tyramine- and putrescine-producing strain Enterococcus faecalis V583 [27], from the ODC+ strain Lactobacillus saerimneri 30A [37], and from the histamine producer Lactobacillus buchneri B301 [38], were used to provide positive controls.
PCR products were separated in 0.8 % (w/v) agarose gels in 1XTAE buffer and visualized after staining with ethidium bromide using a GelDoc 2000 system (Bio-Rad, Hercules, USA). The Gene Ruler DNA ladder mix (Fermentas, Lithuania) was used as molecular weight marker.
Results and discussion
The selection of starter strains with no BA-producing capacity is an important step toward reducing the presence of these toxins in dairy products [22]. In this work, 137 LAB strains, previously isolated from artisanal cheeses made from raw milk, were evaluated for their BA-producing capability.
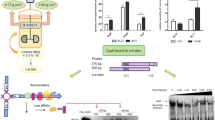
Twenty (14.69 %) of the 137 examined strains were found to possess genes involved in BA production, including four strains of L. brevis, three of L. curvatus, 11 of L. lactis (eight belonging to L. lactis subsp. lactis and three to L. lactis subsp. cremoris) and two of S. thermophilus (Table 1; Fig. 1).
Results of PCR tests for the presence of genes involved in BA production (tdcA, hdcA, odc and aguD-AguA). A representative of each positive species is shown. For L. lactis subsp. lactis and cremoris, a representative of the negative strains is also shown (see text for details). For each BA cluster, the negative (−) and positive (+) controls (E. faecalis V583 for TDC and AGDI, L. buchneri B301 for HDC and L. saerimneri 30A for ODC) are indicated. MW molecular weight standard Gene Ruler (Fermentas), TDC tyramine-producing cluster, HDC histamine-producing cluster, ODC putrescine-producing cluster (via the ornithine decarboxylase pathway), AGDI putrescine-producing cluster (via the agmatine deiminase pathway)
Eighteen (13.1 %) of the tested strains showed the capacity to produce at least one BA in a supplemented medium. These corresponded to all the strains in which the presence of genes involved in BA production was detected by PCR, except for the two S. thermophilus strains (see below). Six strains (4.4 %), four L. brevis and two L. curvatus strains, produced both tyramine and putrescine.
Similar percentages of BA-producing strains have been reported by other authors [19, 39]. During their analysis of dairy isolates, Bunkova et al. [19] found 20 % of the strains tested to produce tyramine and to be positive for the tdcA gene. In some studies, higher percentages of BA producers have been reported [39, 40], but in most of these investigations, strains of Enterococcus were analyzed. The capacity to produce BA is widespread among enterococci and has been shown as a species-specific trait in some enterococcal species [27, 41], thus increasing the occurrence of BA-producing strains. It was, therefore, decided not to include enterococcal strains in the present work.
All of the strains that gave a positive PCR result for the presence of genes involved in BA production were able to synthesize the corresponding BA (Table 1), except for two strains of S. thermophilus. Both of these strains possessed the histidine decarboxylase gene hdcA, but no histamine was found in the culture supernatant after 24 or even after 48 h of culture in M17-H. This might be due to a non-functionality of the HDC cluster or because the conditions assayed were not optimal for histamine production in these strains since the production of BA can be affected by different cultures conditions [11]. Certainly, some authors report that all S. thermophilus strains with the capacity to produce histamine from histidine produce small amounts of histamine in broth but not in milk. [18, 42]. In any event, the present work highlights a good correlation between the results of molecular and functional analysis of BA-producing capacity. All the BA-producing strains returned positive PCR results, indicating that this culture-independent technique is suitable for assessing this property in potential LAB starter strains [28].
Even though two S. thermophilus strains were negative for the in vivo production of BA, their possession of genes involved in BA production must be seen as a risk. These genes could be horizontally transferred to other LAB present in the culture or dairy product [43–45], conferring the ability to produce histamine upon them.
Of the 137 strains tested, seven produced tyramine from tyrosine in broth, and were positive for tdcA in PCR tests (Table 1). All these strains belonged to L. brevis or L. curvatus. Tyramine-producing strains of these species have been isolated from cheeses by other authors [44, 46, 47]. In L. brevis, tyramine production has been described as a strain-level trait—perhaps horizontally acquired [44, 48]. For L. curvatus, there are insufficient data to confirm whether it is a species- or strain-dependent trait. The majority of L. curvatus strains isolated from meat, however, were reported to be tyramine producers [49–51]. All the present tyramine producers, independent of their species, were ‘strong tyramine producers’ (Table 2). L. curvatus strains have been described as strong tyramine producers by other authors [47], showing high conversion rates in broth media supplemented with tyrosine. L. brevis has also been described as a strong tyramine producer, although different media and conditions were assayed and variation in tyramine production capacity was observed [52].
None of the tested strains was able to produce β-phenylethylamine under the present assay conditions. No specific phenylalanine decarboxylases have been described, but several authors have reported that certain tyrosine decarboxylases can use phenylalanine as an alternative substrate, converting it into β-phenylethylamine [53]. In the present work, only the E. faecalis positive control was able to produce β-phenylethylamine in medium supplemented with tyrosine (data not shown).
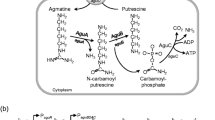
Putrescine is produced from arginine via a decarboxylation and a deimination reaction [11, 54]. However, the order of these reactions can differ, and, depending on that order, two different pathways are recognized: (1) the ornithine decarboxylation pathway (ODC, in which arginine is first deiminated to form ornithine, which is then decarboxylated to form putrescine) and (2) the agmatine deimination pathway (AGDI, in which arginine is first decarboxylated to form agmatine, which is then deiminated to form putrescine) [11, 54]. To distinguish which pathway was being used, the tested strains were grown in media supplemented with ornithine or agmatine. No strain produced putrescine from ornithine. Although the ODC pathway has been described in several LAB strains, including strains of L. brevis [25, 43], it is not a pathway commonly used by dairy bacteria [11, 54, 55]. Thus, the lack of strains with ODC pathway capacity among those tested in the present work was expected. Seventeen strains of the 137 examined were, however, able to produce putrescine from agmatine (Table 1). Putrescine is the most commonly found BA in dairy products [14]. It is not surprising, therefore, that the largest number of BA-producing strains detected should be putrescine producers. It is important to highlight that all the putrescine producers detected in the present survey have the AGDI and not the ODC pathway. Although the prevalence of the AGDI pathway in dairy strains has been previously suggested [11], a test to determine the presence of the AGDI pathway is not usually done. In fact, as far as we are aware, this is the first study to include screening for the AGDI pathway when testing for BA-producing capacity in dairy LAB.
The production of putrescine via the AGDI pathway has, however, been described in L. brevis of non-dairy origin by other authors [48, 56]. All the present strains of L. brevis shown to be putrescine producers were also tyramine producers. It has been suggested that, in this species, the AGDI genetic determinants are linked to those of the TDC pathway, producing a locus of acid resistance mechanisms probably acquired by horizontal gene transfer [43, 48]. Two of the three L. curvatus strains tested produced putrescine from agmatine and also returned positive PCR results (Table 1); both strains were also able to produce tyramine. As in L. brevis, BA-producing capacities of these two strains have been related to acid resistance. The corresponding genes have been described as lying adjacent to one another in the chromosome of some dairy isolates of L. curvatus [43].
Among the L. lactis strains tested, i.e., of both subspecies lactis and cremoris, 11 were shown to produce putrescine from agmatine. Such a capacity had already been reported for some L. lactis strains [17], and putrescine-producing L. lactis can be found in large numbers in cheeses with high putrescine concentrations [55]. Not all the L. lactis strains tested were able to produce putrescine, although the capacity to produce it from agmatine has been described as a species-level trait [17]. Traditionally, BA-producing pathways have been thought horizontally acquired [44, 48]. The present L. lactis strains unable to synthesize putrescine may have lost this capacity during their use in the dairy environment. Putrescine would negatively affect acidification and/or final flavor, and such putrescine-producing strains would have been avoided [17]. The loss of this capacity seems to have occurred in two ways: (1) via the loss of the AGDI pathway genes, as shown for strains of L. lactis subsp. cremoris, and (2) the inactivation of the cluster by an insertion element (IS) in L. lactis subsp. lactis strains [17]. To differentiate between putrescine and non-putrescine producers, a specific PCR test is available [17] in which non-putrescine-producing L. lactis subsp. cremoris strains do not render a PCR band, while L. lactis subsp. lactis non-putrescine-producing strains do, although the amplification product is 1 kb larger than expected due to the presence of an IS element. In the present work, none of the non-putrescine-producing strains of L. lactis subsp. cremoris was associated with any positive amplification, while those of L. lactis subsp. lactis rendered the expected enlarged amplicon (Fig. 1).
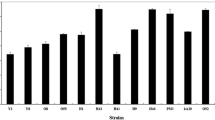
Variation in the efficiency of putrescine production was observed among the producing strains of L. lactis; this allowed their classification as ‘strong’ or ‘medium putrescine producers’ (Table 2). Variation in the capacity to produce putrescine from agmatine has been described before among L. lactis subsp. cremoris strains [31]. In the present work, however, the greatest variation was observed among the L. lactis subsp. lactis producers (Table 2).
One of the most effective measures for reducing the presence of BA in dairy products is the use of starter cultures that have been properly tested and selected as non-BA producers [22]. The present results show that both culture-dependent and culture-independent methods are appropriate for ruling out BA-producing strains for use as starters and adjunct cultures. The culture-independent methods based on PCR testing detected not only BA producers but also non-producer strains that possessed genes involved in BA production; these pose a risk since they might be spread by horizontal gene transfer.
There is increasing interest in the use of autochthonous LAB strains in starter and adjunct cultures for producing dairy products with PGI status [8, 57]. Strains intended for use in their manufacture should be systematically monitored for BA production capacity to avoid the accumulation of these toxins and thus produce safer dairy products.
Conclusions
This work shows that some of the strains belonging to the species most frequently used in the manufacture of dairy products can produce BA, highlighting the importance of adequately selecting indigenous strains for inclusion in starter and adjunct cultures. The prevalence of putrescine-producing strains (which use the AGDI pathway) is noteworthy. The literature contains little on this, even though putrescine is one of the commonest BA in dairy products and the AGDI pathway is the main route of its synthesis. Tests for the presence of the AGDI pathway should be included when examining the BA production capability in dairy strains. As shown in this work, the capacity to produce BA can be tested by either chromatographic or molecular methods, although PCR testing affords many advantages.
References
Parente E, Cogan TM (2004) Starter cultures: general aspects. In: Fox PF, McSweeney PLH, Cogan TM, Guinee TP (eds) Cheese: chemistry, physics and microbiology, vol 1, 3rd edn. Elsevier, Oxford, pp 123–147
Poveda JM, Cabezas L, McSweeney PLH (2004) Free amino acid content of Manchego cheese manufactured with different starter cultures and changes throughout ripening. Food Chem 84(2):213–218
Coeuret V, Dubernet S, Bernardeau M, Gueguen M, Vernoux JP (2003) Isolation, characterisation and identification of lactobacilli focusing mainly on cheeses and other dairy products. Lait 83(4):269–306
Antonsson M, Molin G, Ardo Y (2003) Lactobacillus strains isolated from Danbo cheese as adjunct cultures in a cheese model system. Int J Food Microbiol 85(1–2):159–169
Pisano MB, Patrignani F, Cosentino S, Guerzoni ME, Franz CMAP, Holzapfel WH (2011) Diversity and functional properties of Lactobacillus plantarum-group strains isolated from Italian cheese products. Dairy Sci Technol 91(1):65–76
Nieto-Arribas P, Sesena S, Poveda JM, Palop L, Cabezas L (2010) Genotypic and technological characterization of Leuconostoc isolates to be used as adjunct starters in Manchego cheese manufacture. Food Microbiol 27(1):85–93
Nieto-Arribas P, Poveda JM, Sesena S, Palop L, Cabezas L (2009) Technological characterization of Lactobacillus isolates from traditional Manchego cheese for potential use as adjunct starter cultures. Food Control 20(12):1092–1098
Capozzi V, Spano G (2011) Food microbial biodiversity and “microbes of protected origin”. Front Microbiol 2:237. doi:10.3389/Fmicb.2011.00237
Florez AB, Lopez-Diaz TM, Alvarez-Martin P, Mayo B (2006) Microbial characterisation of the traditional Spanish blue-veined Cabrales cheese: identification of dominant lactic acid bacteria. Eur Food Res Technol 223(4):503–508
Smit G, Smit BA, Engels WJM (2005) Flavour formation by lactic acid bacteria and biochemical flavour profiling of cheese products. FEMS Microbiol Rev 29(3):591–610
Linares DM, Del Rio B, Ladero V, Martinez N, Fernandez M, Martin MC, Alvarez MA (2012) Factors influencing biogenic amines accumulation in dairy products. Front Microbiol 3:180
Ladero V, Calles-Enríquez M, Fernández M, Alvarez MA (2010) Toxicological effects of dietary biogenic amines. Curr Nutr Food Sci 6(2):145–156
ten Brink B, Damink C, Joosten HMLJ, Tveld JHJHI (1990) Occurrence and formation of biologically-active amines in foods. Int J Food Microbiol 11(1):73–84
Fernandez M, Linares DM, del Rio B, Ladero V, Alvarez MA (2007) HPLC quantification of biogenic amines in cheeses: correlation with PCR-detection of tyramine-producing microorganisms. J Dairy Res 74(3):276–282
Spano G, Russo P, Lonvaud-Funel A, Lucas P, Alexandre H, Grandvalet C, Coton E, Coton M, Barnavon L, Bach B, Rattray F, Bunte A, Magni C, Ladero V, Alvarez M, Fernandez M, Lopez P, de Palencia PF, Corbi A, Trip H, Lolkema JS (2010) Biogenic amines in fermented foods. Eur J Clin Nutr 64:S95–S100
Ladero V, Fernandez M, Cuesta I, Alvarez MA (2010) Quantitative detection and identification of tyramine-producing enterococci and lactobacilli in cheese by multiplex qPCR. Food Microbiol 27(7):933–939
Ladero V, Rattray FP, Mayo B, Martin MC, Fernandez M, Alvarez MA (2011) Sequencing and transcriptional analysis of the biosynthesis gene cluster of putrescine-producing Lactococcus lactis. Appl Environ Microbiol 77(18):6409–6418
Calles-Enriquez M, Eriksen BH, Andersen PS, Rattray FP, Johansen AH, Fernandez M, Ladero V, Alvarez MA (2010) Sequencing and transcriptional analysis of the Streptococcus thermophilus histamine biosynthesis gene cluster: factors that affect differential hdcA expression. Appl Environ Microbiol 76(18):6231–6238
Bunkova L, Bunka F, Hlobilova M, Vanatkova Z, Novakova D, Drab V (2009) Tyramine production of technological important strains of Lactobacillus, Lactococcus and Streptococcus. Eur Food Res Technol 229(3):533–538
Ladero V, Fernandez M, Alvarez MA (2009) Effect of post-ripening processing on the histamine and histamine-producing bacteria contents of different cheeses. Int Dairy J 19(12):759–762
Novella-Rodriguez S, Veciana-Nogues MT, Roig-Sagues AX, Trujillo-Mesa AJ, Vidal-Carou MC (2002) Influence of starter and nonstarter on the formation of biogenic amine in goat cheese during ripening. J Dairy Sci 85(10):2471–2478
EFSA Panel on Biological Hazards (BIOHAZ) (2011) Scientific opinion on risk based control of biogenic amine formation in fermented foods. EFSA J 9(10):2393–2486
Bover-Cid S, Holzapfel WH (1999) Improved screening procedure for biogenic amine production by lactic acid bacteria. Int J Food Microbiol 53(1):33–41
Garcia-Moruno E, Carrascosa AV, Munoz R (2005) A rapid and inexpensive method for the determination of biogenic amines from bacterial cultures by thin-layer chromatography. J Food Prot 68(3):625–629
Coton M, Romano A, Spano G, Ziegler K, Vetrana C, Desmarais C, Lonvaud-Funel A, Lucas P, Coton E (2010) Occurrence of biogenic amine-forming lactic acid bacteria in wine and cider. Food Microbiol 27(8):1078–1085
Landete JM, de las Rivas B, Marcobal A, Munoz R (2007) Molecular methods for the detection of biogenic amine-producing bacteria on foods. Int J Food Microbiol 117(3):258–269
Ladero V, Fernandez M, Calles-Enriquez M, Sanchez-Llana E, Canedo E, Martin MC, Alvarez MA (2012) Is the production of the biogenic amines tyramine and putrescine a species-level trait in enterococci? Food Microbiol 30(1):132–138
Bhardwaj A, Gupta H, Iyer R, Kumar N, Malik RK (2009) Tyramine-producing enterococci are equally detected on tyramine production medium, by quantification of tyramine by HPLC, or by tdc gene-targeted PCR. Dairy Sci Technol 89(6):601–611
Lopez S, Mayo B (1997) Identification and characterization of homofermentative mesophilic Lactobacillus strains isolated from artisan starter-free cheeses. Lett Appl Microbiol 25(4):233–238
Estepar J, Sanchez MD, Alonso L, Mayo B (1999) Biochemical and microbiological characterization of artisanal ‘Penamellera’ cheese: analysis of its indigenous lactic acid bacteria. Int Dairy J 9(10):737–746
Linares DM, del Rio B, Ladero V, Redruello B, Martin MC, Fernandez M, Alvarez MA (2013) The putrescine biosynthesis pathway in Lactococcus lactis is transcriptionally regulated by carbon catabolic repression, mediated by CcpA. Int J Food Microbiol 165(1):43–50
Redruello B, Ladero V, Cuesta I, Alvarez-Buylla JR, Martin MC, Fernandez M, Alvarez MA (2013) A fast, reliable, ultra high performance liquid chromatography method for the simultaneous determination of amino acids, biogenic amines and ammonium ions in cheese, using diethyl ethoxymethylenemalonate as a derivatising agent. Food Chem 139(1–4):1029–1035
Lucas P, Lonvaud-Funel A (2002) Purification and partial gene sequence of the tyrosine decarboxylase of Lactobacillus brevis IOEB 9809. FEMS Microbiol Lett 211(1):85–89
Le Jeune C, Lonvaud-Funel A, ten Brink B, Hofstra H, van der Vossen JM (1995) Development of a detection system for histidine decarboxylating lactic acid bacteria based on DNA probes, PCR and activity test. J Appl Bacteriol 78(3):316–326
Marcobal A, De las Rivas B, Moreno-Arribas MV, Munoz R (2005) Multiplex PCR method for the simultaneous detection of histamine-, tyramine-, and putrescine-producing lactic acid bacteria in foods. J Food Prot 68(4):874–878
Ruiz-Barba JL, Maldonado A, Jimenez-Diaz R (2005) Small-scale total DNA extraction from bacteria and yeast for PCR applications. Anal Biochem 347(2):333–335
Romano A, Trip H, Campbell-Sills H, Bouchez O, Sherman D, Lolkema JS, Lucas PM (2013) Genome sequence of Lactobacillus saerimneri 30a (formerly Lactobacillus sp. strain 30a), a reference lactic acid bacterium strain producing biogenic amines. Genome Announc 1(1):e00097-12
Martin MC, Fernandez M, Linares DM, Alvarez MA (2005) Sequencing, characterization and transcriptional analysis of the histidine decarboxylase operon of Lactobacillus buchneri. Microbiology 151:1219–1228
Pircher A, Bauer F, Paulsen P (2007) Formation of cadaverine, histamine, putrescine and tyramine by bacteria isolated from meat, fermented sausages and cheeses. Eur Food Res Technol 226(1–2):225–231
Lorencova E, Bunkova L, Matoulkova D, Drab V, Pleva P, Kuban V, Bunka F (2012) Production of biogenic amines by lactic acid bacteria and bifidobacteria isolated from dairy products and beer. Int J Food Sci Technol 47(10):2086–2091
Capozzi V, Ladero V, Beneduce L, Fernandez M, Alvarez M, Benoit B, Laurent B, Grieco F, Spano G (2011) Isolation and characterization of tyramine-producing Enterococcus faecium strains from red wine. Food Microbiol 28(3):434–439
Rossi F, Gardini F, Rizzotti L, La Gioia F, Tabanelli G, Torriani S (2011) Quantitative analysis of histidine decarboxylase gene (hdcA) transcription and histamine production by Streptococcus thermophilus PRI60 under conditions relevant to cheese making. Appl Environ Microbiol 77(8):2817–2822
Romano A, Ladero V, Alvarez MA, Lucas PM (2014) Putrescine production via the ornithine decarboxylation pathway improves the acid stress survival of Lactobacillus brevis and is part of a horizontally transferred acid resistance locus. Int J Food Microbiol 175:14–19
Coton E, Coton M (2009) Evidence of horizontal transfer as origin of strain to strain variation of the tyramine production trait in Lactobacillus brevis. Food Microbiol 26(1):52–57
Lucas PM, Wolken WAM, Claisse O, Lolkema JS, Lonvaud-Funel A (2005) Histamine-producing pathway encoded on an unstable plasmid in Lactobacillus hilgardii 0006. Appl Environ Microbiol 71(3):1417–1424
Komprda T, Burdychova R, Dohnal V, Cwikova O, Sladkova P, Dvorackova H (2008) Tyramine production in Dutch-type semi-hard cheese from two different producers. Food Microbiol 25(2):219–227
Bunkova L, Bunka F, Mantlova G, Cablova A, Sedlacek I, Svec P, Pachlova V, Kracmar S (2010) The effect of ripening and storage conditions on the distribution of tyramine, putrescine and cadaverine in Edam-cheese. Food Microbiol 27(7):880–888
Lucas PM, Blancato VS, Claisse O, Magni C, Lolkema JS, Lonvaud-Funel A (2007) Agmatine deiminase pathway genes in Lactobacillus brevis are linked to the tyrosine decarboxylation operon in a putative acid resistance locus. Microbiology 153:2221–2230
Aymerich T, Martin B, Garriga M, Vidal-Carou MC, Bover-Cid S, Hugas M (2006) Safety properties and molecular strain typing of lactic acid bacteria from slightly fermented sausages. J Appl Microbiol 100(1):40–49
Bover-Cid S, Hugas M, Izquierdo-Pulido M, Vidal-Carou MC (2001) Amino acid-decarboxylase activity of bacteria isolated from fermented pork sausages. Int J Food Microbiol 66(3):185–189
Pereira CI, Crespo MTB, Romao MVS (2001) Evidence for proteolytic activity and biogenic amines production in Lactobacillus curvatus and L. homohiochii. Int J Food Microbiol 68(3):211–216
Landete JM, Pardo I, Ferrer S (2007) Tyramine and phenylethylamine production among lactic acid bacteria isolated from wine. Int J Food Microbiol 115(3):364–368
Marcobal A, de las Rivas B, Munoz R (2006) First genetic characterization of a bacterial beta-phenylethylamine biosynthetic enzyme in Enterococcus faecium RM58. FEMS Microbiol Lett 258(1):144–149
Wunderlichova L, Bunkova L, Koutny M, Jancova P, Bunka F (2014) Formation, degradation, and detoxification of putrescine by foodborne bacteria: a review. Compr Rev Food Sci Food Saf 13(5):1012–1030
Ladero V, Canedo E, Perez M, Martin MC, Fernandez M, Alvarez MA (2012) Multiplex qPCR for the detection and quantification of putrescine-producing lactic acid bacteria in dairy products. Food Control 27(2):307–313
Arena MP, Romano A, Capozzi V, Beneduce L, Ghariani M, Grieco F, Lucas P, Spano G (2011) Expression of Lactobacillus brevis IOEB 9809 tyrosine decarboxylase and agmatine deiminase genes in wine correlates with substrate availability. Lett Appl Microbiol 53(4):395–402
Tristezza M, Vetrano C, Bleve G, Spano G, Capozzi V, Logrieco A, Mita G, Grieco F (2013) Biodiversity and safety aspects of yeast strains characterized from vineyards and spontaneous fermentations in the Apulia Region, Italy. Food Microbiol 36(2):335–342
Acknowledgments
This work was funded by the Ministry of Economy and Competitiveness, Spain (AGL2013-45431-R), the Fundación para el Fomento en Asturias de la Investigación Científica Aplicada y la Tecnología (FICYT), cofunded by FEDER (GRUPIN14-137) and the INIA (RM2011-00005-00-00). The authors thank Adrian Burton for language assistance.
Conflict of interest
None.
Ethical standard
This study does not involve animal or human subjects.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ladero, V., Martín, M.C., Redruello, B. et al. Genetic and functional analysis of biogenic amine production capacity among starter and non-starter lactic acid bacteria isolated from artisanal cheeses. Eur Food Res Technol 241, 377–383 (2015). https://doi.org/10.1007/s00217-015-2469-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00217-015-2469-z